
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jorge Pereira | + 3787 word(s) | 3787 | 2021-11-26 05:20:26 | | | |
| 2 | Peter Tang | Meta information modification | 3787 | 2021-11-30 02:30:56 | | |
Video Upload Options
Oral diseases (ODs) are highly prevalent affecting over 3.5 billion people, particularly in low- and middle-income countries (LMICs). Saliva is often described as the "mirror of the body" and so different omics approaches such us proteomics, metabolomics and more recently, volatomics are being employed to explore the potential of this biofluid towards the non-invasive diagnosis of ODs.
1. Introduction
2. Physiology of Saliva
Saliva Composition and Production
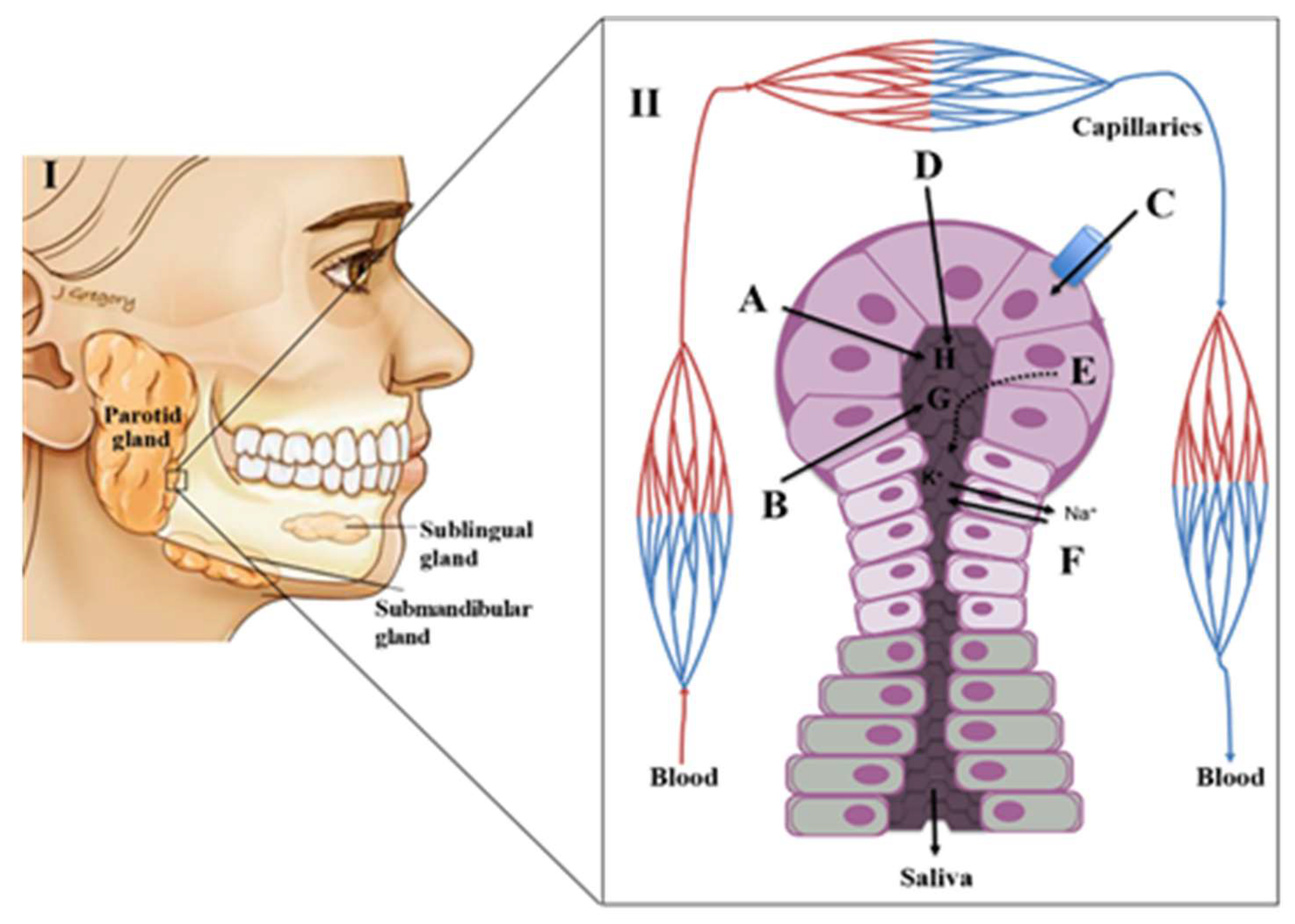
3. Putative Salivary Biomarkers for Oral Diseases
3.1. Gingivitis and Periodontal Disease
3.2. Dental Caries
3.3. Oral Cancer
3.4. Oral Potentially Malignant Disorders (OPMD)
3.5. Burning Mouth Syndrome
3.6. Recurrent Aphthous Ulceration (RAS)
4. Salivary Volatomics
|
Condition Putative Volatile Biomarker |
Metabolic Context |
Ref |
|---|---|---|
|
Periodontal disease |
||
|
pyridine and three methylpyridine isomers (picolines) |
detected in patients but not in controls |
[60] |
|
hydrogen sulphide |
oral bacteria infection |
|
|
methyl mercaptan |
oral bacteria infection |
|
|
Halitosis |
||
|
dimethyl disulphide |
oral bacterial infection |
[60] |
|
dimethyl disulphide, carbon disulphide, VSCs |
drug-related metabolism |
[73] |
|
VSCs |
microbial degradation products of the sulphur-containing amino acids cysteine, cystine and methionine |
[60] |
|
VSCs |
augmented levels detected upon anxiety challenge |
[74] |
|
VSCs, aliphatic amines, branched chain fatty acids, indole and phenol |
oral bacteria metabolism |
[59] |
|
Putrescine, cadaverine, histamine, tyramine, indole, skatole, mercaptans and sulphides |
microbial metabolism of proteinaceous substrates |
[75] |
|
2,3-butanedione; 2,3-pentanedione; Phenol; pyrrole; indole and dimethyl disulphide |
bacterial metabolism of lipids and carbohydrates |
[62] |
|
indole and skatole |
bacterial fermentation products of tryptophan |
[60] |
|
phenol and p-cresol |
bacterial putrefaction metabolites of phenolic amino acids |
|
|
Oral candidiasis |
||
|
3-methyl-2-butanone and styrene |
Candida albicans infection |
[70] |
|
p-xylene, 2-octanone, 2-heptanone and n-butyl acetate |
Candida krusei infection |
|
|
Oral cancer |
||
|
1,4-dichlorobenzene; 1,2-decanediol; 2,5-di-tert-butylphenol and E-3-decen-2-ol |
identified in head and neck cancer cohorts |
[76] |
|
Dietary origin |
||
|
2-heptanone, benzaldehyde, dodecanal, 2-butyl-1-octanol, allyl isothiocyanate |
examples of ketones, aldehydes, alcohols, esters and VSCs obtained from our diet |
[62] |
|
Oxidative stress |
||
|
hexanal and nonanal |
general markers for oxidative damage (endogenously produced from membrane lipid oxidation) |
[62] |
|
Environmental contaminants (air pollutants) |
||
|
long-chain alkane derivatives (hexane, octane and undecane); aromatic compounds (as benzene, toluene, xylenes and styrene) |
common air pollutants found in saliva |
[60] |
References
- Peres, M.A.; Macpherson, L.M.D.; Weyant, R.J.; Daly, B.; Venturelli, R.; Mathur, M.R.; Listl, S.; Celeste, R.K.; Guarnizo-Herreno, C.C.; Kearns, C.; et al. Oral diseases: A global public health challenge. Lancet 2019, 394, 249–260.
- Dawes, C.; Wong, D.T.W. Role of Saliva and Salivary Diagnostics in the Advancement of Oral Health. J. Dent. Res. 2019, 98, 133–141.
- Deepa, T.; Thirrunavukkarasu, N. Saliva as a potential diagnostic tool. Indian J. Med. Sci. 2010, 64, 293–306.
- Katsani, K.R.; Sakellari, D. Saliva proteomics updates in biomedicine. J. Biol. Res. 2019, 26, 17.
- Bellagambi, F.G.; Lomonaco, T.; Salvo, P.; Vivaldi, F.; Hangouët, M.; Ghimenti, S.; Biagini, D.; Di Francesco, F.; Fuoco, R.; Errachid, A. Saliva sampling: Methods and devices. An overview. Tractrends Anal. Chem. 2020, 124, 115781.
- Farnaud, S.J.; Kosti, O.; Getting, S.J.; Renshaw, D. Saliva: Physiology and diagnostic potential in health and disease. Sci. World J. 2010, 10, 434–456.
- Gardner, A.; Carpenter, G.; So, P.W. Salivary metabolomics: From diagnostic biomarker discovery to investigating biological function. Metabolites 2020, 10, 47.
- Aps, J.K.; Martens, L.C. Review: The physiology of saliva and transfer of drugs into saliva. Forensic Sci. Int. 2005, 150, 119–131.
- Papacosta, E.; Nassis, G.P. Saliva as a tool for monitoring steroid, peptide and immune markers in sport and exercise science. J. Sci. Med. Sport 2011, 14, 424–434.
- Edgar, W.; O’Mullane, D.; Dawes, C. Saliva and Oral Health; British Dental Association: London, UK, 2004; Volume 146.
- Sugimoto, M.; Saruta, J.; Matsuki, C.; To, M.; Onuma, H.; Kaneko, M.; Soga, T.; Tomita, M.; Tsukinoki, K. Physiological and environmental parameters associated with mass spectrometry-based salivary metabolomic profiles. Metabolomics 2013, 9, 454–463.
- Ishikawa, S.; Sugimoto, M.; Kitabatake, K.; Tu, M.; Sugano, A.; Yamamori, I.; Iba, A.; Yusa, K.; Kaneko, M.; Ota, S.; et al. Effect of timing of collection of salivary metabolomic biomarkers on oral cancer detection. Amino Acids 2017, 49, 761–770.
- Lamy, E.; Mau, M. Saliva proteomics as an emerging, non-invasive tool to study livestock physiology, nutrition and diseases. J. Proteom. 2012, 75, 4251–4258.
- Gröschl, M. Saliva: A reliable sample matrix in bioanalytics. Bioanalysis 2017, 9, 655–668.
- Proctor, G.B. The physiology of salivary secretion. Periodontol. 2000 2016, 70, 11–25.
- Pereira, J.A.M.; Taware, R.; Porto-Figueira, P.; Rapole, S.; Câmara, J.S. Chapter 29–The salivary volatome in breast cancer. In Precision Medicine for Investigators, Practitioners and Providers; Faintuch, J., Faintuch, S., Eds.; Academic Press: Cambridge, MA, USA, 2020; pp. 301–307.
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424.
- Pihlstrom, B.L.; Michalowicz, B.S.; Johnson, N.W. Periodontal diseases. Lancet 2005, 366, 1809–1820.
- Gonçalves, L.D.R.; Soares, M.R.; Nogueira, F.C.S.; Garcia, C.H.S.; Camisasca, D.R.; Domont, G.; Feitosa, A.C.R.; Pereira, D.A.; Zingali, R.B.; Alves, G. Analysis of the salivary proteome in gingivitis patients. J. Periodontal Res. 2011, 46, 599–606.
- Chan, H.H.; Rahim, Z.H.A.; Jessie, K.; Hashim, O.H.; Taiyeb-Ali, T.B. Salivary proteins associated with periodontitis in patients with type 2 diabetes mellitus. Int. J. Mol. Sci. 2012, 13, 4652–4654.
- Gonçalves, L.D.R.; Soares, M.R.; Nogueira, F.C.S.; Garcia, C.; Camisasca, D.R.; Domont, G.; Feitosa, A.C.R.; Pereira, D.d.A.; Zingali, R.B.; Alves, G. Comparative proteomic analysis of whole saliva from chronic periodontitis patients. J. Proteom. 2010, 73, 1334–1341.
- Rangé, H.; Léger, T.; Huchon, C.; Ciangura, C.; Diallo, D.; Poitou, C.; Meilhac, O.; Bouchard, P.; Chaussain, C. Salivary proteome modifications associated with periodontitis in obese patients. J. Clin. Periodontol. 2012, 39, 799–806.
- Wu, Y.; Shu, R.; Luo, L.J.; Ge, L.H.; Xie, Y.F. Initial comparison of proteomic profiles of whole unstimulated saliva obtained from generalized aggressive periodontitis patients and healthy control subjects. J. Periodontal Res. 2009, 44, 636–644.
- Bostanci, N.; Selevsek, N.; Wolski, W.; Grossmann, J.; Bao, K.; Wahlander, A.; Trachsel, C.; Schlapbach, R.; Oztürk, V.Ö.; Afacan, B.; et al. Targeted proteomics guided by label-free quantitative proteome analysis in saliva reveal transition signatures from health to periodontal disease. Mol. Cell. Proteom. 2018, 17, 1392–1409.
- Sorsa, T.; Gursoy, U.K.; Nwhator, S.; Hernandez, M.; Tervahartiala, T.; Leppilahti, J.; Gursoy, M.; Könönen, E.; Emingil, G.; Pussinen, P.J.; et al. Analysis of matrix metalloproteinases, especially MMP-8, in gingival creviclular fluid, mouthrinse and saliva for monitoring periodontal diseases. Periodontol. 2000 2016, 70, 142–163.
- Emecen-Huja, P.; Hasan, I.; Miller, C.S. Biologic markers of failing implants. Dent. Clin. North Am. 2015, 59, 179–194.
- Gursoy, U.K.; Kononen, E.; Pussinen, P.J.; Tervahartiala, T.; Hyvarinen, K.; Suominen, A.L.; Uitto, V.J.; Paju, S.; Sorsa, T. Use of host- and bacteria-derived salivary markers in detection of periodontitis: A cumulative approach. Dis. Markers 2011, 30, 299–305.
- Kaczynski, T.; Wronski, J.; Gluszko, P.; Kryczka, T.; Miskiewicz, A.; Gorski, B.; Radkowski, M.; Strzemecki, D.; Grieb, P.; Gorska, R. Salivary interleukin 6, interleukin 8, interleukin 17A, and tumour necrosis factor alpha levels in patients with periodontitis and rheumatoid arthritis. Cent. Eur. J. Immunol. 2019, 44, 269–276.
- Pitts, N.B.; Zero, D.T.; Marsh, P.D.; Ekstrand, K.; Weintraub, J.A.; Ramos-Gomez, F.; Tagami, J.; Twetman, S.; Tsakos, G.; Ismail, A. Dental caries. Nat. Rev. Dis. Primers 2017, 3, 17030.
- Zhang, C.Z.; Cheng, X.Q.; Li, J.Y.; Zhang, P.; Yi, P.; Xu, X.; Zhou, X.D. Saliva in the diagnosis of diseases. Int. J. Oral Sci. 2016, 8, 133–137.
- Fiorillo, L.; Cervino, G.; Laino, L.; D’Amico, C.; Mauceri, R.; Tozum, T.F.; Gaeta, M.; Cicciu, M. Porphyromonas gingivalis, Periodontal and Systemic Implications: A Systematic Review. Dent. J. (Basel) 2019, 7, 114.
- Chaturvedi, A.K.; Udaltsova, N.; Engels, E.A.; Katzel, J.A.; Yanik, E.L.; Katki, H.A.; Lingen, M.W.; Silverberg, M.J. Oral leukoplakia and risk of progression to oral cancer: A population-based cohort study. J. Natl. Cancer Inst. 2019.
- Zotti, F.; Nocini, R.; Capocasale, G.; Fior, A.; Peretti, M.; Albanese, M. Malignant transformation evidences of Oral Lichen Planus: When the time is of the essence. Oral Oncol. 2020, 104594.
- Sugimoto, M.; Wong, D.T.; Hirayama, A.; Soga, T.; Tomita, M. Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics 2010, 6, 78–95.
- Ishikawa, S.; Sugimoto, M.; Kitabatake, K.; Sugano, A.; Nakamura, M.; Kaneko, M.; Ota, S.; Hiwatari, K.; Enomoto, A.; Soga, T.; et al. Identification of salivary metabolomic biomarkers for oral cancer screening. Sci. Rep. 2016, 6.
- Ishikawa, S.; Sugimoto, M.; Edamatsu, K.; Sugano, A.; Kitabatake, K.; Iino, M. Discrimination of oral squamous cell carcinoma from oral lichen planus by salivary metabolomics. Oral Dis. 2020, 26, 35–42.
- Wei, J.; Xie, G.; Zhou, Z.; Shi, P.; Qiu, Y.; Zheng, X.; Chen, T.; Su, M.; Zhao, A.; Jia, W. Salivary metabolite signatures of oral cancer and leukoplakia. Int. J. Cancer 2011, 129, 2207–2217.
- Wang, Q.; Gao, P.; Wang, X.; Duan, Y. The early diagnosis and monitoring of squamous cell carcinoma via saliva metabolomics. Sci. Rep. 2014, 4.
- Lohavanichbutr, P.; Zhang, Y.; Wang, P.; Gu, H.; Nagana Gowda, G.A.; Djukovic, D.; Buas, M.F.; Raftery, D.; Chen, C. Salivary metabolite profiling distinguishes patients with oral cavity squamous cell carcinoma from normal controls. PLoS ONE 2018, 13.
- Balkwill, F.; Mantovani, A. Inflammation and cancer: Back to Virchow? Lancet 2001, 357, 539–545.
- Aziz, S.; Ahmed, S.S.; Ali, A.; Khan, F.A.; Zulfiqar, G.; Iqbal, J.; Khan, A.A.; Shoaib, M. Salivary Immunosuppressive Cytokines IL-10 and IL-13 Are Significantly Elevated in Oral Squamous Cell Carcinoma Patients. Cancer Invest. 2015, 33, 318–328.
- Dowling, P.; Wormald, R.; Meleady, P.; Henry, M.; Curran, A.; Clynes, M. Analysis of the saliva proteome from patients with head and neck squamous cell carcinoma reveals differences in abundance levels of proteins associated with tumour progression and metastasis. J. Proteom. 2008, 71, 168–175.
- Honarmand, M.; Farhad-Mollashahi, L.; Nakhaee, A.; Nehi, M. Salivary levels of ErbB2 and CEA in oral squamous cell carcinoma patients. Asian Pac. J. Cancer Prev. 2016, 17, 77–80.
- Jou, Y.J.; Lin, C.D.; Lai, C.H.; Tang, C.H.; Huang, S.H.; Tsai, M.H.; Chen, S.Y.; Kao, J.Y.; Lin, C.W. Salivary zinc finger protein 510 peptide as a novel biomarker for detection of oral squamous cell carcinoma in early stages. Clin. Chim. Acta 2011, 412, 1357–1365.
- Hu, S.; Arellano, M.; Boontheung, P.; Wang, J.; Zhou, H.; Jiang, J.; Elashoff, D.; Wei, R.; Loo, J.A.; Wong, D.T. Salivary proteomics for oral cancer biomarker discovery. Clin. Cancer Res. 2008, 14, 6246–6252.
- Yu, J.S.; Chen, Y.T.; Chiang, W.F.; Hsiao, Y.C.; Chu, L.J.; Seei, L.C.; Wu, C.S.; Tu, H.T.; Chen, H.W.; Chen, C.C.; et al. Saliva protein biomarkers To detect oral squamous cell carcinoma in a high-risk population in Taiwan. Proc. Natl. Acad. Sci. USA 2016, 113, 11549–11554.
- Sivadasan, P.; Gupta, M.K.; Sathe, G.; Sudheendra, H.V.; Sunny, S.P.; Renu, D.; Hari, P.S.; Gowda, H.; Suresh, A.; Kuriakose, M.A.; et al. Salivary proteins from dysplastic leukoplakia and oral squamous cell carcinoma and their potential for early detection. J. Proteom. 2020, 212, 103574.
- Dos Santos, E.S.; Ramos, J.C.; Normando, A.G.C.; Mariano, F.V.; Paes Leme, A.F. Epigenetic alterations in salivary gland tumors. Oral Dis. 2019.
- Li, C.; Ye, J.; Zhang, Z.; Gong, Z.; Lin, Z.; Ding, M. Long non-coding RNA RBM5-AS1 promotes the aggressive behaviors of oral squamous cell carcinoma by regulation of miR-1285-3p/YAP1 axis. Biomed. Pharm. 2020, 123, 109723.
- Park, N.J.; Zhou, H.; Elashoff, D.; Henson, B.S.; Kastratovic, D.A.; Abemayor, E.; Wong, D.T. Salivary microRNA: Discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer Res. 2009, 15, 5473–5477.
- Lodi, G.; Carrozzo, M.; Furness, S.; Thongprasom, K. Interventions for treating oral lichen planus: A systematic review. Br. J. Derm. 2012, 166, 938–947.
- Lorenzo-Pouso, A.I.; Pérez-Sayáns, M.; Bravo, S.B.; López-Jornet, P.; García-Vence, M.; Alonso-Sampedro, M.; Carballo, J.; García-García, A.; Zalewska, A. Protein-Based Salivary Profiles as Novel Biomarkers for Oral Diseases. Dis. Markers 2018, 2018.
- Yang, L.L.; Liu, X.Q.; Liu, W.; Cheng, B.; Li, M.T. Comparative analysis of whole saliva proteomes for the screening of biomarkers for oral lichen planus. Inflamm. Res. 2006, 55, 405–407.
- Souza, M.M.; Florezi, G.P.; Nico, M.M.S.; de Paula, F.; Paula, F.M.; Lourenço, S.V. Salivary proteomics in lichen planus: A relationship with pathogenesis? Oral Dis. 2018, 24, 784–792.
- Camisasca, D.R.; da Rós Gonçalves, L.; Soares, M.R.; Sandim, V.; Nogueira, F.C.S.; Garcia, C.H.S.; Santana, R.; de Oliveira, S.P.; Buexm, L.A.; de Faria, P.A.S.; et al. A proteomic approach to compare saliva from individuals with and without oral leukoplakia. J. Proteom. 2017, 151, 43–52.
- Bender, S.D. Burning Mouth Syndrome. Dent. Clin. North Am. 2018, 62, 585–596.
- Ji, E.H.; Diep, C.; Liu, T.; Li, H.; Merrill, R.; Messadi, D.; Hu, S. Potential protein biomarkers for burning mouth syndrome discovered by quantitative proteomics. Mol. Pain 2017, 13.
- Li, Y.; Wang, D.; Zeng, C.; Liu, Y.; Huang, G.; Mei, Z. Salivary metabolomics profile of patients with recurrent aphthous ulcer as revealed by liquid chromatography–tandem mass spectrometry. J. Int. Med. Res. 2018, 46, 1052–1062.
- Amann, A.; de Lacy Costello, B.; Miekisch, W.; Schubert, J.; Buszewski, B.; Pleil, J.; Ratcliffe, N.; Risby, T. The human volatilome: Volatile organic compounds (VOCs) in exhaled breath, skin emanations, urine, feces and saliva. J. Breath Res. 2014, 8, 034001.
- Milanowski, M.; Pomastowski, P.; Ligor, T.; Buszewski, B. Saliva–Volatile Biomarkers and Profiles. Crit. Rev. Anal. Chem. 2017, 47, 251–266.
- de Lacy Costello, B.; Amann, A.; Al-Kateb, H.; Flynn, C.; Filipiak, W.; Khalid, T.; Osborne, D.; Ratcliffe, N.M. A review of the volatiles from the healthy human body. J. Breath Res. 2014, 8, 014001.
- Al-Kateb, H.; de Lacy Costello, B.; Ratcliffe, N. An investigation of volatile organic compounds from the saliva of healthy individuals using headspace-trap/GC-MS. J. Breath Res. 2013, 7, 036004.
- Soini, H.A.; Klouckova, I.; Wiesler, D.; Oberzaucher, E.; Grammer, K.; Dixon, S.J.; Xu, Y.; Brereton, R.G.; Penn, D.J.; Novotny, M.V. Analysis of volatile organic compounds in human saliva by a static sorptive extraction method and gas chromatography-mass spectrometry. J. Chem. Ecol. 2010, 36, 1035–1042.
- Malathi, N.; Mythili, S.; Vasanthi, H.R. Salivary diagnostics: A brief review. ISRN Dent. 2014, 2014, 158786.
- Pfaffe, T.; Cooper-White, J.; Beyerlein, P.; Kostner, K.; Punyadeera, C. Diagnostic potential of saliva: Current state and future applications. Clin. Chem. 2011, 57, 675–687.
- Pedersen, A.M.L.; Sorensen, C.E.; Proctor, G.B.; Carpenter, G.H.; Ekstrom, J. Salivary secretion in health and disease. J. Oral Rehabil. 2018, 45, 730–746.
- Krishnan, K.; Chen, T.; Paster, B.J. A practical guide to the oral microbiome and its relation to health and disease. Oral Dis. 2017, 23, 276–286.
- Kakoei, S.; Barkhori, F.; Mirzazadeh, A.; Mohammadi, M.; Gholamhoseinian, A. Influence of menstrual cycle and salivary ß-estradiol on volatile sulfur compound. J. Oral Health Oral Epidemiol. 2012, 1, 5.
- Boots, A.W.; Smolinska, A.; van Berkel, J.J.; Fijten, R.R.; Stobberingh, E.E.; Boumans, M.L.; Moonen, E.J.; Wouters, E.F.; Dallinga, J.W.; Van Schooten, F.J. Identification of microorganisms based on headspace analysis of volatile organic compounds by gas chromatography-mass spectrometry. J. Breath Res. 2014, 8, 027106.
- Hertel, M.; Hartwig, S.; Schutte, E.; Gillissen, B.; Preissner, R.; Schmidt-Westhausen, A.M.; Paris, S.; Kastner, I.; Preissner, S. Identification of signature volatiles to discriminate Candida albicans, glabrata, krusei and tropicalis using gas chromatography and mass spectrometry. Mycoses 2016, 59, 117–126.
- Hertel, M.; Preissner, R.; Gillissen, B.; Schmidt-Westhausen, A.M.; Paris, S.; Preissner, S. Detection of signature volatiles for cariogenic microorganisms. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 235–244.
- Apatzidou, A.D.; Bakirtzoglou, E.; Vouros, I.; Karagiannis, V.; Papa, A.; Konstantinidis, A. Association between oral malodour and periodontal disease-related parameters in the general population. Acta Odontol. Scand. 2013, 71, 189–195.
- Torsten, M.; Gomez-Moreno, G.; Aguilar-Salvatierra, A. Drug-related oral malodour (halitosis): A literature review. Eur. Rev. Med. Pharm. Sci. 2017, 21, 4930–4934.
- Calil, C.M.; Marcondes, F.K. Influence of anxiety on the production of oral volatile sulfur compounds. Life Sci. 2006, 79, 660–664.
- Milella, L. The Negative Effects of Volatile Sulphur Compounds. J. Vet. Dent. 2015, 32, 99–102.
- Taware, R.; Taunk, K.; Pereira, J.A.M.; Shirolkar, A.; Soneji, D.; Camara, J.S.; Nagarajaram, H.A.; Rapole, S. Volatilomic insight of head and neck cancer via the effects observed on saliva metabolites. Sci. Rep. 2018, 8, 17725.
- Cavaco, C.; Pereira, J.A.M.; Taunk, K.; Taware, R.; Rapole, S.; Nagarajaram, H.; Camara, J.S. Screening of salivary volatiles for putative breast cancer discrimination: An exploratory study involving geographically distant populations. Anal. Bioanal. Chem. 2018, 410, 4459–4468.
- Cavaco, C.; Perestrelo, R.; Silva, C.; Aveiro, F.; Pereira, J.; Câmara, J. Establishment of the Saliva Volatomic Profile as an Exploratory and Non-Invasive Strategy to Find Potential Breast Cancer Biomarkers; Int. Labmate Ltd.: Hertfordshire, UK, 2014.





