
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Hiroaki Konishi | + 1914 word(s) | 1914 | 2021-11-16 08:52:34 | | | |
| 2 | Conner Chen | Meta information modification | 1914 | 2021-12-01 09:45:52 | | |
Video Upload Options
MicroRNAs (miRNAs), short-chain RNAs of 18–22 nt chain length, are expressed in all vertebrates and control tissue development, differentiation, cell growth, and apoptosis in non-cancer and cancer cells. Additionally, miRNAs target mRNAs on the basis of their sequence and decrease protein production by inhibiting mRNA translation or destroying the mRNAs, thereby controlling cellular homeostasis. miRNA expression is controlled through DNA modification, such as by methylation and transcriptional factors through the signal-transduction pathway. miRNAs are transcribed as pri-mature from DNA and are then processed by the Drosha complex, thus generating pre-miRNAs. These pre-miRNAs are transported by exportin 5 from the nucleus to the cytoplasm and further processed by Dicer to form the double-stranded miRNA RNA-induced silencing complex (RISC). RISCs involving single-strand miRNAs, comprising Ago, Dicer, and trans-activation-responsive RNA-binding protein (TRBP) 2, are directed to the mRNA targets, thus regulating protein expressions. To date, the bioinformatics databases TargetScan (http://www.targetscan.org/vert_72/, 26 September 2021) and miRTarBase (http://mirtarbase.cuhk.edu.cn/php/index.php, 26 September 2021) have been used to predict the interaction between miRNAs and target mRNAs.
1. RNA-Binding Proteins
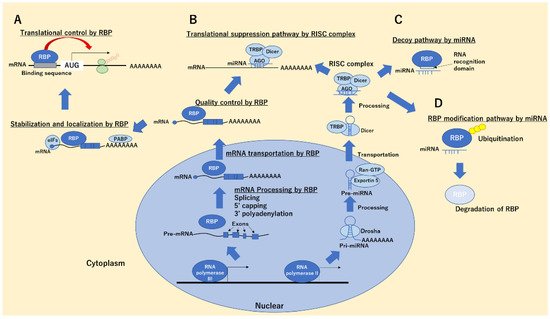
2. RBPs in Cancer Progression and Suppression
3. miRNA–RBP Binding Functions in Cancer Cells
| miRNA | Type of Pathway | Target | Type of Cancer |
Function | Reference |
|---|---|---|---|---|---|
| miR-328 | Decoy | hnRNP E2-CEBPα mRNA | Leukemic blasts | Differentiation | Eiring AM, Cell 2010 |
| Canonical | PIM1 mRNA | Decreased survival | |||
| miR-29 | Decoy | HuR-A20 mRNA | Sarcoma | Differentiation | Balkhi MY, Sci signal, 2013 |
| miR-26a, -584 |
Decoy | hnRNP A1-CDK6 mRNA | Colorectal cancer | Cell growth suppression | Konishi H, Biochem Biophys Res Commun. 2015 |
| miR-574-3p | Decoy | hnRNP L-VEGFA mRNA | Myeloid cells | Inhibition of cell proliferation | Yao P, Nucleic Acids Research, 2017 |
| Canonical | EP300 mRNA | ||||
| miR-574-5p | Decoy | CUGBP1-mPGES-1 mRNA | Lung tumor | Cell growth promotion | Saul MJ, FASEB J, 2019 |
| miR-18a | Degradation | hnRNP A1 | Colorectal cancer | Apoptosis induction | Fujiya M, Oncogene, 2014 |
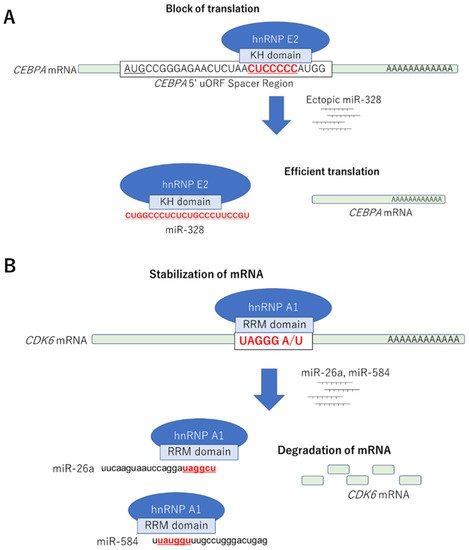
4. Strategies for Identifying Interactions between miRNAs and RBPs
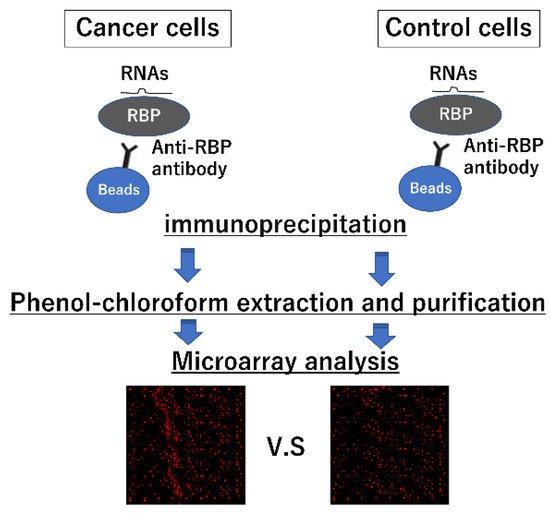
| miRs with Greater than 4-Fold Expression Compared to the Isotype Control IgG | |||
|---|---|---|---|
| Name | ID | Ratio (hnRNP A1/IgG) | LOG2ratio |
| hsa-miR-29a-3p | MIMAT0000086 | 11.49 | 3.52 |
| hsa-miR-26a-5p | MIMAT0000082 | 11.37 | 3.51 |
| hsa-miR-584-5p | MIMAT0003249 | 9.93 | 3.31 |
| hsa-miR-107 | MIMAT0000104 | 9.73 | 3.28 |
| hsa-miR-106b-5p | MIMAT0000680 | 8.99 | 3.17 |
| hsa-miR-1229-5p | MIMAT0022942 | 8.88 | 3.15 |
| hsa-miR-29b-3p | MIMAT0000100 | 8.07 | 3.01 |
| hsa-miR-194-5p | MIMAT0000460 | 8.07 | 3.01 |
| hsa-miR-142-3p | MIMAT0000434 | 7.97 | 2.99 |
| hsa-miR-18a-5p | MIMAT0000072 | 7.93 | 2.99 |
| hsa-let-7c-5p | MIMAT0000064 | 7.31 | 2.87 |
| hsa-miR-16-5p | MIMAT0000069 | 7.22 | 2.85 |
| hsa-miR-500a-3p | MIMAT0002871 | 6.96 | 2.8 |
| hsa-miR-200b-3p | MIMAT0000318 | 6.89 | 2.78 |
| hsa-miR-19a-3p | MIMAT0000073 | 6.55 | 2.71 |
| hsa-miR-222-3p | MIMAT0000279 | 6.4 | 2.68 |
| hsa-let-7b-5p | MIMAT0000063 | 5.98 | 2.58 |
| hsa-miR-23a-3p | MIMAT0000078 | 5.97 | 2.58 |
| hsa-let-7d-5p | MIMAT0000065 | 5.85 | 2.55 |
| hsa-miR-431-3p | MIMAT0004757 | 5.7 | 2.51 |
| hsa-miR-200c-3p | MIMAT0000617 | 5.62 | 2.49 |
| hsa-miR-23b-3p | MIMAT0000418 | 5.27 | 2.4 |
| hsa-miR-27b-3p | MIMAT0000419 | 5.23 | 2.39 |
| hsa-miR-19b-3p | MIMAT0000074 | 5.05 | 2.34 |
| hsa-miR-103a-3p | MIMAT0000101 | 4.95 | 2.31 |
| hsa-miR-1246 | MIMAT0005898 | 4.73 | 2.24 |
| hsa-let-7a-5p | MIMAT0000062 | 4.66 | 2.22 |
| hsa-miR-20a-5p | MIMAT0000075 | 4.5 | 2.17 |
| hsa-miR-27a-3p | MIMAT0000084 | 4.49 | 2.17 |
| hsa-miR-141-3p | MIMAT0000432 | 4.32 | 2.11 |
| hsa-miR-21-5p | MIMAT0000076 | 4.25 | 2.09 |
| hsa-miR-17-5p | MIMAT0000070 | 4.24 | 2.08 |
| hsa-miR-106a-5p | MIMAT0000103 | 4.15 | 2.05 |
| hsa-miR-20b-5p | MIMAT0001413 | 4.11 | 2.04 |
References
- Ortiz-Sánchez, P.; Orero, M.V.; López-Olañeta, M.M.; Larrasa-Alonso, J.; Sanchez-Cabo, F.; Martí-Gómez, C.; Camafeita, E.; Gómez-Salinero, J.M.; Hernández, L.R.; Nielsen, P.J.; et al. Loss of SRSF3 in Cardiomyocytes Leads to Decapping of Contraction-Related mRNAs and Severe Systolic Dysfunction. Circ. Res. 2019, 125, 170–183.
- Zheng, M.; Chen, X.; Cui, Y.; Li, W.; Dai, H.; Yue, Q.; Zhang, H.; Zheng, Y.; Guo, X.; Zhu, H. TULP2, a New RNA-Binding Protein, Is Required for Mouse Spermatid Differentiation and Male Fertility. Front. Cell Dev. Biol. 2021, 9.
- Tollenaere, M.; Tiedje, C.; Rasmussen, S.; Nielsen, J.C.; Vind, A.C.; Blasius, M.; Batth, T.S.; Mailand, N.; Olsen, J.; Gaestel, M.; et al. GIGYF1/2-Driven Cooperation between ZNF598 and TTP in Posttranscriptional Regulation of Inflammatory Signaling. Cell Rep. 2019, 26, 3511–3521.e4.
- Ando, K.; Fujiya, M.; Konishi, H.; Ueno, N.; Inaba, Y.; Moriichi, K.; Ikuta, K.; Tanabe, H.; Ohtake, T.; Kohgo, Y. Heterogeneous Nuclear Ribonucleoprotein A1 Improves the Intestinal Injury by Regulating Apoptosis Through Trefoil Factor 2 in Mice with Anti-CD3–induced Enteritis. Inflamm. Bowel Dis. 2015, 21, 1541–1552.
- Guil, S.; Caceres, J. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nat. Struct. Mol. Biol. 2007, 14, 591–596.
- Tsialikas, J.; Romer-Seibert, J. LIN28: Roles and regulation in development and beyond. Development 2015, 142, 2397–2404.
- O’Brien, K.; Breyne, K.; Ughetto, S.; Laurent, L.C.; Breakefield, X.O. RNA delivery by extracellular vesicles in mammalian cells and its applications. Nat. Rev. Mol. Cell Biol. 2020, 21, 585–606.
- Kang, D.; Lee, Y.; Lee, J.-S. RNA-Binding Proteins in Cancer: Functional and Therapeutic Perspectives. Cancers 2020, 12, 2699.
- Zhou, J.; Jiang, H.; Yuan, T.; Zhou, G.; Li, X.; Wen, K. High hnRNP AB expression is associated with poor prognosis in patients with colorectal cancer. Oncol. Lett. 2019, 18, 6459–6468.
- Carpenter, B.; McKay, M.; Dundas, S.R.; Lawrie, L.C.; Telfer, C.; Murray, G.I. Heterogeneous nuclear ribonucleoprotein K is over expressed, aberrantly localised and is associated with poor prognosis in colorectal cancer. Br. J. Cancer 2006, 95, 921–927.
- Takahashi, K.; Fujiya, M.; Konishi, H.; Murakami, Y.; Iwama, T.; Sasaki, T.; Kunogi, T.; Sakatani, A.; Ando, K.; Ueno, N.; et al. Heterogenous Nuclear Ribonucleoprotein H1 Promotes Colorectal Cancer Progression through the Stabilization of mRNA of Sphingosine-1-Phosphate Lyase 1. Int. J. Mol. Sci. 2020, 21, 4514.
- Yang, G.; Fu, H.; Zhang, J.; Lu, X.; Yu, F.; Jin, L.; Bai, L.; Huang, B.; Shen, L.; Feng, Y.; et al. RNA-Binding Protein Quaking, a Critical Regulator of Colon Epithelial Differentiation and a Suppressor of Colon Cancer. Gastroenterology 2010, 138, 231–240.e5.
- Lee, H.H.; Son, Y.J.; Lee, W.H.; Park, Y.W.; Chae, S.W.; Cho, W.J.; Kim, Y.M.; Choi, H.-J.; Choi, D.H.; Jung, S.W.; et al. Tristetraprolin regulates expression of VEGF and tumorigenesis in human colon cancer. Int. J. Cancer 2009, 126, 1817–1827.
- Eiring, A.M.; Harb, J.G.; Neviani, P.; Garton, C.; Oaks, J.J.; Spizzo, R.; Liu, S.; Schwind, S.; Santhanam, R.; Hickey, C.J.; et al. miR-328 Functions as an RNA Decoy to Modulate hnRNP E2 Regulation of mRNA Translation in Leukemic Blasts. Cell 2010, 140, 652–665.
- Balkhi, M.Y.; Iwenofu, O.H.; Bakkar, N.; Ladner, K.J.; Chandler, D.S.; Houghton, P.J.; London, C.A.; Kraybill, W.; Perrotti, D.; Croce, C.M.; et al. miR-29 Acts as a Decoy in Sarcomas to Protect the Tumor Suppressor A20 mRNA from Degradation by HuR. Sci. Signal. 2013, 6, ra63.
- Konishi, H.; Fujiya, M.; Ueno, N.; Moriichi, K.; Sasajima, J.; Ikuta, K.; Tanabe, H.; Tanaka, H.; Kohgo, Y. microRNA-26a and -584 inhibit the colorectal cancer progression through inhibition of the binding of hnRNP A1-CDK6 mRNA. Biochem. Biophys. Res. Commun. 2015, 467, 847–852.
- Yao, P.; Wu, J.; Lindner, D.; Fox, P.L. Interplay between miR-574-3p and hnRNP L regulates VEGFA mRNA translation and tumorigenesis. Nucleic Acids Res. 2017, 45, 7950–7964.
- Saul, M.J.; Baumann, I.; Bruno, A.; Emmerich, A.C.; Wellstein, J.; Ottinger, S.M.; Contursi, A.; Dovizio, M.; Donnini, S.; Tacconelli, S.; et al. miR-574-5p as RNA decoy for CUGBP1 stimulates human lung tumor growth by mPGES-1 induction. FASEB J. 2019, 33, 6933–6947.





