| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Annarita Fiorillo | + 2469 word(s) | 2469 | 2021-11-17 11:01:23 | | | |
| 2 | Peter Tang | -3 word(s) | 2466 | 2021-11-25 04:03:24 | | |
Video Upload Options
Leishmaniasis, Chagas disease, and human African trypanosomiasis (HAT), also known as sleeping sickness, are vector borne zoonosis that affect millions of people worldwide and lead to the death of about 100,000 humans per year. Among several molecular targets proposed, trypanothione reductase (TR) is of particular interest for its critical role in controlling the parasite’s redox homeostasis.
1. Introduction
2. Relevant Structural Features of TR
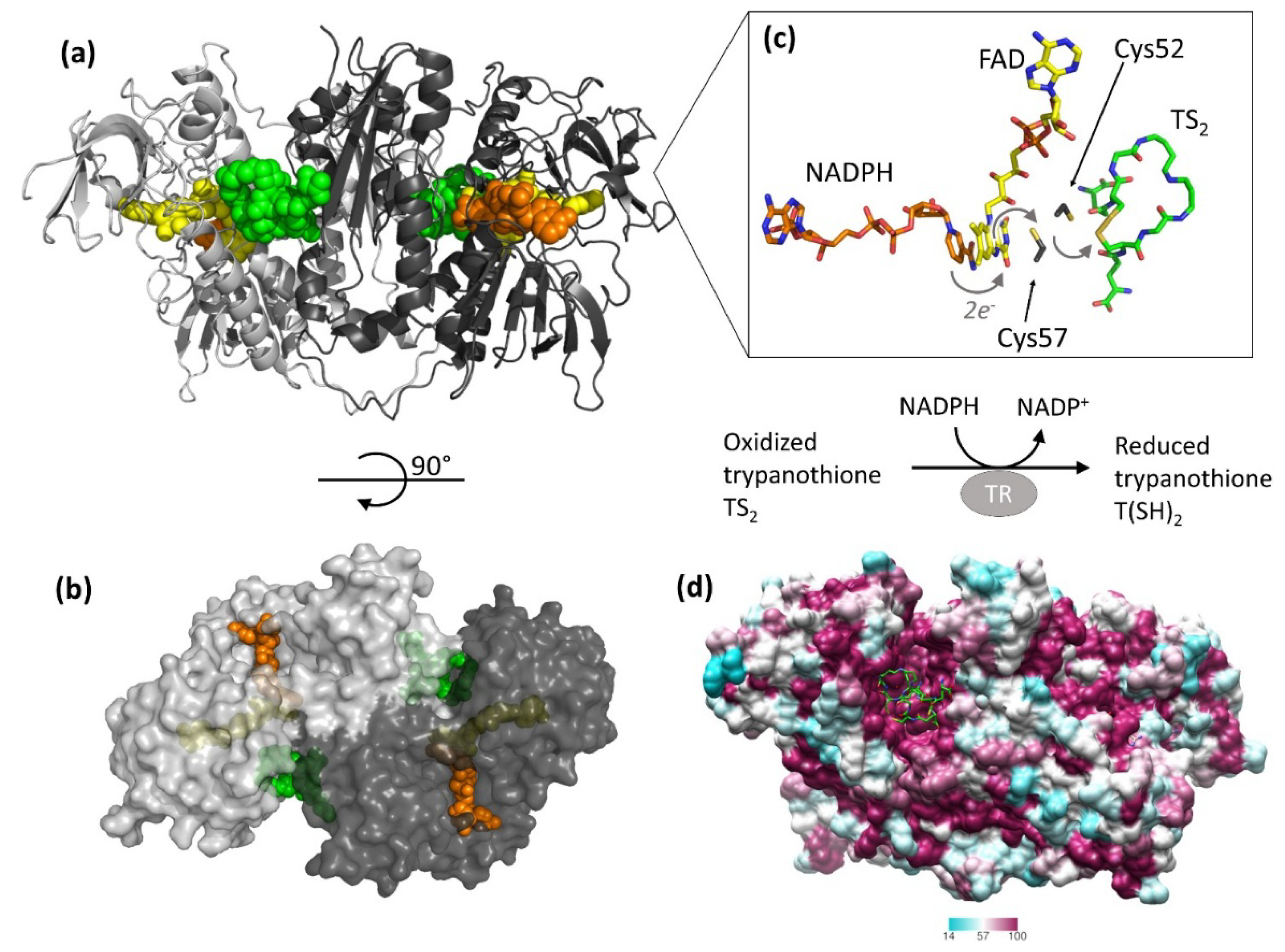
3. Off-Target Evaluation: Comparison with Glutathione Reductase (GR)
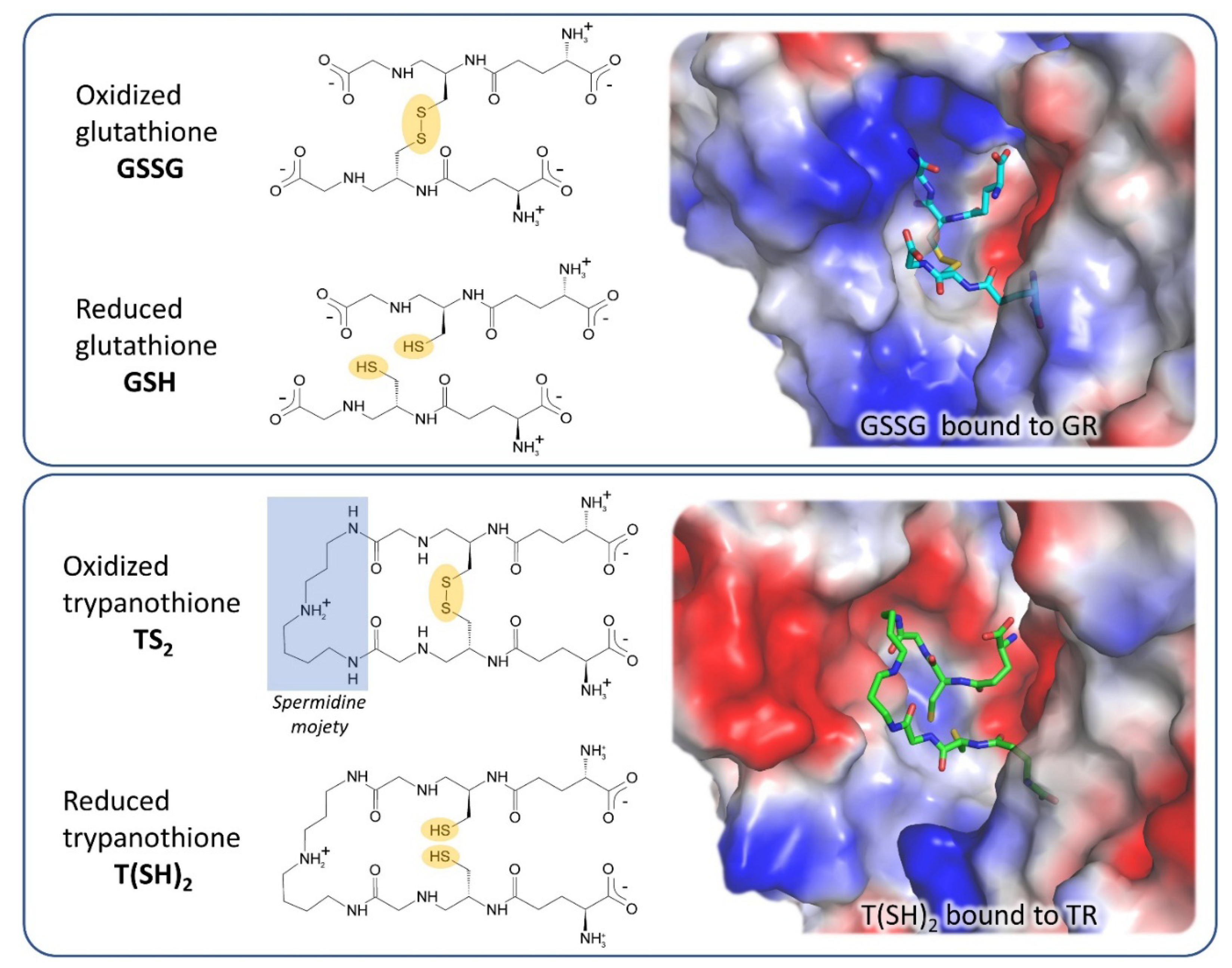
4. Structural Characterization of TR Inhibitors
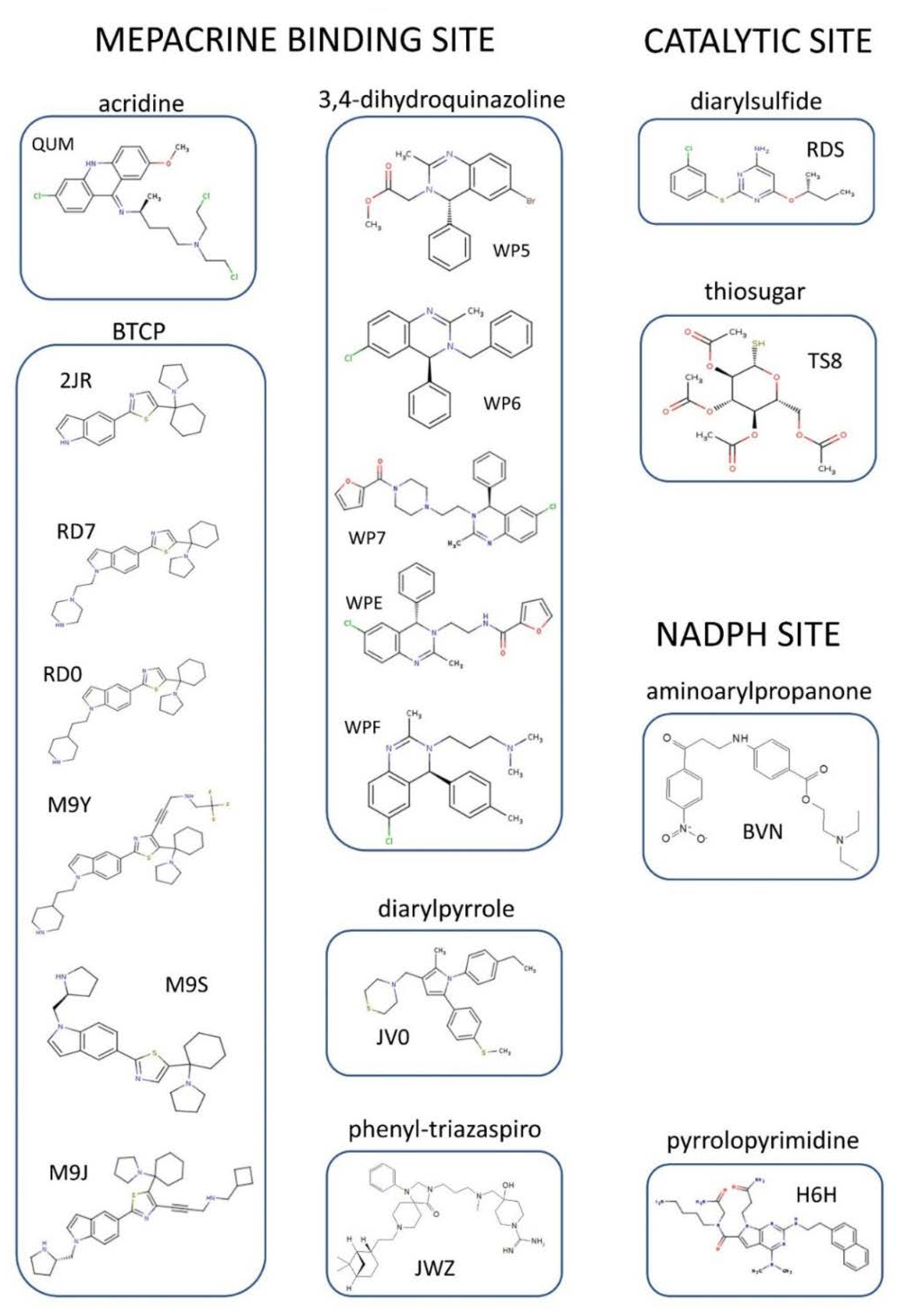
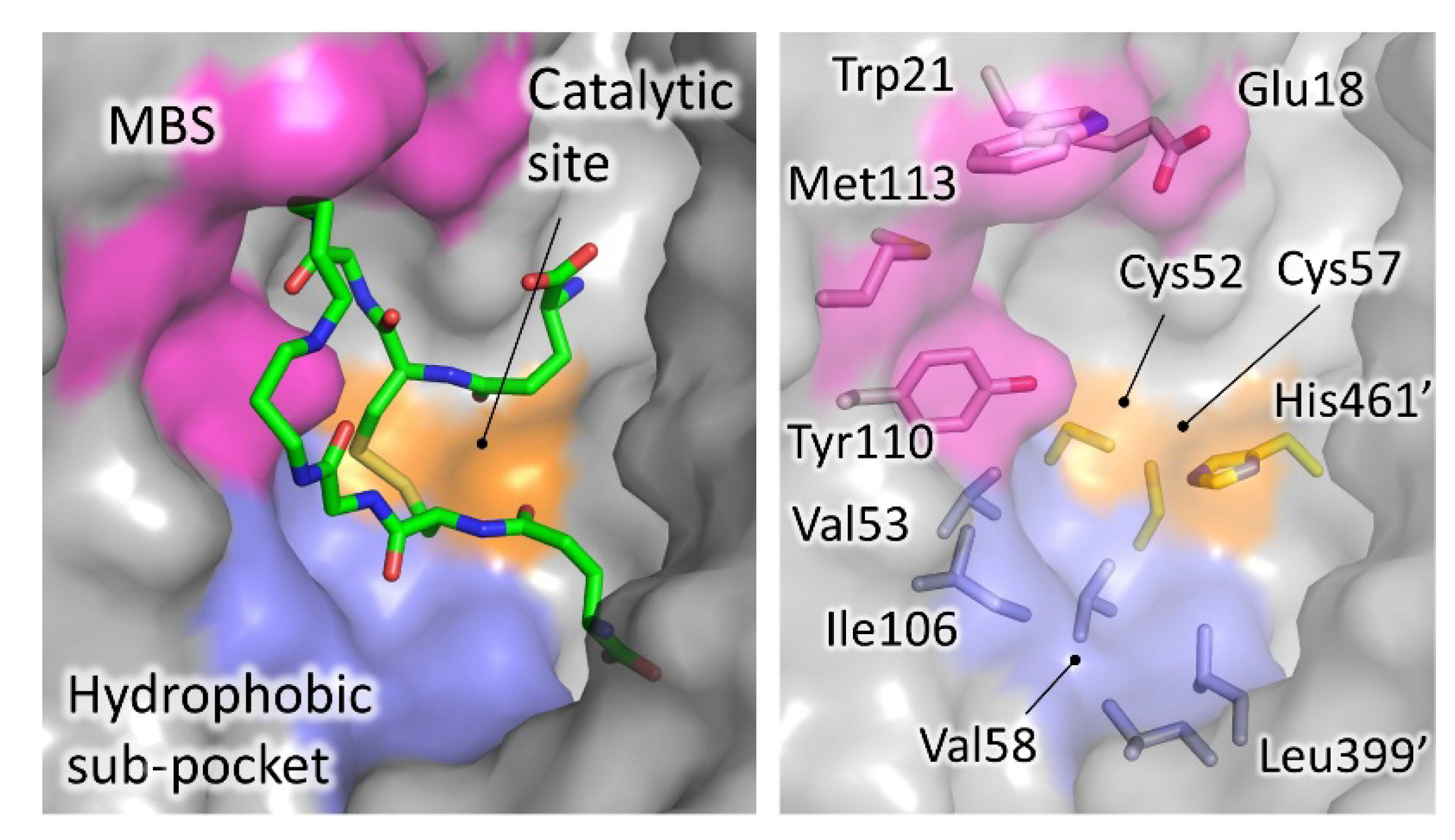
|
Site |
Scaffold |
PDB Code |
Source |
Inhibitor PDB ID (Paper ID a) |
Potency b |
Reference |
|---|---|---|---|---|---|---|
|
MBS |
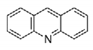
Acridine |
Not available |
Tc |
(Quinacrine or mepacrine) |
Ki: 25 μM |
Jacoby, 1996 |
|
1gxf |
Tb |
QUM (Quin. mustard) |
Irreversible inhibition |
Saravanamuthu, 2004 |
||
|
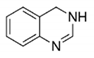
3,4-dihydro quinazoline |
2wp5 |
Tb |
WP5 (1a) |
IC50: 6.8 μM |
Patterson, 2011 |
|
|
2wp6 |
Tb |
WP6 (6a) |
IC50: 0.93 μM |
|||
|
2wpc |
Tb |
WP7 (13e) |
IC50: 0.42 μM |
|||
|
2wpe |
Tb |
WPE (11e) |
IC50: 0.86 μM |
|||
|
2wpf |
Tb |
WPF (29a) |
IC50: 0.23 μM |
|||
|
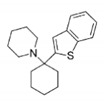
BTCP |
4nev |
Tb |
2JR (10a) |
Ki: 12 μM Inh. [%] b: 43 |
Persch, 2014 |
|
|
4new |
Tc |
2JR (10a) |
Ki: 4 μM Inh. [%] b: 79 |
|||
|
6btl |
Tb |
RD7 (18) |
Ki: 3.8 μM Inh. [%] c: 80 |
De Gasparo, 2018 |
||
|
6bu7 |
Tb |
RD0 (19) |
Ki: 6.4 μM Inh. [%] c: 78 |
|||
|
6oez |
Tb |
M9J ((+)-2) |
Ki: 73 nM |
De Gasparo, 2019 |
||
|
6oey |
Tb |
M9S ((+)-4)) |
Ki: 2.1 μM |
|||
|
6oex |
Tb |
M9Y (5) |
Ki: 1.5 μM |
|||
|
diarylpyrrole |
4apn (B) |
Li |
JV0 (1) |
Ki: 4.6 μM IC50: 13.8 μM |
Baiocco, 2013 |
|
|
phenyl- triazaspiro |
6br5 |
Tb |
JWZ (1) |
IC50: 5.7 µM |
Turcano, 2020 |
|
|
Pyrrolopyrimi- dine |
6i7n (B) |
Li |
H6H (2f) |
IC50: 52.2 µM |
Revuelto, 2019 |
|
|
Catalytic site |
diaryl sulfide |
5ebk |
Li |
RDS (RDS 777) |
Ki: 0.25 µM |
Saccoliti, 2017 |
|
Catalytic site/cysteines |
Metal/thiosugar |
2yau |
Li |
AU-TS8 (auranofin) |
Ki: 0.15 µM |
Ilari, 2012 |
|
Catalytic cysteines |
Metal |
2w0h |
Li |
SB |
Ki: 1.5 µM |
Baiocco, 2009 |
|
2x50 |
Li |
AG |
Ki (Ag1): 500 nM Ki (Ag0): 50 nM |
Baiocco, 2010 |
||
|
NADPH-cavity |
3-amino-1-arylpropan-1-one |
6er5 |
Li |
BVN (3) |
IC50: 12.4 µM |
Turcano, 2018 |
a identification code of the inhibitor as reported in the original paper. b Ki, IC50 and/or percentage of inhibition are reported when available in literature. c Percent inhibition by 40 mM inhibitor in the presence of 40 mM dithiol trypanothione (TS2).
References
- World Health Organization. WHO Publishes On-line Key Information about Leishmaniasis. Available online: https://www.who.int/news-room/fact-sheets/detail/leishmaniasis (accessed on 30 March 2020).
- World Health Organization. WHO Publishes On-line Key Information about Chagas Disease. Available online: https://www.who.int/news-room/fact-sheets/detail/chagas-disease-(american-trypanosomiasis) (accessed on 30 March 2020).
- World Health Organization. WHO Publishes On-line Key Information about HAT. Available online: https://www.who.int/news-room/fact-sheets/detail/trypanosomiasis-human-african-(sleeping-sickness) (accessed on 30 March 2020).
- Field, M.C.; Horn, D.; Fairlamb, A.H.; Ferguson, M.A.; Gray, D.W.; Read, K.D.; De Rycker, M.; Torrie, L.S.; Wyatt, P.G.; Wyllie, S.; et al. Anti-trypanosomatid drug discovery: An ongoing challenge and a continuing need. Nat. Rev. Microbiol. 2017, 15, 217–231.
- Sorci, G.; Faivre, B. Inflammation and oxidative stress in vertebrate host-parasite systems. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 2009, 364, 71–83.
- Kraeva, N.; Horáková, E.; Kostygov, A.Y.; Kořený, L.; Butenko, A.; Yurchenko, V.; Lukeš, J. Catalase in Leishmaniinae: With me or against me? Infect. Genet. Evol. 2017, 50, 121–127.
- El-Sayed, N.M.; Myler, P.J.; Bartholomeu, D.C.; Nilsson, D.; Aggarwal, G.; Tran, A.N.; Ghedin, E.; Worthey, E.A.; Delcher, A.L.; Blandin, G.; et al. The genome sequence of Trypanosoma cruzi, etiologic agent of Chagas disease. Science 2005, 309, 409–415.
- Ivens, A.C.; Peacock, C.S.; Worthey, E.A.; Murphy, L.; Aggarwal, G.; Berriman, M.; Sisk, E.; Rajandream, M.A.; Adlem, E.; Aert, R.; et al. The genome of the kinetoplastid parasite, Leishmania major. Science 2005, 309, 436–442.
- Ilari, A.; Fiorillo, A.; Genovese, I.; Colotti, G. Polyamine-trypanothione pathway: An update. Future Med. Chem. 2017, 9, 61–77.
- Frearson, J.A.; Wyatt, P.G.; Gilbert, I.H.; Fairlamb, A.H. Target assessment for antiparasitic drug discovery. Trends Parasitol. 2007, 23, 589–595.
- Tovar, J.; Cunningham, M.L.; Smith, A.C.; Croft, S.L.; Fairlamb, A.H. Down-regulation of Leishmania donovani trypanothione reductase by heterologous expression of a trans-dominant mutant homologue: Effect on parasite intracellular survival. Proc. Natl. Acad. Sci. USA 1998, 95, 5311–5316.
- Krieger, S.; Schwarz, W.; Ariyanayagam, M.R.; Fairlamb, A.H.; Krauth-Siegel, R.L.; Clayton, C. Trypanosomes lacking trypanothione reductase are avirulent and show increased sensitivity to oxidative stress. Mol. Microbiol. 2000, 35, 542–552.
- Cunningham, M.L.; Fairlamb, A.H. Trypanothione reductase from Leishmania donovani. Purification, characterisation and inhibition by trivalent antimonials. Eur. J. Biochem. 1995, 230, 460–468.
- Baiocco, P.; Colotti, G.; Franceschini, S.; Ilari, A. Molecular basis of antimony treatment in leishmaniasis. J. Med. Chem. 2009, 52, 2603–2612.
- Ilari, A.; Genovese, I.; Fiorillo, F.; Battista, T.; De Ionna, I.; Fiorillo, A.; Colotti, G. Toward a Drug Against All Kinetoplastids: From LeishBox to Specific and Potent Trypanothione Reductase Inhibitors. Mol. Pharm. 2018, 15, 3069–3078.
- Beig, M.; Oellien, F.; Garoff, L.; Noack, S.; Krauth-Siegel, R.L.; Selzer, P.M. Trypanothione reductase: A target protein for a combined in vitro and in silico screening approach. PLoS Negl. Trop. Dis. 2015, 9, e0003773.
- Holloway, G.A.; Charman, W.N.; Fairlamb, A.H.; Brun, R.; Kaiser, M.; Kostewicz, E.; Novello, P.M.; Parisot, J.P.; Richardson, J.; Street, I.P.; et al. Trypanothione reductase high-throughput screening campaign identifies novel classes of inhibitors with antiparasitic activity. Antimicrob. Agents Chemother. 2009, 53, 2824–2833.
- Maccari, G.; Jaeger, T.; Moraca, F.; Biava, M.; Flohé, L.; Botta, M. A fast virtual screening approach to identify structurally diverse inhibitors of trypanothione reductase. Bioorg. Med. Chem. Lett. 2011, 21, 5255–5258.
- Martyn, D.C.; Jones, D.C.; Fairlamb, A.H.; Clardy, J. High-throughput screening affords novel and selective trypanothione reductase inhibitors with anti-trypanosomal activity. Bioorg. Med. Chem. Lett. 2007, 17, 1280–1283.
- Perez-Pineiro, R.; Burgos, A.; Jones, D.C.; Andrew, L.C.; Rodriguez, H.; Suarez, M.; Fairlamb, A.H.; Wishart, D.S. Development of a novel virtual screening cascade protocol to identify potential trypanothione reductase inhibitors. J. Med. Chem. 2009, 52, 1670–1680.
- Richardson, J.L.; Nett, I.R.; Jones, D.C.; Abdille, M.H.; Gilbert, I.H.; Fairlamb, A.H. Improved tricyclic inhibitors of trypanothione reductase by screening and chemical synthesis. ChemMedChem 2009, 4, 1333–1340.
- Salmon-Chemin, L.; Buisine, E.; Yardley, V.; Kohler, S.; Debreu, M.A.; Landry, V.; Sergheraert, C.; Croft, S.L.; Krauth-Siegel, R.L.; Davioud-Charvet, E. 2- and 3-substituted 1,4-naphthoquinone derivatives as subversive substrates of trypanothione reductase and lipoamide dehydrogenase from Trypanosoma cruzi: Synthesis and correlation between redox cycling activities and in vitro cytotoxicity. J. Med. Chem. 2001, 44, 548–565.
- Turcano, L.; Torrente, E.; Missineo, A.; Andreini, M.; Gramiccia, M.; Di Muccio, T.; Genovese, I.; Fiorillo, A.; Harper, S.; Bresciani, A.; et al. Identification and binding mode of a novel Leishmania Trypanothione reductase inhibitor from high throughput screening. PLoS Negl. Trop. Dis. 2018, 12, e0006969.
- Chacón-Vargas, K.F.; Nogueda-Torres, B.; Sánchez-Torres, L.E.; Suarez-Contreras, E.; Villalobos-Rocha, J.C.; Torres-Martinez, Y.; Lara-Ramirez, E.E.; Fiorani, G.; Krauth-Siegel, R.L.; Bolognesi, M.L.; et al. Trypanocidal Activity of Quinoxaline 1,4 Di-N-oxide Derivatives as Trypanothione Reductase Inhibitors. Molecules 2017, 22, 220.
- Colotti, G.; Saccoliti, F.; Gramiccia, M.; Di Muccio, T.; Prakash, J.; Yadav, S.; Dubey, V.K.; Vistoli, G.; Battista, T.; Mocci, S.; et al. Structure-guided approach to identify a novel class of anti-leishmaniasis diaryl sulfide compounds targeting the trypanothione metabolism. Amino Acids 2019, 52, 247–259.
- Da Paixão, V.G.; Pita, S.S.D.R. In silico identification and evaluation of new Trypanosoma cruzi trypanothione reductase (TcTR) inhibitors obtained from natural products database of the Bahia semi-arid region (NatProDB). Comput. Biol. Chem. 2019, 79, 36–47.
- De Gasparo, R.; Halgas, O.; Harangozo, D.; Kaiser, M.; Pai, E.F.; Krauth-Siegel, R.L.; Diederich, F. Targeting a Large Active Site: Structure-Based Design of Nanomolar Inhibitors of Trypanosoma brucei Trypanothione Reductase. Chemistry 2019, 25, 11416–11421.
- Jacomini, A.P.; Silva, M.J.V.; Silva, R.G.M.; Gonçalves, D.S.; Volpato, H.; Basso, E.A.; Paula, F.R.; Nakamura, C.V.; Sarragiotto, M.H.; Rosa, F.A. Synthesis and evaluation against Leishmania amazonensis of novel pyrazolopyridazinone-N-acylhydrazone-(bi)thiophene hybrids. Eur. J. Med. Chem. 2016, 124, 340–349.
- Jagu, E.; Pomel, S.; Diez-Martinez, A.; Rascol, E.; Pethe, S.; Loiseau, P.M.; Labruère, R. Synthesis and antikinetoplastid evaluation of bis(benzyl)spermidine derivatives. Eur. J. Med. Chem. 2018, 150, 655–666.
- Ortalli, M.; Ilari, A.; Colotti, G.; De Ionna, I.; Battista, T.; Bisi, A.; Gobbi, S.; Rampa, A.; Di Martino, R.M.C.; Gentilomi, G.A.; et al. Identification of chalcone-based antileishmanial agents targeting trypanothione reductase. Eur. J. Med. Chem. 2018, 152, 527–541.
- Pandey, R.K.; Kumbhar, B.V.; Srivastava, S.; Malik, R.; Sundar, S.; Kunwar, A.; Prajapati, V.K. Febrifugine analogues as Leishmania donovani trypanothione reductase inhibitors: Binding energy analysis assisted by molecular docking, ADMET and molecular dynamics simulation. J. Biomol. Struct. Dyn. 2017, 35, 141–158.
- Saccoliti, F.; Angiulli, G.; Pupo, G.; Pescatori, L.; Madia, V.N.; Messore, A.; Colotti, G.; Fiorillo, A.; Scipione, L.; Gramiccia, M.; et al. Inhibition of Leishmania infantum trypanothione reductase by diaryl sulfide derivatives. J. Enzyme. Inhib. Med. Chem. 2017, 32, 304–310.
- Zimmermann, L.A.; de Moraes, M.H.; da Rosa, R.; de Melo, E.B.; Paula, F.R.; Schenkel, E.P.; Steindel, M.; Bernardes, L.S.C. Synthesis and SAR of new isoxazole-triazole bis-heterocyclic compounds as analogues of natural lignans with antiparasitic activity. Bioorg. Med. Chem. 2018, 26, 4850–4862.
- Bailey, S.; Smith, K.; Fairlamb, A.H.; Hunter, W.N. Substrate interactions between trypanothione reductase and N1-glutathionylspermidine disulphide at 0.28-nm resolution. Eur. J. Biochem. 1993, 213, 67–75.
- Baiocco, P.; Poce, G.; Alfonso, S.; Cocozza, M.; Porretta, G.C.; Colotti, G.; Biava, M.; Moraca, F.; Botta, M.; Yardley, V.; et al. Inhibition of Leishmania infantum trypanothione reductase by azole-based compounds: A comparative analysis with its physiological substrate by X-ray crystallography. ChemMedChem 2013, 8, 1175–1183.
- Jones, D.C.; Ariza, A.; Chow, W.H.; Oza, S.L.; Fairlamb, A.H. Comparative structural, kinetic and inhibitor studies of Trypanosoma brucei trypanothione reductase with T. cruzi. Mol. Biochem. Parasitol. 2010, 169, 12–19.
- Patterson, S.; Alphey, M.S.; Jones, D.C.; Shanks, E.J.; Street, I.P.; Frearson, J.A.; Wyatt, P.G.; Gilbert, I.H.; Fairlamb, A.H. Dihydroquinazolines as a novel class of Trypanosoma brucei trypanothione reductase inhibitors: Discovery, synthesis, and characterization of their binding mode by protein crystallography. J. Med. Chem. 2011, 54, 6514–6530.
- Fairlamb, A.H.; Cerami, A. Metabolism and functions of trypanothione in the Kinetoplastida. Annu. Rev. Microbiol. 1992, 46, 695–729.
- Stoll, V.S.; Simpson, S.J.; Krauth-Siegel, R.L.; Walsh, C.T.; Pai, E.F. Glutathione reductase turned into trypanothione reductase: Structural analysis of an engineered change in substrate specificity. Biochemistry 1997, 36, 6437–6447.
- Persch, E.; Bryson, S.; Todoroff, N.K.; Eberle, C.; Thelemann, J.; Dirdjaja, N.; Kaiser, M.; Weber, M.; Derbani, H.; Brun, R.; et al. Binding to large enzyme pockets: Small-molecule inhibitors of trypanothione reductase. ChemMedChem 2014, 9, 1880–1891.
- Revuelto, A.; Ruiz-Santaquiteria, M.; de Lucio, H.; Gamo, A.; Carriles, A.A.; Gutiérrez, K.J.; Sánchez-Murcia, P.A.; Hermoso, J.A.; Gago, F.; Camarasa, M.J.; et al. Pyrrolopyrimidine vs. Imidazole-Phenyl-Thiazole Scaffolds in Nonpeptidic Dimerization Inhibitors of Leishmania infantum Trypanothione Reductase. ACS Infect. Dis. 2019, 5, 873–891.