
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Pritha Majumder | + 3847 word(s) | 3847 | 2021-11-08 05:12:19 | | | |
| 2 | Peter Tang | Meta information modification | 3847 | 2021-11-22 05:45:43 | | |
Video Upload Options
The intrinsic cellular heterogeneity and molecular complexity of the mammalian nervous system relies substantially on the dynamic nature and spatiotemporal patterning of gene expression. These features of gene expression are achieved in part through mechanisms involving various epigenetic processes such as DNA methylation, post-translational histone modifications, and non-coding RNA activity, amongst others. In concert, another regulatory layer by which RNA bases and sugar residues are chemically modified enhances neuronal transcriptome complexity. Similar RNA modifications in other systems collectively constitute the cellular epitranscriptome that integrates and impacts various physiological processes. The epitranscriptome is dynamic and is reshaped constantly to regulate vital processes such as development, differentiation and stress responses. Perturbations of the epitranscriptome can lead to various pathogenic conditions, including cancer, cardiovascular abnormalities and neurological diseases. These RNA modifications modulate the stability, transport and, most importantly, translation of RNA.
1. Introduction
2. RNA Metabolism-Associated Neurological Disease Mechanisms
2.1. mRNA Splicing
2.2. mRNA Alternative Polyadenylation
2.3. mRNA Transport and Translation
2.4. mRNA Stability
2.5. miRNA Biogenesis
|
Disease Type |
Altered RNA Metabolism Pathway |
RBP(s) Involved |
Mechanisms |
Neurological Disease(s) |
References |
|---|---|---|---|---|---|
|
Neuro developmental diseases |
Splicing, Translation |
CPEB4 |
Missplicing of CPEB4 causes reduced inclusion of a neuron-specific microexon, leading to diminished expression of the Cpeb4 transcript that activates translation of mRNAs via polyadenylation under normal conditions |
ASD |
[89] |
|
Splicing Translation, mRNA stability, miRNA biogenesis |
RBFOX1, RBFOX2 (RBM9), RBFOX3 (Neun) |
RBFOX1 binds to the 3′-UTR of its target mRNAs and regulates:
RBFOX2 and RBFOX3 (repression)
Altered splicing of RBFOX family proteins impairs their control of gene expression |
ASD |
||
|
Transport, Translation |
FMRP |
CGG repeat expansion beyond 200 (>200) at the 5′-UTR of FMR1 affects protein expression, resulting in dysregulated spatio-temporal transport/translation of dendritic mRNAs |
FXS |
[54] |
|
|
APA |
NUDT21 |
Elevated amount of NUDT21, a subunit of pre-mRNA cleavage factor Im, due to copy number variation causes abnormal usage of polyadenylation sites, resulting in the generation of an inefficiently translated long isoform of MeCP2 protein. |
Neuropsychia tric disease |
[41] |
|
|
Neuro degenerative diseases |
Splicing |
PRPF8 |
Mutated Huntingtin (HTT) traps PRPF8 (a splicing factor) to cause CREB1 mis-splicing |
HD |
[15] |
|
Translation |
HTT |
Mutant HTT stalls ribosomes |
HD |
[62] |
|
|
Splicing |
MBNL family proteins |
RNA corresponding to expanded microsatellite repeats in DMPK traps MBNL-family proteins, impairing their normal function in splicing |
DM |
[93] |
|
|
Translation |
ATAXIN-2 |
CAG expansion in the reading frame of ATAXIN-2 causes loss of protein function that, under normal conditions, acts as an mRNA translation activator via polyadenylation |
SCA2, ALS |
[65] |
|
|
RAN Translation, Abnormal RNA structure (RNA foci) |
Matrin-3 |
GGGGCC repeat expansion mutation in the C9orf72 gene causes sequestration of Matrin-3 at the RNA foci and RAN translated peptides and loss of function of Matrin-3 |
FTLD, ALS |
[94] |
|
|
mRNA stability, Splicing, Translation |
nELAVL |
nELAVL regulates disease-specific splicing of the pre-mRNAs Picalm and Bin1 by incorporating exons 13 and 6a, respectively. The proteins corresponding to these spliced isoforms have been implicated in trafficking of amyloid precursor protein |
AD |
[95] |
|
|
Transport, Translation, miRNA biogenesis |
TDP-43 |
|
FTLD, ALS |
||
|
Transport, Translation |
FUS |
Normal FUS functions such as axonal transport/translation of mRNAs are adversely impacted in diseased neurons. Under disease conditions, the altered intracellular localization of FUS disrupts its functions as an RBP |
FTLD, ALS |
[99] |
|
|
Splicing, miRNA biogenesis |
hnRNPs, MBNL1 |
mRNA corresponding to shorter CGG repeat expansions (<200) in the 5′UTR of FMR1 sequester many RBPs, e.g., hnRNPs and MBNL1 |
Fragile X-associated tremor/ataxia syndrome (FXTAS) |
[100] |
|
|
APA |
α-synuclein |
Presence of an extended 3′-UTR region in α-synuclein transcript impacts accumulation of α-synuclein protein that is redirected away from synaptic terminals towards mitochondria |
PD |
[45] |
UTR—untranslated region; hnRNPs—heterogenous nuclear ribonuleoproteins.
2.6. Roles for RBPs in RNA Metabolism and Neurological Diseases
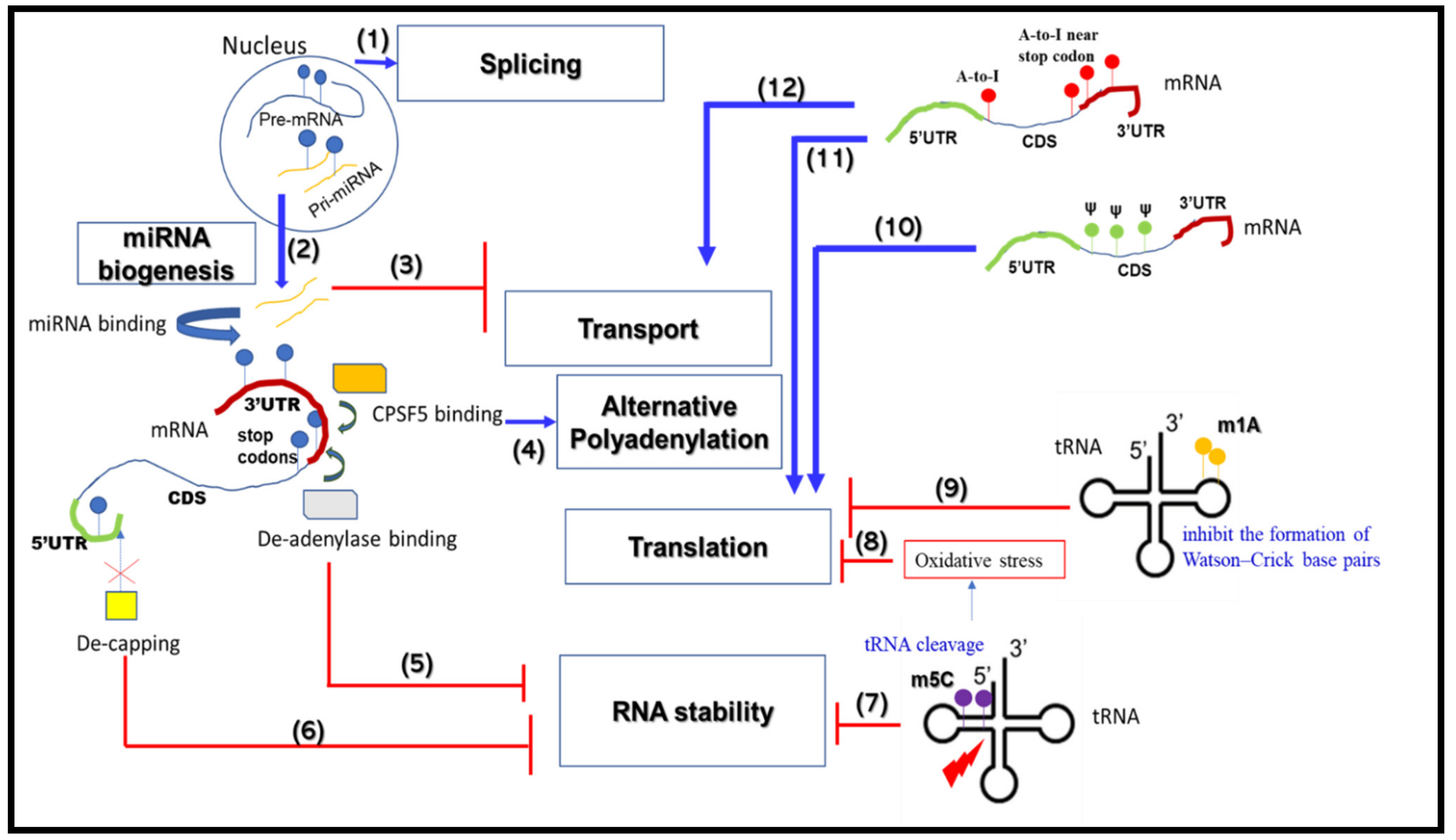
3. RNA Modifications that Change RNA Metabolic Processes
References
- Liu, E.Y.; Cali, C.P.; Lee, E.B. RNA metabolism in neurodegenerative disease. Dis. Model Mech. 2017, 10, 509–518.
- Prashad, S.; Gopal, P.P. RNA-binding proteins in neurological development and disease. RNA Biol. 2021, 18, 972–987.
- Yano, M.; Hayakawa-Yano, Y.; Okano, H. RNA regulation went wrong in neurodevelopmental disorders: The example of Msi/Elavl RNA binding proteins. Int. J. Dev. Neurosci. 2016, 55, 124–130.
- Nussbacher, J.K.; Tabet, R.; Yeo, G.W.; Lagier-Tourenne, C. Disruption of RNA Metabolism in Neurological Diseases and Emerging Therapeutic Interventions. Neuron 2019, 102, 294–320.
- Schieweck, R.; Ninkovic, J.; Kiebler, M.A. RNA-binding proteins balance brain function in health and disease. Physiol. Rev. 2021, 101, 1309–1370.
- Vuong, C.K.; Black, D.L.; Zheng, S. The neurogenetics of alternative splicing. Nat. Rev. Neurosci. 2016, 17, 265–281.
- Chabot, B.; Shkreta, L. Defective control of pre-messenger RNA splicing in human disease. J. Cell Biol. 2016, 212, 13–27.
- Faustino, N.A.; Cooper, T.A. Pre-mRNA splicing and human disease. Genes Dev. 2003, 17, 419–437.
- Mills, J.D.; Janitz, M. Alternative splicing of mRNA in the molecular pathology of neurodegenerative diseases. Neurobiol. Aging 2012, 33, 1012.e11–1012.e24.
- Li, D.; McIntosh, C.S.; Mastaglia, F.L.; Wilton, S.D.; Aung-Htut, M.T. Neurodegenerative diseases: A hotbed for splicing defects and the potential therapies. Transl. Neurodegener. 2021, 10, 16.
- Sanders, S.J.; Schwartz, G.B.; Farh, K.K. Clinical impact of splicing in neurodevelopmental disorders. Genome Med. 2020, 12, 36.
- Thacker, S.; Sefyi, M.; Eng, C. Alternative splicing landscape of the neural transcriptome in a cytoplasmic-predominant Pten expression murine model of autism-like Behavior. Transl. Psychiatry 2020, 10, 380.
- Dick, F.; Nido, G.S.; Alves, G.W.; Tysnes, O.B.; Nilsen, G.H.; Dolle, C.; Tzoulis, C. Differential transcript usage in the Parkinson’s disease brain. PLoS Genet. 2020, 16, e1009182.
- Perrone, B.; La Cognata, V.; Sprovieri, T.; Ungaro, C.; Conforti, F.L.; Ando, S.; Cavallaro, S. Alternative Splicing of ALS Genes: Misregulation and Potential Therapies. Cell Mol. Neurobiol. 2020, 40, 1–14.
- Arnold, E.S.; Ling, S.C.; Huelga, S.C.; Lagier-Tourenne, C.; Polymenidou, M.; Ditsworth, D.; Kordasiewicz, H.B.; McAlonis-Downes, M.; Platoshyn, O.; Parone, P.A.; et al. ALS-linked TDP-43 mutations produce aberrant RNA splicing and adult-onset motor neuron disease without aggregation or loss of nuclear TDP-43. Proc. Natl. Acad. Sci. USA 2013, 110, E736–E745.
- Schilling, J.; Broemer, M.; Atanassov, I.; Duernberger, Y.; Vorberg, I.; Dieterich, C.; Dagane, A.; Dittmar, G.; Wanker, E.; van Roon-Mom, W.; et al. Deregulated Splicing Is a Major Mechanism of RNA-Induced Toxicity in Huntington’s Disease. J. Mol. Biol. 2019, 431, 1869–1877.
- Aikawa, T.; Watanabe, T.; Miyazaki, T.; Mikuni, T.; Wakamori, M.; Sakurai, M.; Aizawa, H.; Ishizu, N.; Watanabe, M.; Kano, M.; et al. Alternative splicing in the C-terminal tail of Cav2.1 is essential for preventing a neurological disease in mice. Hum. Mol. Genet. 2017, 26, 3094–3104.
- Tian, B.; Manley, J.L. Alternative polyadenylation of mRNA precursors. Nat. Rev. Mol. Cell Biol. 2017, 18, 18–30.
- Ren, F.; Zhang, N.; Zhang, L.; Miller, E.; Pu, J.J. Alternative Polyadenylation: A new frontier in post transcriptional regulation. Biomark. Res. 2020, 8, 67.
- Di Giammartino, D.C.; Nishida, K.; Manley, J.L. Mechanisms and consequences of alternative polyadenylation. Mol. Cell 2011, 43, 853–866.
- Tian, B.; Manley, J.L. Alternative cleavage and polyadenylation: The long and short of it. Trends Biochem. Sci. 2013, 38, 312–320.
- Derti, A.; Garrett-Engele, P.; Macisaac, K.D.; Stevens, R.C.; Sriram, S.; Chen, R.; Rohl, C.A.; Johnson, J.M.; Babak, T. A quantitative atlas of polyadenylation in five mammals. Genome Res. 2012, 22, 1173–1183.
- Hoque, M.; Ji, Z.; Zheng, D.; Luo, W.; Li, W.; You, B.; Park, J.Y.; Yehia, G.; Tian, B. Analysis of alternative cleavage and polyadenylation by 3′ region extraction and deep sequencing. Nat. Methods 2013, 10, 133–139.
- Shi, Y. Alternative polyadenylation: New insights from global analyses. RNA 2012, 18, 2105–2117.
- Zhang, H.; Lee, J.Y.; Tian, B. Biased alternative polyadenylation in human tissues. Genome Biol. 2005, 6, R100.
- Liu, D.; Brockman, J.M.; Dass, B.; Hutchins, L.N.; Singh, P.; McCarrey, J.R.; MacDonald, C.C.; Graber, J.H. Systematic variation in mRNA 3′-processing signals during mouse spermatogenesis. Nucleic Acids Res. 2007, 35, 234–246.
- Nam, J.W.; Rissland, O.S.; Koppstein, D.; Abreu-Goodger, C.; Jan, C.H.; Agarwal, V.; Yildirim, M.A.; Rodriguez, A.; Bartel, D.P. Global analyses of the effect of different cellular contexts on microRNA targeting. Mol. Cell 2014, 53, 1031–1043.
- An, J.J.; Gharami, K.; Liao, G.Y.; Woo, N.H.; Lau, A.G.; Vanevski, F.; Torre, E.R.; Jones, K.R.; Feng, Y.; Lu, B.; et al. Distinct role of long 3′ UTR BDNF mRNA in spine morphology and synaptic plasticity in hippocampal neurons. Cell 2008, 134, 175–187.
- Berkovits, B.D.; Mayr, C. Alternative 3′ UTRs act as scaffolds to regulate membrane protein localization. Nature 2015, 522, 363–367.
- Luo, W.; Ji, Z.; Pan, Z.; You, B.; Hoque, M.; Li, W.; Gunderson, S.I.; Tian, B. The conserved intronic cleavage and polyadenylation site of CstF-77 gene imparts control of 3′ end processing activity through feedback autoregulation and by U1 snRNP. PLoS Genet. 2013, 9, e1003613.
- Braz, S.O.; Cruz, A.; Lobo, A.; Bravo, J.; Moreira-Ribeiro, J.; Pereira-Castro, I.; Freitas, J.; Relvas, J.B.; Summavielle, T.; Moreira, A. Expression of Rac1 alternative 3′ UTRs is a cell specific mechanism with a function in dendrite outgrowth in cortical neurons. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 685–694.
- Jereb, S.; Hwang, H.W.; Van Otterloo, E.; Govek, E.E.; Fak, J.J.; Yuan, Y.; Hatten, M.E.; Darnell, R.B. Differential 3′ Processing of Specific Transcripts Expands Regulatory and Protein Diversity Across Neuronal Cell Types. Elife 2018, 7, e34042.
- Yang, Y.; Paul, A.; Bach, T.N.; Huang, Z.J.; Zhang, M.Q. Single-cell alternative polyadenylation analysis delineates GABAergic neuron types. BMC Biol. 2021, 19, 144.
- Grassi, E.; Santoro, R.; Umbach, A.; Grosso, A.; Oliviero, S.; Neri, F.; Conti, L.; Ala, U.; Provero, P.; DiCunto, F.; et al. Choice of Alternative Polyadenylation Sites, Mediated by the RNA-Binding Protein Elavl3, Plays a Role in Differentiation of Inhibitory Neuronal Progenitors. Front. Cell Neurosci. 2018, 12, 518.
- Coutinho, A.M.; Oliveira, G.; Katz, C.; Feng, J.; Yan, J.; Yang, C.; Marques, C.; Ataide, A.; Miguel, T.S.; Borges, L.; et al. MECP2 coding sequence and 3′UTR variation in 172 unrelated autistic patients. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2007, 144, 475–483.
- Hoffbuhr, K.; Devaney, J.M.; LaFleur, B.; Sirianni, N.; Scacheri, C.; Giron, J.; Schuette, J.; Innis, J.; Marino, M.; Philippart, M.; et al. MeCP2 mutations in children with and without the phenotype of Rett syndrome. Neurology 2001, 56, 1486–1495.
- Newnham, C.M.; Hall-Pogar, T.; Liang, S.; Wu, J.; Tian, B.; Hu, J.; Lutz, C.S. Alternative polyadenylation of MeCP2: Influence of cis-acting elements and trans-acting factors. RNA Biol. 2010, 7, 361–372.
- Shibayama, A.; Cook, E.H., Jr.; Feng, J.; Glanzmann, C.; Yan, J.; Craddock, N.; Jones, I.R.; Goldman, D.; Heston, L.L.; Sommer, S.S. MECP2 structural and 3′-UTR variants in schizophrenia, autism and other psychiatric diseases: A possible association with autism. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2004, 128, 50–53.
- Tassone, F.; De Rubeis, S.; Carosi, C.; La Fata, G.; Serpa, G.; Raske, C.; Willemsen, R.; Hagerman, P.J.; Bagni, C. Differential usage of transcriptional start sites and polyadenylation sites in FMR1 premutation alleles. Nucleic Acids Res. 2011, 39, 6172–6185.
- Szkop, K.J.; Cooke, P.I.C.; Humphries, J.A.; Kalna, V.; Moss, D.S.; Schuster, E.F.; Nobeli, I. Dysregulation of Alternative Poly-adenylation as a Potential Player in Autism Spectrum Disorder. Front. Mol. Neurosci. 2017, 10, 279.
- Gennarino, V.A.; Alcott, C.E.; Chen, C.A.; Chaudhury, A.; Gillentine, M.A.; Rosenfeld, J.A.; Parikh, S.; Wheless, J.W.; Roeder, E.R.; Horovitz, D.D.; et al. NUDT21-spanning CNVs lead to neuropsychiatric disease and altered MeCP2 abundance via alternative polyadenylation. Elife 2015, 4, e10782.
- Habib, R.; Noureen, N.; Nadeem, N. Decoding Common Features of Neurodegenerative Disorders: From Differentially Expressed Genes to Pathways. Curr. Genomics 2018, 19, 300–312.
- Noori, A.; Mezlini, A.M.; Hyman, B.T.; Serrano-Pozo, A.; Das, S. Systematic review and meta-analysis of human transcriptomics reveals neuroinflammation, deficient energy metabolism, and proteostasis failure across neurodegeneration. Neurobiol. Dis. 2021, 149, 105225.
- Patel, R.; Brophy, C.; Hickling, M.; Neve, J.; Furger, A. Alternative cleavage and polyadenylation of genes associated with protein turnover and mitochondrial function are deregulated in Parkinson’s, Alzheimer’s and ALS disease. BMC Med. Genomics 2019, 12, 60.
- Rhinn, H.; Qiang, L.; Yamashita, T.; Rhee, D.; Zolin, A.; Vanti, W.; Abeliovich, A. Alternative alpha-synuclein transcript usage as a convergent mechanism in Parkinson’s disease pathology. Nat. Commun. 2012, 3, 1084.
- Jenal, M.; Elkon, R.; Loayza-Puch, F.; van Haaften, G.; Kuhn, U.; Menzies, F.M.; Oude Vrielink, J.A.; Bos, A.J.; Drost, J.; Rooijers, K.; et al. The poly(A)-binding protein nuclear 1 suppresses alternative cleavage and polyadenylation sites. Cell 2012, 149, 538–553.
- Akbalik, G.; Schuman, E.M. Molecular biology. mRNA, live and unmasked. Science 2014, 343, 375–376.
- Nagano, S.; Jinno, J.; Abdelhamid, R.F.; Jin, Y.; Shibata, M.; Watanabe, S.; Hirokawa, S.; Nishizawa, M.; Sakimura, K.; Onodera, O.; et al. TDP-43 transports ribosomal protein mRNA to regulate axonal local translation in neuronal axons. Acta Neuropathol. 2020, 140, 695–713.
- Shigeoka, T.; Koppers, M.; Wong, H.H.; Lin, J.Q.; Cagnetta, R.; Dwivedy, A.; de Freitas Nascimento, J.; van Tartwijk, F.W.; Strohl, F.; Cioni, J.M.; et al. On-Site Ribosome Remodeling by Locally Synthesized Ribosomal Proteins in Axons. Cell Rep. 2019, 29, 3605–3619.e10.
- Richter, J.D.; Bassell, G.J.; Klann, E. Dysregulation and restoration of translational homeostasis in fragile X syndrome. Nat. Rev. Neurosci. 2015, 16, 595–605.
- Bagni, C.; Zukin, R.S. A Synaptic Perspective of Fragile X Syndrome and Autism Spectrum Disorders. Neuron 2019, 101, 1070–1088.
- Chu, J.F.; Majumder, P.; Chatterjee, B.; Huang, S.L.; Shen, C.J. TDP-43 Regulates Coupled Dendritic mRNA Transport-Translation Processes in Co-operation with FMRP and Staufen1. Cell Rep. 2019, 29, 3118–3133.e6.
- Majumder, P.; Chu, J.F.; Chatterjee, B.; Swamy, K.B.; Shen, C.J. Co-regulation of mRNA translation by TDP-43 and Fragile X Syndrome protein FMRP. Acta Neuropathol. 2016, 132, 721–738.
- Wang, E.T.; Taliaferro, J.M.; Lee, J.A.; Sudhakaran, I.P.; Rossoll, W.; Gross, C.; Moss, K.R.; Bassell, G.J. Dysregulation of mRNA Localization and Translation in Genetic Disease. J. Neurosci. 2016, 36, 11418–11426.
- Shukla, S.; Parker, R. Hypo- and Hyper-Assembly Diseases of RNA-Protein Complexes. Trends Mol. Med. 2016, 22, 615–628.
- Buchan, J.R.; Parker, R. Eukaryotic stress granules: The ins and outs of translation. Mol. Cell 2009, 36, 932–941.
- Wang, P.Y.; Lin, Y.M.; Wang, L.H.; Kuo, T.Y.; Cheng, S.J.; Wang, G.S. Reduced cytoplasmic MBNL1 is an early event in a brain-specific mouse model of myotonic dystrophy. Hum. Mol. Genet. 2017, 26, 2247–2257.
- Goodman, L.D.; Bonini, N.M. Repeat-associated non-AUG (RAN) translation mechanisms are running into focus for GGGGCC-repeat associated ALS/FTD. Prog. Neurobiol. 2019, 183, 101697.
- Ishiguro, T.; Nagai, Y.; Ishikawa, K. Insight Into Spinocerebellar Ataxia Type 31 (SCA31) From Drosophila Model. Front. Neurosci. 2021, 15, 472.
- Banez-Coronel, M.; Ayhan, F.; Tarabochia, A.D.; Zu, T.; Perez, B.A.; Tusi, S.K.; Pletnikova, O.; Borchelt, D.R.; Ross, C.A.; Margolis, R.L.; et al. RAN Translation in Huntington Disease. Neuron 2015, 88, 667–677.
- Zu, T.; Gibbens, B.; Doty, N.S.; Gomes-Pereira, M.; Huguet, A.; Stone, M.D.; Margolis, J.; Peterson, M.; Markowski, T.W.; Ingram, M.A.; et al. Non-ATG-initiated translation directed by microsatellite expansions. Proc. Natl. Acad. Sci. USA 2011, 108, 260–265.
- Eshraghi, M.; Karunadharma, P.P.; Blin, J.; Shahani, N.; Ricci, E.P.; Michel, A.; Urban, N.T.; Galli, N.; Sharma, M.; Ramirez-Jarquin, U.N.; et al. Mutant Huntingtin stalls ribosomes and represses protein synthesis in a cellular model of Huntington disease. Nat. Commun. 2021, 12, 1461.
- Akiyama, T.; Suzuki, N.; Ishikawa, M.; Fujimori, K.; Sone, T.; Kawada, J.; Funayama, R.; Fujishima, F.; Mitsuzawa, S.; Ikeda, K.; et al. Aberrant axon branching via Fos-B dysregulation in FUS-ALS motor neurons. EBioMedicine 2019, 45, 362–378.
- Imperatore, J.A.; McAninch, D.S.; Valdez-Sinon, A.N.; Bassell, G.J.; Mihailescu, M.R. FUS Recognizes G Quadruplex Structures Within Neuronal mRNAs. Front. Mol. Biosci. 2020, 7, 6.
- Inagaki, H.; Hosoda, N.; Tsuiji, H.; Hoshino, S.I. Direct evidence that Ataxin-2 is a translational activator mediating cytoplasmic polyadenylation. J. Biol. Chem. 2020, 295, 15810–15825.
- Montalbano, M.; McAllen, S.; Cascio, F.L.; Sengupta, U.; Garcia, S.; Bhatt, N.; Ellsworth, A.; Heidelman, E.A.; Johnson, O.D.; Doskocil, S.; et al. TDP-43 and Tau Oligomers in Alzheimer’s Disease, Amyotrophic Lateral Sclerosis, and Frontotemporal Dementia. Neurobiol. Dis. 2020, 146, 105130.
- Houseley, J.; Tollervey, D. The many pathways of RNA degradation. Cell 2009, 136, 763–776.
- Chen, C.Y.; Ezzeddine, N.; Shyu, A.B. Messenger RNA half-life measurements in mammalian cells. Methods Enzymol. 2008, 448, 335–357.
- Burow, D.A.; Umeh-Garcia, M.C.; True, M.B.; Bakhaj, C.D.; Ardell, D.H.; Cleary, M.D. Dynamic regulation of mRNA decay during neural development. Neural Dev. 2015, 10, 11.
- Porter, R.S.; Jaamour, F.; Iwase, S. Neuron-specific alternative splicing of transcriptional machineries: Implications for neurodevelopmental disorders. Mol. Cell Neurosci. 2018, 87, 35–45.
- Lee, S.; Wei, L.; Zhang, B.; Goering, R.; Majumdar, S.; Wen, J.; Taliaferro, J.M.; Lai, E.C. ELAV/Hu RNA binding proteins determine multiple programs of neural alternative splicing. PLoS Genet. 2021, 17, e1009439.
- DeBoer, E.M.; Azevedo, R.; Vega, T.A.; Brodkin, J.; Akamatsu, W.; Okano, H.; Wagner, G.C.; Rasin, M.R. Prenatal deletion of the RNA-binding protein HuD disrupts postnatal cortical circuit maturation and behavior. J. Neurosci. 2014, 34, 3674–3686.
- Berto, S.; Usui, N.; Konopka, G.; Fogel, B.L. ELAVL2-regulated transcriptional and splicing networks in human neurons link neurodevelopment and autism. Hum. Mol. Genet. 2016, 25, 2451–2464.
- Cha, I.J.; Lee, D.; Park, S.S.; Chung, C.G.; Kim, S.Y.; Jo, M.G.; Kim, S.Y.; Lee, B.H.; Lee, Y.S.; Lee, S.B. Ataxin-2 Dysregulation Triggers a Compensatory Fragile X Mental Retardation Protein Decrease in Drosophila C4da Neurons. Mol. Cells 2020, 43, 870–879.
- Scheckel, C.; Drapeau, E.; Frias, M.A.; Park, C.Y.; Fak, J.; Zucker-Scharff, I.; Kou, Y.; Haroutunian, V.; Ma’ayan, A.; Buxbaum, J.D.; et al. Regulatory consequences of neuronal ELAV-like protein binding to coding and non-coding RNAs in human brain. Elife 2016, 5, e10421.
- Ostrowski, L.A.; Hall, A.C.; Mekhail, K. Ataxin-2: From RNA Control to Human Health and Disease. Genes 2017, 8, 157.
- Alkallas, R.; Fish, L.; Goodarzi, H.; Najafabadi, H.S. Inference of RNA decay rate from transcriptional profiling highlights the regulatory programs of Alzheimer’s disease. Nat. Commun. 2017, 8, 909.
- Costessi, L.; Porro, F.; Iaconcig, A.; Muro, A.F. TDP-43 regulates beta-adducin (Add2) transcript stability. RNA Biol. 2014, 11, 1280–1290.
- Wang, H.; Taguchi, Y.H.; Liu, X. Editorial: miRNAs and Neurological Diseases. Front. Neurol. 2021, 12, 662373.
- Guo, L.; Yin, M.; Wang, Y. CREB1, a direct target of miR-122, promotes cell proliferation and invasion in bladder cancer. Oncol. Lett. 2018, 16, 3842–3848.
- Jasinska, M.; Milek, J.; Cymerman, I.A.; Leski, S.; Kaczmarek, L.; Dziembowska, M. miR-132 Regulates Dendritic Spine Structure by Direct Targeting of Matrix Metalloproteinase 9 mRNA. Mol. Neurobiol. 2016, 53, 4701–4712.
- Rey, R.; Suaud-Chagny, M.F.; Dorey, J.M.; Teyssier, J.R.; d’Amato, T. Widespread transcriptional disruption of the microRNA biogenesis machinery in brain and peripheral tissues of individuals with schizophrenia. Transl. Psychiatry 2020, 10, 376.
- McKeever, P.M.; Schneider, R.; Taghdiri, F.; Weichert, A.; Multani, N.; Brown, R.A.; Boxer, A.L.; Karydas, A.; Miller, B.; Robertson, J.; et al. MicroRNA Expression Levels Are Altered in the Cerebrospinal Fluid of Patients with Young-Onset Alzheimer’s Disease. Mol. Neurobiol. 2018, 55, 8826–8841.
- Riancho, J.; Vazquez-Higuera, J.L.; Pozueta, A.; Lage, C.; Kazimierczak, M.; Bravo, M.; Calero, M.; Gonalezalez, A.; Rodriguez, E.; Lleo, A.; et al. MicroRNA Profile in Patients with Alzheimer’s Disease: Analysis of miR-9-5p and miR-598 in Raw and Exosome Enriched Cerebrospinal Fluid Samples. J. Alzheimers Dis. 2017, 57, 483–491.
- Rajgor, D. Macro roles for microRNAs in neurodegenerative diseases. Noncoding RNA Res. 2018, 3, 154–159.
- Rinchetti, P.; Rizzuti, M.; Faravelli, I.; Corti, S. MicroRNA Metabolism and Dysregulation in Amyotrophic Lateral Sclerosis. Mol. Neurobiol. 2018, 55, 2617–2630.
- Dong, X.; Cong, S. The Emerging Role of microRNAs in Polyglutamine Diseases. Front. Mol. Neurosci. 2019, 12, 156.
- Lopez Castel, A.; Overby, S.J.; Artero, R. MicroRNA-Based Therapeutic Perspectives in Myotonic Dystrophy. Int. J. Mol. Sci. 2019, 20, 5600.
- Parras, A.; Anta, H.; Santos-Galindo, M.; Swarup, V.; Elorza, A.; Nieto-Gonzalez, J.L.; Pico, S.; Hernandez, I.H.; Diaz-Hernandez, J.I.; Belloc, E.; et al. Autism-like phenotype and risk gene mRNA deadenylation by CPEB4 mis-splicing. Nature 2018, 560, 441–446.
- Carreira-Rosario, A.; Bhargava, V.; Hillebrand, J.; Kollipara, R.K.; Ramaswami, M.; Buszczak, M. Repression of Pumilio Protein Expression by Rbfox1 Promotes Germ Cell Differentiation. Dev. Cell 2016, 36, 562–571.
- Lee, J.A.; Damianov, A.; Lin, C.H.; Fontes, M.; Parikshak, N.N.; Anderson, E.S.; Geschwind, D.H.; Black, D.L.; Martin, K.C. Cytoplasmic Rbfox1 Regulates the Expression of Synaptic and Autism-Related Genes. Neuron 2016, 89, 113–128.
- Wei, C.; Xiao, R.; Chen, L.; Cui, H.; Zhou, Y.; Xue, Y.; Hu, J.; Zhou, B.; Tsutsui, T.; Qiu, J.; et al. RBFox2 Binds Nascent RNA to Globally Regulate Polycomb Complex 2 Targeting in Mammalian Genomes. Mol. Cell 2016, 62, 982.
- Lee, J.E.; Cooper, T.A. Pathogenic mechanisms of myotonic dystrophy. Biochem. Soc. Trans. 2009, 37 Pt 6, 1281–1286.
- Ramesh, N.; Daley, E.L.; Gleixner, A.M.; Mann, J.R.; Kour, S.; Mawrie, D.; Anderson, E.N.; Kofler, J.; Donnelly, C.J.; Kiskinis, E.; et al. RNA dependent suppression of C9orf72 ALS/FTD associated neurodegeneration by Matrin-3. Acta Neuropathol. Commun. 2020, 8, 177.
- Tan, M.S.; Yu, J.T.; Tan, L. Bridging integrator 1 (BIN1): Form, function, and Alzheimer’s disease. Trends Mol. Med. 2013, 19, 594–603.
- Alami, N.H.; Smith, R.B.; Carrasco, M.A.; Williams, L.A.; Winborn, C.S.; Han, S.S.W.; Kiskinis, E.; Winborn, B.; Freibaum, B.D.; Kanagaraj, A.; et al. Axonal transport of TDP-43 mRNA granules is impaired by ALS-causing mutations. Neuron 2014, 81, 536–543.
- Coyne, A.N.; Yamada, S.B.; Siddegowda, B.B.; Estes, P.S.; Zaepfel, B.L.; Johannesmeyer, J.S.; Lockwood, D.B.; Pham, L.T.; Hart, M.P.; Cassel, J.A.; et al. Fragile X protein mitigates TDP-43 toxicity by remodeling RNA granules and restoring translation. Hum. Mol. Genet. 2015, 24, 6886–6898.
- Ling, S.C.; Albuquerque, C.P.; Han, J.S.; Lagier-Tourenne, C.; Tokunaga, S.; Zhou, H.; Cleveland, D.W. ALS-associated mutations in TDP-43 increase its stability and promote TDP-43 complexes with FUS/TLS. Proc. Natl. Acad. Sci. USA 2010, 107, 13318–13323.
- Lopez-Erauskin, J.; Tadokoro, T.; Baughn, M.W.; Myers, B.; McAlonis-Downes, M.; Chillon-Marinas, C.; Asiaban, J.N.; Artates, J.; Bui, A.T.; Vetto, A.P.; et al. ALS/FTD-Linked Mutation in FUS Suppresses Intra-axonal Protein Synthesis and Drives Disease Without Nuclear Loss-of-Function of FUS. Neuron 2018, 100, 816–830.e7.
- Sellier, C.; Rau, F.; Liu, Y.; Tassone, F.; Hukema, R.K.; Gattoni, R.; Schneider, A.; Richard, S.; Willemsen, R.; Elliott, D.J.; et al. Sam68 sequestration and partial loss of function are associated with splicing alterations in FXTAS patients. EMBO J. 2010, 29, 1248–1261.
- Conlon, E.G.; Manley, J.L. RNA-binding proteins in neurodegeneration: Mechanisms in aggregate. Genes Dev. 2017, 31, 1509–1528.
- Kelaini, S.; Chan, C.; Cornelius, V.A.; Margariti, A. RNA-Binding Proteins Hold Key Roles in Function, Dysfunction, and Disease. Biology 2021, 10, 366.
- Jung, Y.; Goldman, D. Role of RNA modifications in brain and behavior. Genes Brain Behav. 2018, 17, e12444.
- Sanchez-Vasquez, E.; Alata Jimenez, N.; Vazquez, N.A.; Strobl-Mazzulla, P.H. Emerging role of dynamic RNA modifications during animal development. Mech. Dev. 2018, 154, 24–32.





