
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Li Zhang | + 3564 word(s) | 3564 | 2021-11-08 02:57:21 | | | |
| 2 | Conner Chen | Meta information modification | 3564 | 2021-11-22 02:02:20 | | |
Video Upload Options
According to the latest Global Cancer Statistics, there are about 19.3 million new cancer cases worldwide and nearly 10 million people died of cancer in 2020. As the largest threat to human life, the early detection of cancer is an effective way to reduce its mortality. In addition, heavy metal poisoning and biological toxins also seriously endanger human health, and their detection methods still have some shortcomings. Against this backdrop, biosensors have been developed by integrating modern biotechnology and advanced physical technology. Biosensors are devices that are used for the rapid and sensitive detection of substances at the molecular level. The basic unit of the biosensor includes the identification element, transducer and detector, etc. The components of organisms with molecular recognition capabilities or the organism itself can be used as recognition elements.
1. Working Principle for the Detection of Functional Nucleic-Acid Biosensors
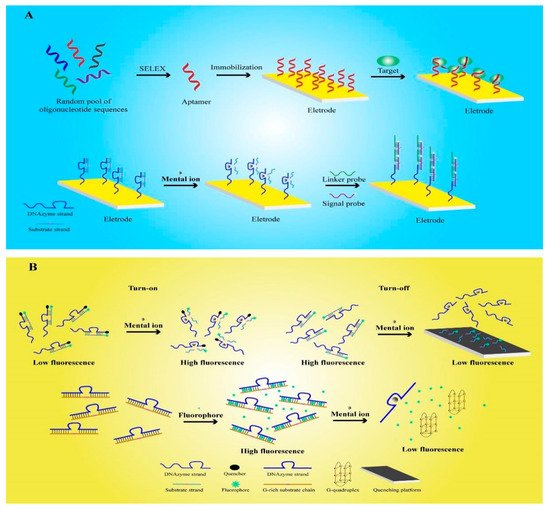
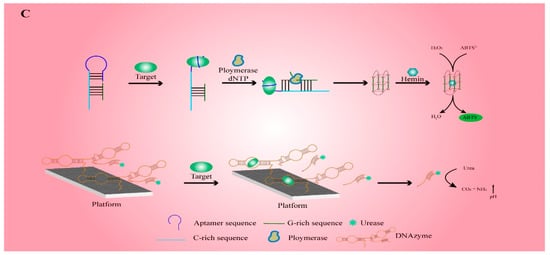
2. Electrochemical Functional Nucleic-Acid Biosensors
3. Fluorescent Functional Nucleic-Acid Biosensors
4. Colorimetric Functional Nucleic-Acid Biosensors
5. Nanotechnology in Functional Nucleic-Acid Biosensors
5.1. DNA Hydrogel
5.2. Metal Nanomaterials
References
- Yin, X.-B. Functional nucleic acids for electrochemical and electrochemiluminescent sensing applications. TrAC Trends Anal. Chem. 2012, 33, 81–94.
- Li, X.-M.; Ju, H.-Q.; Ding, C.-F.; Zhang, S.-S. Nucleic acid biosensor for detection of hepatitis B virus using 2,9-dimethyl-1,10-phenanthroline copper complex as electrochemical indicator. Anal. Chim. Acta 2007, 582, 158–163.
- Wang, C.-F.; Sun, X.-Y.; Su, M.; Wang, Y.-P.; Lv, Y.-K. Electrochemical biosensors based on antibody, nucleic acid and enzyme functionalized graphene for the detection of disease-related biomolecules. Analyst 2020, 145, 1550–1562.
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510.
- Zhang, X.; Feng, Y.; Duan, S.; Su, L.; Zhang, J.; He, F. Mycobacterium tuberculosis strain H37Rv electrochemical sensor mediated by aptamer and AuNPs–DNA. ACS Sens. 2019, 4, 849–855.
- Chung, S.; Moon, J.-M.; Choi, J.; Hwang, H.; Shim, Y.-B. Magnetic force assisted electrochemical sensor for the detection of thrombin with aptamer-antibody sandwich formation. Biosens. Bioelectron. 2018, 117, 480–486.
- Beiranvand, S.; Azadbakht, A. Electrochemical switching with a DNA aptamer-based electrochemical sensor. Mater. Sci. Eng. C 2017, 76, 925–933.
- Teengam, P.; Siangproh, W.; Tuantranont, A.; Henry, C.S.; Vilaivan, T.; Chailapakul, O. Electrochemical paper-based peptide nucleic acid biosensor for detecting human papillomavirus. Anal. Chim. Acta 2017, 952, 32–40.
- Raina, D.; Kosugi, M.; Ahmad, R.; Panchamoorthy, G.; Rajabi, H.; Alam, M.; Shimamura, T.; Shapiro, G.I.; Supko, J.; Kharbanda, S. Dependence on the MUC1-C oncoprotein in non–small cell lung cancer cells. Mol. Cancer Ther. 2011, 10, 806–816.
- Besmer, D.M.; Curry, J.M.; Roy, L.D.; Tinder, T.L.; Sahraei, M.; Schettini, J.; Hwang, S.-I.; Lee, Y.Y.; Gendler, S.J.; Mukherjee, P. Pancreatic ductal adenocarcinoma mice lacking mucin 1 have a profound defect in tumor growth and metastasis. Cancer Res. 2011, 71, 4432–4442.
- Wu, P.; Gao, Y.; Zhang, H.; Cai, C. Aptamer-guided silver–gold bimetallic nanostructures with highly active surface-enhanced raman scattering for specific detection and near-infrared photothermal therapy of human breast cancer cells. Anal. Chem. 2012, 84, 7692–7699.
- Song, J.; Zhou, Y.; Chen, B.; Lou, W.; Gu, J. Development of electrochemical aptamer biosensor for tumor marker MUC1 determination. Int. J. Electrochem. Sci. 2017, 12, 5618–5627.
- Cao, H.; Ye, D.; Zhao, Q.; Luo, J.; Zhang, S.; Kong, J. A novel aptasensor based on MUC-1 conjugated CNSs for ultrasensitive detection of tumor cells. Analyst 2014, 139, 4917–4923.
- Sun, X.; Li, Y. Ga2O3 and GaN semiconductor hollow spheres. Angew. Chem. Int. Ed. 2004, 43, 3827–3831.
- Karpik, A.E.; Crulhas, B.P.; Rodrigues, C.B.; Castro, G.R.; Pedrosa, V.A. Aptamer-based biosensor developed to monitor MUC1 released by prostate cancer cells. Electroanalysis 2017, 29, 2246–2253.
- Day, E.S.; Riley, R.S.; Billingsley, M.M. Antibody-Nanoparticle Conjugates to Enhance the Sensitivity of ELISA-Based Detection Methods; Public Library of Science: Newark, NJ, USA, 2017.
- Jacobs, M.V.; Snijders, P.; Van Den Brule, A.; Helmerhorst, T.; Meijer, C.; Walboomers, J. A general primer GP5+/GP6 (+)-mediated PCR-enzyme immunoassay method for rapid detection of 14 high-risk and 6 low-risk human papillomavirus genotypes in cervical scrapings. J. Clin. Microbiol. 1997, 35, 791–795.
- Li, X.; Yang, J.; Xie, J.; Jiang, B.; Yuan, R.; Xiang, Y. Cascaded signal amplification via target-triggered formation of aptazyme for sensitive electrochemical detection of ATP. Biosens. Bioelectron. 2018, 102, 296–300.
- Ji, R.; Niu, W.; Chen, S.; Xu, W.; Ji, X.; Yuan, L.; Zhao, H.; Geng, M.; Qiu, J.; Li, C. Target-inspired Pb2+-dependent DNAzyme for ultrasensitive electrochemical sensor based on MoS2-AuPt nanocomposites and hemin/G-quadruplex DNAzyme as signal amplifier. Biosens. Bioelectron. 2019, 144, 111560.
- Liao, X.; Luo, J.; Wu, J.; Fan, T.; Yao, Y.; Gao, F.; Qian, Y. A sensitive DNAzyme-based electrochemical sensor for Pb2+ detection with platinum nanoparticles decorated TiO2/α-Fe2O3 nanocomposite as signal labels. J. Electroanal. Chem. 2018, 829, 129–137.
- Xiao, Y.; Rowe, A.A.; Plaxco, K.W. Electrochemical detection of parts-per-billion lead via an electrode-bound DNAzyme assembly. J. Am. Chem. Soc. 2007, 129, 262–263.
- Tang, S.; Lu, W.; Gu, F.; Tong, P.; Yan, Z.; Zhang, L. A novel electrochemical sensor for lead ion based on cascade DNA and quantum dots amplification. Electrochim. Acta 2014, 134, 1–7.
- Tagar, Z.A.; Memon, N.; Agheem, M.H.; Junejo, Y.; Hassan, S.S.; Kalwar, N.H.; Khattak, M.I. Selective, simple and economical lead sensor based on ibuprofen derived silver nanoparticles. Sens. Actuators B Chem. 2011, 157, 430–437.
- Li, F.; Yang, L.; Chen, M.; Li, P.; Tang, B. A selective amperometric sensing platform for lead based on target-induced strand release. Analyst 2013, 138, 461–466.
- Liu, J.; Lu, Y. Adenosine-dependent assembly of aptazyme-functionalized gold nanoparticles and its application as a colorimetric biosensor. Anal. Chem. 2004, 76, 1627–1632.
- Wang, X.X.; Zhu, L.J.; Li, S.T.; Zhang, Y.Z.; Liu, S.Y.; Huang, K.L.; Xu, W.T. Fluorescent Functional Nucleic Acid: Principles, Properties and Applications in Bioanalyzing. TrAC Trends Anal. Chem. 2021, 141, 116292.
- Lee, J.; Lin, L.; Li, Y. Functional nucleic acids for fluorescence-based biosensing applications. In Advanced Fluorescence Reporters in Chemistry and Biology III; Springer: Cham, Switzerland, 2011; pp. 201–221.
- Li, J.; Lu, Y. A highly sensitive and selective catalytic DNA biosensor for lead ions. J. Am. Chem. Soc. 2000, 122, 10466–10467.
- Guo, Y.; Li, J.; Zhang, X.; Tang, Y. A sensitive biosensor with a DNAzyme for lead (II) detection based on fluorescence turn-on. Analyst 2015, 140, 4642–4647.
- Zhang, H.; Ruan, Y.; Lin, L.; Lin, M.; Zeng, X.; Xi, Z.; Fu, F. A turn-off fluorescent biosensor for the rapid and sensitive detection of uranyl ion based on molybdenum disulfide nanosheets and specific DNAzyme. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2015, 146, 1–6.
- Yue, G.; Huang, D.; Luo, F.; Guo, L.; Qiu, B.; Lin, Z.; Chen, G. Highly selective fluorescence sensor for hydrogen sulfide based on the Cu (II)-dependent DNAzyme. J. Lumin. 2019, 207, 369–373.
- Ji, X.; Wang, Z.; Niu, S.; Ding, C. DNAzyme-functionalized porous carbon nanospheres serve as a fluorescent nanoprobe for imaging detection of microRNA-21 and zinc ion in living cells. Microchim. Acta 2020, 187, 1–9.
- Torabi, S.-F.; Wu, P.; McGhee, C.E.; Chen, L.; Hwang, K.; Zheng, N.; Cheng, J.; Lu, Y. In vitro selection of a sodium-specific DNAzyme and its application in intracellular sensing. Proc. Natl. Acad. Sci. USA 2015, 112, 5903–5908.
- Saran, R.; Liu, J. A silver DNAzyme. Anal. Chem. 2016, 88, 4014–4020.
- Zhou, W.; Vazin, M.; Yu, T.; Ding, J.; Liu, J. In Vitro Selection of Chromium-Dependent DNAzymes for Sensing Chromium (III) and Chromium (VI); UWSpace: Waterloo, ON, Canada, 2016.
- Huang, P.-J.J.; Liu, J. Rational evolution of Cd2+-specific DNAzymes with phosphorothioate modified cleavage junction and Cd2+ sensing. Nucleic Acids Res. 2015, 43, 6125–6133.
- Huang, P.-J.J.; Lin, J.; Cao, J.; Vazin, M.; Liu, J. Ultrasensitive DNAzyme beacon for lanthanides and metal speciation. Anal. Chem. 2014, 86, 1816–1821.
- Huang, P.-J.J.; Vazin, M.; Liu, J. In vitro selection of a new lanthanide-dependent DNAzyme for ratiometric sensing lanthanides. Anal. Chem. 2014, 86, 9993–9999.
- Chen, Q.; Guo, Q.; Chen, Y.; Pang, J.; Fu, F.; Guo, L. An enzyme-free and label-free fluorescent biosensor for small molecules by G-quadruplex based hybridization chain reaction. Talanta 2015, 138, 15–19.
- Lipps, H.J.; Rhodes, D. G-quadruplex structures: In vivo evidence and function. Trends Cell Biol. 2009, 19, 414–422.
- Zhua, P.; Zhanga, Y.; Xub, S.; Zhanga, X. G-quadruplex-assisted enzyme strand recycling for amplified label-free fluorescent detection of UO2 2. Chin. Chem. Lett. 2019, 30, 58–62.
- Zhan, S.; Wu, Y.; Luo, Y.; Liu, L.; He, L.; Xing, H.; Zhou, P. Label-free fluorescent sensor for lead ion detection based on lead (II)-stabilized G-quadruplex formation. Anal. Biochem. 2014, 462, 19–25.
- Sun, X.; Li, Q.; Xiang, J.; Wang, L.; Zhang, X.; Lan, L.; Xu, S.; Yang, F.; Tang, Y. Novel fluorescent cationic benzothiazole dye that responds to G-quadruplex aptamer as a novel K+ sensor. Analyst 2017, 142, 3352–3355.
- Ma, G.; Yu, Z.; Zhou, W.; Li, Y.; Fan, L.; Li, X. Investigation of Na+ and K+ Competitively Binding with a G-Quadruplex and Discovery of a Stable K+–Na+-Quadruplex. J. Phys. Chem. B 2019, 123, 5405–5411.
- Xu, L.; Chen, Y.; Zhang, R.; Gao, T.; Zhang, Y.; Shen, X.; Pei, R. A highly Sensitive Turn-on Fluorescent Sensor for Ba2+ Based on G-Quadruplexes. J. Fluoresc. 2017, 27, 569–574.
- Wang, M.; Wang, W.; Kang, T.-S.; Leung, C.-H.; Ma, D.-L. Development of an Iridium (III) complex as a G-quadruplex probe and its application for the G-quadruplex-based luminescent detection of picomolar insulin. Anal. Chem. 2016, 88, 981–987.
- Zhu, Q.; Liu, L.; Xing, Y.; Zhou, X. Duplex functional G-quadruplex/NMM fluorescent probe for label-free detection of lead (II) and mercury (II) ions. J. Hazard. Mater. 2018, 355, 50–55.
- Hoang, M.; Huang, P.-J.J.; Liu, J. G-quadruplex DNA for fluorescent and colorimetric detection of thallium (I). Acs Sens. 2016, 1, 137–143.
- Chen, Q.; Zuo, J.; Chen, J.; Tong, P.; Mo, X.; Zhang, L.; Li, J. A label-free fluorescent biosensor for ultratrace detection of terbium (III) based on structural conversion of G-quadruplex DNA mediated by ThT and terbium (III). Biosens. Bioelectron. 2015, 72, 326–331.
- Taghdisi, S.M.; Danesh, N.M.; Ramezani, M.; Abnous, K. A new amplified fluorescent aptasensor based on hairpin structure of G-quadruplex oligonucleotide-Aptamer chimera and silica nanoparticles for sensitive detection of aflatoxin B1 in the grape juice. Food Chem. 2018, 268, 342–346.
- Michalet, X.; Pinaud, F.; Bentolila, L.; Tsay, J.; Doose, S.; Li, J.; Sundaresan, G.; Wu, A.; Gambhir, S.; Weiss, S. Quantum dots for live cells, in vivo imaging, and diagnostics. Science 2005, 307, 538–544.
- Chu, T.C.; Shieh, F.; Lavery, L.A.; Levy, M.; Richards-Kortum, R.; Korgel, B.A.; Ellington, A.D. Labeling tumor cells with fluorescent nanocrystal–aptamer bioconjugates. Biosens. Bioelectron. 2006, 21, 1859–1866.
- Lim, S.Y.; Shen, W.; Gao, Z. Carbon quantum dots and their applications. Chem. Soc. Rev. 2015, 44, 362–381.
- Hamd-Ghadareh, S.; Salimi, A.; Fathi, F.; Bahrami, S. An amplified comparative fluorescence resonance energy transfer immunosensing of CA125 tumor marker and ovarian cancer cells using green and economic carbon dots for bio-applications in labeling, imaging and sensing. Biosens. Bioelectron. 2017, 96, 308–316.
- Chen, S.; Yuan, R.; Chai, Y.; Xu, Y.; Min, L.; Li, N. A new antibody immobilization technique based on organic polymers protected Prussian blue nanoparticles and gold colloidal nanoparticles for amperometric immunosensors. Sens. Actuators B Chem. 2008, 135, 236–244.
- Wu, L.; Sha, Y.; Li, W.; Wang, S.; Guo, Z.; Zhou, J.; Su, X.; Jiang, X. One-step preparation of disposable multi-functionalized g-C3N4 based electrochemiluminescence immunosensor for the detection of CA125. Sens. Actuators B Chem. 2016, 226, 62–68.
- Zhu, D.; Liu, B.; Wei, G. Two-Dimensional Material-Based Colorimetric Biosensors: A Review. Biosensors 2021, 11, 259.
- Li, D.; Cheng, W.; Yan, Y.; Zhang, Y.; Yin, Y.; Ju, H.; Ding, S. A colorimetric biosensor for detection of attomolar microRNA with a functional nucleic acid-based amplification machine. Talanta 2016, 146, 470–476.
- Ali, M.M.; Wolfe, M.; Tram, K.; Gu, J.; Filipe, C.D.; Li, Y.; Brennan, J.D. A DNAzyme-Based Colorimetric Paper Sensor for Helicobacter pylori. Angew. Chem. 2019, 131, 10012–10016.
- Jiang, J.; Chen, Y.; Shi, J.; Song, C.; Zhang, J.; Wang, K. Population attributable burden of Helicobacter pylori-related gastric cancer, coronary heart disease, and ischemic stroke in China. Eur. J. Clin. Microbiol. Infect. Dis. 2017, 36, 199–212.
- Talebi Bezmin Abadi, A. Helicobacter pylori: Emergence of a Superbug. Front. Med. 2014, 1, 34.
- Backert, S.; Clyne, M. Pathogenesis of Helicobacter pylori infection. Helicobacter 2011, 16, 19–25.
- Ferwana, M.; Abdulmajeed, I.; Alhajiahmed, A.; Madani, W.; Firwana, B.; Hasan, R.; Altayar, O.; Limburg, P.J.; Murad, M.H.; Knawy, B. Accuracy of urea breath test in Helicobacter pylori infection: Meta-analysis. World J. Gastroenterol. WJG 2015, 21, 1305.
- Li, J.; Mo, L.; Lu, C.-H.; Fu, T.; Yang, H.-H.; Tan, W. Functional nucleic acid-based hydrogels for bioanalytical and biomedical applications. Chem. Soc. Rev. 2016, 45, 1410–1431.
- Zhang, L.; Lei, J.; Liu, L.; Li, C.; Ju, H. Self-assembled DNA hydrogel as switchable material for aptamer-based fluorescent detection of protein. Anal. Chem. 2013, 85, 11077–11082.
- Zhou, L.; Sun, N.; Xu, L.; Chen, X.; Cheng, H.; Wang, J.; Pei, R. Dual signal amplification by an “on-command” pure DNA hydrogel encapsulating HRP for colorimetric detection of ochratoxin A. Rsc Adv. 2016, 6, 114500–114504.
- Mao, X.; Chen, G.; Wang, Z.; Zhang, Y.; Zhu, X.; Li, G. Surface-immobilized and self-shaped DNA hydrogels and their application in biosensing. Chem. Sci. 2018, 9, 811–818.
- Xu, W.; He, W.; Du, Z.; Zhu, L.; Huang, K.; Lu, Y.; Luo, Y. Functional nucleic acid nanomaterials: Development, properties, and applications. Angew. Chem. Int. Ed. 2021, 60, 6890–6918.
- Fakude, C.T.; Arotiba, O.A.; Mabuba, N. Electrochemical aptasensing of cadmium (II) on a carbon black-gold nano-platform. J. Electroanal. Chem. 2020, 858, 113796.
- Yao, X.; Chadan Chen, L.C.; Wei, X.; Cui, H.; Xu, H.; Fan, H. A Novel PCB77 Electrochemical Sensor Based on Nano-functionalized Electrode and Selected Aptamer. J. New Mater. Electrochem. Syst. 2020, 23, 66–70.
- Yin, B.-C.; Ma, J.-L.; Le, H.-N.; Wang, S.; Xu, Z.; Ye, B.-C. A new mode to light up an adjacent DNA-scaffolded silver probe pair and its application for specific DNA detection. Chem. Commun. 2014, 50, 15991–15994.
- Sun, K.; Chang, Y.; Zhou, B.; Wang, X.; Liu, L. Gold nanoparticles-based electrochemical method for the detection of protein kinase with a peptide-like inhibitor as the bioreceptor. Int. J. Nanomed. 2017, 12, 1905.
- Flajolet, M.; He, G.; Heiman, M.; Lin, A.; Nairn, A.C.; Greengard, P. Regulation of Alzheimer’s disease amyloid-β formation by casein kinase I. Proc. Natl. Acad. Sci. USA 2007, 104, 4159–4164.
- Wang, M.; Lin, Z.; Liu, Q.; Jiang, S.; Liu, H.; Su, X. DNA-hosted copper nanoclusters/graphene oxide based fluorescent biosensor for protein kinase activity detection. Anal. Chim. Acta 2018, 1012, 66–73.





