Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Li, X. PPR Proteins in Plant Growth and Development. Encyclopedia. Available online: https://encyclopedia.pub/entry/15530 (accessed on 25 May 2026).
Li X. PPR Proteins in Plant Growth and Development. Encyclopedia. Available at: https://encyclopedia.pub/entry/15530. Accessed May 25, 2026.
Li, Xiulan. "PPR Proteins in Plant Growth and Development" Encyclopedia, https://encyclopedia.pub/entry/15530 (accessed May 25, 2026).
Li, X. (2021, October 29). PPR Proteins in Plant Growth and Development. In Encyclopedia. https://encyclopedia.pub/entry/15530
Li, Xiulan. "PPR Proteins in Plant Growth and Development." Encyclopedia. Web. 29 October, 2021.
Copy Citation
Pentatricopeptide repeat (PPR) proteins are characterized by the presence of tandem arrays of a degenerate 35-amino-acid repeat motif, PPR motif. Based on the types of motif and their arrangement, PPR proteins are divided into two classes, P and PLS. P-class proteins only contain canonical P-motifs with 35 amino acids, whereas PLS-class proteins consist of P-, L- (35 or 36 amino acids), and S- (31 or 32 amino acids) motifs forming tandemly repeated PLS triplets.
PPR protein
cytoplasmic male sterility
seed development
1. Introduction
Pentatricopeptide repeat (PPR) proteins are characterized by the presence of tandem arrays of a degenerate 35-amino-acid repeat motif, PPR motif [1]. Based on the types of motif and their arrangement, PPR proteins are divided into two classes, P and PLS. P-class proteins only contain canonical P-motifs with 35 amino acids, whereas PLS-class proteins consist of P-, L- (35 or 36 amino acids), and S- (31 or 32 amino acids) motifs forming tandemly repeated PLS triplets [2]. Many of the PLS-class proteins are carboxyl terminally extended with highly conserved E, E+, or DYW domains. Thus, PLS-class proteins can be further divided into PLS, E, E+, and DYW subclasses according to the domains identified in carboxyl terminal [3].
PPR proteins are sequence-specific RNA-binding proteins that are mostly localized to mitochondria and/or chloroplasts, where they are involved in RNA post-transcriptional processing [4] (Figure 1A). In general, the P-class PPR proteins mediate diverse aspects of RNA processing in plant organelles, while the PLS-class PPR proteins mainly function in RNA editing [5]. Mutations in these PPR protein-coding genes lead to the dysfunction of mitochondria and/or chloroplasts, thereby resulting in growth retardation, pollen abortion, and seed development defects in plants [4], indicating the important roles of PPR proteins in plant growth and development (Figure 1B).
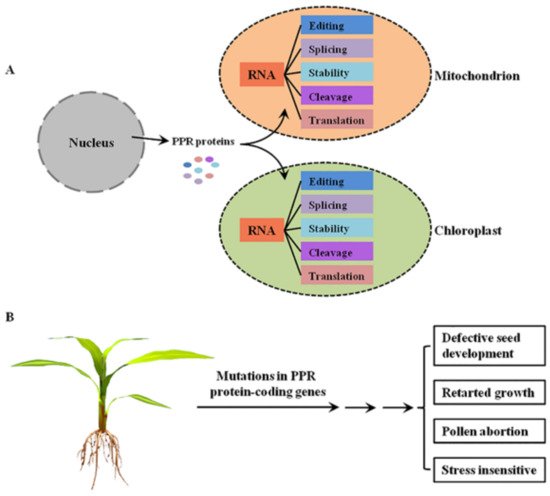
Figure 1. The functions of PPR proteins in plants. (A) The molecular functions of PPR proteins in plant mitochondria and chloroplasts. PPR proteins are encoded by nuclear genes, translated in the cytoplasm, and then imported into mitochondrion or chloroplast to mediate multiple steps of RNA processing. (B) The main growing and developmental phenotypes of plant mutants of PPR protein-coding genes.
2. Functions of PPR Proteins in Cytoplasmic Male Sterility
Cytoplasmic male sterility (CMS) is a maternally inherited trait that presents a defect in the production of viable pollen. CMS is widespread in higher plants, and has been widely used in the production of hybrid seeds and utilization of heterosis in many crop species [6][7]. Plant CMS is usually caused by mutations, rearrangements, or recombinations of mitochondrial DNA, and in many instances, male fertility can be restored specifically by restorer-of-fertility (Rf) genes in the nuclear genome [8][9]. To date, more than ten Rf genes have been cloned and functionally characterized in various crop species, and the majority of them were found to encode PPR protein, including Rf1a [10], Rf1b [11], Rf3 [12], Rf4 [13], Rf5 [14], Rf6 [15] in rice; Rfo [16], PPR-B [17], RsRf3-4 [18], Rfk1 [19] in radish; Rf1 [20], Rf2 [21] in sorghum; Rfp [22], Rfn [23] in rapeseed; Rf-PPR592 [24] in petunia; BrRfp1 [25] in Chinese cabbage; and Rfm1 [26] in barley. With the exception of the sorghum Rf1 [20] and the barely Rfm1 [26], the PPR-type Rf genes encode PPR proteins belonging to P class.
Members of P-class PPR proteins mostly function in various post-transcriptional process of organellar RNAs, such as RNA splicing, RNA stabilization, RNA cleavage, and translation [4][5]. Proteins encoded by Rf genes usually target mitochondria and act as fertility restorers by suppressing the production of mitochondrial CMS-inducing proteins [6][11]; however, the exact molecular mechanisms underlying fertility restoration by RF proteins is presently unclear. The PPR-type RF proteins have been proposed to rescue fertility by regulating the expression of CMS-conferring genes through the similar molecular mechanisms as that of other PPR proteins. In most of the CMS systems, PPR-type RF proteins bind to the mitochondrial CMS-conferring transcripts and promote their cleavage or degradation [27]. For example, rice RF1A and RF1B proteins, respectively encoded by Rf1a and Rf1b genes, have been considered to restore CMS by processing mitochondrial orf79 transcript via different mechanisms. RF1A directly binds to and cleaves the atp6-orf79 transcript at the intercistronic region, whereas RF1B promotes the rapid degradation of the atp6-orf79 transcript [10][11]. The protein encoded by rice Rf4 gene was reported to suppress WA352-mediated male sterility by reducing WA352 transcript levels [13]. Two rice RF proteins, Rfp and Rfn, are involved in transcript cleavage of orf224 and orf222, respectively [22][23].
In some CMS systems, PPR-type RF proteins restore the male fertility by impeding the translation or post-translational processing of mitochondrial CMS-inducing proteins. Studies on radish PPR-B revealed that PPR-B protein does not function through cleavage or degradation of the orf138 mRNA, but rather block its translation by inhibiting either the association with or the progression of mitochondrial ribosomes on the orf138 mRNA [17]. Similarly, rice RF3 protein does not affect the abundance of WA352 transcript but suppresses the accumulation of WA352 protein [12]. In addition, Rf1 from sorghum and Rfm1 from barely encode PPR proteins belonging to PLS class. As PLS-class PPR proteins almost exclusively play a role in RNA editing, it is possible that sorghum Rf1 and barely Rfm1 restore pollen fertility by editing S-orf transcripts or other target RNAs [8][26].
CMS was not only an ideal model system to study the interaction between mitochondrial and nuclear genomes but also a useful genetic tool for breeding to exploit hybrid vigor in crops [27]. Although PPR proteins are involved in the restoration of male fertility, functions of most PPR proteins are still obscure. Therefore, exploring the functions of PPR proteins will contribute to understanding the CMS mechanism and improving molecular breeding in crops.
3. Functions of PPR Proteins in Seed Development
In angiosperm plants, seed development starts with double fertilization of egg and central cells with two sperm cells, which leads to the formation of a diploid embryo and a triploid endosperm, and develops into mature seeds comprising three structures: maternal coat, embryo, and endosperm [28]. Development of embryo and endosperm is well correlated and regulated by numerous distinct proteins involved in many important physiological processes [29], including cell growth, RNA transcription and post-transcriptional processing, etc.
In recent years, more and more genetic and biochemical studies have shown that PPR proteins play important roles in seed development of higher plants, and loss-of-function of these PPR proteins usually leads to defects in embryogenesis and/or endosperm development [4][30]. According to the phenotypic expression, seed mutants can be divided into four major classes: empty pericarp (emp), embryo specific (emb), defective kernel (dek), and small kernel (smk). A detailed summary of the maize and Arabidopsis seed mutants caused by the functional defects of PPR proteins is provided in Table 1.
Table 1. Selected functionally characterized PPR proteins essential for seed development in maize and Arabidopsis.
| Species | Subcellular Localization |
Mutant Phenotype |
Protein Name |
PPR Class |
Function(s) | References |
|---|---|---|---|---|---|---|
| Maize | Mitochondrion | dek | DEK2 | P | RNA splicing, nad1 intron 1 | [31] |
| Mitochondrion | dek | DEK10 | PLS | RNA editing, nad3-61, 62, and cox2-550 | [32] | |
| Mitochondrion | dek | DEK35 | P | RNA splicing, nad4 intron 1 | [33] | |
| Mitochondrion | dek | DEK36 | PLS | RNA editing, atp4-59, nad7-383, and ccmFN-302 | [34] | |
| Mitochondrion | dek | DEK37 | P | RNA splicing, nad2 intron 1 | [35] | |
| Mitochondrion | dek | DEK39 | PLS | RNA editing, nad3-247 and nad3-275 | [36] | |
| Mitochondrion | dek | DEK40 | PLS | RNA editing, cox3-314, nad2-26, and nad5-1916 | [37] | |
| Mitochondrion | dek | DEK41/ DEK43 |
P | RNA splicing, nad4 intron 1 and 3 | [38][39] | |
| Mitochondrion | dek | DEK46 | PLS | RNA editing, D5-C22 of nad7 intron 3 and 4 | [40] | |
| Mitochondrion | dek | DEK53 | PLS | RNA editing, multiples sites | [41] | |
| Mitochondrion | dek | DEK55 | PLS | RNA splicing, nad4 intron 1 and 3;RNA editing, multiple sites | [42] | |
| Mitochondrion | dek | DEK605 | PLS | RNA editing, nad1-608 | [43] | |
| Mitochondrion | smk | SMK1 | PLS | RNA editing, nad7-836 | [44] | |
| Mitochondrion | smk | SMK4 | PLS | RNA editing, cox1-1489 | [45] | |
| Mitochondrion | smk | SMK6 | PLS | RNA editing, nad1-740, nad4L-110, nad7-739, and mttB-138, 139 | [46] | |
| Mitochondrion | smk | ZmSMK9 | P | RNA splicing, nad5 intron 1 and 4 | [47] | |
| Mitochondrion | smk | PPR2263 | PLS | RNA editing, nad5-1550 and cob-908 | [48] | |
| Mitochondrion | smk | MPPR6 | P | Translation, rps3 mRNA | [49] | |
| Mitochondrion | dek/smk | PPR20 | P | RNA splicing, nad2 intron 3 | [50] | |
| Mitochondrion | smk | PPR78 | P | RNA stabilization, nad5 mature mRNA | [51] | |
| Mitochondrion | emp | EMP4 | P | Expression of mitochondrial transcripts | [52] | |
| Mitochondrion | emp | EMP5 | PLS | RNA editing, multiple sites | [53] | |
| Mitochondrion | emp | EMP7 | PLS | RNA editing, ccmFN-1553 | [54] | |
| Mitochondrion | emp | EMP8 | P | RNA splicing, nad1 intron 4, nad2 intron 1, and nad4 intron 1 | [55] | |
| Mitochondrion | emp | EMP9 | PLS | RNA editing, ccmB-43 and rps4-335 | [56] | |
| Mitochondrion | emp | EMP10 | P | RNA splicing, nad2 intron 1 | [57] | |
| Mitochondrion | emp | EMP11 | P | RNA splicing, nad1 intron 1, 2, 3, and 4 | [58] | |
| Mitochondrion | emp | EMP12 | P | RNA splicing, nad2 intron 1, 2, and 4 | [59] | |
| Chloroplast | smk | qKW9 | PLS | RNA editing, NdhB-246 | [60] | |
| Mitochondrion | emp | EMP16 | P | RNA splicing, nad2 intron 4 | [61] | |
| Mitochondrion | emp | EMP17 | PLS | RNA editing, ccmFC-799 and nad2-677 | [62] | |
| Mitochondrion | emp | EMP18 | PLS | RNA editing, atp6-635 and cox2-449 | [63] | |
| Mitochondrion | emp | EMP21 | PLS | RNA editing, multiple sites | [64] | |
| Mitochondrion | emp | EMP32 | P | RNA splicing, nad7 intron 2 | [65] | |
| Mitochondrion | emp | EMP602 | P | RNA splicing, nad4 intron 1 and 3 | [66] | |
| Mitochondrion | emp | EMP603 | P | RNA splicing, nad1 intron 2 | [67] | |
| Mitochondrion | emp | PPR14 | P | RNA splicing, nad2 intron 3, nad7 intron 1 and 2 | [68] | |
| Mitochondrion | emp | PPR18 | P | RNA splicing, nad4 intron 1 | [69] | |
| Mitochondrion | emp | PPR27 | PLS | RNA editing, multiple sites | [70] | |
| Mitochondrion | emp | PPR101 | P | RNA splicing, nad5 intron 1 and 2 | [71] | |
| Mitochondrion | emp | PPR231 | P | RNA splicing, nad5 intron 1, 2, 3 and nad2 intron 3 | [71] | |
| Mitochondrion | emp | PPR-SMR1 | P | RNA splicing, multiple introns | [72] | |
| Chloroplast | emb | PPR8522 | P | RNA transcription, nearly all chloroplast-encoded genes | [73] | |
| Chloroplast | emb | EMB-7L | P | RNA splicing, multiple introns | [74] | |
| Arabidopsis | Mitochondrion | dek | OTP43 | P | RNA splicing, nad1 intron 1 | [75] |
| Mitochondrion | smk | PPR19 | P | RNA stabilization, nad1 mature mRNA | [76] | |
| Mitochondrion | emp | BLX | PLS | RNA editing, multiple sites; RNA splicing, nad1 intron 4 and nad2 intron 1 | [77] | |
| Chloroplast | emb | AtPPR2 | P | RNA translation | [78] | |
| Chloroplast | emb | ECD2 | P | RNA splicing, ndhA, ycf3 intron 1, rps12 intron 2 and clpp intron 2 | [79] | |
| Chloroplast | emb | EMB2654 | P | RNA splicing, rps12 intron 1 | [80] | |
| Mitochondrion | emb | EMB2794 | P | RNA splicing, nad2 intron 2 | [81] | |
| Nucleus | emb | GRP23 | P | RNA transcription | [82] | |
| Mitochondrion | emb | MID1 | P | RNA splicing, nad2 intron 1 | [83] | |
| Chloroplast | emb | PMD3 | P | RNA splicing, trnA, ndhB, and clpP-1 | [84] |
Most PPR proteins identified to date are targeted to mitochondria and/or chloroplasts [4]. The disruption of PPR proteins localized to chloroplasts usually results in the emb mutants that are defective in embryogenesis, but relatively normal in endosperm development. For instance, PPR8522 [73] and EMB-7L [74] in maize and GRP23 [82] in Arabidopsis are necessary for embryogenesis, and their mutations lead to arrested embryo development at the transition stage, resulting in an embryo-lethal phenotype. For mitochondrion-targeted PPR proteins, their disruptions mostly cause diverse seed development mutants, including smk, dek, and emp, with different degrees of defects in embryo and endosperm. The loss-of-function of SMK1 [44], SMK4 [45], SMK6 [46], ZmSMK9 [47], PPR2263 [48], and MPPR6 [49] in maize and PPR19 [76] in Arabidopsis arrests both embryo and endosperm development, resulting in smk phenotypes. Some characterized mitochondrion-targeted PPR proteins, such as DEK2 [31], DEK10 [32], DEK35 [33], DEK36 [34], DEK37 [35], DEK39 [36], DEK40 [37], DEK41/43 [38][39], DEK46 [40], and DEK53 [41] in maize, are necessary for seed development, and their disruptions result in dek mutants with arrested development of both the embryo and endosperm at an early stage. Meanwhile, many PPR proteins are targeted to mitochondria and function in development of both embryo and endosperm, and mutations in their encoding genes arrest embryo and endosperm development at early stages and even result in embryo lethality. For example, Emp5 encodes a mitochondrion-targeted DYW-type PPR protein, the emp5 mutants exhibit abortion of embryo and endosperm development at early stages in maize [53]. Loss-of-function of the mitochondrial P-type PPR protein EMP10 severely disturbs embryo and endosperm development, resulting in empty pericarp or papery seeds in maize [57]. Additionally, the P-type protein PPR5 was recently identified as a regulator required for endosperm development in rice, the ppr5 mutants develop small starch grains [85].
References
- Small, I.D.; Peeters, N. The PPR motif-a TPR-related motif prevalent in plant organellar proteins. Trends Biochem. Sci. 2000, 25, 45–47.
- Lurin, C.; Andrés, C.; Aubourg, S.; Bellaoui, M.; Bitton, F.; Bruyère, C.; Caboche, M.; Debast, C.; Gualberto, J.; Hoffmann, B.; et al. Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 2004, 16, 2089–2103.
- Cheng, S.F.; Gutmann, B.; Zhong, X.; Ye, Y.T.; Fisher, M.F.; Bai, F.Q.; Castleden, I.; Song, Y.; Song, B.; Huang, J.Y.; et al. Redefining the structural motifs that determine RNA binding and RNA editing by pentatricopeptide repeat proteins in land plants. Plant J. 2016, 85, 532–547.
- Barkan, A.; Small, I. Pentatricopeptide repeat proteins in plants. Annu. Rev. Plant Biol. 2014, 65, 415–442.
- Schmitz-Linneweber, C.; Small, I. Pentatricopeptide repeat proteins: A socket set for organelle gene expression. Trends Plant Sci. 2008, 13, 663–670.
- Chase, C.D. Cytoplasmic male sterility: A window to the world of plant mitochondrial-nuclear interactions. Trends Genet. 2007, 23, 81–90.
- Bohra, A.; Jha, U.C.; Adhimoolam, P.; Bisht, D.; Singh, N.P. Cytoplasmic male sterility (CMS) in hybrid breeding in field crops. Plant Cell Rep. 2016, 35, 967–993.
- Linke, B.; Börner, T. Mitochondrial effects on flower and pollen development. Mitochondrion 2005, 5, 389–402.
- Eckardt, N.A. Cytoplasmic male sterility and fertility restoration. Plant Cell 2006, 18, 515–517.
- Wang, Z.H.; Zou, Y.J.; Li, X.Y.; Zhang, Q.Y.; Chen, L.T.; Wu, H.; Su, D.H.; Chen, Y.L.; Guo, J.X.; Luo, D.; et al. Cytoplasmic male sterility of rice with Boro II cytoplasm is caused by a cytotoxic peptide and is restored by two related PPR motif genes via distinct modes of mRNA silencing. Plant Cell 2006, 18, 676–687.
- Kazama, T.; Nakamura, T.; Watanabe, M.; Sugita, M.; Toriyama, K. Suppression mechanism of mitochondrial ORF79 accumulation by Rf1 protein in BT-type cytoplasmic male sterile rice. Plant J. 2008, 55, 619–628.
- Luo, D.P.; Xu, H.; Liu, Z.L.; Guo, J.X.; Li, H.Y.; Chen, L.T.; Fang, C.; Zhang, Q.Y.; Bai, M.; Yao, N.; et al. A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice. Nat. Genet. 2013, 45, 573–577.
- Tang, H.; Luo, D.; Zhou, D.; Zhang, Q.; Tian, D.; Zheng, X.; Chen, L.; Liu, Y.G. The rice restorer Rf4 for wild-abortive cytoplasmic male sterility encodes a mitochondrial-localized PPR protein that functions in reduction of WA352 transcripts. Mol. Plant 2014, 7, 1497–1500.
- Hu, J.; Wang, K.; Huang, W.C.; Liu, G.; Gao, Y.; Wang, J.M.; Huang, Q.; Ji, Y.X.; Qin, X.J.; Wan, L.; et al. The rice pentatricopeptide repeat protein RF5 restores fertility in Hong-Lian cytoplasmic male-sterile lines via a complex with the glycine-rich protein GRP162. Plant Cell 2012, 24, 109–122.
- Huang, W.C.; Yu, C.C.; Hu, J.; Wang, L.L.; Dan, Z.W.; Zhou, W.; He, C.L.; Zeng, Y.F.; Yao, G.X.; Qi, J.Z.; et al. Pentatricopeptide-repeat family protein RF6 functions with hexokinase 6 to rescue rice cytoplasmic male sterility. Proc. Natl. Acad. Sci. USA 2015, 112, 14984–14989.
- Brown, G.G.; Formanová, N.; Jin, H.; Wargachuk, R.; Dendy, C.; Patil, P.; Laforest, M.; Zhang, J.F.; Cheung, W.Y.; Landry, B.S. The radish Rfo restorer gene of Ogura cytoplasmic male sterility encodes a protein with multiple pentatricopeptide repeats. Plant J. 2003, 35, 262–272.
- Uyttewaal, M.; Arnal, N.; Quadrado, M.; Martin-Canadell, A.; Vrielynck, N.; Hiard, S.; Gherbi, H.; Bendahmane, A.; Budar, F.; Mireau, H. Characterization of Raphanus sativus pentatricopeptide repeat proteins encoded by the fertility restorer locus for Ogura cytoplasmic male sterility. Plant Cell 2008, 20, 3331–3345.
- Wang, Z.W.; De Wang, C.D.; Mei, S.Y.; Gao, L.; Zhou, Y.; Wang, T. An insertion-deletion at a pentatricopeptide repeat locus linked to fertility transition to cytoplasmic male sterility in radish (Raphanus sativus L.). Mol. Breed. 2015, 35, 108–112.
- Imai, R.; Koizuka, N.; Fujimoto, H.; Hayakawa, T.; Sakai, T.; Imamura, J. Delimitation of the fertility restorer locus Rfk1 to a 43-kb contig in Kosena radish (Raphanus sativus L.). Mol. Genet. Genom. 2003, 269, 388–394.
- Klein, R.R.; Klein, P.E.; Mullet, J.E.; Minx, P.; Rooney, W.L.; Schertz, K.F. Fertility restorer locus Rf1 of sorghum (Sorghum bicolor L.) encodes a pentatricopeptide repeat protein not present in the colinear region of rice chromosome 12. Theor. Appl. Genet. 2005, 111, 994–1012.
- Madugula, P.; Uttam, A.G.; Tonapi, V.A.; Ragimasalawada, M. Fine mapping of Rf2, a major locus controlling pollen fertility restoration in sorghum A1 cytoplasm, encodes a PPR gene and its validation through expression analysis. Plant Breed. 2018, 137, 148–161.
- Liu, Z.; Yang, Z.; Wang, X.; Li, K.; An, H.; Liu, J.; Yang, G.S.; Fu, T.; Yi, B.; Hong, D.F. A mitochondria-targeted PPR protein restores pol cytoplasmic male sterility by reducing orf224 transcript levels in oilseed rape. Mol. Plant 2016, 9, 1082–1084.
- Liu, Z.; Dong, F.M.; Wang, X.; Wang, T.; Su, R.; Hong, D.F.; Yang, G.S. A pentatricopeptide repeat protein restores nap cytoplasmic male sterility in Brassica napus. J. Exp. Bot. 2017, 68, 4115–4123.
- Gillman, J.D.; Bentolila, S.; Hanson, M.R. The petunia restorer of fertility protein is part of a large mitochondrial complex that interacts with transcripts of the CMS-associated locus. Plant J. 2007, 49, 217–227.
- Zhang, H.M.; Wu, J.Q.; Dai, Z.H.; Qin, M.L.; Hao, L.Y.; Ren, Y.J.; Li, Q.F.; Zhang, L.G. Allelism analysis of BrRfp locus in different restorer lines and map-based cloning of a fertility restorer gene, BrRfp1, for pol CMS in Chinese cabbage (Brassica rapa L.). Theor. Appl. Genet. 2017, 130, 539–547.
- Rizzolatti, C.; Bury, P.; Tatara, E.; Pin, P.A.; Rodde, N.; Bergès, H.; Budar, F.; Mireau, H.; Gielen, J.J.L. Map-based cloning of the fertility restoration locus Rfm1 in cultivated barley (Hordeum vulgare). Euphytica 2017, 213, 276–287.
- Chen, L.T.; Liu, Y.G. Male sterility and fertility restoration in crops. Annu. Rev. Plant Biol. 2014, 65, 579–606.
- Goldberg, R.B.; de Paiva, G.; Yadegari, R. Plant embryogenesis: Zygote to seed. Science 1994, 266, 605–614.
- Linkies, A.; Graeber, K.; Knight, C.; Leubner-Metzger, G. The evolution of seeds. New Phytol. 2010, 186, 817–831.
- Dai, D.W.; Ma, Z.Y.; Song, R.T. Maize kernel development. Mol. Breed. 2021, 41, 2.
- Qi, W.W.; Yang, Y.; Feng, X.Z.; Zhang, M.L.; Song, R.T. Mitochondrial function and maize kernel development requires Dek2, a pentatricopeptide repeat protein involved in nad1 mRNA splicing. Genetics 2017, 205, 239–249.
- Qi, W.W.; Tian, Z.R.; Lu, L.; Chen, X.Z.; Chen, X.Z.; Zhang, W.; Song, R. Editing of mitochondrial transcripts nad3 and cox2 by Dek10 is essential for mitochondrial function and maize plant development. Genetics 2017, 205, 1489–1501.
- Chen, X.Z.; Feng, F.; Qi, W.W.; Xu, L.M.; Yao, D.S.; Wang, Q.; Song, R.T. Dek35 encodes a PPR protein that affects cis-splicing of mitochondrial nad4 intron 1 and seed development in maize. Mol. Plant 2017, 10, 427–441.
- Wang, G.; Zhong, M.Y.; Shuai, B.L.; Song, J.D.; Zhang, J.; Han, L.; Ling, H.L.; Tang, Y.P.; Wang, G.F.; Song, R.T. E+ subgroup PPR protein defective kernel 36 is required for multiple mitochondrial transcripts editing and seed development in maize and Arabidopsis. New Phytol. 2017, 214, 1563–1578.
- Dai, D.W.; Luan, S.C.; Chen, X.Z.; Wang, Q.; Feng, Y.; Zhu, C.G.; Qi, W.W.; Song, R.T. Maize Dek37 encodes a P-type PPR protein that affects cis-splicing of mitochondrial nad2 intron 1 and seed development. Genetics 2018, 208, 1069–1082.
- Li, X.J.; Gu, W.; Sun, S.L.; Chen, Z.L.; Chen, J.; Song, W.B.; Zhao, H.M.; Lai, J.S. Defective Kernel 39 encodes a PPR protein required for seed development in maize. J. Integr. Plant Biol. 2018, 60, 45–64.
- Ren, R.C.; Lu, X.D.; Zhao, Y.J.; Wei, Y.M.; Wang, L.L.; Zhang, L.; Zhang, W.T.; Zhang, C.Y.; Zhang, X.S.; Zhao, X.Y. Pentatricopeptide repeat protein DEK40 is required for mitochondrial function and kernel development in maize. J. Exp. Bot. 2019, 70, 6163–6179.
- Ren, R.C.; Wang, L.L.; Zhang, L.; Zhao, Y.J.; Wu, J.W.; Wei, Y.M.; Zhang, X.S.; Zhao, X.Y. DEK43 is a P-type pentatricopeptide repeat (PPR) protein responsible for the cis-splicing of nad4 in maize mitochondria. J. Integr. Plant Biol. 2020, 62, 299–313.
- Zhu, C.G.; Jin, G.P.; Fang, P.; Zhang, Y.; Feng, X.Z.; Tang, Y.P.; Qi, W.W.; Song, R.T. Maize pentatricopeptide repeat protein DEK41 affects cis-splicing of mitochondrial nad4 intron 3 and is required for normal seed development. J. Exp. Bot. 2019, 70, 3795–3808.
- Xu, C.H.; Song, S.; Yang, Y.Z.; Lu, F.; Zhang, M.D.; Sun, F.; Jia, R.X.; Song, R.L.; Tan, B.C. DEK46 performs C-to-U editing of a specific site in mitochondrial nad7 introns that is critical for intron splicing and seed development in maize. Plant J. 2020, 103, 1767–1782.
- Dai, D.W.; Jin, L.F.; Huo, Z.Z.; Yan, S.M.; Ma, Z.Y.; Qi, W.W.; Song, R.T. Maize pentatricopeptide repeat protein DEK53 is required for mitochondrial RNA editing at multiple sites and seed development. J. Exp. Bot. 2020, 71, 6246–6261.
- Ren, R.C.; Yan, X.W.; Zhao, Y.J.; Wei, Y.M.; Lu, X.D.; Zang, J.; Wu, J.W.; Zheng, G.M.; Ding, X.H.; Zhang, X.S.; et al. The novel E-subgroup pentatricopeptide repeat protein DEK55 is responsible for RNA editing at multiple sites and for the splicing of nad1 and nad4 in maize. BMC Plant Biol. 2020, 20, 553.
- Fan, K.J.; Peng, Y.X.; Ren, Z.J.; Li, D.L.; Zhen, S.H.; Hey, S.; Cui, Y.; Fu, J.J.; Gu, R.L.; Wang, J.H.; et al. Maize defective kernel605 encodes a canonical DYW-Type PPR protein that edits a conserved site of nad1 and is essential for seed nutritional quality. Plant Cell Physiol. 2020, 61, 1954–1966.
- Li, X.J.; Zhang, Y.F.; Hou, M.M.; Sun, F.; Shen, Y.; Xiu, Z.H.; Wang, X.M.; Chen, Z.L.; Sun, S.S.M.; Small, I.; et al. Small kernel 1 encodes a pentatricopeptide repeat protein required for mitochondrial nad7 transcript editing and seed development in maize (Zea mays) and rice (Oryza sativa). Plant J. 2014, 79, 797–809.
- Wang, H.C.; Sayyed, A.; Liu, X.Y.; Yang, Y.Z.; Sun, F.; Wang, Y.; Wang, M.D.; Tan, B.C. SMALL KERNEL4 is required for mitochondrial cox1 transcript editing and seed development in maize. J. Integr. Plant Biol. 2020, 62, 777–792.
- Ding, S.; Liu, X.Y.; Wang, H.C.; Wang, Y.; Tang, J.J.; Yang, Y.Z.; Tan, B.C. SMK6 mediates the C-to-U editing at multiple sites in maize mitochondria. J. Plant Physiol. 2019, 240, 152992.
- Pan, Z.Y.; Liu, M.; Xiao, Z.Y.; Ren, X.M.; Zhao, H.L.; Gong, D.M.; Liang, K.; Tan, Z.D.; Shao, Y.Q.; Qiu, F.Z. ZmSMK9, a pentatricopeptide repeat protein, is involved in the cis-splicing of nad5, kernel development and plant architecture in maize. Plant Sci. 2019, 288, 110205.
- Sosso, D.; Mbelo, S.; Vernoud, V.; Gendrot, G.; Dedieu, A.; Chambrier, P.; Dauzat, M.; Heurtevin, L.; Guyon, V.; Takenaka, M.; et al. PPR2263, a DYW-subgroup pentatricopeptide repeat protein, is required for mitochondrial nad5 and cob transcript editing, mitochondrion biogenesis, and maize growth. Plant Cell 2012, 24, 676–691.
- Manavski, N.; Guyon, V.; Meurer, J.; Wienand, U.; Brettschneider, R. An essential pentatricopeptide repeat protein facilitates 5′ maturation and translation initiation of rps3 mRNA in maize mitochondria. Plant Cell 2012, 24, 3087–3105.
- Yang, Y.Z.; Ding, S.; Wang, Y.; Wang, H.C.; Liu, X.Y.; Sun, F.; Xu, C.H.; Liu, B.H.; Tan, B.C. PPR20 is required for the cis-splicing of mitochondrial nad2 intron 3 and seed development in maize. Plant Cell Physiol. 2020, 61, 370–380.
- Zhang, Y.F.; Suzuki, M.; Sun, F.; Tan, B.C. The mitochondrion-targeted PENTATRICOPEPTIDE REPEAT78 protein is required for nad5 mature mRNA stability and seed development in maize. Mol. Plant 2017, 10, 1321–1333.
- Gutierrez-Marcos, J.F.; Dal Pra, M.; Giulini, A.; Costa, L.M.; Gavazzi, G.; Cordelier, S.; Sellam, O.; Tatout, C.; Paul, W.; Perez, P.; et al. Empty pericarp4 encodes a mitochondrion-targeted pentatricopeptide repeat protein necessary for seed development and plant growth in maize. Plant Cell 2007, 19, 196–210.
- Liu, Y.J.; Xiu, Z.H.; Meeley, R.; Tan, B.C. Empty pericarp5 encodes a pentatricopeptide repeat protein that is required for mitochondrial RNA editing and seed development in maize. Plant Cell 2013, 25, 868–883.
- Sun, F.; Wang, X.M.; Bonnard, G.; Shen, Y.; Xiu, Z.H.; Li, X.J.; Gao, D.H.; Zhang, Z.H.; Tan, B.C. Empty pericarp7 encodes a mitochondrial E-subgroup pentatricopeptide repeat protein that is required for ccmFN editing, mitochondrial function and seed development in maize. Plant J. 2015, 84, 283–295.
- Sun, F.; Zhang, X.Y.; Shen, Y.; Wang, H.C.; Liu, R.; Wang, X.M.; Gao, D.H.; Yang, Y.Z.; Liu, Y.W.; Tan, B.C. The pentatricopeptide repeat protein EMPTY PERICARP8 is required for the splicing of three mitochondrial introns and seed development in maize. Plant J. 2018, 95, 919–932.
- Yang, Y.Z.; Ding, S.; Wang, H.C.; Sun, F.; Huang, W.L.; Song, S.; Xu, C.H.; Tan, B.C. The pentatricopeptide repeat protein EMP9 is required for mitochondrial ccmB and rps4 transcript editing, mitochondrial complex biogenesis and seed development in maize. New Phytol. 2017, 214, 782–795.
- Cai, M.J.; Li, S.Z.; Sun, F.; Sun, Q.; Zhao, H.L.; Ren, X.M.; Zhao, Y.X.; Tan, B.C.; Zhang, Z.X.; Qiu, F.Z. Emp10 encodes a mitochondrial PPR protein that affects the cis-splicing of nad2 intron 1 and seed development in maize. Plant J. 2017, 91, 132–144.
- Ren, X.M.; Pan, Z.Y.; Zhao, H.L.; Zhao, J.L.; Cai, M.J.; Li, J.; Zhang, Z.X.; Qiu, F.Z. EMPTY PERICARP11 serves as a factor for splicing of mitochondrial nad1 intron and is required to ensure proper seed development in maize. J. Exp. Bot. 2017, 68, 4571–4581.
- Sun, F.; Xiu, Z.H.; Jiang, R.C.; Liu, Y.W.; Zhang, X.Y.; Yang, Y.Z.; Li, X.J.; Zhang, X.; Wang, Y.; Tan, B.C. The mitochondrial pentatricopeptide repeat protein EMP12 is involved in the splicing of three nad2 introns and seed development in maize. J. Exp. Bot. 2019, 70, 963–972.
- Huang, J.; Lu, G.; Liu, L.; Raihan, M.S.; Xu, J.T.; Jian, L.M.; Zhao, L.X.; Tran, T.M.; Zhang, Q.H.; Liu, J.; et al. The kernel size-related quantitative trait locus qKW9 encodes a pentatricopeptide repeat protein that affects photosynthesis and grain filling. Plant Physiol. 2020, 183, 1696–1709.
- Xiu, Z.H.; Sun, F.; Shen, Y.; Zhang, X.Y.; Jiang, R.C.; Bonnard, G.; Zhang, J.H.; Tan, B.C. EMPTY PERICARP16 is required for mitochondrial nad2 intron 4 cis-splicing, complex I assembly and seed development in maize. Plant J. 2016, 85, 507–519.
- Wang, Y.; Liu, X.Y.; Huang, Z.Q.; Li, Y.Y.; Yang, Y.Z.; Sayyed, A.; Sun, F.; Gu, Z.Q.; Wang, X.M.; Tan, B.C. PPR-DYW protein EMP17 is required for mitochondrial RNA editing, complex III biogenesis, and seed development in maize. Front. Plant Sci. 2021, 12, 693272.
- Li, X.L.; Huang, W.L.; Yang, H.H.; Jiang, R.C.; Sun, F.; Wang, H.C.; Zhao, J.; Xu, C.H.; Tan, B.C. EMP18 functions in mitochondrial atp6 and cox2 transcript editing and is essential to seed development in maize. New Phytol. 2019, 221, 896–907.
- Wang, Y.; Liu, X.Y.; Yang, Y.Z.; Huang, J.; Sun, F.; Lin, J.S.; Gu, Z.Q.; Sayyed, A.; Xu, C.H.; Tan, B.C. Empty Pericarp21 encodes a novel PPR-DYW protein that is required for mitochondrial RNA editing at multiple sites, complexes I and V biogenesis, and seed development in maize. PLoS Genet. 2019, 15, e1008305.
- Yang, Y.Z.; Ding, S.; Liu, X.Y.; Tang, J.J.; Wang, Y.; Sun, F.; Xu, C.H.; Tan, B.C. EMP32 is required for the cis-splicing of nad7 intron 2 and seed development in maize. RNA Biol. 2021, 18, 499–509.
- Ren, Z.J.; Fan, K.J.; Fang, T.; Zhang, J.J.; Yang, L.; Wang, J.H.; Wang, G.Y.; Liu, Y.J. Maize Empty pericarp602 encodes a P-type PPR protein that is essential for seed development. Plant Cell Physiol. 2019, 60, 1734–1746.
- Fan, K.J.; Ren, Z.J.; Zhang, X.F.; Liu, Y.; Fu, J.J.; Qi, C.L.; Tatar, W.; Rasmusson, A.G.; Wang, G.Y.; Liu, Y.J. The pentatricopeptide repeat protein EMP603 is required for the splicing of mitochondrial Nad1 intron 2 and seed development in maize. J. Exp. Bot. 2021, erab339.
- Wang, H.C.; Chen, Z.L.; Yang, Y.Z.; Sun, F.; Ding, S.; Li, X.L.; Xu, C.H.; Tan, B.C. PPR14 interacts with PPR-SMR1 and CRM protein Zm-mCSF1 to facilitate mitochondrial intron splicing in maize. Front. Plant Sci. 2020, 11, 732.
- Liu, R.; Cao, S.K.; Sayyed, A.; Xu, C.H.; Sun, F.; Wang, X.M.; Tan, B.C. The mitochondrial pentatricopeptide repeat protein PPR18 is required for the cis-splicing of nad4 intron 1 and essential to seed development in maize. Int. J. Mol. Sci. 2020, 21, 4047.
- Liu, R.; Cao, S.K.; Sayyed, A.; Yang, H.H.; Zhao, J.; Wang, X.M.; Jia, R.X.; Sun, F.; Tan, B.C. The DYW-subgroup pentatricopeptide repeat protein PPR27 functions on editing of multiple mitochondrial transcripts and interacts with ZmMORF1 in maize. J. Exp. Bot. 2020, 71, 5495–5505.
- Yang, H.H.; Xiu, Z.H.; Wang, L.; Cao, S.K.; Li, X.L.; Sun, F.; Tan, B.C. Two pentatricopeptide repeat proteins are required for the splicing of nad5 introns in maize. Front. Plant Sci. 2020, 11, 732.
- Chen, Z.L.; Wang, H.C.; Shen, J.Y.; Sun, F.; Wang, M.D.; Xu, C.H.; Tan, B.C. PPR-SMR1 is required for the splicing of multiple mitochondrial introns, interacts with Zm-mCSF1, and is essential for seed development in maize. J. Exp. Bot. 2019, 70, 5245–5258.
- Sosso, D.; Canut, M.; Gendrot, G.; Dedieu, A.; Chambrier, P.; Barkan, A.; Consonni, G.; Rogowsky, P.M. PPR8522 encodes a chloroplast-targeted pentatricopeptide repeat protein necessary for maize embryogenesis and vegetative development. J. Exp. Bot. 2012, 63, 5843–5857.
- Yuan, N.N.; Wang, J.C.; Zhou, Y.; An, D.; Xiao, Q.; Wang, W.Q.; Wu, Y.R. EMB-7L is required for embryogenesis and plant development in maize involved in RNA splicing of multiple chloroplast genes. Plant Sci. 2019, 287, 110203.
- De Longevialle, A.F.; Meyer, E.H.; Andres, C.; Taylor, N.L.; Lurin, C.; Millar, A.H.; Small, I.D. The pentatricopeptide repeat gene OTP43 is required for trans-splicing of the mitochondrial nad1 intron 1 in Arabidopsis thaliana. Plant Cell 2007, 19, 3256–3265.
- Lee, K.; Han, J.H.; Park, Y.I.; Colas des Francs-Small, C.; Small, I.; Kang, H. The mitochondrial pentatricopeptide repeat protein PPR19 is involved in the stabilization of NADH dehydrogenase 1 transcripts and is crucial for mitochondrial function and Arabidopsis thaliana development. New Phytol. 2017, 215, 202–216.
- Sun, Y.; Huang, J.Y.; Zhong, S.; Gu, H.Y.; He, S.; Qu, L.J. Novel DYW-type pentatricopeptide repeat (PPR) protein BLX controls mitochondrial RNA editing and splicing essential for early seed development of Arabidopsis. J. Genet. Genom. 2018, 45, 155–168.
- Lu, Y.Q.; Li, C.; Wang, H.; Chen, H.; Berg, H.; Xia, Y.J. AtPPR2, an Arabidopsis pentatricopeptide repeat protein, binds to plastid 23S rRNA and plays an important role in the first mitotic division during gametogenesis and in cell proliferation during embryogenesis. Plant J. 2011, 67, 13–25.
- Wang, X.W.; An, Y.Q.; Qi, Z.; Xiao, J.W. PPR protein early chloroplast development 2 is essential for chloroplast development at the early stage of Arabidopsis development. Plant Sci. 2021, 308, 110908.
- Aryamanesh, N.; Ruwe, H.; Sanglard, L.V.P.; Eshraghi, L.; Bussell, J.D.; Howell, K.A.; Small, I.; Colas des Francs-Small, C. The pentatricopeptide repeat protein EMB2654 is essential for trans-splicing of a chloroplast small ribosomal subunit transcript. Plant Physiol. 2017, 173, 1164–1176.
- Marchetti, F.; Cainzos, M.; Shevtsov, S.; Córdoba, J.P.; Sultan, L.D.; Brennicke, A.; Takenaka, M.; Pagnussat, G.; Ostersetzer-Biran, O.; Zabaleta, E. Mitochondrial pentatricopeptide repeat protein, EMB2794, plays a pivotal role in NADH dehydrogenase subunit nad2 mRNA maturation in Arabidopsis thaliana. Plant Cell Physiol. 2020, 61, 1080–1094.
- Ding, Y.H.; Liu, N.Y.; Tang, Z.S.; Liu, J.; Yang, W.C. Arabidopsis GLUTAMINE-RICH PROTEIN23 is essential for early embryogenesis and encodes a novel nuclear PPR motif protein that interacts with RNA polymerase II subunit III. Plant Cell 2006, 18, 815–830.
- Zhao, P.; Wang, F.; Li, N.; Shi, D.Q.; Yang, W.C. Pentatricopeptide repeat protein MID1 modulates nad2 intron 1 splicing and Arabidopsis development. Sci Rep. 2020, 10, 2008.
- Zhang, J.; Xiao, J.W.; Li, Y.Q.; Su, B.D.; Xu, H.M.; Shan, X.Y.; Song, C.W.; Xie, J.B.; Li, R.L. PDM3, a pentatricopeptide repeat-containing protein, affects chloroplast development. J. Exp. Bot. 2017, 68, 5615–5627.
- Zhang, L.; Qi, Y.Z.; Wu, M.M.; Zhao, L.; Zhao, Z.C.; Lei, C.L.; Hao, Y.Y.; Yu, X.W.; Sun, Y.L.; Zhang, X.; et al. Mitochondrion-targeted PENTATRICOPEPTIDE REPEAT5 is required for cis-splicing of nad4 intron 3 and endosperm development in rice. Crop J. 2021, 9, 282–296.
More
Information
Subjects:
Agriculture, Dairy & Animal Science
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.0K
Revisions:
2 times
(View History)
Update Date:
01 Nov 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





