
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Ioannis Petrakis | + 691 word(s) | 691 | 2021-10-20 09:50:40 | | | |
| 2 | Dean Liu | Meta information modification | 691 | 2021-10-20 11:35:52 | | |
Video Upload Options
Heat-shock proteins (HSPs) are molecular chaperones participating primarily in protein folding preventing protein degradation and subsequent cellular distress.
1. Introduction
Heat-shock proteins (HSPs) are molecular chaperones participating primarily in protein folding preventing protein degradation and subsequent cellular distress [1]. HSPs are regulated through heat-shock factor 1(HSF-1) [2]. In the steady state HSF-1 is bound to HSP90 or HSP70 [3][4]. Upon stressful signals HSF-1 dissociates from HSPs and translocates into the nucleus where it stimulates HSP expression [1]. HSPs can be exposed to the immune system through tissue necrosis and the resultant cellular debris, via organized release of exosomes/endosomes, or through their presence on the cellular membrane [5][6][7]. Their evolutionary conservation can elicit interspecies immune recognition [8]. The resulting immune response can be either immunoregulatory or immunostimulatory [9][10]. Furthermore specific HSP domains as well as certain HSP isoforms and their client proteins induce a differential autoimmune response.
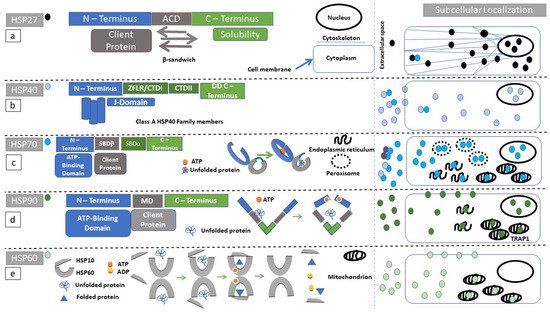
2. Structure and Subcellular Localization of the Small HSP Family
References
- Calderwood, S.K.; Repasky, E.A.; Neckers, L.; Hightower, L.E. The IXth CSSI international symposium on heat shock proteins in biology and medicine: Stress responses in health and disease: Alexandria Old Town, Alexandria, Virginia, November 10–13, 2018. Cell Stress Chaperones 2019, 24, 1–6.
- Calderwood, S.K.; Hightower, L.E. Report on the VIIth International Symposium on Heat Shock Proteins in Biology & Medicine. Cell Stress Chaperones 2015, 20, 213–216.
- Krakowiak, J.; Zheng, X.; Patel, N.; Feder, Z.A.; Anandhakumar, J.; Valerius, K.; Gross, D.S.; Khalil, A.S.; Pincus, D. Hsf1 and Hsp70 constitute a two-component feedback loop that regulates the yeast heat shock response. Elife 2018, 7.
- Kmiecik, S.W.; Le Breton, L.; Mayer, M.P. Feedback regulation of heat shock factor 1 (Hsf1) activity by Hsp70-mediated trimer unzipping and dissociation from DNA. EMBO J. 2020, 39, e104096.
- Tukaj, S. Heat Shock Protein 70 as a Double Agent Acting Inside and Outside the Cell: Insights into Autoimmunity. Int. J. Mol. Sci. 2020, 21.
- Calderwood, S.K.; Stevenson, M.A.; Murshid, A. Heat shock proteins, autoimmunity, and cancer treatment. Autoimmune Dis. 2012, 2012, 486069.
- Thaxton, J.E.; Liu, B.; Zheng, P.; Liu, Y.; Li, Z. Deletion of CD24 impairs development of heat shock protein gp96-driven autoimmune disease through expansion of myeloid-derived suppressor cells. J. Immunol. 2014, 192, 5679–5686.
- Winfield, J.B. Stress proteins, arthritis, and autoimmunity. Arthritis Rheum. 1989, 32, 1497–1504.
- Rajaiah, R.; Moudgil, K.D. Heat-shock proteins can promote as well as regulate autoimmunity. Autoimmun. Rev. 2009, 8, 388–393.
- Zuo, D.; Subjeck, J.; Wang, X.Y. Unfolding the Role of Large Heat Shock Proteins: New Insights and Therapeutic Implications. Front. Immunol. 2016, 7, 75.
- Kampinga, H.H.; Hageman, J.; Vos, M.J.; Kubota, H.; Tanguay, R.M.; Bruford, E.A.; Cheetham, M.E.; Chen, B.; Hightower, L.E. Guidelines for the nomenclature of the human heat shock proteins. Cell Stress Chaperones 2009, 14, 105–111.
- Hino, M.; Kurogi, K.; Okubo, M.A.; Murata-Hori, M.; Hosoya, H. Small heat shock protein 27 (HSP27) associates with tubulin/microtubules in HeLa cells. Biochem. Biophys. Res. Commun. 2000, 271, 164–169.
- Vos, M.J.; Kanon, B.; Kampinga, H.H. HSPB7 is a SC35 speckle resident small heat shock protein. Biochim. Biophys. Acta 2009, 1793, 1343–1353.
- Batulan, Z.; Pulakazhi Venu, V.K.; Li, Y.; Koumbadinga, G.; Alvarez-Olmedo, D.G.; Shi, C.; O’Brien, E.R. Extracellular Release and Signaling by Heat Shock Protein 27: Role in Modifying Vascular Inflammation. Front. Immunol. 2016, 7, 285.
- Haslbeck, M.; Weinkauf, S.; Buchner, J. Small heat shock proteins: Simplicity meets complexity. J. Biol. Chem. 2019, 294, 2121–2132.
- Charmpilas, N.; Kyriakakis, E.; Tavernarakis, N. Small heat shock proteins in ageing and age-related diseases. Cell Stress Chaperones 2017, 22, 481–492.
- Reddy, V.S.; Madala, S.K.; Trinath, J.; Reddy, G.B. Extracellular small heat shock proteins: Exosomal biogenesis and function. Cell Stress Chaperones 2018, 23, 441–454.
- Kostenko, S.; Johannessen, M.; Moens, U. PKA-induced F-actin rearrangement requires phosphorylation of Hsp27 by the MAPKAP kinase MK5. Cell. Signal. 2009, 21, 712–718.
- Mainz, A.; Peschek, J.; Stavropoulou, M.; Back, K.C.; Bardiaux, B.; Asami, S.; Prade, E.; Peters, C.; Weinkauf, S.; Buchner, J.; et al. The chaperone alphaB-crystallin uses different interfaces to capture an amorphous and an amyloid client. Nat. Struct. Mol. Biol. 2015, 22, 898–905.
- Ehrnsperger, M.; Hergersberg, C.; Wienhues, U.; Nichtl, A.; Buchner, J. Stabilization of proteins and peptides in diagnostic immunological assays by the molecular chaperone Hsp25. Anal. Biochem. 1998, 259, 218–225.
- Rosenzweig, R.; Nillegoda, N.B.; Mayer, M.P.; Bukau, B. The Hsp70 chaperone network. Nat. Rev. Mol. Cell. Biol. 2019, 20, 665–680.
- Zhang, K.; Ezemaduka, A.N.; Wang, Z.; Hu, H.; Shi, X.; Liu, C.; Lu, X.; Fu, X.; Chang, Z.; Yin, C.C. A novel mechanism for small heat shock proteins to function as molecular chaperones. Sci. Rep. 2015, 5, 8811.
- Kampinga, H.H.; Brunsting, J.F.; Stege, G.J.; Konings, A.W.; Landry, J. Cells overexpressing Hsp27 show accelerated recovery from heat-induced nuclear protein aggregation. Biochem. Biophys. Res. Commun. 1994, 204, 1170–1177.
- Cashikar, A.G.; Duennwald, M.; Lindquist, S.L. A chaperone pathway in protein disaggregation. Hsp26 alters the nature of protein aggregates to facilitate reactivation by Hsp104. J. Biol. Chem. 2005, 280, 23869–23875.





