| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Amira Abubakar | + 3680 word(s) | 3680 | 2021-10-09 11:35:36 | | | |
| 2 | Vivi Li | Meta information modification | 3680 | 2021-10-13 03:23:28 | | |
Video Upload Options
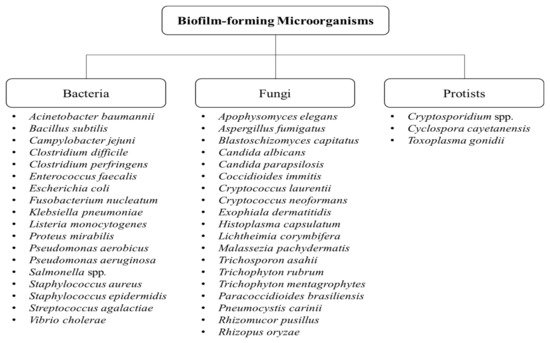
Biofilms comprising aggregates of microorganisms or multicellular communities have been a major issue as they cause resistance against antimicrobial agents and biofouling. To date, numerous biofilm-forming microorganisms have been identified, which have been shown to result in major effects including biofouling and biofilm-related infections. Quorum sensing (which describes the cell communication within biofilms) plays a vital role in the regulation of biofilm formation and its virulence. As such, elucidating the various mechanisms responsible for biofilm resistance (including quorum sensing) will assist in developing strategies to inhibit and control the formation of biofilms in nature. Employing biological control measures (such as the use of bioactive compounds) in targeting biofilms is of great interest since they naturally possess antimicrobial activity among other favorable attributes and can also possibly act as potent antibiofilm agents.
1. Introduction
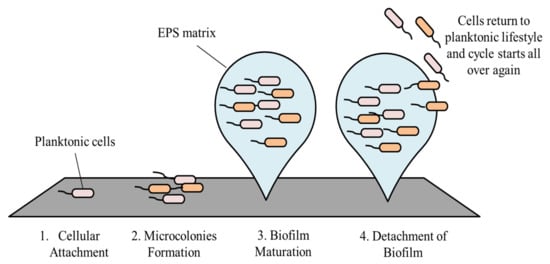
2. Stages in Biofilm Formation and Its Development
2.1. Cellular Attachment
2.2. Microcolonies Formation
2.3. Biofilm Maturation
2.4. Detachment of Biofilm
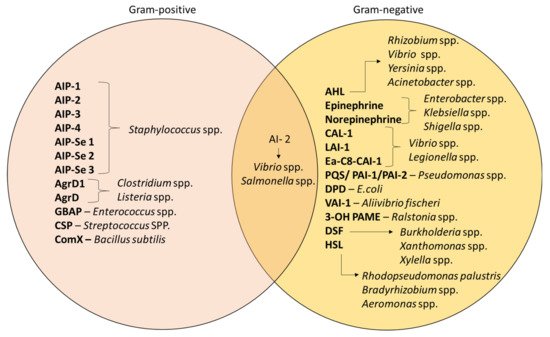
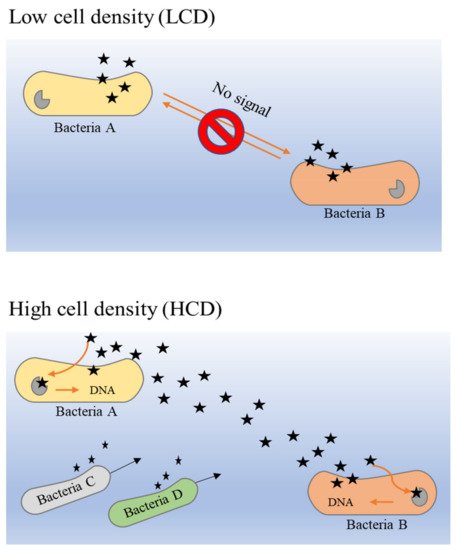
3. Quorum Sensing in Biofilm Formation
4. Consequences of Biofilm Formation
4.1. Biofouling
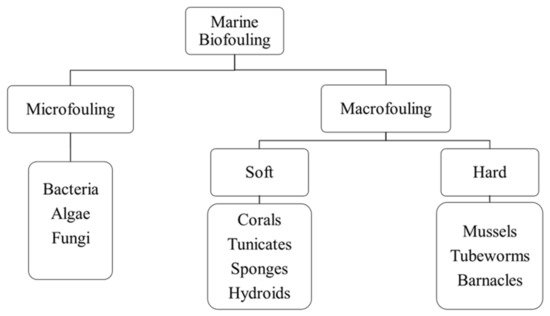
4.1.1. Biofouling in Marine Industries
4.1.2. Biofouling in Food and Beverage Industries
| Type of Food Industry | Prominent Bacteria | Effects | References |
|---|---|---|---|
| Dairy Industry | L. monocytogenes S. typhimurium and S. enteritidis E. coli (STEC) B. cereus |
Gastroenteritis or listeriosis Gastroenteritis Enterohemorrhagic gastroenteritis or hemolytic uremic syndrome (HUS) Gastroenteritis or occasionally acute liver failure |
[84][86][90][93][95] |
| Poultry Industry | S. enterica C. jejuni and C. coli |
Gastroenteritis or septicemia Enterocolitis or gastroenteritis |
[84][87][92] |
| Meat Industry | E. coli O157:H7 L. Monocytogenes Salmonella spp. |
Hemorrhagic colitis or thrombotic thrombocytopenic purpura (TTP) Gastroenteritis or listeriosis Salmonellosis |
[82][88][91][94][96] |
| Fish and Seafood Industry | Vibrio cholerae Aeromonas spp. Pseudomonas spp. |
Cholera or gastroenteritis Epizootic ulcerative syndrome (EUS) |
[83][85][86][89] |
4.1.3. Biofouling in Medical Industries
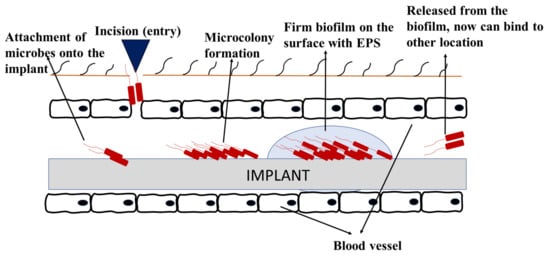
4.2. Biofilm-Related Infections
4.2.1. Periodontitis
4.2.2. Rhinosinusitis
4.2.3. Cystic Fibrosis (CF)
References
- Armbruster, C.R.; Parsek, M.R. New insight into the early stages of biofilm formation. Proc. Natl. Acad. Sci. USA 2018, 115, 4317–4319.
- Vasudevan, R. Biofilms: Microbial cities of scientific significance. J. Microbiol. Exp. 2014, 1, 84–98.
- Tasneem, U.; Yasin, N.; Nisa, I.; Shah, F.; Rasheed, U.; Momin, F.; Zaman, S.; Qasim, M. Biofilm producing bacteria: A serious threat to public health in developing countries. J. Food Sci. Nutr. 2018, 1, 25–31.
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–663.
- Karygianni, L.; Ren, Z.; Koo, H.; Thurnheer, T. Biofilm Matrixome: Extracellular components in structured microbial communities. Trends Microbiol. 2020, 28, 668–681.
- Costa-Orlandi, C.B.; Sardi, J.; Pitangui, N.S.; de Oliveira, H.C.; Scozorni, L.; Galeane, M.C.; Medina-Alarcón, K.P.; Melo, W.; Marcelino, M.Y.; Braz, J.D.; et al. Fungal biofilms and polymicrobial diseases. J. Fungi 2017, 3, 22.
- Bonaventura, G.D.; Piccolomini, R.; Paludi, D.; D’Orio, V.; Vergara, A.; Conter, M.; Ianieri, A. Influence of temperature on biofilm formation by Listeria monocytogenes on various food-contact surfaces: Relationship with motility and cell surface hydrophobicity. J. Appl. Microbiol. 2008, 104, 1552–1561.
- Hostacká, A.; Ciznar, I.; Mária Štefkoviková, M. Temperature and pH Affect the Production of Bacterial Biofilm. Folia Microbiol. 2010, 55, 75–78.
- Singh, R.; Shivaprakash, M.R.; Chakrabarti, A. Biofilm formation by zygomycetes: Quantification, structure and matrix composition. Microbiology 2011, 157, 2611–2618.
- Fanning, S.; Mitchell, A.P. Fungal biofilms. PLoS Pathog. 2012, 8, e1002585.
- D’Urzo, N.; Martinelli, M.; Pezzicoli, A.; De Cesare, V.; Pinto, V.; Margarit, I.; Telford, J.L.; Maione, D. Acidic pH strongly enhances in vitro biofilm formation by a subset of hypervirulent ST-17 Streptococcus agalactiae strains. Appl. Environ. Microbiol. 2014, 80, 2176–2185.
- Sarah, L.S.; Pryjma, M.; Gaynor, E.C. Flagella-mediated adhesion and extracellular DNA release contribute to biofilm formation and stress tolerance of Campylobacter jejuni. PLoS ONE 2014, 9, e106063.
- Zhang, W.B.; Seminara, A.; Suaris, M.; Brenner, M.P.; Weitz, D.A.; Angelini, T.E. Nutrient depletion in Bacillus subtilis biofilms triggers matrix production. New J. Phys. 2014, 16, 15028.
- Luo, L.M.; Wu, L.J.; Xiao, Y.L.; Zhao, D.; Chen, Z.X.; Kang, M.; Zhang, Q.; Xie, Y. Enhancing pili assembly and biofilm formation in Acinetobacter baumannii ATCC19606 using non-native acyl-homoserine lactones. BMC Microbiol. 2015, 15, 62.
- Maldarelli, G.A.; Piepenbrink, K.H.; Scott, A.J.; Freiberg, J.A.; Song, Y.; Achermann, Y.; Ernst, R.K.; Shirtliff, M.E.; Sundberg, E.J.; Donnenberg, M.S.; et al. Type IV pili promote early biofilm formation by Clostridium difficile. Pathog. Dis. 2016, 74, 1–10.
- Kirchhoff, L.; Olsowski, M.; Zilmans, K.; Dittmer, S.; Haase, G.; Sedlacek, L.; Steinmann, E.; Buer, J.; Rath, P.M.; Steinmann, J. Biofilm formation of the black yeast-like fungus Exophiala dermatitidis and its susceptibility to antiinfective agents. Sci. Rep. 2017, 7, 42886.
- Jamal, M.; Ahmad, W.; Andleeb, S.; Jalil, F.; Imran, M.; Nawaz, M.A.; Hussain, T.; Ali, M.; Rafiq, M.; Kamil, M.A. Bacterial biofilm and associated infections. J. Chin. Med. Assoc. 2018, 81, 7–11.
- Khatoon, Z.; McTiernan, C.; Suuronen, E.; Mah, T.; Alarcon, E. Bacterial biofilm formation on implantable devices and approaches to its treatment and prevention. Heliyon 2018, 4, e01067.
- Sharma, D.; Misba, L.; Khan, A.U. Antibiotics versus biofilm: An emerging battleground in microbial communities. Antimicrob. Resist. Infect. Control 2019, 8, 1–10.
- Yin, W.; Wang, Y.; Liu, L.; He, J. Biofilms: The microbial “protective clothing” in extreme environments. Int. J. Mol. Sci. 2019, 20, 3423.
- Rumbaugh, K.P.; Sauer, K. Biofilm dispersion. Nat. Rev. Microbiol. 2020, 18, 571–586.
- Domenech, M.; Ramos-Sevillano, E.; García, E.; Moscoso, M.; Yuste, J. Biofilm formation avoids complement immunity and phagocytosis of Streptococcus pneumoniae. Infect. Immun. 2013, 81, 2606–2615.
- Larsen, T.; Fiehn, N.E. Dental biofilm infections—An update. J. Pathol. Microbiol. Immunol. 2017, 125, 376–384.
- Crouzet, M.; Le Senechal, C.; Brözel, V.S.; Costaglioli, P.; Barthe, C.; Bonneu, M.; Garbay, B.; Vilain, S. Exploring early steps in biofilm formation: Set-up of an experimental system for molecular studies. BMC Microbiol. 2014, 14, 253.
- Palmer, J.; Flint, S.; Brooks, J. Bacterial cell attachment, the beginning of a biofilm. J. Ind. Microbiol. Biotechnol. 2007, 34, 577–588.
- Petrova, O.E.; Sauer, K. Sticky situations: Key components that control bacterial surface attachment. J. Bacteriol. 2012, 194, 2413–2425.
- Caiazza, N.C.; O’Toole, G.A. SadB is required for the transition from reversible to irreversible attachment during biofilm formation by Pseudomonas aeruginosa PA14. J. Bacteriol. 2004, 186, 4476–4485.
- Rabin, N.; Zheng, Y.; Opoku, T.C.; Du, Y.X.; Bonsu, F.; Sintim, H.O. Biofilm formation mechanisms and targets for developing antibiofilm agents. Future Med. Chem. 2015, 7, 493–512.
- Flemming, H.; Neu, T.; Wozniak, D. The EPS matrix: The “house of biofilm cells”. J. Bacteriol. 2007, 189, 7945–7947.
- Drescher, K.; Dunkel, J.; Nadell, C.D.; Teeffelen, S.; Grnja, I.; Wingreen, N.S.; Stone, H.A.; Bassler, B.L. Architectural transitions in Vibrio cholerae biofilms at single-cell resolution. Proc. Natl. Acad. Sci. USA 2016, 113, E2066–E2072.
- Bowen, W.; Burne, R.; Wu, H.; Koo, H. Oral biofilms: Pathogens, matrix, and polymicrobial interactions in microenvironments. Trends Microbiol. 2018, 26, 229–242.
- Hobley, L.; Harkins, C.; MacPhee, C.; Stanley-Wall, N. Giving structure to the biofilm matrix: An overview of individual strategies and emerging common themes. FEMS Microbiol. Rev. 2015, 39, 649–669.
- Preda, V.G.; Săndulescu, O. Communication is the key: Biofilms, quorum sensing, formation and prevention. Discoveries 2019, 7, e100.
- Saxena, P.; Joshi, Y.; Rawat, K.; Bisht, R. Biofilms: Architecture, resistance, quorum sensing and control mechanisms. Indian J. Microbiol. 2019, 59, 3–12.
- Wilking, J.N.; Zaburdaev, V.; De Volder, M.; Losick, R.; Brenner, M.P.; Weitz, D.A. Liquid transport facilitated by channels in Bacillus subtilis biofilms. Proc. Natl. Acad. Sci. USA 2013, 110, 848–852.
- Parsek, M.; Singh, P. Bacterial biofilms: An emerging link to disease pathogenesis. Annu. Rev. Microbiol. 2003, 57, 677–701.
- Chua, S.L.; Liu, Y.; Yam, J.K.H.; Chen, Y.C.; Veiborg, R.M.; Tan, B.G.C.; Kjelleberg, S.; Tolker-Nielsen, T.; Givskov, M.; Yang, L. Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles. Nat. Commun. 2014, 5, 4462.
- Kaplan, J.B. Biofilm dispersal: Mechanisms, clinical implications, and potential therapeutic uses. J. Dent. Res. 2010, 89, 205–218.
- Fleming, D.; Rumbaugh, K.P. Approaches to dispersing medical biofilms. Microorganisms 2017, 5, 15.
- Kostakioti, M.; Hadjifrangiskou, M.; Hultgren, S.J. Bacterial biofilms: Development, dispersal, and therapeutic strategies in the dawn of the postantibiotic era. Cold Spring Harb. Perspect. Med. 2013, 3, a010306.
- Guilhen, C.; Forestier, C.; Balestrino, D. Biofilm dispersal: Multiple elaborate strategies for dissemination of bacteria with unique properties. Mol. Microbiol. 2017, 105, 188–210.
- Pirrone, M.; Pinciroli, R.; Berra, L. Microbiome, biofilms, and pneumonia in the ICU. Curr. Opin. Infect. Diseases 2016, 29, 160–166.
- Shrikant, P.; Chandrajit, L. Quorum sensing: An imperative longevity weapon in bacteria. Afr. J. Microbiol. Res. 2018, 12, 96–104.
- Tommonaro, G. Quorum Sensing: Molecular Mechanism and Biotechnological Application; Elsevier: Amsterdam, The Netherlands, 2019; pp. 3–22.
- Zhang, J.; Feng, T.; Wang, J.; Wang, Y.; Zhang, X. The mechanisms and applications of quorum sensing (QS) and quorum quenching (QQ). J. Ocean Univ. China 2019, 18, 1427–1442.
- Mangwani, N.; Dash, H.; Chauhan, A.; Das, S. Bacterial quorum sensing: Functional features and potential applications in biotechnology. J. Mol. Microbiol. Biotechnol. 2012, 22, 215–227.
- Terwagne, M.; Mirabella, A.; Lemaire, J.; Deschamps, C.; De Bolle, X.; Letesson, J. Quorum sensing and self-quorum quenching in the intracellular pathogen brucellamelitensis. PLoS ONE 2013, 8, e82514.
- Papenfort, K.; Bassler, B. Quorum sensing signal–response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 2016, 14, 576–588.
- Rémy, B.; Mion, S.; Plener, L.; Elias, M.; Chabrière, E.; Daudé, D. Interference in bacterial quorum sensing: A biopharmaceutical perspective. Front. Pharmacol. 2018, 9, 203.
- Moreno-Gámez, S.; Sorg, R.; Domenech, A.; Kjos, M.; Weissing, F.; van Doorn, G.; Veening, J. Quorum sensing integrates environmental cues, cell density and cell history to control bacterial competence. Nat. Commun. 2017, 8, 854.
- Ng, W.; Bassler, B. Bacterial quorum-sensing network architectures. Annu. Rev. Genet. 2009, 43, 197–222.
- Hence, B.; Schuster, M. Core principles of bacterial autoinducer systems. Microbiol. Mol. Biol. Rev. 2015, 79, 153–169.
- Rutherford, S.; Bassler, B. Bacterial quorum sensing: Its role in virulence and possibilities for its control. Cold Spring Harb. Perspect. Med. 2012, 2, a012427.
- Trajtenberg, F.; Albanesi, D.; Ruétalo, N.; Botti, H.; Mechaly, A.; Nieves, M.; Aguilar, P.; Cybulski, L.; Larrieux, N.; de Mendoza, D.; et al. Allosteric activation of bacterial response regulators: The role of the cognate histidine kinase beyond phosphorylation. mBio 2014, 5, e02105-14.
- Della Sala, G.; Teta, R.; Esposito, G.; Costantino, V. The chemical language of gram-negative bacteria. Quor. Sens. 2019, 3–28.
- Jacobi, C.; Grundler, S.; Hsieh, C.; Frick, J.; Adam, P.; Lamprecht, G.; Autenrieth, I.; Gregor, M.; Malfertheiner, P. Quorum sensing in the probiotic bacterium Escherichia coli Nissle 1917 (Mutaflor)—evidence that furanosyl borate diester (AI-2) is influencing the cytokine expression in the DSS colitis mouse model. Gut Pathog. 2012, 4, 8.
- Asfour, H. Anti-quorum sensing natural compounds. J. Microsc. Ultrastruct. 2018, 6, 1.
- Monnet, V.; Juillard, V.; Gardan, R. Peptide conversations in Gram-positive bacteria. Crit. Rev. Microbiol. 2016, 42, 339–351.
- Schuster, M.; Joseph Sexton, D.; Diggle, S.; Peter Greenberg, E. Acyl-homoserine lactone quorum sensing: From evolution to application. Annu. Rev. Microbiol. 2013, 67, 43–63.
- Fuqua, C.; Winans, S.; Greenberg, E. Census and consensus in bacterial ecosystems: The LuxR-LuxI family of quorum-sensing transcriptional regulators. Annu. Rev. Microbiol. 1996, 50, 727–751.
- Zhou, S.; Zhang, A.; Yin, H.; Chu, W. Bacillus sp. QSI-1 modulate quorum sensing signals reduce aeromonas hydrophila level and alter gut microbial community structure in fish. Front. Cell. Infect. Microbiol. 2016, 6, 184.
- Achari, G.; Ramesh, R. Characterization of bacteria degrading 3-hydroxy palmitic acid methyl ester (3OH-PAME), a quorum sensing molecule of Ralstonia solanacearum. Lett. Appl. Microbiol. 2015, 60, 447–455.
- Flemming, H.C.; Murthy, P.S.; Venkatesan, R.; Cooksey, K. Marine and Industrial Biofouling; Springer: Los Angeles, CA, USA, 2009.
- Plouguerné, E.; Hellio, C.; Deslandes, E.; Véron, B.; Stiger-Pouvreau, V. Anti-microfouling activities in extracts of two invasive algae: Grateloupia turuturu and Sargassum muticum. Bot. Mar. 2008, 51, 202–208.
- Flemming, H. Microbial biofouling: Unsolved problems, insufficient approaches, and possible solutions. In Springer Series on Biofilms; Springer: Berlin/Heidelberg, Germany, 2011; pp. 81–109.
- Cloete, E.; Molobela, I.; Van Der Merwe, A.; Richards, M. Biofilms in the food and beverage industries: An introduction. In Biofilms in the Food and Beverage Industries; Woodhead Publishing: Sawston, UK, 2009; pp. 3–41.
- Bixler, G.; Bhushan, B. Biofouling: Lessons from nature. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2012, 370, 2381–2417.
- Gu, J. Biofouling and Prevention. In Handbook of Environmental Degradation of Materials; Elsevier: Amsterdam, The Netherlands, 2012; pp. 243–282.
- Dobretsov, S.; Dahms, H.; Qian, P. Inhibition of biofouling by marine microorganisms and their metabolites. Biofouling 2006, 22, 43–54.
- Dobretsov, S. Inhibition and induction of marine biofouling by biofilms. In Springer Series on Biofilms; Springer: Berlin/Heidelberg, Germany, 2008; pp. 293–313.
- Bressy, C.; Lejars, M. Marine fouling: An overview. J. Ocean Technol. 2014, 9, 19–28.
- Daal, L.; de Vos, F.; Soons, J.; de Vries, T. Membrane technologies for water treatment and reuse in the power industries. In Advances in Membrane Technologies for Water Treatment; Woodhead Publishing: Sawston, UK, 2015; pp. 605–624.
- De Carvalho, C. Marine biofilms: A successful microbial strategy with economic implications. Front. Mar. Sci. 2018, 5, 5.
- Doble, M.; Venkatesan, R.; Vijaya Kumar, N.; Kumar, R. Polymers in a Marine Environment; Smithers Information: Shawbury, UK, 2014.
- Salta, M.; Chambers, L.; Wharton, J.; Wood, R.; Briand, J.F.; Blache, Y.; Stokes, K.R. Marine fouling organisms and their use in antifouling bioassays. In Proceedings of the EUROCORR, Nice, France, 6–9 September 2009.
- Lebret, K.; Thabard, M.; Hellio, C. Algae as marine fouling organisms: Adhesion damage and prevention. In Advances in Marine Antifouling Coatings and Technologies; Woodhead Publishing: Sawston, UK, 2009; pp. 80–112.
- Gordon, D.P.; Mawatari, S.F. Atlas of marine-fouling Bryozoa of New-Zealand ports and harbours. Misc. Publ. N. Z. Oceanogr. Inst. 1992, 107, 1–52.
- Baciocco, A.J. ‘Keynote address’. In Marine Biodeterioration: An Interdisciplinary Study; Costlow, J.D., Tipper, R.C., Eds.; Naval Institute Press: Annapolis, MD, USA, 1984; pp. 9–21.
- Munk, T.; Kane, D.; Yebra, D. The effects of corrosion and fouling on the performance of ocean-going vessels: A naval architectural perspective. In Advances in Marine Antifouling Coatings and Technologies; Woodhead Publishing: Sawston, UK, 2009; pp. 148–176.
- Moreira, J.; Gomes, L.; Simões, M.; Melo, L.; Mergulhão, F. The impact of material properties, nutrient load and shear stress on biofouling in food industries. Food Bioprod. Process. 2015, 95, 228–236.
- Verran, J. Biofouling in food processing: Biofilm or bio transfer potential? Food Bioprod. Process. 2002, 80, 292–298.
- Gule, N.; Begum, N.; Klumperman, B. Advances in biofouling mitigation: A review. Crit. Rev. Environ. Sci. Technol. 2016, 46, 535–555.
- Mizan, M.; Jahid, I.; Ha, S. Microbial biofilms in seafood: A food-hygiene challenge. Food Microbiol. 2015, 49, 41–55.
- Galié, S.; García-Gutiérrez, C.; Miguélez, E.; Villar, C.; Lombó, F. Biofilms in the food industry: Health aspects and control methods. Front. Microbiol. 2018, 9, 898.
- Bridier, A.; Sanchez-Vizuete, P.; Guilbaud, M.; Piard, J.; Naïtali, M.; Briandet, R. Biofilm-associated persistence of food-borne pathogens. Food Microbiol. 2015, 45, 167–178.
- Srey, S.; Jahid, I.; Ha, S. Biofilm formation in food industries: A food safety concern. Food Control 2013, 31, 572–585.
- Umaraw, P.; Prajapati, A.; Verma, A.; Pathak, V.; Singh, V. Control of campylobacter in poultry industry from farm to poultry processing unit: A review. Crit. Rev. Food Sci. Nutr. 2015, 57, 659–665.
- Wang, H.; Ding, S.; Dong, Y.; Ye, K.; Xu, X.; Zhou, G. Biofilm formation of salmonella serotypes in simulated meat processing environments and its relationship to cell characteristics. J. Food Prot. 2013, 76, 1784–1789.
- Rajkowski, K. Biofilms in fish processing. In Biofilms in the Food and Beverage Industries; Woodhead Publishing: Sawston, UK, 2009; pp. 499–516.
- Hickey, C.; Sheehan, J.; Wilkinson, M.; Auty, M. Growth and location of bacterial colonies within dairy foods using microscopy techniques: A review. Front. Microbiol. 2015, 6, 99.
- Jessen, B.; Lammert, L. Biofilm and disinfection in meat processing plants. Int. Biodeterior. Biodegrad. 2003, 51, 265–269.
- Marotta, F.; Garofolo, G.; Di Donato, G.; Aprea, G.; Platone, I.; Cianciavicchia, S.; Alessiani, A.; Di Giannatale, E. Population diversity of campylobacter jejuni poultry and its dynamic of contamination in chicken meat. BioMed Res. Int. 2015, 2015, 859845.
- Ehling-Schulz, M.; Frenzel, E.; Gohar, M. Food–bacteria interplay: Pathometabolism of emetic Bacillus cereus. Front. Microbiol. 2015, 6, 704.
- Gamazo, C.; Solano, C.; Lasa, I. Biofilm formation by Salmonella in food processing environments. In Biofilms in the Food and Beverage Industries; Woodhead Publishing: Sawston, UK, 2009; pp. 226–249.
- Linscott, A. Food-borne illnesses. Clin. Microbiol. Newsl. 2011, 33, 41–45.
- Giaouris, E.; Heir, E.; Hébraud, M.; Chorianopoulos, N.; Langsrud, S.; Møretrø, T.; Habimana, O.; Desvaux, M.; Renier, S.; Nychas, G. Attachment and biofilm formation by foodborne bacteria in meat processing environments: Causes, implications, role of bacterial interactions and control by alternative novel methods. Meat Sci. 2014, 97, 298–309.
- Pozo, J.L.D. Biofilm-related disease. Expert Rev. Anti-Infect. Ther. 2017, 16, 51–65.
- Darouiche, R. Treatment of infections associated with surgical implants. N. Engl. J. Med. 2004, 350, 1422–1429.
- LoVetri, K.; Gawande, P.V.; Yakandawala, N.; Madhyastha, S. Biofouling and anti-fouling of medical devices. In Biofouling: Types, Impact and Anti-Fouling; Nova Science Pub Inc.: Hauppauge, NY, USA, 2010; pp. 105–128.
- Trinidad, A.; Ibanez, A.; Gomez, D.; Garcia-Berrocal, J.R.; Ramirez-Camacho, R. Application of environmental scanning electron microscopy for study of biofilms in medical devices. Microsc. Sci. Technol. Appl. Educ. 2010, 1, 204–210.
- Mukherji, R.; Patil, A.; Prabhune, A. Role of extracellular proteases in biofilm disruption of gram positive bacteria with special emphasis on Staphylococcus aureus biofilms. Enz. Eng. 2015, 4.
- Harding, J.; Reynolds, M. Combating medical device fouling. Trends Biotechnol. 2014, 32, 140–146.
- Williams, C.; Ramage, G. Fungal Biofilms in Human Disease. Biofilm-Based Healthcare-Associated Infections, 2nd ed.; Springer: Cham, Switzerland, 2015.
- Strindberg, L.Z. The dependence of the results of pulp therapy on certain factors. Acta Odontol. Scand. 1956, 14, 1–75.
- Torabinejad, M.; Walton, R.E.; Fouad, A.F. Endodontics: Principles and Practice, 5th ed.; Elsevier: Beijing, China, 2015.
- Williams, C.; Rajendran, R.; Ramage, G. Aspergillus Biofilms in Human Disease. Fungal Biofilms and Related Infections, 3rd ed.; Springer: Cham, Switzerland, 2016.
- Toretto, S.; Pignataro, L. The Role of Biofilms in Upper Respiratory Tract Infections. Infections of the Ears, Nose, Throat, and Sinuses; Springer: Cham, Switzerland, 2018.
- Schurmann, M.; Oppel, F.; Gottaschalk, M.; Buker, B.; Jantos, C.A.; Knabbe, C.; Hutten, A.; Kaltschmidt, B.; Kaltschmidt, C.; Sudhoff, H. The therapeutic effect of 1, 8-cineol on pathogenic bacteria species present in chronic rhinosinusitis. Front. Microbiol. 2019, 10, 1–12.
- Brussow, H. Pseudomonas biofilms, cystic fibrosis, and phage: A silver lining? mBio 2012, 3, 1–2.
- McDaniel, C.T.; Panmanee, W.; Hassett, D.J. An Overview of Infections in Cystic Fibrosis Airways and the Role of Environmental Conditions on Pseudomonas aeruginosa Biofilm Formation and Viability. Cystic Fibrosis in the Light of New Research; Intech Open: Rijeka, Croatia, 2015.
- Romling, U.; Fiedler, B.; Bosshammer, J.; Grothues, D.; Greipel, J.; von der Hardt, H.; Tummler, B. Epidemiology of chronic pseudomonas aeruginosa infections in cystic fibrosis. J. Infect. Dis. 1994, 170, 1616–1621.
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322.