NCAM2 governs neuronal morphogenesis and axodendritic architecture, and controls important neuron-specific processes such as neuronal differentiation, synaptogenesis and memory formation. In the adult brain, NCAM2 is highly expressed in dendritic spines, and it regulates synaptic plasticity and learning processes. NCAM2’s functions are related to its ability to adapt to the external inputs of the cell and to modify the cytoskeleton accordingly. Different studies show that NCAM2 interacts with proteins involved in cytoskeleton stability and proteins that regulate calcium influx, which could also modify the cytoskeleton.
1. Introduction
The mammalian NCAM family has two members: NCAM1 and NCAM2
[1][2]. The two proteins have similar ectodomains, each of them containing five immunoglobulin domains and two fibronectin type III domains
[3],
Figure 1. NCAM1 was the first cell adhesion molecule identified in the nervous system and has been extensively studied. The different isoforms of NCAM1 (180KDa, 140KDa and 120KDa) have the same extracellular domain, but different transmembrane and cytosolic domains as a result of alternative splicing of the gene. The NCAM180 isoform is mostly expressed in neurons in the adult brain; NCAM140, in neurons in both embryonic and adult stages; and NCAM120, mostly in glia in adult stages.
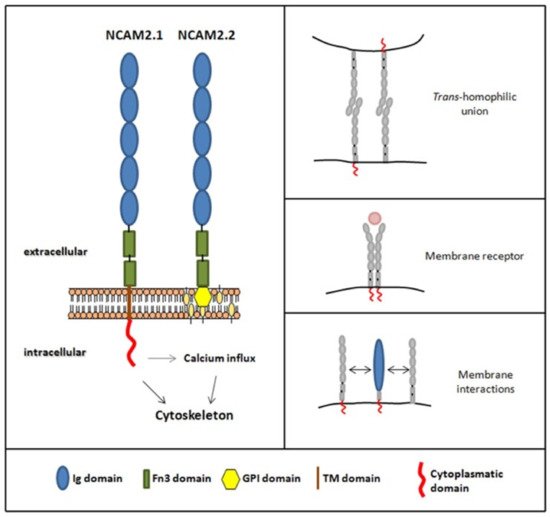
Figure 1. Schematic representation of NCAM2 protein and isoforms. Both isoforms have an extracellular part with five immunoglobulin domains (blue) and two fibronectin type III domains (green). NCAM2.1 isoform bears a transmembrane domain (brown) and a cytoplasmic domain (red), whereas NCAM2.2 isoform is glycosylphosphatidylinositol, GPI, anchored (yellow). It has been described that NCAM2 protein could do different type of interactions, such as trans-homophilic unions, membrane interactions and could also act as a membrane receptor.
In contrast to the wealth of knowledge currently available for NCAM1, NCAM2 has been less extensively studied and therefore its functions are still poorly understood, relegating this member of the protein family to a dark area for many years. However, over the last few years, a number of studies have revealed the importance of NCAM2 in neuronal morphogenesis, synapse formation and synaptic plasticity, with some of these functions being related to calcium dynamics and the ability of NCAM2 to modulate the cytoskeleton.
2. NCAM2 Expression and Interactors
The NCAM2 gene is believed to have been originated by genetic duplication of
ncam1, as they are paralog genes
[1][2]. In humans, the NCAM2 gene is located on chromosome 21 (region 21q21.1) and contains 25 exons. In mice, it is located on chromosome 16 (16 C3.3) and contains 19 exons. In both species, and due to an alternative splicing process, two isoforms are transcribed:
ncam2.1 and
ncam2.2. Compared to the mRNA of the
ncam2.1 isoform, which contains a stability element localized to the 3’-UTR region
[4], the mRNA of the
ncam2.2 isoform is more unstable, there is less expression and it is more temporarily restricted
[5]. In addition to the isoform differences at the transcript level, the two resulting proteins are also different: the NCAM2.1 isoform bears transmembrane and cytoplasmic domains, whereas the shorter NCAM2.2 isoform is glycosylphosphatidylinositol, GPI, anchored
[4][5],
Figure 1.
NCAM2 is mostly expressed in numerous central nervous system areas, including the cerebral cortex, the hippocampus and the olfactory system. Several studies showed expression of the murine NCAM2 gene in cortical areas, from embryonic day (E) 14 onwards, with the protein localizing to both dendritic and fiber compartments
[4][6]. At adult stages, there is a switch in NCAM2 cell localization that results in higher levels of the protein being present in synaptic contacts
[7][8].
The NCAM2 protein undergoes different post-translational modifications including palmitoylation, phosphorylation, proteolysis and glycosylation. Firstly, the cytosolic domain of NCAM2.1 contains four cysteine residues that may undergo palmitoylation: C723, C729, C734 and C740
[9]. Palmitoylation is an important modification for proteins to cluster into lipid raft domains
[10][11][12]. Secondly, three phosphorylated residues in the cytoplasmatic tail have been described in murine samples from developing brains: S765, T780, and S786
[13] and the cytosolic tail of NCAM2 is a PEST region—that is, a peptide sequence that is rich in phosphorylatable residues: proline (P), glutamic acid (E), serine (S), and threonine (T)
[14]. Besides, the cytosolic tail of NCAM2.1 presents SH2 and SH3 domains. Many proteins with these domains are involved in cell signaling through the activation of tyrosine kinases, such us Tyrosine-protein kinase Fyn or focal adhesion kinase, FAK. Thirdly, both NCAM2 isoforms present a proteolytic site between amino acids 682 and 701; proteolysis at this site causes the extracellular fragment to be released
[15][16]. Fourthly, the extracellular region has eight N-glycosylation sites which are HNK-1 sequences. It has been reported that HNK-1 sequences and glycosylation present in L1CAM and myelin-associated glycoprotein, MAG, could facilitate migration of neural-crest-derived cells
[17]. Noteworthy, and despite the high structural similarity of NCAM2 and NCAM1, polysialylation modifications have not been reported in NCAM2 proteins.
Few studies have addressed to NCAM2 interactions; these are summarized in
Table 1. NCAM2 establishes homophilic bonds through its first immunoglobulin domain. Such bonds facilitate interactions in
cis, which lead to dimerization of the protein, and in
trans, which facilitate binding between different neurons
[18]. Various studies have described interactions of the extracellular part of NCAM2 with prion protein (Prp), beta-amyloid peptide, fibroblast growth factor receptor (FGFR), epidermal growth factor (EGFR), Beta-site APP cleaving enzyme 1 (BACE1), Nogo and granulin (GRN)
[9][18][19][20][21][22]. Other studies have shown interactions of the cytosolic tail of NCAM2.1 with members of the 14-3-3 protein family; Proto-oncogene tyrosine-protein kinase, c-Src; microtubule associated proteins, MAPs; neurofilaments; NFs, Calcium/calmodulin-dependent protein kinase type II, CaMKII; and F-actin-capping protein, CAPZ
[8][9][16][18][21][22][23],
Figure 1.
Table 1. NCAM2 interactions and post-translational modifications.
| Extracellular Region |
| Interaction |
References |
| Fibroblast Growth Factor Receptor (FGFR) |
[20] |
| Epidermal Growth Factor Receptor (EGFR) |
[19] |
| Nogo |
[9] |
| Granulin |
[9] |
| Prion protein (Prp) |
[21][22] |
| Beta-site APP cleaving enzyme 1 (BACE1) |
[15] |
| Post-translational modification |
|
| N-glycosylation |
[18] |
| Proteolytic cleavage |
[15][16] |
| Intracellular region |
| Interaction |
References |
| Proto-oncogene tyrosine-protein kinase Src |
[23] |
| Calcium/calmodulin-dependent protein kinase type II (CaMKII) |
[8][23] |
| Microtubule-associated protein 2 (MAP2) |
[8] |
| Actin |
[9] |
| Tubulin |
[8] |
| 14-3-3 family proteins |
[8][21][22] |
| Microtubule-associated protein 1B (MAP1B) |
[9] |
| F-actin-capping complex (CAPZ) |
[9] |
| Heat shock cognate 71 protein (HSC70) |
[9] |
| Postranslational modification |
|
| Palmitoylation |
[9][14] |
| Phosphorylation |
[13] |
3. NCAM2 in Neuronal Cell Fate Determination and Differentiation
During neuronal development, the differentiation of a neuron from a postmitotic cell encompasses different processes [24]. Imbalances in the homeostasis of the cytoskeleton dynamics of the postmitotic cell give rise to neuronal polarization, which results in the formation of dendrites and an axonal terminal [25][26][27]. This polarization process is intrinsic to the differentiating neuron but modulated through external signals. Different CAMs are involved in neuronal polarization, act as transducers with the outside, and are key in the overall neuronal development and differentiation processes both in vivo and in vitro [28][29]. Studies with NCAM2 revealed that this protein is essential in the process of neuronal differentiation [8][23]. The distribution of NCAM2 inside the neuron and its interactions with proteins that regulate the cytoskeleton dynamics are key to NCAM2 participating in the establishment of both the dendritic tree and the axonal process.
3.1. NCAM2 Role in Neuronal Migration and Corticogenesis
Brain formation involves a myriad of mechanisms, including temporal and spatial regulations during neuronal development that only occur in vivo
[28]. The study of the mechanisms that regulate the polarization and development of neurons in vitro made it possible to identify proteins that have an implication at the physiological level
[30]. Not only that, but the complexity of in vivo models has demonstrated the relevance of the extracellular matrix and cell adhesion molecules in the overall development of the brain. In this sense, adhesion proteins participate in different processes of neuronal development, such as neuronal migration, positioning, morphogenesis, development of dendritic and axonal compartments, and synaptogenesis
[31][29][32][33]. In vivo, NCAM2 expression progressively increases during neuronal development; NCAM2 is involved in neuronal migration, cell positioning during corticogenesis, neuronal differentiation and synaptogenesis in the olfactory bulb, neocortex and hippocampus. In these regions, NCAM2 participates in the development of dendrites and axons, the establishment of connections, and the regulation of synaptic plasticity
[5][7][34], see
Figure 2.
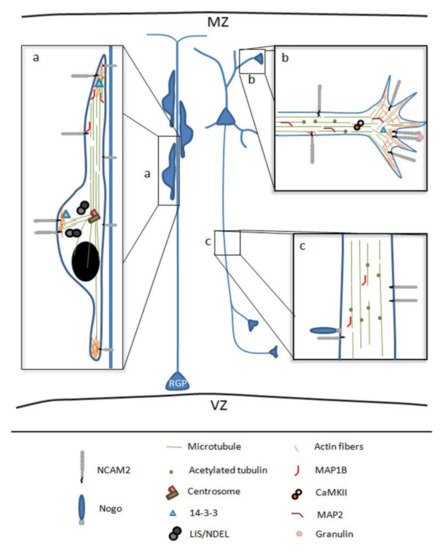
Figure 2. Schematic representation of NCAM2 functions and interactions during brain development. (a) NCAM2 is involved in neuronal migration, NCAM2 protein levels have been associated with an aberrant neuronal migration and layering positioning in the cortex. NCAM2 interacts with different cytoskeletal associated proteins which control cytoskeleton dynamics during neuronal migration. (b,c) NCAM2 has a crucial role during neuronal differentiation interacting with different proteins that are necessary for dendritic tree development and axon elongation. NCAM2 depletion alters the growth cone mobility and microtubules stability. Microtubule associated protein 1B, MAP1B; Microtubule associated protein 2, MAP2; and Calcium/calmodulin-dependent protein kinase type II, CaMKII. Lissencephaly-1/Nuclear distribution protein nudE-like1, LIS/NDEL; radial glia progenitors, RGP; marginal zone, MZ; and ventricular zone, VZ.
3.2. Neuronal Differentiation
NCAM2 is expressed in dendritic and axonal compartments and several studies showed its crucial role in neurite outgrowth, dendrite development and axon elongation.
3.2.1. NCAM2 in Dendritic Tree Development
The activation of NCAM2 produces an increase in the number of filopodia and the length of neurites due to an increase in the intracellular calcium levels, which activate the CaMKII complex
[23]. Conversely, the knockdown of
ncam2 expression during dendrites’ formation causes the retraction of existing neurites and the appearance of new processes from the soma
[8]. These phenomena produce a significant increase in the number of primary dendrites; however, these dendrites are shorter, have more branching points, and (at a qualitative level) they are thinner compared to the control situation.
3.2.2. NCAM2 in Axon Formation and Development
Different mechanisms are involved in the branching and elongation of the axonal terminal, some of which required a certain level of cytoskeleton stability
[35]. In this sense, excessive instability of the microtubule cytoskeleton results in a higher number of branching points
[36]. NCAM2 deficiency increases the number of branches and reduces the maximum length of the main axon, without altering the total length of the axonal shaft
[8]. Moreover, 20% of NCAM2-depleted neurons exhibit two or more axons. At the cytoskeleton level, NCAM2 deficiency reduces the signal of acetylated tubulin in axons. Tubulin acetylation is a posttranslational modification that occurs only when microtubules are polymerized and thus is an indirect measure of microtubule cytoskeleton stability
[37]. Regarding axon elongation, NCAM2 interacts with MAP1B, which binds to the microtubules and regulates their dynamics. The regulation of MAP1B binding to microtubules is crucial during the axon branching process and changes according to the degree of phosphorylation
[38]. In addition, NCAM2 interactions with the light- and medium subunits of neurofilaments have been identified. During development and neuritogenesis, these two subunits form heterodimers and participate in the cytoskeleton dynamics required for axonal growth and transport
[39]. Through direct interaction with these neurofilament subunits, with MAP1B, or through signaling pathways that modulate said stability, NCAM2 has a role in stabilizing microtubule cytoskeleton to ensure a proper axon development in terms of length and branching points.
3.2.3. NCAM2 in Synaptogenesis and Synaptic Plasticity
During brain development, neuronal polarization and synaptogenesis occur in parallel. The process of stabilization of synaptic contacts is dependent on cytoskeleton rearrangement and neuronal activity. Synaptogenesis is highly regulated by cell adhesion molecules, which control the recognition of membranes, allow the formation of synaptic structures and modulate the neuronal maturation through calcium signaling
[40][41][42][43].
In vivo, NCAM2 is detected in presynaptic and postsynaptic compartments and presents trans-homophilic binding between both compartments
[7]—see
Figure 3.
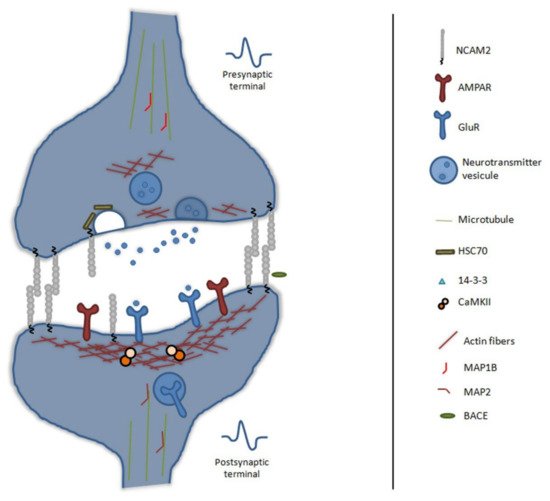
Figure 3. Schematic representation of NCAM2 functions and interactions in synapse maintenance and plasticity. NCAM2 is involved in synapses formation, it is detected in the presynaptic and postsynaptic compartments and undergoes trans-homophilic binding between both compartments. NCAM2 interacts with different scaffold proteins or complexes, which control the shape and dynamics of synapses; such as CaMKII, Actin, 14-3-3 or CAPZ. NCAM2 is involved in synaptic transmission through the neurotransmitters vesicles recycling pathway and the amount of glutamate receptors. Proteolytic cleavage of NCAM2 by BACE1 is important for synapse plasticity and remodeling. Beta-site APP cleaving enzyme 1, BACE1; Microtubule associated protein 1B, MAP1B; Microtubule associated protein 2, MAP2; Calcium/calmodulin-dependent protein kinase type II, CaMKII; F-actin-capping protein, CAPZ; α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptor, AMPAR; glutamate receptor, GluR and Heat shock cognate 71 kDa, HSC70.
In the adult brain, synaptic plasticity is crucial for learning processes and memory tasks, and it causes a restructuration of presynaptic and postsynaptic compartments. In these processes, cell adhesion molecules interact with neurotransmitter receptors and transduce the synaptic inputs to the cytoskeleton structure. NCAM2 is located at the excitatory synapses and is important for glutamate receptor positioning
[16]. The disruption of NCAM2 functions at the cell surface results in the disassembly of glutamatergic synapses and reduces the amount of actin filaments in the spine
[16]. Dendritic spines have an actin cytoskeleton core, which plays a key role in maintaining the shape and plasticity of the spine
[44][45]. NCAM2 interacts with actin, CAPZ complex and 14-3-3.
3.3. NCAM2 in Calcium Signaling and Homeostasis
Relevant functions of CAMs members regarding neuronal development or synaptic plasticity are vehiculated through the calcium signaling
[40]. The activation of different members of CAMs induces changes in calcium concentration that are transduced by different proteins to operate in survival mechanisms, modulate cytoskeleton dynamics and activate gene transcription
[40]. In the case of NCAM2, there are some studies that analyze the relation between the protein and the calcium signaling during neurite growth and synapse formation
[23][46]. In detail, it has been shown that the activation of NCAM2 increases calcium concentration through Scr and VDCC channels. More studies are required to explore other mechanisms, for example, the possibility that NCAM2 increases calcium concentration through FGFR or endoplasmic reticulum (ER) as there is evidence that some NCAM2 partners also interact with the ER. Moreover, the interaction of NCAM2 with FGFR has been previously observed
[40] and it is known that the interaction of different CAMs with FGFR could increase intracellular calcium concentration, promoting neurite outgrowth and dendritic development.
4. NCAM2 in Neuronal Diseases
NCAM2 is expressed during brain development as well as in adulthood and its functions are crucial for the proper cognitive process. Genetic and molecular studies show that changes in NCAM2 expression could be the cause of different pathologies both during brain development and in the adult stages—see Table 2.
Table 2. NCAM2 implications in neurodevelopmental disorders and neurodegenerative diseases.
| Neurodevelopmental Disorder |
| Disorder |
Type of Study |
Implications |
References |
| Autism Spectrum Disorders |
Genetic in humans |
Genetic studies associate alterations and deletions in Ncam2 with ASD. |
[47][48][49] |
| Down Syndrome |
Genetic in humans |
Increased expression in DS patients due to the location of Ncam2 in the 21 chromosome. |
[6][50] |
| Other neurodevelopmental disorders |
Genetic in humans |
Deletions Ncam2 are found in patients with neurodevelpmental disorders. |
[51] |
| Neurodegenerative diseases |
| Disorder |
|
Implications |
References |
| Alzheimer’s Disease |
Genetic in humans |
Alterations in Ncam2 found in AD patients. |
[52][53] |
| |
Experimental with human and mouse samples. |
β-amyloid induces proteolysis of synaptic NCAM2. |
[16] |
| Frontotemporal dementia |
Experimental with mouse tissue samples |
NCAM2 proposed as a candidate receptor for GRN |
[9] |
4.1. Neurodevelopment Diseases
NCAM2 is necessary both for the correct neuronal migration and for the appropriate development of dendritic trees and axonal compartments in cortical neurons and olfactory bulb neurons. Upon NCAM2 knockdown, cortical neurons lacked a clear main apical dendrite and showed numerous primary dendrites arising from any position in the cell body
[8]. This phenotype could explain the implication of NCAM2 in neurodevelopment disorders. Alterations in NCAM2 have been associated with different disorders, such as Down Syndrome or Autism Spectrum Disorders
[47][6][48][49][50]. Due to its location in human chromosome 21, NCAM2 has been proposed as a candidate for the intellectual disability phenotype observed in Down Syndrome
[6][50]. Even though
Ncam2 is located outside the critical region of chromosome 21, it is believed that an increased expression of NCAM2 could negatively affect the nervous system development due to dosage-related effects. In addition, genetic analyses revealed single nucleotide polymorphisms in NCAM2 gene in patients with abnormal neurodevelopment
[51], Marden–Walker syndrome patients
[54] and Autism Spectrum Disorders
[47][48][49]. Not only does NCAM2 regulate neuronal polarization and formation of axo-dendritic compartments, but it also interacts with different cytoskeleton components that are linked to autism disorder
[8][9][55][56][57]. Genetic variations in the NCAM2 gene are less known but could be the cause of intellectual disability. Therefore, one hypothesis is that structural alterations observed in patients with aberrant neurodevelopment and autism could be triggered by cytoskeleton dynamics modifications produced by variations in NCAM2 gene expression.
4.2. Neurodegenerative Diseases
Changes in the density and structure of the spines have been associated with the early stages of neurodegenerative diseases. At a molecular level, changes in the expression and cellular localization of adhesion proteins have been described in these diseases. Genetic studies established a relation between NCAM2 and Alzheimer’s Disease
[52][53]. It has been proposed that the early loss of synapses detected in patients with Alzheimer’s Disease could be the result of the β-amyloid-induced proteolysis of synaptic NCAM2. Due to the pivotal role of NCAM2 in synaptic structures and the mentioned interactions of NCAM2 with the cytoskeleton, the ablation of NCAM2 in synapses could lead to major changes in cytoskeleton structures, thus compromising synaptic viability in Alzheimer’s Disease
[16][58][59][60][61]. The interaction of NCAM2 with Granulin, GRN, one of the principal proteins associated with familial frontotemporal dementia, suggest a plausible role of NCAM2 as a receptor of the protein and a possible implication in frontotemporal dementia
[9][62]. A decrease in NCAM2 levels and a change in NCAM2 localization in the spine could be associated with loss of structure of synapses in the early stages of neurodegenerative diseases.
5. Conclusions
In the development of the nervous system, NCAM2 regulates the molecular recognition that allows the formation of contacts between axonal and dendritic compartments leading to the formation of synapses and neural circuits. In particular, NCAM2 is present in both axonal and dendritic compartments and establishes homophilic junctions and intracellular interactions. These interactions regulate the modification of the cytoskeleton and affect the structure of the dendrite and axon. Different mechanisms are modulated by NCAM2 interactions but these can be divided in two groups: calcium signaling mechanisms and structuring of the cytoskeleton. NCAM2 is essential for the proper development of dendritic and axonal structures; its depletion causes aberrant neuron migration and differentiation and is detected in neurodevelopmental diseases. With these results, we can affirm that changes in NCAM2 alter the neuronal molecular identity, produces alterations in neuronal development and conduces to an aberrant neuronal network formation. Moreover, different CAMs modulate the neuronal differentiation, structure pruning and synaptic stabilization through the activation of gene expression. It is known that calcium influx and CaMKII modulate gene expression. More studies are necessary to identify the possible role of NCAM2 in gene expression (Figure 4).
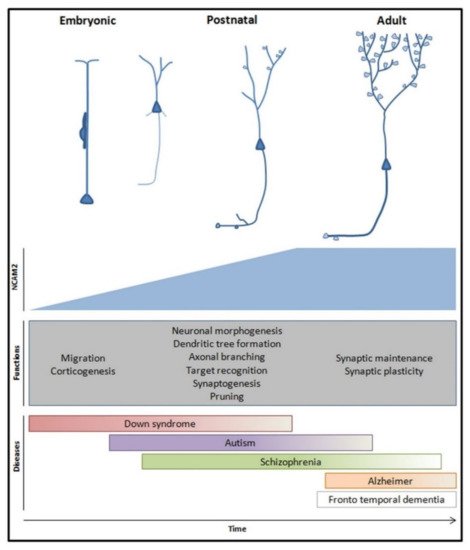
Figure 4. Global representation of NCAM2 physiological functions and pathological implications during brain development and in adult stages. NCAM2 expression increases during embryonic and early postnatal stages and it is maintained in adulthood. NCAM2 participates in the establishment of the neuronal molecular identity which is essential for an amount of processes during neuron development and neuronal network morphogenesis; e.g., neuronal migration, dendritic tree development, synaptogenesis and synaptic plasticity. These functions and genetic studies of neuronal pathologies showed that NCAM2 could be an important target for new therapeutic strategies.
In the adult brain, the neuronal network is less plastic, the dendrites and axons are stable structures and neuronal plasticity is restricted to synapses and adult neurogenesis. For synaptic plasticity to take place, neuronal molecular identity and a proper balance between stabilization of membranes and remodeling of contacts are necessary. NCAM2 is present in membranes in both synaptic compartments: presynaptic and postsynaptic. NCAM2 participates in plasticity processes through the diverse interactions in the membrane and with both presynaptic and postsynaptic scaffold structures. The amount of NCAM2 in the membrane could regulate the synaptic plasticity processes. NCAM2 controls the cytoskeleton structure and dynamics, which proves crucial for the maintenance of the synaptic scaffold and neuronal synapses. Moreover, NCAM2 could participate in local transcription, as it interacts with different transcription complexes. NCAM2 would be controlling the specific transcription of proteins in synapses, a necessary mechanism for synapse remodeling and potentiation. Besides, NCAM2 cleavage by metalloproteinases is another mechanism for synapse remodeling. Some processes controlled by NCAM2 are altered at the early stages of neurodegenerative diseases, such as Alzheimer’s disease. So, NCAM2 could represent a therapeutic target for a new strategy to preserve the synapse structure.