
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Vanessa Soto-Cerrato | + 3731 word(s) | 3731 | 2021-08-17 05:00:08 | | | |
| 2 | Vivi Li | Meta information modification | 3731 | 2021-09-08 08:32:13 | | | | |
| 3 | Conner Chen | Meta information modification | 3731 | 2021-09-22 03:09:39 | | |
Video Upload Options
Autophagy is a tightly regulated catabolic process that facilitates nutrient recycling from damaged organelles and other cellular components through lysosomal degradation. Deregulation of this process has been associated with the development of several pathophysiological processes, such as cancer and neurodegenerative diseases. In cancer, autophagy has opposing roles, being either cytoprotective or cytotoxic. Thus, deciphering the role of autophagy in each tumor context is crucial. Moreover, autophagy has been shown to contribute to chemoresistance in some patients. In this regard, autophagy modulation has recently emerged as a promising therapeutic strategy for the treatment and chemosensitization of tumors, and has already demonstrated positive clinical results in patients.
1. Introduction
2. Therapeutic Strategies Targeting Autophagy
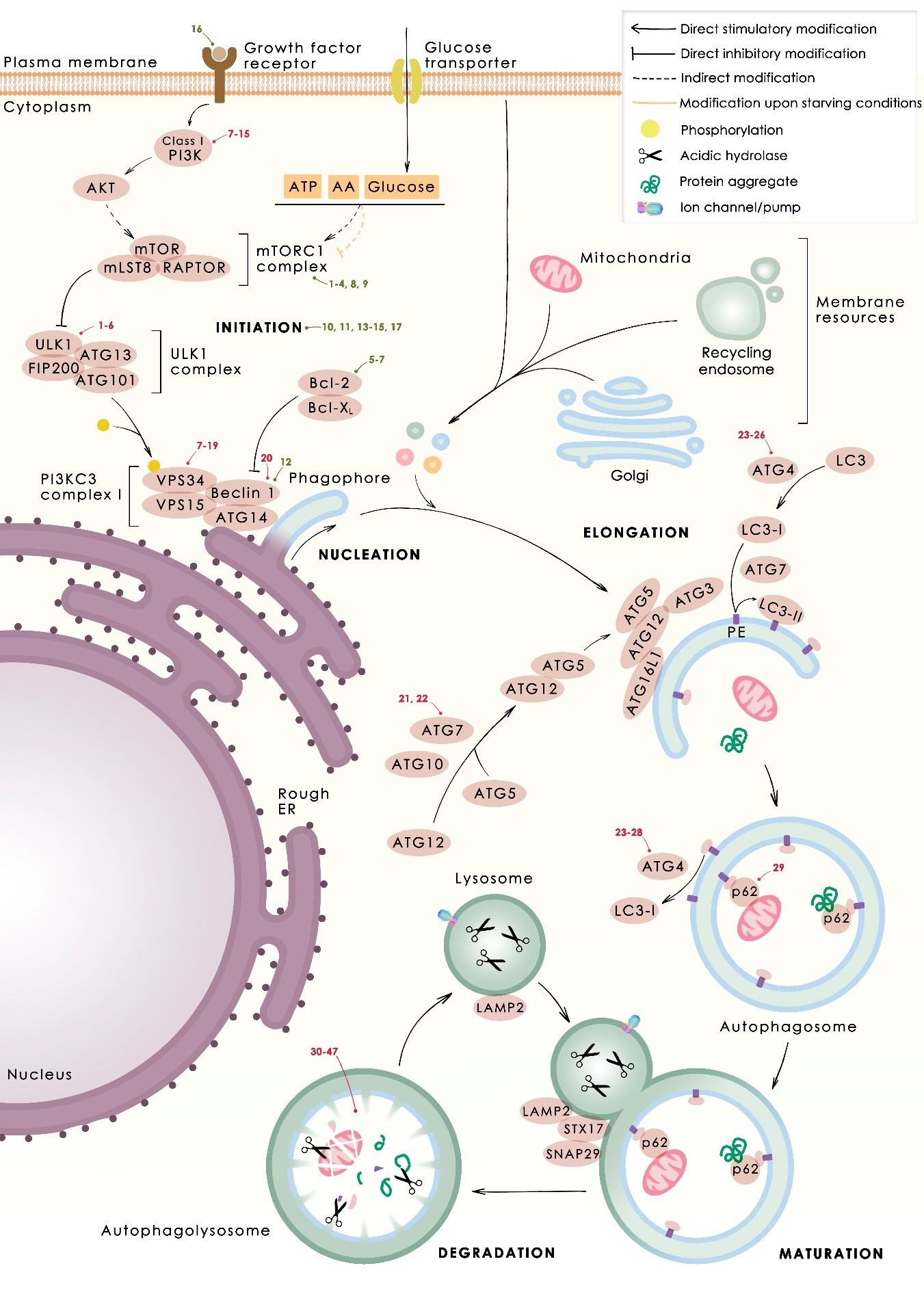
Figure 1. Mechanism of autophagy. The phases of the process of autophagy (nucleation, elongation, maturation and degradation), with the main proteins that participate in each one, are depicted.Autophagy activators (green) and inhibitors (red) are marked where they interfere with the autophagy process. Numbers correspond to those compounds listed in table 1 and 2, respectively.
2.1. Autophagy Stimulation for Cancer Treatment
| Mechanism of Action/Type | Name | Structure | Number in Figure 1 | Refs. |
|---|---|---|---|---|
| mTOR Inhibitors | Rapacmycin | 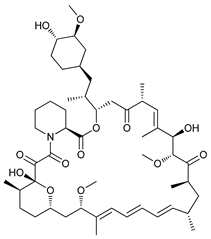 |
1 | [8][9][10][11] |
| Temsirolimus (CCI779) | 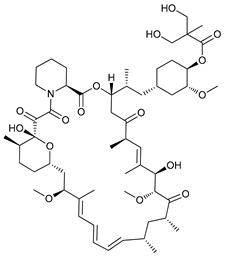 |
2 | [12][13] | |
| Everolimus (RAD001) | 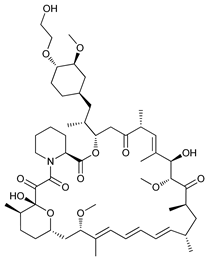 |
3 | [14][15] | |
| AZD8055 | 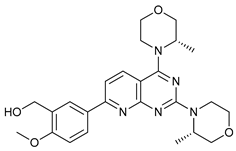 |
4 | [16][17] | |
| BH3 Mimetics | (-)-gossypol (AT-101) | 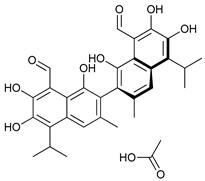 |
5 | [18][19][20][21] |
| Obatoclax (GX15-070) | 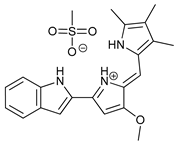 |
6 | [22][23][24] | |
| ABT-737 | 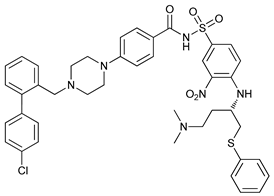 |
7 | [25] | |
| Cannabinoids | Δ9-Tetrahydrocannabinol (THC) | 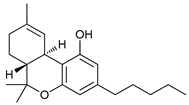 |
8 | [26][27][28] |
| JWH-015 | 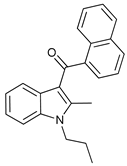 |
9 | [29] | |
| Histone Deacetylase Inhibitors | Suberoylanilide hydroxamic acid (SAHA, Vorinostat) | 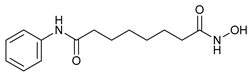 |
10 | [30] |
| MHY2256 | 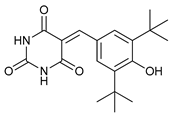 |
11 | [31] | |
| Natural Products | Betulinic acid | 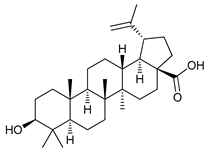 |
12 | [32] |
| Resveratrol | 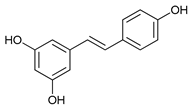 |
13 | [33] | |
| δ-Tocotrienol | 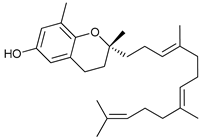 |
14 | [34] | |
| Curcumin | 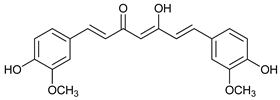 |
15 | [35] | |
| Others | Lapatinib | 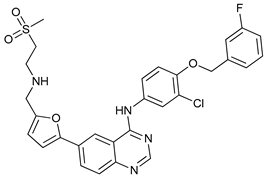 |
16 | [36][37] |
| APO866 | 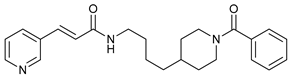 |
17 | [38] |
2.1.1. mTOR Inhibitors
2.1.2. BH3 Mimetics
2.1.3. Cannabinoids
2.1.4. Histone Deacetylase Inhibitors (HDACIs)
2.1.5. Natural Products
2.1.6. Others
2.2. Autophagy Inhibition for Cancer Treatment
| Mechanism of Action | Name | Structure | Number in Figure 1 | Refs. | |
|---|---|---|---|---|---|
| ULK Inhibitors | Compound 6 | 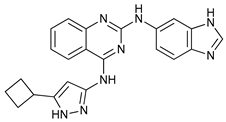 |
1 | [61] | |
| MRT68921 | 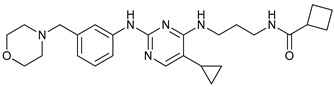 |
2 | [62][63] | ||
| MRT67307 | 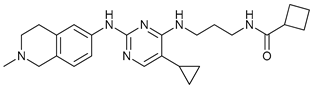 |
3 | [62][63] | ||
| SBI-0206965 | 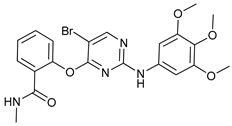 |
4 | [64][65][66][67] | ||
| ULK-100 | 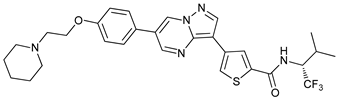 |
5 | [68] | ||
| ULK-101 | 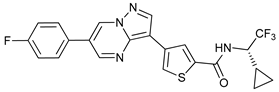 |
6 | [68] | ||
| Pan PI3k Inhibitors | 3MA | 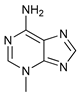 |
7 | [69][70][71] | |
| 3 MA derivatives | 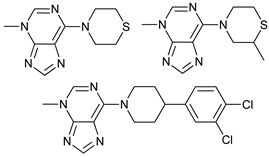 |
8 | [72] | ||
| Wortmannin | 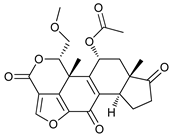 |
9 | [73][74] | ||
| LY294002 | 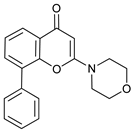 |
10 | [75] | ||
| SF1126 | 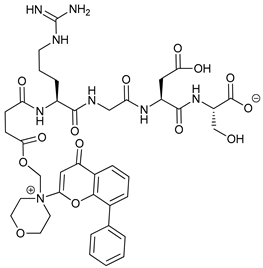 |
11 | [76][77] | ||
| PI103 | 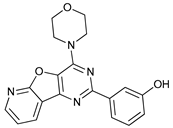 |
12 | [78] | ||
| KU55933 | 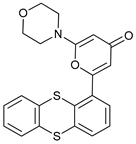 |
13 | [79] | ||
| Gö6976 | 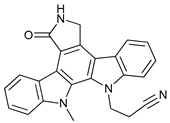 |
14 | [79] | ||
| GSK1059615 | 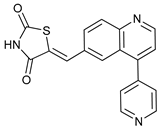 |
15 | [80][81] | ||
| VPS34 (PI3KC3) Inhibitors | SAR405 | 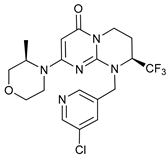 |
16 | [78] | |
| VPS34-IN1 | 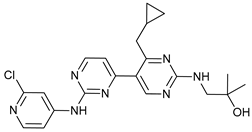 |
17 | [82] | ||
| PIK-III | 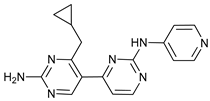 |
18 | [83] | ||
| Compound 31 | 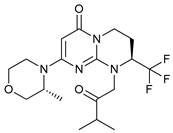 |
19 | [84] | ||
| Spautin-1 | 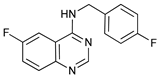 |
20 | [85] | ||
| ATG Inhibitors | ATG7 inhibitor | 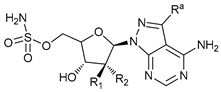 |
21 | WO2018/089786 | |
| ATG7 inhibitor, miR154 | UAGGUUAUCCGUGUUGCCUUCG | 22 | [86] | ||
| NSC185058 | 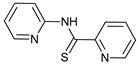 |
23 | [87] | ||
| Tioconazol | 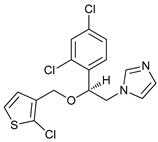 |
24 | [88] | ||
| UAMC-2526 | 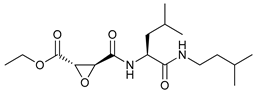 |
25 | [89] | ||
| LV320 | 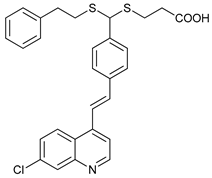 |
26 | [90] | ||
| S130 | 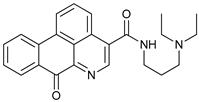 |
27 | [91] | ||
| FMK-9a | 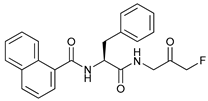 |
28 | [92][93][94] | ||
| Autophagy Formation | Verteporfin | 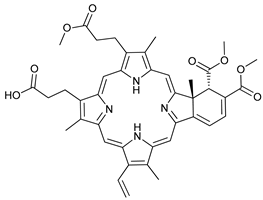 |
29 | [95][96][97][98] | |
| Lysosome Inhibitors | Lysosomotropic Agents | Chloroquine | 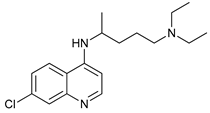 |
30 | [99][100] |
| Hydroxychloroquine | 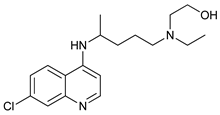 |
31 | [101] | ||
| Lys05 | 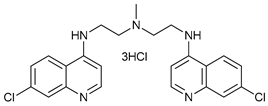 |
32 | [102][103] | ||
| DQ661 | 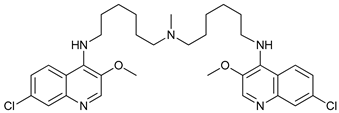 |
33 | [104] | ||
| VATG-027 | 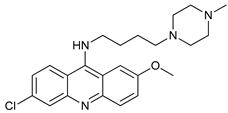 |
34 | [105] | ||
| Mefloquine | 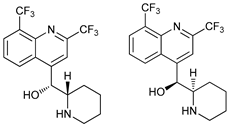 |
35 | [106][107] | ||
| Ganoderma lucidum polysaccharide (GLP) | 36 | [108][109][110] | |||
| Vacuolar H+ ATPase Inhibitors | Bafilomycin A1 | 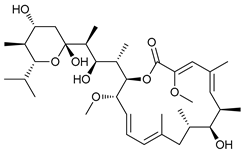 |
37 | [111][112][113] | |
| Ionophores | Tambjamines | 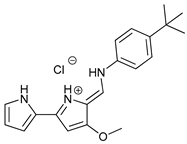 |
38 | [114] | |
| Monensin | 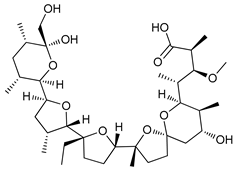 |
39 | [115] | ||
| Squaramides | 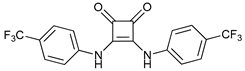 |
40 | [116] | ||
| Inhibition of Autophagosome-Lysosome Fusion | WX8 family | 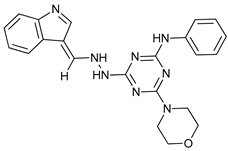 |
41 | [117] | |
| Vacuolin-1 | 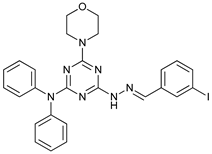 |
42 | [118] | ||
| Desmethylclomipramine | 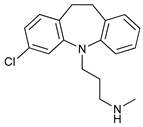 |
43 | [119] | ||
| Acid Protease Inhibitors | Pepstatin A | 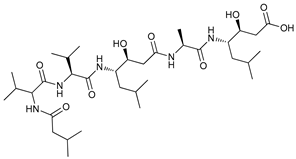 |
44 | [120] | |
| Leupeptin | 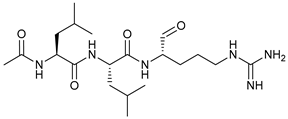 |
45 | [120] | ||
| E64d | 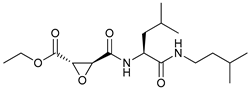 |
46 | [121] | ||
| Others | Nanoparticles | 47 | [122][123][124] | ||
2.2.1. ULK Inhibitors
2.2.2. Pan PI3K Inhibitors
2.2.3. VPS34 (PI3KC3) Complex Inhibitors
2.2.4. ATG inhibitors
2.2.5. Autophagosome Formation Inhibition
2.2.6. Lysosome Inhibitors
References
- Deter, R.L.; Baudhuin, P.; Duve, C. de Participation of lysosomes in cellular autophagy induced in rat liver by glucagon. J. Cell Biol. 1967, 35, 11–16.
- Mizushima, N. Autophagy: Process and function. Genes Dev. 2007, 21, 2861–2873.
- Mizushima, N.; Levine, B.; Cuervo, A.M.; Klionsky, D.J. Autophagy fights disease through cellular self-digestion. Nature 2008, 451, 1069–1075.
- Dikic, I.; Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 2018, 19, 349–364.
- Chude, C.I.; Amaravadi, R.K. Targeting autophagy in cancer: Update on clinical trials and novel inhibitors. Int. J. Mol. Sci. 2017, 18, 1279.
- U.S. National Library of Medicine of Clinical Trials. Available online: https://clinicaltrials.gov/ (accessed on 10 September 2019).
- White, E. The role for autophagy in cancer (White, 2015).pdf. J. Clin. Investig. 2015, 125, 42–46.
- Shi, H.; Zhang, L.; Zhang, C.; Hao, Y.; Zhao, X. Rapamycin may inhibit murine S180 sarcoma growth by regulating the pathways associated with autophagy and cancer stem cells. J. Cancer Res. Ther. 2019, 15, 398–403.
- Lin, X.; Han, L.; Weng, J.; Wang, K.; Chen, T. Rapamycin inhibits proliferation and induces autophagy in human neuroblastoma cells. Biosci. Rep. 2018, 38, 1–8.
- Jiang, R.Y.; Pei, H.L.; Gu, W.D.; Huang, J.; Wang, Z.G. Autophagic inhibitor attenuates rapamycin-induced inhibition of proliferation in cultured A549 lung cancer cells. Eur. Rev. Med. Pharmacol. Sci. 2014, 18, 806–810.
- Xie, Z.G.; Xie, Y.; Dong, Q.R. Inhibition of the mammalian target of rapamycin leads to autophagy activation and cell death of MG63 osteosarcoma cells. Oncol. Lett. 2013, 6, 1465–1469.
- Liu, W.; Huang, S.; Chen, Z.; Wang, H.; Wu, H.; Zhang, D. Temsirolimus, the mTOR inhibitor, induces autophagy in adenoid cystic carcinoma: In vitro and in vivo. Pathol. Res. Pract. 2014, 210, 764–769.
- Singla, M.; Bhattacharyya, S. Autophagy as a potential therapeutic target during epithelial to mesenchymal transition in renal cell carcinoma: An in vitro study. Biomed. Pharmacother. 2017, 94, 332–340.
- Chen, G.; Ding, X.-F.; Bouamar, H.; Pressley, K.; Sun, L.-Z. Everolimus induces G 1 cell cycle arrest through autophagy-mediated protein degradation of cyclin D1 in breast cancer cells. Am. J. Physiol. Physiol. 2019, 317, C244–C252.
- Lui, A.; New, J.; Ogony, J.; Thomas, S.; Lewis-Wambi, J. Everolimus downregulates estrogen receptor and induces autophagy in aromatase inhibitor-resistant breast cancer cells. BMC Cancer 2016, 16, 1–15.
- Chresta, C.M.; Davies, B.R.; Hickson, I.; Harding, T.; Cosulich, S.; Critchlow, S.E.; Vincent, J.P.; Ellston, R.; Jones, D.; Sini, P.; et al. AZD8055 is a potent, selective, and orally bioavailable ATP-competitive mammalian target of rapamycin kinase inhibitor with in vitro and in vivo antitumor activity. Cancer Res. 2010, 70, 288–298.
- Hu, M.; Huang, H.; Zhao, R.; Li, P.; Li, M.; Miao, H.; Chen, N.; Chen, M. AZD8055 induces cell death associated with autophagy and activation of AMPK in hepatocellular carcinoma. Oncol. Rep. 2014, 31, 649–656.
- Benvenuto, M.; Mattera, R.; Masuelli, L.; Taffera, G.; Andracchio, O.; Tresoldi, I.; Lido, P.; Giganti, M.G.; Godos, J.; Modesti, A.; et al. (±)-Gossypol induces apoptosis and autophagy in head and neck carcinoma cell lines and inhibits the growth of transplanted salivary gland cancer cells in BALB/c mice. Int. J. Food Sci. Nutr. 2017, 68, 298–312.
- Benvenuto, M.; Mattera, R.; Sticca, J.I.; Rossi, P.; Cipriani, C.; Giganti, M.G.; Volpi, A.; Modesti, A.; Masuelli, L.; Bei, R. Effect of the BH3 mimetic polyphenol (-)-Gossypol (AT-101) on the in vitro and in vivo growth of malignant mesothelioma. Front. Pharmacol. 2018, 9, 1–13.
- Lan, L.; Appelman, C.; Smith, A.R.; Yu, J.; Larsen, S.; Marquez, R.T.; Liu, H.; Wu, X.; Gao, P.; Roy, A.; et al. Natural product (-)-gossypol inhibits colon cancer cell growth by targeting RNA-binding protein Musashi-1. Mol. Oncol. 2015, 9, 1406–1420.
- Voss, V.; Senft, C.; Lang, V.; Ronellenfitsch, M.W.; Steinbach, J.P.; Seifert, V.; Kögel, D. The pan-Bcl-2 inhibitor (-)-gossypol triggers autophagic cell death in malignant glioma. Mol. Cancer Res. 2010, 8, 1002–1016.
- Sulkshane, P.; Teni, T. BH3 mimetic Obatoclax (GX15-070) mediates mitochondrial stress predominantly via MCL-1 inhibition and induces autophagy-dependent necroptosis in human oral cancer cells. Oncotarget 2017, 8, 60060–60079.
- Basit, F.; Cristofanon, S.; Fulda, S. Obatoclax (GX15-070) triggers necroptosis by promoting the assembly of the necrosome on autophagosomal membranes. Cell Death Differ. 2013, 20, 1161–1173.
- Bonapace, L.; Bornhauser, B.C.; Schmitz, M.; Cario, G.; Ziegler, U.; Niggli, F.K.; Schäfer, B.W.; Schrappe, M.; Stanulla, M.; Bourquin, J.P. Induction of autophagy-dependent necroptosis is required for childhood acute lymphoblastic leukemia cells to overcome glucocorticoid resistance. J. Clin. Investig. 2010, 120, 1310–1323.
- Yao, X.; Li, X.; Zhang, D.; Xie, Y.; Sun, B.; Li, H.; Sun, L.; Zhang, X. B-cell lymphoma 2 inhibitor ABT-737 induces Beclin1- and reactive oxygen species-dependent autophagy in Adriamycin-resistant human hepatocellular carcinoma cells. Tumor Biol. 2017, 39, 1–12.
- Armstrong, J.L.; Hill, D.S.; McKee, C.S.; Hernandez-Tiedra, S.; Lorente, M.; Lopez-Valero, I.; Anagnostou, M.E.; Babatunde, F.; Corazzari, M.; Redfern, C.P.F.; et al. Exploiting cannabinoid-induced cytotoxic autophagy to drive melanoma cell death. J. Investig. Dermatol. 2015, 135, 1629–1637.
- Hernández-Tiedra, S.; Fabriàs, G.; Dávila, D.; Salanueva, Í.J.; Casas, J.; Montes, L.R.; Antón, Z.; García-Taboada, E.; Salazar-Roa, M.; Lorente, M.; et al. Dihydroceramide accumulation mediates cytotoxic autophagy of cancer cells via autolysosome destabilization. Autophagy 2016, 12, 2213–2229.
- Salazar, M.; Carracedo, A.; Salanueva, Í.J.; Hernández-Tiedra, S.; Lorente, M.; Egia, A.; Vázquez, P.; Blázquez, C.; Torres, S.; García, S.; et al. Cannabinoid action induces autophagy-mediated cell death through stimulation of ER stress in human glioma cells. J. Clin. Investig. 2009, 119, 1359–1372.
- Vara, D.; Salazar, M.; Olea-Herrero, N.; Guzmán, M.; Velasco, G.; Díaz-Laviada, I. Anti-tumoral action of cannabinoids on hepatocellular carcinoma: Role of AMPK-dependent activation of autophagy. Cell Death Differ. 2011, 18, 1099–1111.
- Han, H.; Li, J.; Feng, X.; Zhou, H.; Guo, S.; Zhou, W. Autophagy-related genes are induced by histone deacetylase inhibitor suberoylanilide hydroxamic acid via the activation of cathepsin B in human breast cancer cells. Oncotarget 2017, 8, 53352–53365.
- De, U.; Son, J.Y.; Sachan, R.; Park, Y.J.; Kang, D.; Yoon, K.; Lee, B.M.; Kim, I.S.; Moon, H.R.; Kim, H.S. A new synthetic histone deacetylase inhibitor, MHY2256, induces apoptosis and autophagy cell death in endometrial cancer cells via p53 acetylation. Int. J. Mol. Sci. 2018, 19, 2743.
- Zhou, H.; Luo, W.; Zeng, C.; Zhang, Y.; Wang, L.; Yao, W.; Nie, C. PP2A mediates apoptosis or autophagic cell death in multiple myeloma cell lines. Oncotarget 2017, 8, 80770–80789.
- Fu, Y.; Chang, H.; Peng, X.; Bai, Q.; Yi, L.; Zhou, Y.; Zhu, J.; Mi, M. Resveratrol inhibits breast cancer stem-like cells and induces autophagy via suppressing Wnt/β-catenin signaling pathway. PLoS ONE 2014, 9, e102535.
- Fontana, F.; Moretti, R.M.; Raimondi, M.; Marzagalli, M.; Beretta, G.; Procacci, P.; Sartori, P.; Montagnani Marelli, M.; Limonta, P. δ-Tocotrienol induces apoptosis, involving endoplasmic reticulum stress and autophagy, and paraptosis in prostate cancer cells. Cell Prolif. 2019, 52, 1–15.
- Deng, Q.; Liang, L.; Liu, Q.; Duan, W.; Jiang, Y.; Zhang, L. Autophagy is a major mechanism for the dual effects of curcumin on renal cell carcinoma cells. Eur. J. Pharmacol. 2018, 826, 24–30.
- Chen, Y.J.; Chi, C.W.; Su, W.C.; Huang, H.L. Lapatinib induces autophagic cell death and inhibits growth of human hepatocellular carcinoma. Oncotarget 2014, 5, 4845–4854.
- Chen, Y.J.; Fang, L.W.; Su, W.C.; Hsu, W.Y.; Yang, K.C.; Huang, H.L. Lapatinib induces autophagic cell death and differentiation in acute myeloblastic leukemia. Onco. Targets. Ther. 2016, 9, 4453–4464.
- Ginet, V.; Puyal, J.; Rummel, C.; Aubry, D.; Breton, C.; Cloux, A.J.; Majjigapu, S.R.; Sordat, B.; Vogel, P.; Bruzzone, S.; et al. A critical role of autophagy in antileukemia/lymphoma effects of APO866, an inhibitor of NAD biosynthesis. Autophagy 2014, 10, 603–617.
- Hua, H.; Kong, Q.; Zhang, H.; Wang, J.; Luo, T.; Jiang, Y. Targeting mTOR for cancer therapy. J. Hematol. Oncol. 2019, 12, 71.
- Noda, T.; Ohsumi, Y. Tor, a phosphatidylinositol kinase homologue, controls autophagy in yeast. J. Biol. Chem. 1998, 273, 3963–3966.
- Sehgal, S.N.; Baker, H.; Vézina, C. Rapamycin (Ay-22,989), a New Antifungal Antibiotic. II. Fermentation, Isolation and Characterization. J. Antibiot. (Tokyo). 1975, 28, 727–732.
- Vézina, C.; Kudelski, A.; Sehgal, S.N. Rapamycin (AY 22,989) A NEW ANTIFUNGAL ANTIBIOTIC. J. Antibiot. (Tokyo). 1975, 28, 721–726.
- Pattingre, S.; Espert, L.; Biard-Piechaczyk, M.; Codogno, P. Regulation of macroautophagy by mTOR and Beclin 1 complexes. Biochimie 2008, 90, 313–323.
- Benjamin, D.; Colombi, M.; Moroni, C.; Hall, M.N. Rapamycin passes the torch: A new generation of mTOR inhibitors. Nat. Rev. Drug Discov. 2011, 10, 868–880.
- Chiarini, F.; Evangelisti, C.; Lattanzi, G.; McCubrey, J.A.; Martelli, A.M. Advances in understanding the mechanisms of evasive and innate resistance to mTOR inhibition in cancer cells. Biochim. Biophys. Acta. Mol. Cell Res. 2019, 1866, 1322–1337.
- Mao, B.; Gao, S.; Weng, Y.; Zhang, L.; Zhang, L. Design, synthesis, and biological evaluation of imidazo[1,2-b]pyridazine derivatives as mTOR inhibitors. Eur. J. Med. Chem. 2017, 129, 135–150.
- Chen, Y.; Lee, C.H.; Tseng, B.Y.; Tsai, Y.H.; Tsai, H.W.; Yao, C.L.; Tseng, S.H. AZD8055 exerts antitumor effects on colon cancer cells by inhibiting mTOR and cell-cycle progression. Anticancer Res. 2018, 38, 1445–1454.
- Carew, J.S.; Kelly, K.R.; Nawrocki, S.T. Mechanisms of mTOR inhibitor resistance in cancer therapy. Target. Oncol. 2011, 6, 17–27.
- Merino, D.; Kelly, G.L.; Lessene, G.; Wei, A.H.; Roberts, A.W.; Strasser, A. BH3-Mimetic Drugs: Blazing the Trail for New Cancer Medicines. Cancer Cell 2018, 34, 879–891.
- Opydo-Chanek, M.; Gonzalo, O.; Marzo, I. Multifaceted anticancer activity of BH3 mimetics: Current evidence and future prospects. Biochem. Pharmacol. 2017, 136, 12–23.
- Czabotar, P.E.; Lessene, G.; Strasser, A.; Adams, J.M. Control of apoptosis by the BCL-2 protein family: Implications for physiology and therapy. Nat. Rev. Mol. Cell Biol. 2014, 15, 49–63.
- Liang, L.Z.; Ma, B.; Liang, Y.J.; Liu, H.C.; Zhang, T.H.; Zheng, G.S.; Su, Y.X.; Liao, G.Q. Obatoclax induces Beclin 1- and ATG5-dependent apoptosis and autophagy in adenoid cystic carcinoma cells. Oral Dis. 2015, 21, 470–477.
- Koehler, B.C.; Jassowicz, A.; Scherr, A.L.; Lorenz, S.; Radhakrishnan, P.; Kautz, N.; Elssner, C.; Weiss, J.; Jaeger, D.; Schneider, M.; et al. Pan-Bcl-2 inhibitor Obatoclax is a potent late stage autophagy inhibitor in colorectal cancer cells independent of canonical autophagy signaling. BMC Cancer 2015, 15, 1–11.
- Śledziński, P.; Zeyland, J.; Słomski, R.; Nowak, A. The current state and future perspectives of cannabinoids in cancer biology. Cancer Med. 2018, 7, 765–775.
- Costa, L.; Amaral, C.; Teixeira, N.; Correia-Da-Silva, G.; Fonseca, B.M. Cannabinoid-induced autophagy: Protective or death role? Prostaglandins Other Lipid Mediat. 2016, 122, 54–63.
- Mrakovcic, M.; Kleinheinz, J.; Fröhlich, L.F. Histone deacetylase inhibitor-induced autophagy in tumor cells: Implications for p53. Int. J. Mol. Sci. 2017, 18, 1883.
- Newbold, A.; Falkenberg, K.J.; Prince, H.M.; Johnstone, R.W. How do tumor cells respond to HDAC inhibition? FEBS J. 2016, 283, 4032–4046.
- Mrakovcic, M.; Bohner, L.; Hanisch, M.; Fröhlich, L.F. Epigenetic targeting of autophagy via HDAC inhibition in tumor cells: Role of p53. Int. J. Mol. Sci. 2018, 19, 3952.
- Mann, B.S.; Johnson, J.R.; Cohen, M.H.; Justice, R.; Pazdur, R. FDA Approval Summary: Vorinostat for Treatment of Advanced Primary Cutaneous T-Cell Lymphoma. Oncologist 2007, 12, 1247–1252.
- Wang, J.; Ren, X.R.; Piao, H.; Zhao, S.; Osada, T.; Premont, R.T.; Mook, R.A.; Morse, M.A.; Lyerly, H.K.; Chen, W. Niclosamide-induced Wnt signaling inhibition in colorectal cancer is mediated by autophagy. Biochem. J. 2019, 476, 535–546.
- Lazarus, M.B.; Novotny, C.J.; Shokat, K.M. Structure of the human autophagy initiating kinase ULK1 in complex with potent inhibitors. ACS Chem. Biol. 2015, 10, 257–261.
- Skah, S.; Richartz, N.; Duthil, E.; Gilljam, K.M.; Bindesbøll, C.; Naderi, E.H.; Eriksen, A.B.; Ruud, E.; Dirdal, M.M.; Simonsen, A.; et al. cAMP-mediated autophagy inhibits DNA damage-induced death of leukemia cells independent of p53. Oncotarget 2018, 9, 30434–30449.
- Petherick, K.J.; Conway, O.J.L.; Mpamhanga, C.; Osborne, S.A.; Kamal, A.; Saxty, B.; Ganley, I.G. Pharmacological inhibition of ULK1 kinase blocks mammalian target of rapamycin (mTOR)-dependent autophagy. J. Biol. Chem. 2015, 290, 11376–11383.
- Lu, J.; Zhu, L.; Zheng, L.P.; Cui, Q.; Zhu, H.H.; Zhao, H.; Shen, Z.J.; Dong, H.Y.; Chen, S.S.; Wu, W.Z.; et al. Overexpression of ULK1 Represents a Potential Diagnostic Marker for Clear Cell Renal Carcinoma and the Antitumor Effects of SBI-0206965. EBioMedicine 2018, 34, 85–93.
- Dower, C.M.; Bhat, N.; Gebru, M.T.; Chen, L.; Wills, C.A.; Miller, B.A.; Wang, H.-G. Targeted Inhibition of ULK1 Promotes Apoptosis and Suppresses Tumor Growth and Metastasis in Neuroblastoma. Mol. Cancer Ther. 2018, 17, 2365–2376.
- Tang, F.; Hu, P.; Yang, Z.; Xue, C.; Gong, J.; Sun, S.; Shi, L.; Zhang, S.; Li, Z.; Yang, C.; et al. SBI0206965, a novel inhibitor of Ulk1, suppresses non-small cell lung cancer cell growth by modulating both autophagy and apoptosis pathways. Oncol. Rep. 2017, 37, 3449–3458.
- Egan, D.F.; Chun, M.G.H.; Vamos, M.; Zou, H.; Rong, J.; Miller, C.J.; Lou, H.J.; Raveendra-Panickar, D.; Yang, C.-C.; Sheffler, D.J.; et al. Small molecule inhibition of the autophagy kinase ULK1 and identification of ULK1 substrates. Mol. Cell 2015, 59, 285–297.
- Martin, K.R.; Celano, S.L.; Solitro, A.R.; Gunaydin, H.; Scott, M.; O’Hagan, R.C.; Shumway, S.D.; Fuller, P.; MacKeigan, J.P. A Potent and Selective ULK1 Inhibitor Suppresses Autophagy and Sensitizes Cancer Cells to Nutrient Stress. iScience 2018, 8, 74–84.
- Seglen, P.O.; Gordon, P.B. 3-Methyladenine: Specific inhibitor of autophagic/lysosomal protein degradation in isolated rat hepatocytes. Proc. Natl. Acad. Sci. USA 1982, 79, 1889–1892.
- Wu, Y.T.; Tan, H.L.; Shui, G.; Bauvy, C.; Huang, Q.; Wenk, M.R.; Ong, C.N.; Codogno, P.; Shen, H.M. Dual role of 3-methyladenine in modulation of autophagy via different temporal patterns of inhibition on class I and III phosphoinositide 3-kinase. J. Biol. Chem. 2010, 285, 10850–10861.
- Zou, Z.; Zhang, J.; Zhang, H.; Liu, H.; Li, Z.; Cheng, D.; Chen, J.; Liu, L.; Ni, M.; Zhang, Y.; et al. 3-Methyladenine can depress drug efflux transporters via blocking the PI3K-AKT-mTOR pathway thus sensitizing MDR cancer to chemotherapy. J. Drug Target. 2014, 22, 839–848.
- Wu, Y.; Wang, X.; Guo, H.; Zhang, B.; Zhang, X.B.; Shi, Z.J.; Yu, L. Synthesis and screening of 3-MA derivatives for autophagy inhibitors. Autophagy 2013, 9, 595–603.
- Powis, G.; Bonjouklian, R.; Berggren, M.M.; Gallegos, A.; Abraham, R.; Ashendel, C.; Zalkow, L.; Matter, W.F.; Dodge, J.; Grindey, G.; et al. Wortmannin, a Potent and Selective Inhibitor of Phosphatidylinositol-3-kinase. Cancer Res. 1994, 54, 2419–2423.
- Thelen, M.; Wymann, M.P.; Langen, H. Wortmannin binds specifically to 1-phosphatidylinositol 3-kinase while inhibiting guanine nucleotide-binding protein-coupled receptor signaling in neutrophil leukocytes. Proc. Natl. Acad. Sci. USA 1994, 91, 4960–4964.
- Vlahos, C.J.; Matter, W.F.; Hui, K.Y.; Brown, R.F. A Specific Inhibitor of Phosphatidylinositol 3 Kinase, 2-(4-Morpholinyl)-8-phenyl-4H-1-benzopyran-4-one (LY294002). J. Biol. Chem. 1994, 269, 5241–5248.
- Garlich, J.R.; De, P.; Dey, N.; Jing, D.S.; Peng, X.; Miller, A.; Murali, R.; Lu, Y.; Mills, G.B.; Kundra, V.; et al. A vascular targeted pan phosphoinositide 3-kinase inhibitor prodrug, SF1126, with antitumor and antiangiogenic activity. Cancer Res. 2008, 68, 206–215.
- Qin, A.; Li, Y.; Zhou, L.; Xing, C.; Lu, X. Dual PI3K-BRD4 Inhibitor SF1126 Inhibits Colorectal Cancer Cell Growth in Vitro and in Vivo. Cell. Physiol. Biochem. 2019, 52, 758–768.
- Ronan, B.; Flamand, O.; Vescovi, L.; Dureuil, C.; Durand, L.; Fassy, F.; Bachelot, M.F.; Lamberton, A.; Mathieu, M.; Bertrand, T.; et al. A highly potent and selective Vps34 inhibitor alters vesicle trafficking and autophagy. Nat. Chem. Biol. 2014, 10, 1013–1019.
- Farkas, T.; Daugaard, M.; Jäättelä, M. Identification of small molecule inhibitors of phosphatidylinositol 3-kinase and autophagy. J. Biol. Chem. 2011, 286, 38904–38912.
- Xie, J.; Li, Q.; Ding, X.; Gao, Y. GSK1059615 kills head and neck squamous cell carcinoma cells possibly via activating mitochondrial programmed necrosis pathway. Oncotarget 2017, 8, 50814–50823.
- Bei, S.; Li, F.; Li, H.; Li, J.; Zhang, X.; Sun, Q.; Feng, L. Inhibition of gastric cancer cell growth by a PI3K-mTOR dual inhibitor GSK1059615. Biochem. Biophys. Res. Commun. 2019, 511, 13–20.
- Bago, R.; Malik, N.; Munson, M.J.; Prescott, A.R.; Davies, P.; Sommer, E.; Shpiro, N.; Ward, R.; Cross, D.; Ganley, I.G.; et al. Characterization of VPS34-IN1, a selective inhibitor of Vps34, reveals that the phosphatidylinositol 3-phosphate-binding SGK3 protein kinase is a downstream target of class III phosphoinositide 3-kinase. Biochem. J. 2014, 463, 413–427.
- Dowdle, W.E.; Nyfeler, B.; Nagel, J.; Elling, R.A.; Liu, S.; Triantafellow, E.; Menon, S.; Wang, Z.; Honda, A.; Pardee, G.; et al. Selective VPS34 inhibitor blocks autophagy and uncovers a role for NCOA4 in ferritin degradation and iron homeostasis in vivo. Nat. Cell Biol. 2014, 16, 1069–1079.
- Pasquier, B.; El-Ahmad, Y.; Filoche-Rommé, B.; Dureuil, C.; Fassy, F.; Abecassis, P.Y.; Mathieu, M.; Bertrand, T.; Benard, T.; Barrière, C.; et al. Discovery of (2 S)-8-[(3 R)-3-Methylmorpholin-4-yl]-1-(3-methyl-2-oxobutyl)-2-(trifluoromethyl)-3,4-dihydro-2 H -pyrimido[1,2- a ]pyrimidin-6-one: A novel potent and selective inhibitor of Vps34 for the treatment of solid tumors. J. Med. Chem. 2015, 58, 376–400.
- Liu, J.; Xia, H.; Kim, M.; Xu, L.; Li, Y.; Zhang, L.; Cai, Y.; Norberg, H.V.; Zhang, T.; Furuya, T.; et al. Beclin1 Controls the Levels of p53 by Regulating the Deubiquitination Activity of USP10 and USP13. Cell 2011, 147, 223–234.
- Zhang, J.; Mao, S.; Wang, L.; Zhang, W.; Zhang, Z.; Guo, Y.; Wu, Y.; Yi, F.; Yao, X. MicroRNA-154 functions as a tumor suppressor in bladder cancer by directly targeting ATG7. Oncol. Rep. 2019, 41, 819–828.
- Akin, D.; Wang, S.K.; Habibzadegah-Tari, P.; Law, B.; Ostrov, D.; Li, M.; Yin, X.M.; Kim, J.S.; Horenstein, N.; Dunn, W.A. A novel ATG4B antagonist inhibits autophagy and has a negative impact on osteosarcoma tumors. Autophagy 2014, 10, 2021–2035.
- Liu, P.-F.; Hsu, C.-J.; Tseng, H.-H.; Wu, C.-H.; Shu, C.-W.; Tsai, K.-L.; Yang, L.-W.; Tsai, W.-L.; Cheng, J.-S.; Chang, H.-W.; et al. Drug repurposing screening identifies tioconazole as an ATG4 inhibitor that suppresses autophagy and sensitizes cancer cells to chemotherapy. Theranostics 2018, 8, 830–845.
- Kurdi, A.; Cleenewerck, M.; Vangestel, C.; Lyssens, S.; Declercq, W.; Timmermans, J.P.; Stroobants, S.; Augustyns, K.; De Meyer, G.R.Y.; Van Der Veken, P.; et al. ATG4B inhibitors with a benzotropolone core structure block autophagy and augment efficiency of chemotherapy in mice. Biochem. Pharmacol. 2017, 138, 150–162.
- Bosc, D.; Vezenkov, L.; Bortnik, S.; An, J.; Xu, J.; Choutka, C.; Hannigan, A.M.; Kovacic, S.; Loo, S.; Clark, P.G.K.; et al. A new quinoline-based chemical probe inhibits the autophagy-related cysteine protease ATG4B OPEN. Sci. Rep. 2018, 8, 11653.
- Fu, Y.; Hong, L.; Xu, J.; Zhong, G.; Gu, Q.; Gu, Q.; Guan, Y.; Zheng, X.; Dai, Q.; Luo, X.; et al. Discovery of a small molecule targeting autophagy via ATG4B inhibition and cell death of colorectal cancer cells in vitro and in vivo. Autophagy 2019, 15, 295–311.
- Qiu, Z.; Kuhn, B.; Aebi, J.; Lin, X.; Ding, H.; Zhou, Z.; Xu, Z.; Xu, D.; Han, L.; Liu, C.; et al. Discovery of Fluoromethylketone-Based Peptidomimetics as Covalent ATG4B (Autophagin-1) Inhibitors. ACS Med. Chem. Lett. 2016, 7, 802–806.
- Xu, D.; Xu, Z.; Han, L.; Liu, C.; Zhou, Z.; Qiu, Z.; Lin, X.; Tang, G.; Shen, H.; Aebi, J.; et al. Identification of new ATG4B inhibitors based on a novel high-throughput screening platform. SLAS Discov. 2017, 22, 338–347.
- Chu, J.; Fu, Y.; Xu, J.; Zheng, X.; Gu, Q.; Luo, X.; Dai, Q.; Zhang, S.; Liu, P.; Hong, L.; et al. ATG4B inhibitor FMK-9a induces autophagy independent on its enzyme inhibition. Arch. Biochem. Biophys. 2018, 644, 29–36.
- Donohue, E.; Tovey, A.; Vogl, A.W.; Arns, S.; Sternberg, E.; Young, R.N.; Roberge, M. Inhibition of autophagosome formation by the benzoporphyrin derivative verteporfin. J. Biol. Chem. 2011, 286, 7290–7300.
- Liu-Chittenden, Y.; Huang, B.; Shim, J.S.; Chen, Q.; Lee, S.-J.; Anders, R.A.; Liu, J.O.; Pan, D. Genetic and pharmacological disruption of the TEAD-YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 2012, 26, 1300–1305.
- Donohue, E.; Thomas, A.; Maurer, N.; Manisali, I.; Zeisser-Labouebe, M.; Zisman, N.; Anderson, H.J.; Ng, S.S.W.; Webb, M.; Bally, M.; et al. The Autophagy Inhibitor Verteporfin Moderately Enhances the Antitumor Activity of Gemcitabine in a Pancreatic Ductal Adenocarcinoma Model. J. Cancer 2013, 4, 585–596.
- Lui, J.W.; Xiao, S.; Ogomori, K.; Hammarstedt, J.E.; Little, E.C.; Lang, D. The efficiency of verteporfin as a therapeutic option in pre-clinical models of melanoma. J. Cancer 2019, 10, 1–10.
- Poole, B.; Ohkuma, S. Effect of weak bases on the intralysosomal pH in mouse peritoneal macrophages. J. Cell Biol. 1981, 90, 665–669.
- Solomon, V.R.; Lee, H. Chloroquine and its analogs: A new promise of an old drug for effective and safe cancer therapies. Eur. J. Pharmacol. 2009, 625, 220–233.
- Shi, T.T.; Yu, X.X.; Yan, L.J.; Xiao, H.T. Research progress of hydroxychloroquine and autophagy inhibitors on cancer. Cancer Chemother. Pharmacol. 2017, 79, 287–294.
- McAfee, Q.; Zhang, Z.; Samanta, A.; Levi, S.M.; Ma, X.-H.; Piao, S.; Lynch, J.P.; Uehara, T.; Sepulveda, A.R.; Davis, L.E.; et al. Autophagy inhibitor Lys05 has single-agent antitumor activity and reproduces the phenotype of a genetic autophagy deficiency. Proc. Natl. Acad. Sci. USA 2012, 109, 8253–8258.
- Baquero, P.; Dawson, A.; Mukhopadhyay, A.; Kuntz, E.M.; Mitchell, R.; Olivares, O.; Ianniciello, A.; Scott, M.T.; Dunn, K.; Nicastri, M.C.; et al. Targeting quiescent leukemic stem cells using second generation autophagy inhibitors. Leukemia 2019, 33, 981–994.
- Rebecca, V.W.; Nicastri, M.C.; Mclaughlin, N.; Fennelly, C.; Ronghe, A.; Nofal, M.; Lim, C.; Witze, E.; Chude, C.I.; Zhang, G.; et al. A unified approach to targeting the lysosome’s degradative and growth signaling roles. Cancer Discov. 2017, 7, 1266–1283.
- Goodall, M.L.; Wang, T.; Martin, K.R.; Kortus, M.G.; Kauffman, A.L.; Trent, J.M.; Gately, S.; MacKeigan, J.P. Development of potent autophagy inhibitors that sensitize oncogenic BRAF V600E mutant melanoma tumor cells to vemurafenib. Autophagy 2014, 10, 1120–1136.
- Lam Yi, H.; Than, H.; Sng, C.; Cheong, M.A.; Chuah, C.; Xiang, W. Lysosome Inhibition by Mefloquine Preferentially Enhances the Cytotoxic Effects of Tyrosine Kinase Inhibitors in Blast Phase Chronic Myeloid Leukemia. Transl. Oncol. 2019, 12, 1221–1228.
- Sharma, N.; Thomas, S.; Golden, E.B.; Hofman, F.M.; Chen, T.C.; Petasis, N.A.; Schönthal, A.H.; Louie, S.G. Inhibition of autophagy and induction of breast cancer cell death by mefloquine, an antimalarial agent. Cancer Lett. 2012, 326, 143–154.
- Cheng, S.; Sliva, D. Ganoderma lucidum for cancer treatment: We are close but still not there. Integr. Cancer Ther. 2015, 14, 249–257.
- Pan, H.; Wang, Y.; Na, K.; Wang, Y.; Wang, L.; Li, Z.; Guo, C.; Guo, D.; Wang, X. Autophagic flux disruption contributes to Ganoderma lucidum polysaccharide-induced apoptosis in human colorectal cancer cells via MAPK/ERK activation. Cell Death Dis. 2019, 10.
- Wu, K.; Na, K.; Chen, D.; Wang, Y.; Pan, H.; Wang, X. Effects of non-steroidal anti-inflammatory drug-activated gene-1 on Ganoderma lucidum polysaccharides-induced apoptosis of human prostate cancer PC-3 cells. Int. J. Oncol. 2018, 53, 2356–2368.
- Yamamoto, A.; Tagawa, Y.; Yoshimori, T.; Moriyama, Y.; Masaki, R.; Tashiro, Y. Bafilomycin Ai Prevents Maturation of Autophagic Vacuoles by Inhibiting Fusion between Autophagosomesand Lysosomesin Rat HepatomaCell Line. Cell Struct. Funct. 1998, 23, 33–42.
- Bowman, E.J.; Siebers, A.; Altendorf, K. Bafilomycins; A class of inhibitors of membrane ATPases from microorganisms, animal cells, and plant cells. Proc. Natl. Acad. Sci. USA 1988, 85, 7972–7976.
- Mauvezin, C.; Neufeld, T.P. Bafilomycin A1 disrupts autophagic flux by inhibiting both V-ATPase-dependent acidification and Ca-P60A/SERCA-dependent autophagosome-lysosome fusion. Autophagy 2015, 11, 1437–1438.
- Rodilla, A.M.; Korrodi-Gregório, L.; Hernando, E.; Manuel-Manresa, P.; Quesada, R.; Pérez-Tomás, R.; Soto-Cerrato, V. Synthetic tambjamine analogues induce mitochondrial swelling and lysosomal dysfunction leading to autophagy blockade and necrotic cell death in lung cancer. Biochem. Pharmacol. 2017, 126, 23–33.
- Grinde, B. Effect of carboxylic ionophores on lysosomal protein degradation in rat hepatocytes. Exp. Cell Res. 1983, 149, 27–35.
- Busschaert, N.; Park, S.H.; Baek, K.H.; Choi, Y.P.; Park, J.; Howe, E.N.W.; Hiscock, J.R.; Karagiannidis, L.E.; Marques, I.; Félix, V.; et al. A synthetic ion transporter that disrupts autophagy and induces apoptosis by perturbing cellular chloride concentrations. Nat. Chem. 2017, 9, 667–675.
- Sharma, G.; Guardia, C.M.; Roy, A.; Vassilev, A.; Saric, A.; Griner, L.N.; Marugan, J.; Ferrer, M.; Bonifacino, J.S.; DePamphilis, M.L. A family of PIKFYVE inhibitors with therapeutic potential against autophagy-dependent cancer cells disrupt multiple events in lysosome homeostasis. Autophagy 2019, 15, 1694–1718.
- Lu, Y.; Dong, S.; Hao, B.; Li, C.; Zhu, K.; Guo, W.; Wang, Q.; Cheung, K.-H.; Wong, C.W.M.; Wu, W.-T.; et al. Vacuolin-1 potently and reversibly inhibits autophagosome-lysosome fusion by activating RAB5A. Autophagy 2014, 10, 1895–1905.
- Rossi, M.; Munarriz, E.R.; Bartesaghi, S.; Milanese, M.; Dinsdale, D.; Guerra-Martin, M.A.; Bampton, E.T.W.; Glynn, P.; Bonanno, G.; Knight, R.A.; et al. Desmethylclomipramine induces the accumulation of autophagy markers by blocking autophagic flux. J. Cell Sci. 2009, 122, 3330–3339.
- Libby, P.; Goldberg, A.L. Leupeptin, a protease inhibitor, decreases protein degradation in normal and diseased muscles. Science (80-. ). 1978, 199, 534–536.
- Tanida, I.; Minematsu-Ikeguchi, N.; Ueno, T.; Kominami, E. Lysosomal turnover, but not a cellular level, of endogenous LC3 is a marker for autophagy. Autophagy 2005, 1, 84–91.
- Stern, S.T.; Adiseshaiah, P.P.; Crist, R.M. Autophagy and lysosomal dysfunction as emerging mechanisms of nanomaterial toxicity. Part. Fibre Toxicol. 2012, 9, 20.
- Ma, X.; Wu, Y.; Jin, S.; Tian, Y.; Zhang, X.; Zhao, Y.; Yu, L.; Liang, X.J. Gold nanoparticles induce autophagosome accumulation through size-dependent nanoparticle uptake and lysosome impairment. ACS Nano 2011, 5, 8629–8639.
- Cui, Z.; Zhang, Y.; Xia, K.; Yan, Q.; Kong, H.; Zhang, J.; Zuo, X.; Shi, J.; Wang, L.; Zhu, Y.; et al. Nanodiamond autophagy inhibitor allosterically improves the arsenical-based therapy of solid tumors. Nat. Commun. 2018, 9, 4347.
- Hosokawa, N.; Hara, T.; Kaizuka, T.; Kishi, C.; Takamura, A.; Miura, Y.; Iemura, S.; Natsume, T.; Takehana, K.; Yamada, N.; et al. Nutrient-dependent mTORC1 Association with the ULK1–Atg13–FIP200 Complex Required for Autophagy. Mol. Biol. Cell 2009, 20, 1981–1991.
- Chen, S.; Wang, C.; Yeo, S.; Liang, C.C.; Okamoto, T.; Sun, S.; Wen, J.; Guan, J.L. Distinct roles of autophagy-dependent and -independent functions of FIP200 revealed by generation and analysis of a mutant knock-in mouse model. Genes Dev. 2016, 30, 856–869.
- Mercer, C.A.; Kaliappan, A.; Dennis, P.B. A novel, human Atg13 binding protein, Atg101, interacts with ULK1 and is essential for macroautophagy. Autophagy 2009, 5, 649–662.
- Lee, E.J.; Tournier, C. The requirement of uncoordinated 51-like kinase 1 (ULK1) and ULK2 in the regulation of autophagy. Autophagy 2011, 7, 689–695.
- Chaikuad, A.; Koschade, S.E.; Stolz, A.; Zivkovic, K.; Pohl, C.; Shaid, S.; Ren, H.; Lambert, L.J.; Cosford, N.D.P.; Brandts, C.H.; et al. Conservation of structure, function and inhibitor binding in UNC-51-like kinase 1 and 2 (ULK1/2). Biochem. J. 2019, 2, BCJ20190038.
- Yun, M.; Bai, H.Y.; Zhang, J.X.; Rong, J.; Weng, H.W.; Zheng, Z.S.; Xu, Y.; Tong, Z.T.; Huang, X.X.; Liao, Y.J.; et al. ULK1: A promising biomarker in predicting poor prognosis and therapeutic response in human nasopharygeal carcinoma. PLoS ONE 2015, 10, e0117375.
- Zou, Y.; Chen, Z.; He, X.; He, X.; Wu, X.; Chen, Y.; Wu, X.; Wang, J.; Lan, P. High expression levels of unc-51-like kinase 1 as a predictor of poor prognosis in colorectal cancer. Oncol. Lett. 2015, 10, 1583–1588.
- Jiang, S.; Li, Y.; Zhu, Y.H.; Wu, X.Q.; Tang, J.; Li, Z.; Feng, G.K.; Deng, R.; Li, D.D.; Luo, R.Z.; et al. Intensive expression of UNC-51-like kinase 1 is a novel biomarker of poor prognosis in patients with esophageal squamous cell carcinoma. Cancer Sci. 2011, 102, 1568–1575.
- Dite, T.A.; Langendorf, C.G.; Hoque, A.; Galic, S.; Rebello, R.J.; Ovens, A.J.; Lindqvist, L.M.; Ngoei, K.R.W.; Ling, N.X.Y.; Furic, L.; et al. AMP-activated protein kinase selectively inhibited by the type II inhibitor SBI-0206965. J. Biol. Chem. 2018, 293, 8874–8885.
- Lindmo, K. Regulation of membrane traffic by phosphoinositide 3-kinases. J. Cell Sci. 2006, 119, 605–614.
- Liu, P.; Cheng, H.; Roberts, T.M.; Zhao, J.J. Targeting the phophoinositide 3-kinase(PI3K)pathway in cancer. Nat Rev Drug Discov 2009, 8, 627–644.
- Leontieva, O.V.; Blagosklonny, M. V Gerosuppression by pan-mTOR inhibitors. Aging (Albany. NY). 2016, 8, 3535–3551.
- Funderburk, S.F.; Wang, Q.J.; Yue, Z. Public Access NIH Public Access. Trends Cell Biol. 2010, 20, 355–362.
- Valet, C.; Levade, M.; Chicanne, G.; Bilanges, B.; Cabou, C.; Viaud, J.; Gratacap, M.-P.; Gaits-Iacovoni, F.; Vanhaesebroeck, B.; Payrastre, B.; et al. A dual role for the class III PI3K, Vps34, in platelet production and thrombus growth. Blood 2017, 130, 2032–2042.
- Tanida, I.; Sou, Y.S.; Ezaki, J.; Minematsu-Ikeguchi, N.; Ueno, T.; Kominami, E. HsAtg4B/HsApg4B/autophagin-1 cleaves the carboxyl termini of three human Atg8 homologues and delipidates microtubule-associated protein light chain 3- and GABAA receptor-associated protein-phospholipid conjugates. J. Biol. Chem. 2004, 279, 36268–36276.
- Yu, Z.Q.; Ni, T.; Hong, B.; Wang, H.Y.; Jiang, F.J.; Zou, S.; Chen, Y.; Zheng, X.L.; Klionsky, D.J.; Liang, Y.; et al. Dual roles of Atg8—PE deconjugation by Atg4 in autophagy. Autophagy 2012, 8, 883–892.
- Fu, Y.; Huang, Z.; Hong, L.; Lu, J.H.; Feng, D.; Yin, X.M.; Li, M. Targeting ATG4 in cancer therapy. Cancers 2019, 11, 649.
- Tran, E.; Chow, A.; Goda, T.; Wong, A.; Blakely, K.; Rocha, M.; Taeb, S.; Hoang, V.C.; Liu, S.K.; Emmenegger, U. Context-dependent role of ATG4B as target for autophagy inhibition in prostate cancer therapy. Biochem. Biophys. Res. Commun. 2013, 441, 726–731.
- Bortnik, S.; Choutka, C.; Horlings, H.M.; Leung, S.; Baker, J.H.E.; Lebovitz, C.; Dragowska, W.H.; Go, N.E.; Bally, M.B.; Minchinton, A.I.; et al. Identification of breast cancer cell subtypes sensitive to ATG4B inhibition. Oncotarget 2016, 7.
- Donohue, E.; Balgi, A.D.; Komatsu, M.; Roberge, M. Induction of Covalently Crosslinked p62 Oligomers with Reduced Binding to Polyubiquitinated Proteins by the Autophagy Inhibitor Verteporfin. PLoS ONE 2014, 9, e114964.
- Zhang, H.; Ramakrishnan, S.K.; Triner, D.; Centofanti, B.; Maitra, D.; Győrffy, B.; Sebolt-Leopold, J.S.; Dame, M.K.; Varani, J.; Brenner, D.E.; et al. Tumor-selective proteotoxicity of verteporfin inhibits colon cancer progression independently of YAP1. Sci. Signal. 2015, 8, ra98.
- Ota, M.; Sasaki, H. Mammalian Tead proteins regulate cell proliferation and contact inhibition as transcriptional mediators of Hippo signaling. Development 2008, 135, 4059–4069.
- Wei, H.; Wang, F.; Wang, Y.; Li, T.; Xiu, P.; Zhong, J.; Sun, X.; Li, J. Verteporfin suppresses cell survival, angiogenesis and vasculogenic mimicry of pancreatic ductal adenocarcinoma via disrupting the YAP-TEAD complex. Cancer Sci. 2017, 108, 478–487.
- Melles, R.B.; Marmor, M.F. The risk of toxic retinopathy in patients on long-term hydroxychloroquine therapy. JAMA Ophthalmol. 2014, 132, 1453–1460.
- Costedoat-Chalumeau, N.; Hulot, J.-S.; Amoura, Z.; Delcourt, A.; Maisonobe, T.; Dorent, R.; Bonnet, N.; Sablé, R.; Lechat, P.; Wechsler, B.; et al. Cardiomyopathy Related to Antimalarial Therapy with Illustrative Case Report. Cardiology 2007, 107, 73–80.
- Marmor, M.F.; Kellner, U.; Lai, T.Y.Y.; Melles, R.B.; Mieler, W.F.; Lum, F. Recommendations on Screening for Chloroquine and Hydroxychloroquine Retinopathy (2016 Revision). Ophthalmology 2016, 123, 1386–1394.
- Bristol, M.L.; Emery, S.M.; Maycotte, P.; Thorburn, A.; Chakradeo, S.; Gewirtz, D.A. Autophagy inhibition for chemosensitization and radiosensitization in cancer: Do the preclinical data support this therapeutic strategy? J. Pharmacol. Exp. Ther. 2013, 344, 544–552.
- Bongiorno-Borbone, L.; Giacobbe, A.; Compagnone, M.; Eramo, A.; De Maria, R.; Peschiaroli, A.; Melino, G. Anti-tumoral effect of desmethylclomipramine in lung cancer stem cells. Oncotarget 2015, 6, 16926–16938.
- Boya, P.; Casares, N.; Perfettini, J.; Dessen, P.; Larochette, N.; Métivier, D.; Meley, D.; Souquere, S.; Pierron, G.; Codogno, P.; et al. Inhibition of Macroautophagy Triggers Apoptosis. Mol. Cell. Biol. 2005, 25, 1025–1040.
- Verbaanderd, C.; Maes, H.; Schaaf, M.B.; Sukhatme, V.P.; Pantziarka, P.; Sukhatme, V.; Agostinis, P.; Bouche, G. Repurposing drugs in oncology (ReDO)—Chloroquine and hydroxychloroquine as anti-cancer agents. Ecancermedicalscience 2017, 11, 1–35.
- Piao, S.; Ojha, R.; Rebecca, V.W.; Samanta, A.; Ma, X.H.; Mcafee, Q.; Nicastri, M.C.; Buckley, M.; Brown, E.; Winkler, J.D.; et al. ALDH1A1 and HLTF modulate the activity of lysosomal autophagy inhibitors in cancer cells. Autophagy 2017, 13, 2056–2071.
- Maycotte, P.; Aryal, S.; Ct, C.; Thorburn, J. Chloroquine sensitizes breast cancer cells to chemotherapy independent of autophagy. Autophagy 2012, 8, 200–212.
- Eng, C.H.; Wang, Z.; Tkach, D.; Toral-Barza, L.; Ugwonali, S.; Liu, S.; Fitzgerald, S.L.; George, E.; Frias, E.; Cochran, N.; et al. Macroautophagy is dispensable for growth of KRAS mutant tumors and chloroquine efficacy. Proc. Natl. Acad. Sci. USA 2016, 113, 182–187.
- Degtyarev, M.; De Mazière, A.; Orr, C.; Lin, J.; Lee, B.B.; Tien, J.Y.; Prior, W.W.; Van Dijk, S.; Wu, H.; Gray, D.C.; et al. Akt inhibition promotes autophagy and sensitizes PTEN-null tumors to lysosomotropic agents. J. Cell Biol. 2008, 183, 101–116.
- Kwakye-Berko, F.; Meshnick, S.R. Binding of chloroquine to DNA. Mol. Biochem. Parasitol. 1989, 35, 51–55.
- Loehberg, C.R.; Strissel, P.L.; Dittrich, R.; Strick, R.; Dittmer, J.; Dittmer, A.; Fabry, B.; Kalender, W.A.; Koch, T.; Wachter, D.L.; et al. Akt and p53 are potential mediators of reduced mammary tumor growth by Chloroquine and the mTOR inhibitor RAD001. Biochem. Pharmacol. 2012, 83, 480–488.
- Enzenmüller, S.; Gonzalez, P.; Debatin, K.M.; Fulda, S. Chloroquine overcomes resistance of lung carcinoma cells to the dual PI3K/mTOR inhibitor PI103 by lysosome-mediated apoptosis. Anticancer. Drugs 2013, 24, 14–19.
- Kroemer, G.; Jäättelä, M. Lysosomes and autophagy in cell death control. Nat. Rev. Cancer 2005, 5, 886–897.
- Carew, J.S.; Nawrocki, S.T.; Kahue, C.N.; Zhang, H.; Yang, C.; Chung, L.; Houghton, J.A.; Huang, P.; Giles, F.J.; Cleveland, J.L. Targeting autophagy augments the anticancer activity of the histone deacetylase inhibitor SAHAto overcome Bcr-Abl-mediated drug resistance. Blood 2007, 110, 313–322.
- Morgan, M.J.; Fitzwalter, B.E.; Owens, C.R.; Powers, R.K.; Sottnik, J.L.; Gamez, G.; Costello, J.C.; Theodorescu, D.; Thorburn, A. Metastatic cells are preferentially vulnerable to lysosomal inhibition. Proc. Natl. Acad. Sci. USA 2018, 115, E8479–E8488.
- Fennelly, C.; Amaravadi, R.K. Lysosomal Biology in Cancer. Methods Mol Biol 2017, 1594, 293–308.
- Uhlman, A.; Folkers, K.; Liston, J.; Pancholi, H.; Hinton, A. Effects of Vacuolar H(+)-ATPase Inhibition on Activation of Cathepsin B and Cathepsin L Secreted from MDA-MB231 Breast Cancer Cells. Cancer Microenviron. 2017, 10, 49–56.
- Balic, A.; Sørensen, M.D.; Trabulo, S.M.; Sainz, B.; Cioffi, M.; Vieira, C.R.; Miranda-Lorenzo, I.; Hidalgo, M.; Kleeff, J.; Erkan, M.; et al. Chloroquine Targets Pancreatic Cancer Stem Cells via Inhibition of CXCR4 and Hedgehog Signaling. Mol. Cancer Ther. 2014, 13, 1758–1771.
- Maes, H.; Kuchnio, A.; Peric, A.; Moens, S.; Nys, K.; DeBock, K.; Quaegebeur, A.; Schoors, S.; Georgiadou, M.; Wouters, J.; et al. Tumor vessel normalization by chloroquine independent of autophagy. Cancer Cell 2014, 26, 190–206.
- Mulcahy Levy, J.M.; Thompson, J.C.; Griesinger, A.M.; Amani, V.; Donson, A.M.; Birks, D.K.; Morgan, M.J.; Mirsky, D.M.; Handler, M.H.; Foreman, N.K.; et al. Autophagy Inhibition ImprovesChemosensitivity in BRAFV600E Brain Tumors. Cancer Discov 2014, 4, 773–780.





