
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Olena Filchakova | + 2357 word(s) | 2357 | 2021-06-10 08:19:31 | | | |
| 2 | Catherine Yang | Meta information modification | 2357 | 2021-06-22 03:51:21 | | |
Video Upload Options
Acetylcholine is a widely distributed excitatory neurotransmitter. Within the human body, it is present in both branches of the autonomic nervous system: within the parasympathetic system in pre- and postganglionic cells, and within the sympathetic system in preganglionic cells. It is also a neurotransmitter at the periphery within the neuromuscular junction.
1. Introduction
The receptors are characterized by radial symmetry and have pentameric organization, with five subunits arranged radially around a central ion-conducting pore. There are 17 subunits described in vertebrates: α1 to α10, β1 to β4, γ, δ, and ε. α5 and β3 subunits are considered auxiliary subunits, not contributing to the ACh binding site but influencing properties of the receptor [1][2]. α1, β1, δ, ε, and γ are subunits within muscle-type of nAChR, with ε subunits present in adult receptor type, while γ subunit present in fetal receptor.
The orthosteric ligand-binding site is formed at the junction between two neighboring subunits, where loops A, B, and C, from principal subunits and loops D, E, and F, coalesce and form a hydrophobic cage from aromatic residues that provides a space for ACh binding. The aromatic residues protruding to the ligand-binding site are rather conserved; within the muscle α1 subunit, they include Tyr 93 from loop A, Trp 149 from loop B, and Tyr 190 and Tyr 198 from loop C. These aromatic residues within the principal subunit are assisted by aromatic residues of a complementary subunit, such as Trp 57 of the γ subunit [3]. The conserved aromatic residues interact with ligands via cation-π interaction.
The receptor structure was studied at an atomic level. The quest for receptor structure determination was facilitated through studies on acetylcholine-binding protein (AChBP), a soluble multimeric protein secreted by snail glial cells which buffers ACh concentration [4].
nAChRs are widely distributed within the central nervous system (CNS) and peripheral nervous system, as well as outside the nervous system. The most prevalent receptor subtype within the brain is the α4β2-containing receptor, the upregulation of which is observed following chronic nicotine consumption [5]. Existing in two stoichiometric forms ((α4)2(β2)3 and (α4)3(β2)2), α4β2-containing receptors can vary in their sensitivities to ACh [6]. They play critical role within reward pathways, in particular, within nigrostriatal dopaminergic circuitry [7].
Disfunction in nAChRs within neuromuscular junctions can lead to myasthenic syndrome. Within the CNS, abnormal functioning of the receptors can manifest itself as Alzheimer’s disease, schizophrenia, depression, and Parkinson’s disease [8]. Outside the CNS, improper functionality of the receptors can lead to skin disorders such as pemphigus vulgaris and cancer.
Thus, knowledge about the functionality of the receptors is of paramount importance. Ligands targeting the receptors can help decipher physiological mechanisms, as well as prevent pathological processes where receptors are involved [9][10]. Natural products and, in particular, venoms of venomous animals, such as snails, snakes, spiders, and scorpions, contain biologically active compounds targeting nAChRs; thus, they are actively and thoroughly investigated [11].
2. Snail Venoms as a Source of Toxins Acting on Acetylcholine Receptors
Predatory snails within the Conus genus produce peptides— conotoxins that target different ion channels [12]. The peptides with a general structure CC-Xm-C-Xn-C, where C represent cysteines, Xm and Xn—variable number of residues between cysteines—constitute the family of α-conotoxins (α-Ctx). Structurally, α-conotoxins have very rigid three-dimensional structure due to the restraints imposed by disulfide bonds.
α3/5 conotoxins predominantly target muscle subtypes of nAChRs [13]. α-Conotoxins MI, GI, and SIA have been shown to exhibit >10,000-fold selectivity for α/δ over the α/γ site in mammalian receptors. However, the reverse pattern is observed in Torpedo receptors. Conserved Pro 6 and Tyr 12 residues are critical for hydrophobic interaction between the δ subunit and MI [14].
Ac1.1a and Ac1.1b are almost identical α3/5 conotoxins differing in only one residue at position 14 [15] (Figure 1). They exhibit selectivity towards α/δ over α/γ site. The first three residues before the Cys 4 have been shown to lack an effect on binding the target.
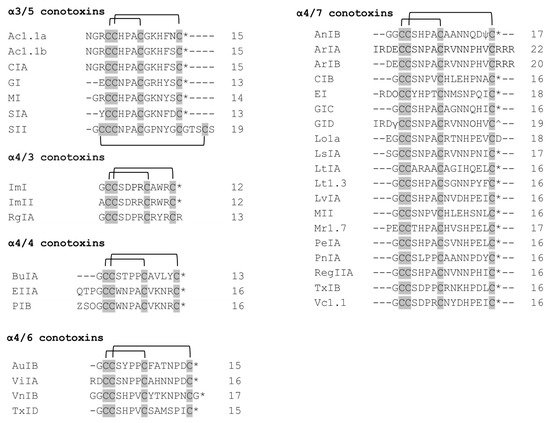
Figure 1. The α-conotoxins with 3/5, 4/4, 4/6, and 4/7 spacing. The cysteines residues are shaded in gray and intrachain disulfide bonds are shown. The number of residues in each toxin is given to the right. * C—terminal carboxamide, ^ C—terminal carboxylate, ψ—sulfated tyrosine, O—hydroxyproline, Z—pyroglutamate.
α-Conotoxin GI blocks muscle-type nAChRs, with selectivity for the α/δ site in mouse muscle receptors [16], and the α/γ site in Torpedo californica electric organ receptors [17]. Pro 5, Gly 8, Arg 9, and Tyr 11 are critical residues for GI to target GI mutant demonstrated a three-fold increase in potency in mouse α1β1δε and a decreased potency in rat neuronal α9α10 nAChR, compared to wildtype GI [18].
NMR structures for both CIA and CIB alpha-conotoxins derived from Conus catus have been studied. α-Conotoxin CIA caused muscle paralysis in fish when tested in vivo [19], while 4/7 α-Conotoxin CIB blocked neuronal type nAChRs.
Overall, there are two conserved regions (residues 2 and 3 in the first cysteine loop, as well as residues 1, 2, and 4 in the second cysteine loop) inherent to α3/5 conotoxins (Figure 1), which include important residues for the binding to muscle-type nAChR. (1) it has Pro residue in place of expected Arg/Lys; (2) it has a long C-terminus. Structure–functional studies on SII are very scarce, except for one that characterized SII as possessing three disulfide bonds and no net positive charge [20].
Asp 5, Pro 6, Arg 7, and Trp 10 are critical residues of the ImI conotoxin, conferring its specificity for the α7 nAChRs [21].
Despite the fact that ImI and ImII conotoxins share a high sequence homology (9 of 12 amino acids are identical) (Figure 1), they were found to target different sites at the α7 nAChR, since only ImI (and not ImII) showed competitive inhibition of the receptor when α-bungarotoxin (α-Bgtx) was co-added [22]. Mutational studies revealed that the amino acid residue at position 6 (Pro in α-Ctx ImI, Arg in α-Ctx ImII) determines the selectivity for a specific site, whereas the amino acid residue in position 9 (Ala in α-Ctx ImI, Arg in α-Ctx ImII) confers each of the toxins its optimal affinity to bind to their corresponding sites at the α7 nAChR [22].
The selectivity and potency of α-conotoxin RgIA for the α9α10 subtype of nAChR is conferred by Arg 9 and Arg 13 residues [23]. It is also interesting to note that [24][6]-dicarba RgIA is able to act specifically at α9α10 nAChRs, losing its effect on N-type calcium channels [25]. This analogue could potentially be used in a clinical trial for eliciting an effect on a single receptor subtype.
α-conotoxin α3β2 nAChRs with a significantly lower IC50 than α4β2 receptors [26]. In fact, there is a 40,000-fold difference between their IC50 values [26]. The five residues of the α6 subunit, Lys 85, Asp 187, Ile 188, Thr 198, and Tyr 205, were determined to be responsible for this effect [27].
It should be noted that α-conotoxin BuIA on nAChRs with β4 subunits have slow off-time when compared to corresponding nAChRs with β2 subunits [26]. This implies that BuIA can discriminate between different β subunits based on the time needed to unblock the receptor.
The α-conotoxin BuIA [T5A; P6O] was found to be selective for α6β4 vs. α6β2 nAChRs [28]. It was mainly achieved by P6O substitution, which resulted in a 2800-fold increase of IC50 at α6/α3β2β3 and only a 6-fold increase at α6/α3β4 [28].
It is interesting to note that, though α-conotoxins EIIA and PIB come from different Conus species, they are almost identical in sequence, except for one residue at the second position. Therefore, it is not surprising that they both have selectivity for muscle-type Pro 5, Ala 6, and Lys 9 residues located in the first and second loops, respectively, which were previously found to be critical for the muscle nAChR selectivity of α3/5 conotoxins [29].
Alanine scanning mutagenesis revealed three residues in the AuIB conotoxin that affect the inhibition of α3β4 nAChRs [30]. Hence, its substitution with alanine disrupted a favorable interaction of AuIB with the receptor. It was elucidated that Pro 6 caused inhibition due to its effect on 3D structure of the peptide, whereas Phe 9 was responsible for the interaction with a two-residue binding pocket (Trp 59 and Lys 61) of the β4 subunit [30].
Structure– activity studies on the ViIA conotoxin revealed that Arg 1 and His 11 are critical residues for binding to the target effectively. (a mutant with additional Leu residue at position 16) produced a 12-fold stronger interaction with α3β2 nAChR when compared to the native peptide. This was explained by hydrophobicity Leu residue conferred to the overall structure of the ViIA conotoxin [31].
The VnIB conotoxin selectively inhibits However, no structure–activity studies on this conopeptide have been carried out so far. It is hypothesized that all of the members of α4/6 conotoxins, including VnIB, exhibit β4 subunit selectivity due to their common 4/6 substructure. The selectivity of VnIB toward α6-containing nAChRs can potentially be explained by the five residues present in the second disulfide-loop, excluding Pro residue (YTKNPN) [32].
There are five residues (His 5, Pro 6, Val 7, Met 11, and Pro 13) of TxID critical for the inhibition of α3β4 and α6/α3β4 nAChRs [33]. [S9A] TxID mutant was found to discriminate between the two receptor subtypes, having a 46-fold higher affinity to α3β4 than to α6/α3β4 nAChRs [33].
Ala 11 and Gln 14 play a role in α3β2 selectivity of AnIB, whereas the C-terminal amidation and sulfation of Tyr 16 are responsible for α7 nAChR binding [34].
ArIB [V11L; V16D] is considered to be the most selective α7 nAChR antagonist to date (IC50 = 1.09 nM) [35].
Important residues conferring EI potency to bind mouse α1β1δε nAChR were found to be His 7, In addition, deletion of Arg1–Asn2–Hyp3 residues are also crucial, since their deletion causes a total loss of activity at muscle receptors [36]. [S13A] EI mutant exhibited increased potency and selectivity for α1β1δε nAChRs [36].
GIC is a very interesting conotoxin from a research standpoint, since it exhibits extreme potency and selectivity towards α3β2 nAChRs (IC50 = 1.1 nM) [37]. His 5 and Gln 13 were found to be the most important residues that confer GIC its potency and selectivity, respectively. In addition, Ala 7, Asn 11, and Asn 12 residues of the peptide were involved in binding to the principal site of the receptor [37].
Ala 10, Val 13 and Val 18 are important residues of the GID conotoxin, conferring it selectivity for α4β2 over α3β2 nAChRs [38]. In particular, GID[V18N] analog exhibits the highest selectivity for α4β2, with no inhibitory effect on the α3β2 subtype [38].
The structure–functional study conducted to the Lo1a conotoxin found that its Asp-18 residue is critical for the selectivity of the peptide for neuronal vs. muscle subtype nAChRs [39]. This was evident from the observation that Lo1a-ΔD and Lo1a-RRR analogues, which either lacked Asp 18 or had it replaced with 3 Arg residues, respectively, acquired affinity for the adult muscle subtype α1β1δε, aside from α7 receptors [39].
Ser-1 and Gly 2 are important residues for the affinity of LsIA to α3β2 and α7 nAChRs [40]. In addition, carboxylation of the C-terminus of LsIA resulted in its selectivity for α3β2 over α7 to increase 9-fold [40].
Selectivity of the Lt1.3 conotoxin for α3β2 receptors is attributed to a small hydrophobic Gly 10 residue, whereas Asn 11, Asn 12, Pro 13, Tyr 14, and Phe 15 residues contribute to the receptor binding [41].
His 5, Pro 6, Ala 7, and His 12 of LvIA are important residues that interact with α3-containing nAChRs [42]. LvIA exhibits unusual selectivity toward α3β2 vs. α6/ α3β2β3 nAChRs due to Asn 9 and Asp 11 residues [43].
Asp 5, Pro 6, and His 12 are the main residues of the MII conotoxin, which confers its potency for binding to the rat α3β2 nAChR subtype [44]. The MII [H9A; L15A] analog of α-conotoxin MII is selective towards α6-containing receptors (IC50 on rat α6/α3β2β3 = 2.4 nM), and can discriminate between α6 and α3 subunits [45].
Mr1.7 [E2A] is potent and selective for the α3β2 nAChR subtype, with no inhibitory effect on other subtypes [46]. However, the substitution of Ser 12 with Ala results in the loss of selectivity to α3β2, with new binding ability to other subtypes (α3β4, α2β4, and α7) [46].
The PeIA conotoxin can be made >15,000-fold more potent on α6/α3β2β3 vs. α3β2 nAChRs via substituting Asn 11 with either Lys or Arg [47]. On the contrary, PeIA-4566, which is a synthetic peptide composed of non-natural amino acids, targets α3β2 over α6/α3β2β3, with a 300-fold difference in potency [48]. V10A; E14N] analog blocks α6/α3β2β3 and α6/α3β4 nicotinic receptors with IC50 values of 223 pM and 65 nM, respectively [49].
Residues from Leu 5 to Pro 13 and Tyr 15 are important residues of PnIA, conferring its inhibitory effect on α7 nAChRs [50]. In addition, the PnIA [A10L] mutant is able to exclusively bind α7 nAChRs [51].
The RegIIA [N11A; N12A] analog selectively blocks α3β4 vs. α3β2 nAChRs, with a 1000-fold difference in potency [52].
TxIB is unique due to its ability to selectively bind α6/ α3β2β3 nAChRs, while lacking any capability to bind to other receptor subtypes [53]. However, structure–functional studies accounting for this have not been conducted yet.
It is generally accepted that α-Ctxs are antagonists of nAChRs, but MrIC (PECCTHPACHVSNPELC), which is a peptide variant of Mr1.7, plays a full agonist role of endogenous human α7 nAChRs in the presence of PNU (positive allosteric modulator of α7 nAChR subtype) [54]. It is suggested that the Pro residue in the N-terminus of the peptide has a role in binding α7 nAChR [54].
It was found that Asp 5 to Arg 7, and Asp 11 to Ile 15, are critical residues for the inhibitory effect of Vc1.1 at the α9α10 nAChR subtype [55]. In addition, mutations at positions 4 and 9 (substitution of Ser 4 with either Lys or Arg and substitution of Asn 9 with either Ala, Leu, or Ile) increased the potency of Vc1.1 at α9α10 nAChRs [55].
Due to the fact that nAChRs are implicated in a range of pathological conditions, α-conotoxins targeting nAChRs pose themselves as attractive candidates for drug development For example, α-conotoxins RgIA and Vc1.1 demonstrate the analgesic effect in animal models of neuropathic pain [56][57][58][59][60][61][62][63]. Such analgesic effect works through targeting α9α10 The mechanism of analgesia mediated by RgIA and Vc1.1 conotoxins is not currently clear, but there are indications that it involves immune-mediated pathways.
RgIA was shown to be potent in reducing mechanical allodynia and oxaliplatin-induced hypersensitivity to thermal and mechanical stimuli [64], as well as accelerating the recovery of nerve damage following chronic constriction injury (CCI) [57]. Other than alleviating neuropathic pain, RgIA was also shown to be able to reduce the severity of dextran sodium sulfate-induced colitis in an animal model [65].
Vc1.1 was shown to reduce mechanical allodynia in rats with diabetic neuropathy induced by streptozotocin, and in rats with CCI [56][61]. Vc1.1 reduced mechanical hyperalgesia in rats with peripheral nerve ligation when the toxin was injected intramuscularly [62].
Another α-conotoxin with a potential therapeutic application due to its antitumor activity is TxID. The toxin inhibited the growth of lung cancer cells [66].
The results from in vitro and animal studies serve as an encouragement for drug development. There are, however, hindrances along the way [67], which stem from the peptidic nature of the toxins, the potential off-target effects, and different potencies on human receptors vs. rodent ones. For example, Vc1.1 was discontinued from phase II human clinical trials due to its low affinity on human α9α10 nAChRs [68]
The α-conotoxins mentioned in the main text are shown, together with their targets and potencies.
References
- Scholze, P.; Huck, S. The A5 Nicotinic Acetylcholine Receptor Subunit Differentially Modulates A4β2* and A3β4* Receptors. Front. Synaptic Neurosci. 2020, 12, 54.
- Boorman, J.P.; Beato, M.; Groot-Kormelink, P.J.; Broadbent, S.D.; Sivilotti, L.G. The Effects of Β3 Subunit Incorporation on the Pharmacology and Single Channel Properties of Oocyte-Expressed Human A3β4 Neuronal Nicotinic Receptors. J. Biol. Chem. 2003, 278, 44033–44040.
- Gharpure, A.; Noviello, C.M.; Hibbs, R.E. Progress in Nicotinic Receptor Structural Biology. Neuropharmacology 2020, 171, 108086.
- Karlin, A. A Touching Picture of Nicotinic Binding. Neuron 2004, 41, 841–842.
- Govind, A.P.; Vezina, P.; Green, W.N. Nicotine-Induced Upregulation of Nicotinic Receptors: Underlying Mechanisms and Relevance to Nicotine Addiction. Biochem. Pharmacol. 2009, 78, 756–765.
- Carbone, A.-L.; Moroni, M.; Groot-Kormelink, P.-J.; Bermudez, I. Pentameric Concatenated (A4)2(Β2)3 and (A4)3(Β2)2 Nicotinic Acetylcholine Receptors: Subunit Arrangement Determines Functional Expression. Br. J. Pharmacol. 2009, 156, 970–981.
- McGranahan, T.M.; Patzlaff, N.E.; Grady, S.R.; Heinemann, S.F.; Booker, T.K. A4β2 Nicotinic Acetylcholine Receptors on Dopaminergic Neurons Mediate Nicotine Reward and Anxiety Relief. J. Neurosci. 2011, 31, 10891.
- Winek, K.; Soreq, H.; Meisel, A. Regulators of Cholinergic Signaling in Disorders of the Central Nervous System. J. Neurochem. 2021.
- Dineley, K.T.; Pandya, A.A.; Yakel, J.L. Nicotinic ACh Receptors as Therapeutic Targets in CNS Disorders. Trends Pharmacol. Sci. 2015, 36, 96–108.
- Quik, M.; Zhang, D.; McGregor, M.; Bordia, T. Alpha7 Nicotinic Receptors as Therapeutic Targets for Parkinson’s Disease. Biochem. Pharmacol. 2015, 97, 399–407.
- Pennington, M.W.; Czerwinski, A.; Norton, R.S. Peptide Therapeutics from Venom: Current Status and Potential. Bioorgan. Med. Chem. 2018, 26, 2738–2758.
- Jin, A.-H.; Muttenthaler, M.; Dutertre, S.; Himaya, S.W.A.; Kaas, Q.; Craik, D.J.; Lewis, R.J.; Alewood, P.F. Conotoxins: Chemistry and Biology. Chem. Rev. 2019, 119, 11510–11549.
- McIntosh, J.M.; Santos, A.D.; Olivera, B.M. Conus Peptides Targeted to Specific Nicotinic Acetylcholine Receptor Subtypes. Annu. Rev. Biochem. 1999, 68, 59–88.
- Jacobsen, R.B.; DelaCruz, R.G.; Grose, J.H.; McIntosh, J.M.; Yoshikami, D.; Olivera, B.M. Critical Residues Influence the Affinity and Selectivity of α-Conotoxin MI for Nicotinic Acetylcholine Receptors. Biochemistry 1999, 38, 13310–13315.
- Liu, L.; Chew, G.; Hawrot, E.; Chi, C.; Wang, C. Two Potent A3/5 Conotoxins from Piscivorous Conus Achatinus. Acta Biochim. Biophys. Sin. 2007, 39, 438–444.
- Groebe, D.R.; Dumm, J.M.; Levitan, E.S.; Abramson, S.N. Alpha-Conotoxins Selectively Inhibit One of the Two Acetylcholine Binding Sites of Nicotinic Receptors. Mol. Pharmacol. 1995, 48, 105.
- Groebe, D.R.; Gray, W.R.; Abramson, S.N. Determinants Involved in the Affinity of α-Conotoxins GI and SI for the Muscle Subtype of Nicotinic Acetylcholine Receptors. Biochemistry 1997, 36, 6469–6474.
- Ning, J.; Li, R.; Ren, J.; Zhangsun, D.; Zhu, X.; Wu, Y.; Luo, S. Alanine-Scanning Mutagenesis of α-Conotoxin GI Reveals the Residues Crucial for Activity at the Muscle Acetylcholine Receptor. Mar. Drugs 2018, 16, 507.
- Giribaldi, J.; Wilson, D.; Nicke, A.; El Hamdaoui, Y.; Laconde, G.; Faucherre, A.; Moha Ou Maati, H.; Daly, N.L.; Enjalbal, C.; Dutertre, S. Synthesis, Structure and Biological Activity of CIA and CIB, Two α-Conotoxins from the Predation-Evoked Venom of Conus Catus. Toxins 2018, 10, 222.
- Ramilo, C.A.; Zafaralla, G.C.; Nadasdi, L.; Hammerland, L.G.; Yoshikami, D.; Gray, W.R.; Kristipati, R.; Ramachandran, J.; Miljanich, G. Novel .Alpha.- and .Omega.-Conotoxins and Conus Striatus Venom. Biochemistry 1992, 31, 9919–9926.
- Quiram, P.A.; Sine, S.M. Structural Elements in α-Conotoxin ImI Essential for Binding to Neuronal A7 Receptors. J. Biol. Chem. 1998, 273, 11007–11011.
- Ellison, M.; McIntosh, J.M.; Olivera, B.M. α-Conotoxins ImI and ImII: Similar A7 nicotinic receptor antagonists act at different sites. J. Biol. Chem. 2003, 278, 757–764.
- Ellison, M.; Feng, Z.-P.; Park, A.J.; Zhang, X.; Olivera, B.M.; McIntosh, J.M.; Norton, R.S. α-RgIA, a Novel Conotoxin That Blocks the A9α10 NAChR: Structure and Identification of Key Receptor-Binding Residues. J. Mol. Biol. 2008, 377, 1216–1227.
- Millar, N.S.; Gotti, C. Diversity of Vertebrate Nicotinic Acetylcholine Receptors. Neuropharmacology 2009, 56, 237–246.
- Chhabra, S.; Belgi, A.; Bartels, P.; van Lierop, B.J.; Robinson, S.D.; Kompella, S.N.; Hung, A.; Callaghan, B.P.; Adams, D.J.; Robinson, A.J.; et al. Dicarba Analogues of α-Conotoxin RgIA. Structure, Stability, and Activity at Potential Pain Targets. J. Med. Chem. 2014, 57, 9933–9944.
- Azam, L.; Dowell, C.; Watkins, M.; Stitzel, J.A.; Olivera, B.M.; McIntosh, J.M. α-Conotoxin BuIA, a Novel Peptide from Conus Bullatus, Distinguishes among Neuronal Nicotinic Acetylcholine Receptors. J. Biol. Chem. 2005, 280, 80–87.
- Kim, H.-W.; McIntosh, J.M. A6 NAChR Subunit Residues That Confer α-Conotoxin BuIA Selectivity. FASEB J. 2012, 26, 4102–4110.
- Azam, L.; Maskos, U.; Changeux, J.-P.; Dowell, C.D.; Christensen, S.; De Biasi, M.; McIntosh, J.M. α-Conotoxin BuIA[T5A;P6O]: A Novel Ligand That Discriminates between A6ß4 and A6ß2 Nicotinic Acetylcholine Receptors and Blocks Nicotine-Stimulated Norepinephrine Release. FASEB J. 2010, 24, 5113–5123.
- Quinton, L.; Servent, D.; Girard, E.; Molgó, J.; Le Caer, J.-P.; Malosse, C.; Haidar, E.A.; Lecoq, A.; Gilles, N.; Chamot-Rooke, J. Identification and Functional Characterization of a Novel α-Conotoxin (EIIA) from Conus Ermineus. Anal. Bioanal. Chem. 2013, 405, 5341–5351.
- Grishin, A.A.; Cuny, H.; Hung, A.; Clark, R.J.; Brust, A.; Akondi, K.; Alewood, P.F.; Craik, D.J.; Adams, D.J. Identifying Key Amino Acid Residues That Affect α-Conotoxin AuIB Inhibition of A3β4 Nicotinic Acetylcholine Receptors. J. Biol. Chem. 2013, 288, 34428–34442.
- Li, L.; Liu, N.; Ding, R.; Wang, S.; Liu, Z.; Li, H.; Zheng, X.; Dai, Q. A Novel 4/6-Type Alpha-Conotoxin ViIA Selectively Inhibits NAchR A3β2 Subtype. Acta Biochim. Biophys. Sin. 2015, 47, 1023–1028.
- van Hout, M.; Valdes, A.; Christensen, S.B.; Tran, P.T.; Watkins, M.; Gajewiak, J.; Jensen, A.A.; Olivera, B.M.; McIntosh, J.M. α-Conotoxin VnIB from Conus Ventricosus Is a Potent and Selective Antagonist of A6β4* Nicotinic Acetylcholine Receptors. Neuropharmacology 2019, 157, 107691.
- Wu, Y.; Zhangsun, D.; Zhu, X.; Kaas, Q.; Zhangsun, M.; Harvey, P.J.; Craik, D.J.; McIntosh, J.M.; Luo, S. α-Conotoxin [S9A]TxID Potently Discriminates between A3β4 and A6/A3β4 Nicotinic Acetylcholine Receptors. J. Med. Chem. 2017, 60, 5826–5833.
- Loughnan, M.L.; Nicke, A.; Jones, A.; Adams, D.J.; Alewood, P.F.; Lewis, R.J. Chemical and Functional Identification and Characterization of Novel Sulfated α-Conotoxins from the Cone Snail Conus Anemone. J. Med. Chem. 2004, 47, 1234–1241.
- Whiteaker, P.; Christensen, S.; Yoshikami, D.; Dowell, C.; Watkins, M.; Gulyas, J.; Rivier, J.; Olivera, B.M.; McIntosh, J.M. Discovery, Synthesis, and Structure Activity of a Highly Selective A7 Nicotinic Acetylcholine Receptor Antagonist. Biochemistry 2007, 46, 6628–6638.
- Ning, J.; Ren, J.; Xiong, Y.; Wu, Y.; Zhangsun, M.; Zhangsun, D.; Zhu, X.; Luo, S. Identification of Crucial Residues in α-Conotoxin EI Inhibiting Muscle Nicotinic Acetylcholine Receptor. Toxins 2019, 11, 603.
- Lin, B.; Xu, M.; Zhu, X.; Wu, Y.; Liu, X.; Zhangsun, D.; Hu, Y.; Xiang, S.-H.; Kasheverov, I.E.; Tsetlin, V.I.; et al. From Crystal Structure of α-Conotoxin GIC in Complex with Ac-AChBP to Molecular Determinants of Its High Selectivity for A3β2 NAChR. Sci. Rep. 2016, 6, 22349.
- Banerjee, J.; Yongye, A.B.; Chang, Y.-P.; Gyanda, R.; Medina-Franco, J.L.; Armishaw, C.J. Design and Synthesis of α-Conotoxin GID Analogues as Selective A4β2 Nicotinic Acetylcholine Receptor Antagonists. Biopolymers 2014, 102, 78–87.
- Lebbe, E.K.M.; Peigneur, S.; Maiti, M.; Devi, P.; Ravichandran, S.; Lescrinier, E.; Ulens, C.; Waelkens, E.; D’Souza, L.; Herdewijn, P.; et al. Structure-Function Elucidation of a New α-Conotoxin, Lo1a, from Conus Longurionis. J. Biol. Chem. 2014, 289, 9573–9583.
- Inserra, M.C.; Kompella, S.N.; Vetter, I.; Brust, A.; Daly, N.L.; Cuny, H.; Craik, D.J.; Alewood, P.F.; Adams, D.J.; Lewis, R.J. Isolation and Characterization of α-Conotoxin LsIA with Potent Activity at Nicotinic Acetylcholine Receptors. Biochem. Pharmacol. 2013, 86, 791–799.
- Chen, J.; Liang, L.; Ning, H.; Cai, F.; Liu, Z.; Zhang, L.; Zhou, L.; Dai, Q. Cloning, Synthesis and Functional Characterization of a Novel α-Conotoxin Lt1.3. Mar. Drugs 2018, 16, 112.
- Luo, S.; Zhangsun, D.; Schroeder, C.I.; Zhu, X.; Hu, Y.; Wu, Y.; Weltzin, M.M.; Eberhard, S.; Kaas, Q.; Craik, D.J.; et al. A Novel A4/7-Conotoxin LvIA from Conus Lividus That Selectively Blocks A3β2 vs. A6/A3β2β3 Nicotinic Acetylcholine Receptors. FASEB J. 2014, 28, 1842–1853.
- Xu, M.; Zhu, X.; Yu, J.; Yu, J.; Luo, S.; Wang, X. The Crystal Structure of Ac-AChBP in Complex with α-Conotoxin LvIA Reveals the Mechanism of Its Selectivity towards Different NAChR Subtypes. Protein Cell 2017, 8, 675–685.
- Everhart, D.; Cartier, G.E.; Malhotra, A.; Gomes, A.V.; McIntosh, J.M.; Luetje, C.W. Determinants of Potency on Alpha-Conotoxin MII, a Peptide Antagonist of Neuronal Nicotinic Receptors. Biochemistry 2004, 43, 2732–2737.
- McIntosh, J.M.; Azam, L.; Staheli, S.; Dowell, C.; Lindstrom, J.M.; Kuryatov, A.; Garrett, J.E.; Marks, M.J.; Whiteaker, P. Analogs of Alpha-Conotoxin MII Are Selective for Alpha6-Containing Nicotinic Acetylcholine Receptors. Mol. Pharmacol. 2004, 65, 944–952.
- Wang, S.; Zhao, C.; Liu, Z.; Wang, X.; Liu, N.; Du, W.; Dai, Q. Structural and Functional Characterization of a Novel α-Conotoxin Mr1.7 from Conus Marmoreus Targeting Neuronal NAChR A3β2, A9α10 and A6/A3β2β3 Subtypes. Mar. Drugs 2015, 13, 3259–3275.
- Hone, A.J.; Ruiz, M.; Scadden, M.; Christensen, S.; Gajewiak, J.; Azam, L.; McIntosh, J.M. Positional Scanning Mutagenesis of α-Conotoxin PeIA Identifies Critical Residues That Confer Potency and Selectivity for A6/A3β2β3 and A3β2 Nicotinic Acetylcholine Receptors. J. Biol. Chem. 2013, 288, 25428–25439.
- Hone, A.J.; Fisher, F.; Christensen, S.; Gajewiak, J.; Larkin, D.; Whiteaker, P.; McIntosh, J.M. PeIA-5466: A Novel Peptide Antagonist Containing Non-Natural Amino Acids That Selectively Targets A3β2 Nicotinic Acetylcholine Receptors. J. Med. Chem. 2019, 62, 6262–6275.
- Hone, A.J.; Scadden, M.; Gajewiak, J.; Christensen, S.; Lindstrom, J.; McIntosh, J.M. α-Conotoxin PeIA[S9H,V10A,E14N] Potently and Selectively Blocks A6β2β3 versus A6β4 Nicotinic Acetylcholine Receptors. Mol. Pharmacol. 2012, 82, 972–982.
- Luo, S.; Nguyen, T.A.; Cartier, G.E.; Olivera, B.M.; Yoshikami, D.; McIntosh, J.M. Single-Residue Alteration in α-Conotoxin PnIA Switches Its NAChR Subtype Selectivity. Biochemistry 1999, 38, 14542–14548.
- Hogg, R.C.; Hopping, G.; Alewood, P.F.; Adams, D.J.; Bertrand, D. Alpha-Conotoxins PnIA and [A10L]PnIA Stabilize Different States of the Alpha7-L247T Nicotinic Acetylcholine Receptor. J. Biol. Chem. 2003, 278, 26908–26914.
- Kompella, S.N.; Hung, A.; Clark, R.J.; Marí, F.; Adams, D.J. Alanine Scan of α-Conotoxin RegIIA Reveals a Selective A3β4 Nicotinic Acetylcholine Receptor Antagonist. J. Biol. Chem. 2015, 290, 1039–1048.
- Luo, S.; Zhangsun, D.; Zhu, X.; Wu, Y.; Hu, Y.; Christensen, S.; Harvey, P.J.; Akcan, M.; Craik, D.J.; McIntosh, J.M. Characterization of a Novel α-Conotoxin TxID from Conus Textile That Potently Blocks Rat A3β4 Nicotinic Acetylcholine Receptors. J. Med. Chem. 2013, 56, 9655–9663.
- Jin, A.-H.; Vetter, I.; Dutertre, S.; Abraham, N.; Emidio, N.B.; Inserra, M.; Murali, S.S.; Christie, M.J.; Alewood, P.F.; Lewis, R.J. MrIC, a Novel α-Conotoxin Agonist in the Presence of PNU at Endogenous A7 Nicotinic Acetylcholine Receptors. Biochemistry 2014, 53, 1–3.
- Halai, R.; Clark, R.J.; Nevin, S.T.; Jensen, J.E.; Adams, D.J.; Craik, D.J. Scanning Mutagenesis of Alpha-Conotoxin Vc1.1 Reveals Residues Crucial for Activity at the Alpha9alpha10 Nicotinic Acetylcholine Receptor. J. Biol. Chem. 2009, 284, 20275–20284.
- McIntosh, J.M.; Absalom, N.; Chebib, M.; Elgoyhen, A.B.; Vincler, M. Alpha9 Nicotinic Acetylcholine Receptors and the Treatment of Pain. Biochem. Pharmacol. 2009, 78, 693–702.
- Di Cesare Mannelli, L.; Cinci, L.; Micheli, L.; Zanardelli, M.; Pacini, A.; McIntosh, J.M.; Ghelardini, C. α-Conotoxin RgIA Protects against the Development of Nerve Injury-Induced Chronic Pain and Prevents Both Neuronal and Glial Derangement. Pain 2014, 155, 1986–1995.
- Hone, A.J.; Servent, D.; McIntosh, J.M. A9-Containing Nicotinic Acetylcholine Receptors and the Modulation of Pain. Br. J. Pharmacol. 2018, 175, 1915–1927.
- Mohammadi, S.; Christie, M.J. A9-Nicotinic Acetylcholine Receptors Contribute to the Maintenance of Chronic Mechanical Hyperalgesia, but Not Thermal or Mechanical Allodynia. Mol. Pain 2014, 10, 1744–8069.
- Mohammadi, S.A.; Christie, M.J. Conotoxin Interactions with A9α10-NAChRs: Is the A9α10-Nicotinic Acetylcholine Receptor an Important Therapeutic Target for Pain Management? Toxins 2015, 7, 3916–3932.
- Vincler, M.; McIntosh, J.M. Targeting the A9α10 Nicotinic Acetylcholine Receptor to Treat Severe Pain. Expert Opin. Ther. Targets 2007, 11, 891–897.
- Satkunanathan, N.; Livett, B.; Gayler, K.; Sandall, D.; Down, J.; Khalil, Z. Alpha-Conotoxin Vc1.1 Alleviates Neuropathic Pain and Accelerates Functional Recovery of Injured Neurones. Brain Res. 2005, 1059, 149–158.
- Romero, H.K.; Christensen, S.B.; Di Cesare Mannelli, L.; Gajewiak, J.; Ramachandra, R.; Elmslie, K.S.; Vetter, D.E.; Ghelardini, C.; Iadonato, S.P.; Mercado, J.L.; et al. Inhibition of A9α10 Nicotinic Acetylcholine Receptors Prevents Chemotherapy-Induced Neuropathic Pain. Proc. Natl. Acad. Sci. USA 2017, 114, E1825–E1832.
- Christensen, S.B.; Hone, A.J.; Roux, I.; Kniazeff, J.; Pin, J.-P.; Upert, G.; Servent, D.; Glowatzki, E.; McIntosh, J.M. RgIA4 Potently Blocks Mouse A9α10 NAChRs and Provides Long Lasting Protection against Oxaliplatin-Induced Cold Allodynia. Front. Cell. Neurosci. 2017, 11, 219.
- AlSharari, S.D.; Toma, W.; Mahmood, H.M.; Michael McIntosh, J.; Imad Damaj, M. The A9α10 Nicotinic Acetylcholine Receptors Antagonist α-Conotoxin RgIA Reverses Colitis Signs in Murine Dextran Sodium Sulfate Model. Eur. J. Pharmacol. 2020, 883, 173320.
- Qian, J.; Liu, Y.-Q.; Sun, Z.-H.; Zhangsun, D.-T.; Luo, S.-L. Identification of Nicotinic Acetylcholine Receptor Subunits in Different Lung Cancer Cell Lines and the Inhibitory Effect of Alpha-Conotoxin TxID on Lung Cancer Cell Growth. Eur. J. Pharmacol. 2019, 865, 172674.
- Bertrand, D.; Terry, A.V.J. The Wonderland of Neuronal Nicotinic Acetylcholine Receptors. Biochem. Pharmacol. 2018, 151, 214–225.
- Del Bufalo, A.; Cesario, A.; Salinaro, G.; Fini, M.; Russo, P. Alpha9 Alpha10 Nicotinic Acetylcholine Receptors as Target for the Treatment of Chronic Pain. Curr. Pharm. Des. 2014, 20, 6042–6047.





