Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Cucu, D. Exosomal microRNAs Fuctions in HCC. Encyclopedia. Available online: https://encyclopedia.pub/entry/10060 (accessed on 05 May 2026).
Cucu D. Exosomal microRNAs Fuctions in HCC. Encyclopedia. Available at: https://encyclopedia.pub/entry/10060. Accessed May 05, 2026.
Cucu, Dana. "Exosomal microRNAs Fuctions in HCC" Encyclopedia, https://encyclopedia.pub/entry/10060 (accessed May 05, 2026).
Cucu, D. (2021, May 25). Exosomal microRNAs Fuctions in HCC. In Encyclopedia. https://encyclopedia.pub/entry/10060
Cucu, Dana. "Exosomal microRNAs Fuctions in HCC." Encyclopedia. Web. 25 May, 2021.
Copy Citation
Hepatocellular carcinoma (HCC) is the fifth most common cancer worldwide and the second most common cause of cancer-related death globally. This type of liver cancer is frequently detected at a late stage by current biomarkers because of the high clinical and biological heterogeneity of HCC tumours. From a plethora of molecules and cellular compounds, small nanoparticles with an endosomal origin are valuable cancer biomarkers or cargos for novel treatments. Despite their small sizes, in the range of 40–150 nm, these particles are delimited by a lipid bilayer membrane with a specific lipid composition and carry functional information—RNA, proteins, miRNAs, long non-coding RNAs (lncRNAs), or DNA fragments.
hepatocellular carcinoma
exosomes
miRNA
1. Introduction
Hepatocellular carcinoma (HCC), also known as primary liver cancer, is one of the most severe malignancies worldwide. Recently, the Global Cancer Observatory (GLOBOCAN) reports 840,000 HCC cases and 780,000 individuals per year diagnosed with HCC [1]. Risk factors for HCC include age, hepatitis B or C, cirrhosis, genetic disorders, contaminants, excessive drinking, and smoking. Recently, non-alcoholic fatty liver disease (NAFLD), and its severe form, non-alcoholic steatohepatitis (NASH), were identified as major causes of HCC in western countries, and as being among the main indications for liver transplant after HCV infection [2][3]. NAFLD has been associated with a 2.6-fold increased risk of HCC occurrence [4][5]. Similarly, a study by the US Veterans Health Administration on a group of 1500 patients with HCC showed that HCC patients with NAFLD had a five-fold higher risk of developing HCC in the absence of cirrhosis, compared with HCC and HCV patients [6].
Currently, the first-line treatment for small (≤2 cm) HCC tumours is radiofrequency ablation (RFA), which results in reduced disruption and rapid healing, and has comparable effects to surgical resection. An alternative first-line therapy for intermediate-stage patients is transcatheter arterial chemoembolization (TACE), but due to the compensatory effects of vascular proliferation after hypoxia, the treatment efficacy remains unsatisfactory [3]. Sorafenib, the main therapeutic agent for advanced HCC, can substantially extend the average survival time of patients, but it can also enhance tumour progression by inducing drug resistance [7].
The progression of HCC involves the dynamic tumour microenvironment—consisting of fibroblasts, endothelial cells, cancer stem cells, myeloid cells, and related soluble cytokines—impeding early diagnosis of the disease. Therefore, biomarkers and HCC targets are continuously investigated in order to address these unmet needs in clinical practice.
To date, as a non-invasive biomarker in the diagnosis of HCC, clinicians use α-fetoprotein (AFP) most frequently; des-gamma-carboxy prothrombin (DCP) and glypican-3 (GPC3) also serve as non-invasive biomarkers. However, clinical practice revealed that AFP has low sensitivity and specificity for early-stage HCC, as around 50% of patients exhibit negative expression [8]. Better alternatives are imaging procedures, such as magnetic resonance imaging (MRI) and computed tomography (CT), although small tumours cannot be identified using these methods [9]. Histopathological biopsies would complete these diagnosis tools, but they are limited by their invasiveness and the high negative rate of their results [10].
In the last decade, extensive data have described small (nm) extracellular vesicles (EVs) named exosomes as carriers of various molecules—especially microRNAs (miRNA)—and important players in immune and cell–cell communication processes. Hepatic cells generate exosomes in the liver identified in parenchymal cells (hepatocytes), non-parenchymal immune cells (macrophages, dendritic, and T/B natural killer (NK) cells), and non-parenchymal hepatic stromal cells (stellate cells) [11]. The hepatitis B virus (HBV) and hepatitis C virus (HCV) use exosomes to spread viral RNA complexes to neighbouring human liver cells. Furthermore, these particles stimulate nonspecific immune responses through plasmacytoid dendritic cells (PDCs), and suppress unique immune responses by T cells [12]. Due to their creative mechanism of transmitting effector molecules and signals between cells—including RNA, proteins, miRNAs, long non-coding RNAs (lncRNAs), and DNA fragments [13]—exosome EVs have attracted the interest of the scientific community as putative biomarkers, as well as possible delivery vehicles in HCC.
Because the specific isolation of exosomes is challenging, the term exosomes has been indiscriminately used in original and review articles for a heterogeneous population of extracellular vesicles that includes exosomes and other type of vesicles. However, the most consistent results present exosomes as small noncoding RNAs with 19–22 nucleotides and vehicles for miRNAs. We base this review on reports that specify the exosomal nature of miRNAs, and we will use the term hereafter. The miRNAs bind to the 3′-UTR of pre-mRNAs during the post-transcriptional process, and directly target mRNAs for degradation. Today, modern next-generation sequencing and microarray technologies frequently identify miRNAs in the diagnosis of malignant tumours. Several reports of exosomal miRNAs have recently indicated that these particles are better biomarkers for the diagnosis and treatment of HCC than their serum-free-derived counterparts [14]. Exosomal miRNAs exhibit long-term survival due to their better stability in exosomes than in circulating cell-free miRNAs [15]. While exosomes have been explored for several years, we have only recently begun to understand the biological functions of exosomal miRNAs, particularly in HCC.
2. Biological Characteristics of Exosomes
2.1. Exosome Biogenesis and Secretion
Cells release different types of extracellular vesicles (EVs)—including exosomes, microvesicles, apoptotic bodies, ectosomes, and membrane particles—which can be isolated from bodily fluids and from the culture media of the secreting cells [16][17]. According to their biogenesis and size, EVs are divided into three main categories: microvesicles/ectosomes (100 nm–1 µm), released through the plasma membrane; exosomes (30–150 nm), derived from multivesicular bodies (MVBs); and apoptotic bodies (100 nm–5 µm), resulting from cellular apoptosis [18].
Exosomes originating from intracellular endosomes are heterogeneous intraluminal vesicles (ILVs) secreted into the extracellular space. The process of exosome biogenesis has 3 stages: (1) the formation of endocytic vesicles from the plasma membrane; (2) the inward budding of membrane endosomal vesicles, resulting in MVBs; and (3) MVB fusion with the plasma membrane and exosome release (Figure 1).
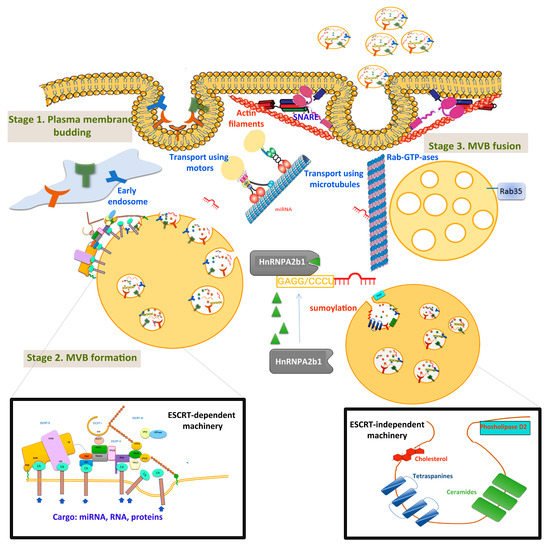
Figure 1. Extracellular vesicles’ biogenesis, transport, and membrane fusion. The first stage, membrane budding, integrates receptors that are transferred into early endosomes. MVB formation (stage 2) and sorting take place in the late exosome. Detailed ESCRT-dependent and ESCRT-independent pathways are presented in insets. MVBs are transported via microtubules and motor proteins, and fuse with the cellular membrane using SNARE, synaptotagmin proteins, and actin filaments. The pathway proposed for miRNA sorting using sumoylated HnNPA2b1 protein is also illustrated.
Briefly, in the first stage, the plasma membrane budding creates the early endosome, and then the endocytic vesicles mature into a late endosome.
In the second stage, membrane inward budding forms intraluminal vesicles (ILVs), which accumulate in the late endosomes and became MVBs. Two pathways mediate MVB formation: the ESCRT (endosomal sorting complexes required for transport)-dependent and ESCRT-independent pathways. The ESCRT consists of four soluble multiprotein complexes—ESCRT-0, ESCRT-I, ESCRT-II, and ESCRT-III—and the VPS4 (vacuolar protein sorting-associated protein 4) complex. Special attention was given to the molecular mechanisms of ESCRT-III/VPS4-dependent membrane remodelling, important in sending membrane budding away from the cytoplasm [19]. Particularly in HCC tissues, one group reported significant downregulation of VPS4A protein expression. This profile is associated with TNM stage, tumour size, tumour capsule integrity, and regional lymph node metastasis [20]. Ectopic overexpression of VPS4A repressed the growth, colony formation, migration, and invasion of HCC cells, whereas the regulation of VPS4A altered the release and uptake of exosomal miRNAs in HCC cells.
ESCRT-independent mechanisms for the formation of ILVs, MVBs, and exosomes include ceramides, cholesterol, D2 phospholipases, and tetraspanins [21][22]. MVBs can be either degraded in lysosomes or released into the extracellular space as exosomes. In the latter case, MVBs move to the cell membrane using cytoskeletal proteins (such as actin and microtubules) [23], motor proteins (dynein, quinine, and myosin) [24], and molecular transducers (some GTPases) [25].
Much effort has been put to unravel the mechanisms of sorting proteins, miRNAs, and other cargos into ILVs, and the process is still not fully understood. A proposed mechanism is sumoylated ribonucleoprotein (HnRNPA2b1), which specifically recognises and internalizes miRNAs in exosomes through binding to specific short motifs [26].
Finally, in the third stage, MVBs fuse with the cell membrane, releasing exosomes into the extracellular space. This process is regulated by the SNARE (soluble NSF-attachment protein receptors) protein complex [27], syntaxin 1A [28], the synaptotagmin protein family, Wnt proteins [29], and Ca2+ ions [30]. Rab GTPases—such as Rab7, Rab27a/b, Rab11, and Rab35—mediate the anchoring, docking, and secretion of MVBs to the cell membrane [31][32]. Particularly, Rab GTPases contribute to the secretion of exosomes in HCC cells. One study used HOX transcript antisense RNA (HOTAIR)—a 2158 nucleotide lncRNA transcribed from the HOXC locus [33]. Over-expressed HOTAIR in HCC cells upregulated Rab35 and facilitated the transport of MVBs to the plasma membrane.
2.2. Exosome Composition
The nanospheric membrane structure of exosomes resembles that of their parental cells. Some databases (ExoCarta, Vesiclepedia, and EVpedia) contain information about EV composition and procedures for the isolation and purification of these vesicles. The continuous updating of these databases makes them crucial tools for improving the understanding of EVs. ExoCarta’s database gathers the results of exosome composition [36], while Vesiclepedia is more inclusive and provides important information on all categories of extracellular vesicles, constantly updated by the scientific community [37]. EVpedia, a community web portal, includes studies of non-mammalian EVs of different sizes [38]. According to ExoCarta, more than 8000 proteins and 194 lipids are now associated with exosomes [36].
The lipid bilayer resembles the plasma membrane of the cell of origin, containing different types of lipid complexes: sphingomyelin, phosphatidylcholine, phosphatidylethanolamine, phosphatidylserine, monosialotetrahexosylganglioside (GM3), and phosphatidylinositol [39]. Sphingomyelin and GM3 are responsible for the exosome’s rigidity, while phosphatidylserine is expressed on the exosome membrane by different types of phospholipid transport enzymes [40]. Other lipids that support exosome biogenesis are cholesterol, ceramides, and phosphoglycerides.
The exosomal membrane contains the following proteins: integrins, differentiation clusters (CD), tetraspanins, and even the major histocompatibility complex (MHC)-II. Like their parental cells, exosomes contain intracellular proteins, such as the heat shock proteins (Hsp70 and Hsp90), fusion and membrane transport proteins (GTPases, annexins, and flotillins) and tetraspanins (CD9, CD63, CD81, and CD82). These proteins facilitate fusion, cell migration, cell–cell adhesion, and cellular signalling. Adhesion molecules such as integrins mediate cellular binding to the extracellular matrix and to the targeted cells [41]. Exosome-associated proteins include thrombospondin, CD55, CD59, lactadherin, ALIX (ALG-2-interacting protein X), and TSG101 [42], which are incorporated into exosomes during their biogenesis and serve as cargos for cell–cell communication.
Exosomes have tissue-specific molecular content, including DNA, mRNA, miRNA, HSP, and Ras proteins (Rab) [43].
The miRNAs from the cells of origin are to date the most studied constituents as biomarkers. One of the reasons for this is that in different cell types, miRNA species are significantly more abundant in exosomes than in the cell of origin [44]. These findings point toward a specific sorting mechanism of cargo molecules into the exosomes.
Using microarray experiments, previous reports have shown that exosomes derived from mast cells contain a unique set of 1300 mRNAs and 100 miRNAs absent from the cell of origin. In this study, mRNA transcripts from exosomes were translated into the recipient cell [45]. Other reports demonstrated that B cells infected with the Epstein–Barr virus (EBV) secrete mature EBV-miRNA via exosomes. Functional EBV-miRNAs such as BHRF1 and BART are transported to uninfected primary immature monocyte-derived dendritic cells (MoDCs). In these recipient cells, miRNAs trigger functional translational repression of the Epstein–Barr virus latent membrane protein 1 (LMP1) and C-X-C motif chemokine 1 (CXCL11/ITAC) [46].
The calcium-dependent phospholipid-binding proteins (e.g., annexins) play equally important roles in the regulation of RNA loading in exosomes. Annexin-2, the most abundant among annexins, recognizes the 3’-UTR sequence elements of the MYC gene, and participates in miRNA packaging in exosomes [47][48][49]. Specific exosomal proteins—such as ALIX, TSG101, flotillin 1, HSP70, CD9, CD81, and CD63—contribute to the protein profile characterization and identification of exosomes obtained through various isolation methods [34].
Exosomes are difficult to isolate and characterize because of their small size, which technically impedes scientists from distinguishing them from other cellular particles. In the following subsection, we review several techniques that are widely used, along with their advantages and disadvantages.
2.3. Exosome Isolation and Characterization
The most common exosome extraction method is ultracentrifugation, which allows the separation of large particles and cell debris using a centrifugal force of 200–100,000× g [50][51]. However, from the cell culture medium, researchers often extract cell aggregates and other particles [36]. Additionally, this procedure requires a long time, advanced equipment, and high sample volumes. To improve these shortcomings, exosomes are separated in a sucrose density gradient, or with optimal density gradient medium iodixanol solution (OptiPrep), leading to better purification and increased exosome volumes [52][53].
Other common approaches use monoclonal antibodies. Common choices are CD63, CD81, CD82, and CD9 antibodies, epithelial cell adhesion molecules (EpCAMs), and Ras-related protein 5 (Rab5a). These molecules are immobilized under different conditions and combined with magnetic beads and matrix chromatography. To separate them, scientists use plates and microfluidic devices [53]. The drawback of this strategy is that non-exosomal vesicles carrying the antigen also bind to the antibody, thereby reducing the purity of the isolated exosomes. Exosome isolation with ultrafiltration is an appropriate approach, shorter in time than ultracentrifugation, and without the need for specific equipment. Another method of exosome separation is high-efficiency liquid chromatography, which requires advanced technology and results in a sample with high purity [54].
In conclusion, techniques to isolate and purify exosomes are not rigorously adjusted. Therefore, the best approach is to combine isolation with an accurate method for particle characterization. One method is labelling exosome markers—including integrins, tetraspanins (CD81, CD9, and CD63), TSG101, ALIX, and cell adhesion molecules (integrins)—with specific antibodies. Lipid complexes are also sometimes used [43]. Flow cytometry, NTA (nanoparticle tracking analysis), DLS (dynamic light scattering), Western blot, mass spectrometry, and TEM (transmission electron microscopy) are also used in combination with those methods mentioned so far [55].
In conclusion, we cannot identify a gold standard technique for exosome isolation and purification. In the field, several methods employed in combination create a reliable picture of the presence of these particles in bodily fluids. While the isolation and characterization of exosomes are techniques commonly used for all cancers, the function of exosomes, especially miRNAs, is specific to each type of malignancy. The next subsections are dedicated solely to the role of miRNAs in HCC.
3. Exosomal miRNA Functions in HCC
3.1. Correlation of Exosomal miRNAs with the Clinicopathological Features and Prognosis of HCC
The current incidence and mortality rates of HCC tend to be high. The five-year survival rate for HCC-diagnosed patients is 6%. This unfavourable prognosis stems from the aggressive progression of tumours and high recurrence rates [56][57]. Therefore, early diagnosis is the most important prerequisite for the success of HCC treatment. Exosomes from various sources have different molecular expression profiles, important when comparing the exosome content of patients with those extracted from healthy individuals.
The population of exosomes is relatively simple and stable, transporting different functional molecules—such as proteins, miRNAs, mRNAs, and DNA—to the targeted cells through circulation. Out of all of the above, miRNA species are currently the most investigated molecules as putative biomarkers in HCC therapy, because of their specific signatures and correlation with the presence of disease or with clinical features such as tumour size, disease staging, overall survival, and disease recurrence. Analysis of exosomal miRNA species (mainly from serum) is useful for early detection and assessment of disease progression, without the need for tumour biopsy, which is a major advantage in the diagnosis of liver cancer.
Based on the published data, the role of miRNAs specific to exosomes can be divided into (1) miRNAs specific to HCC patients, (2) miRNAs that significantly correlate with the early stages of the disease, and (3) miRNAs that correlate with tumour staging. Changes in the expression levels of all of these miRNA species derived from exosomes have implications for the diagnosis of hepatic disorders.
Table 1 illustrates different miRNA species extracted from the serum of HCC patients and identified as being clinically significant. miRNA quantification was performed with qRT-PCR. As illustrated, species of miRNA are either over-expressed—as indicated for miR-224, miR-21, miR-210-3p, miR-93, miR-92b, miR-155, and miR-665—or under-expressed—as for miR-718, miR-744, miR-9-3p, and miR-125b—in HCC patients. To this end, we lack explanations about the high variation of miRNA species in HCC patients, which downsizes their specificity for this disease. The crucial message is that exosomal miRNAs are different or better expressed than those extracted from serum, and that some of them have expression correlated with tumour stage or clinical parameters.
Table 1. Exosomal miRNAs with clinical significance in HCC.
| miR Species | Expression Profile in HCC Down- or Upregulated | Exosome Isolation Methods | Groups and Sample Size | Normalized (Reference miRs/Internal Control) | Clinical Significance | References |
|---|---|---|---|---|---|---|
| miR-224 | Up | Total Exosome Isolation Kit | HCC (n = 89) and healthy controls (n = 50) | let-7i, let-7g, and let-7d | Detection, prognosis, recurrence, and therapuetic target | [58] |
| miR-21 | Total Exosome Isolation Reagent (serum) | HCC (n = 30), CHB (n = 30), and healthy controls (n = 30) | U6 | Detection and diagnosis | [59] | |
| miR-93 | Total Exosome Isolation kit | HCC (n = 85) and healthy controls (n = 23) | miR-16 | Detection, prognosis, and recurrence | [60] | |
| miR-1247-5p | Ultracentrifugation | HCC patients without lung metastasis (n = 90), HCC patients suffering lung metastasis (n = 20), and healthy controls (n = 25) | 18S | Detection, diagnosis, and therapuetic target | [61] | |
| miR-92b | ExoQuick Exosome Precipitation Solution | Non-HCC (n = 26), HCC no recurrence (n = 28), early HCC recurrence (n = 43), and late HCC recurrence (n = 22) | Synthetic C. elegans miR-39 | Detection, prognosis, and recurrence | [62] | |
| miR-210-3p | Ultracentrifugation | HCC (n = 104) and healthy controls (n = 60) | cel-miR-67 (NC67) | Detection, diagnosis, and therapuetic target | [63] | |
| miR-155 | ExoQuick Exosome Precipitation Solution (System Biosciences) | HCC (n = 40): high level miR-155 of HCC (n = 20) and low level miR-155 of HCC (n = 20) | Not mentioned | Prognosis and recurrence | [64] | |
| miR-665 | ExoQuick TM Kit (System Biosciences) | HCC (n = 30) and healthy controls (n = 10) | U6 | Detection and prognosis | [65] | |
| miR-718 | Down | Ultracentrifugation | Patients with HCC who underwent living donor liver transplantation (LDLT) (n = 59) | Synthetic C. elegans miR-39 | Detection, prognosis, recurrence, and therapeutic target | [66] |
| miR-122 | ExoQuick | Samples collected before and after TACE treatment of HCC patients (n = 75). | cel-miR-39 | Detection and prognosis | [67] | |
| miR-638 | Total Exosome Isolation kit | HCC (n = 126) and healthy controls (n = 21) | miR-16 | Detection, prognosis, and recurrence | [68] | |
| mi-125b | ExoQuick Exosome Precipitation Solution | Group 1: CHB (n = 30), LC (liver cirrhosis) (n = 30), and HCC (n = 30). Group 2: HCC (n = 128) | cel-miR-39 | Detection, prognosis, and recurrence | [69] | |
| miR-9-3p | Ultracentrifugation | HCC (n = 30) and healthy controls (n = 10) | not mentioned | Potential treatment/therapeutic target | [70] | |
| miR-744 | Ultracentrifugation | Group 1 (serum):HCC patients (n = 10) and healthy controls (n = 10). Group 2 (tissue): HCC patients (n = 68) and normal liver tissue (n = 52) | U6 | Detection, diagnosis, and therapeutic target | [71] |
The main endpoints in HCC investigation are overall survival (OS) and disease-free survival (DFS); therefore, a direct connection between exosomal miRNAs and patients’ outcomes would be a valuable asset for clinical research.
Table 2 summarizes statistical correlation between the expression of different miRNAs and clinical data. We selected the reports in which the expression levels of miRNAs were obtained following ROC curve analysis, and the diagnostic values were investigated using the area under the curve (AUC) and p-values.
Table 2. Exosomal miRNAs with statistically significant expression with respect to clinical data.
| No. | miR Species | Diagnostic Values (AUC Values, Area under the Curve) | Correlation with Prognosis and/or with Clinicopathological Features (p Values) | References | |||
|---|---|---|---|---|---|---|---|
| AUC Values (>0.7) | p Values | Large Tumour Size (>3 cm or 5 cm) | Advanced Tumour Stage (III/IV) | Kaplan–Meier Curve Analysis: OS and DFS | |||
| 1 | miR-224 | 0.910 (95% CI: 0.84–0.98) | p < 0.001 | p < 0.001 | p < 0.001 | p < 0.01 | [58] |
| 2 | miR-93 | 0.825 (95% CI: 0.730–0.919) | p < 0.0001 | p = 0.047 | p = 0.006 | p = 0.046 | [60] |
| 3 | miR-92b | 0.702 (95% CI: 0.576–0.828) | p = 0.004 | ND | ND | ND | [62] |
| 4 | miR-665 | ND | p = 0.0042 (<5 cm) | p = 0.0276 | p < 0.05 | [65] | |
| 5 | miR-718 | ND | p = 0.04 | p = 0.026 | p = 0.0002 (DFS) | [66] | |
| 6 | miR-125b | 0.739 (95% CI: 0.648–0.830) | p = 0.048 | p = 0.11 | p = 0.011 | p < 0.001 (DFS and OS) | [69] |
| 7 | miR-194 | 0.738 (95% CI: 0.638–0.838) | p = 0.0001 | p = 0.013 | p > 0.05 | ND | [72] |
| miR-17-5p | 0.850 (95% CI: 0.764–0.936) | p = 0.0001 | p = 0.047 | p > 0.05 | ND | ||
| miR-106a | 0.704 (95% CI: 0.534–0.873) | p = 0.016 | p = 0.035 | p > 0.05 | p = 0.041 | ||
| 8 | miR-10b-5p | 0.968 (95% CI: 0.85–0.99) | p < 0.0001 | ND | not statistically significant | [73] | |
| miR-215-5p | 0.936 (95% CI: 0.80–0.99) | p < 0.0001 | ND | p < 0.01 | p = 0.02 (DFS) | ||
| 9 | miR-595 | 0.92 (95% CI: 0.86–0.97) | p < 0.0001 | ND | p = 0.007 | ND | [74] |
Three miRNAs—miR-125b, miR-665, and miR-638—are associated with improved OS rates [65][68][69], whereas exosomal miR-10b-5p overexpression significantly correlates with disease-free survival in patients with HCC [73]. Despite improvements in therapies, the recurrence rates of HCC after surgical resection remain ≥10%. Thus, exploiting exosomal miRNAs (expressed pre- and post-surgery) as predictive markers for recurrence pursues the unmet clinical need.
The expression level of exosomal miR-92b in the serum of HCC patients was significantly higher than that of the control group (non-HCC) [62]. If the levels of exosomal miR-92b continue to be upregulated, premature recurrence is induced. This result identifies exosomal miR-92b as a prognostic biomarker of post-transplant HCC recurrence. Sugimachi et al. showed using microarray analysis that miR-718 is a potential biomarker for predicting HCC recurrence after surgery. The authors reported different expression levels in patients with and without HCC recurrence [66].
Many studies reported low expression of miR-155 in the plasma of HCC patients compared with that of healthy individuals [75][76][77]. Others detected overexpressed miR-155 in preoperative plasma, and found it to be significantly correlated with early recurrence in patients with HCC [64].
A different approach correlates exosomal miRNA expression with tumour staging, which allows for the selection of the most suitable biomarkers for early diagnosis. Overexpression of miR-665 in plasma exosomes correlates with tumour size, invasion, and staging [65], whereas miR-21 expression strongly correlates with tumour stage and cirrhosis. Because exosomal miR-519d and miR-494 are upregulated in HCC compared with liver cirrhosis patients, they are considered to be independent diagnostic biomarkers [70]. In addition, the level of exosomal miR-224 was significantly higher in patients with large tumours (>3 cm) and advanced tumour stages (III/IV) [58]. Similarly, Shi et al. showed that low levels of exosomal miR-638 in HCC were significantly correlated with tumour size (>5 cm) and advanced staging (III/IV) [68].
To gain a better understanding of intercellular communication, and ultimately improve therapeutic strategies for HCC, researchers have recently performed in vitro studies using HCC cell cultures treated with exosomes derived from other cells. HCC cells (Huh7 and SMMC 7721) proliferate significantly less when co-cultured with high concentrations of miR-638 extracted from serum exosomes [68]. Clinical data support these results: low levels of miR-638 extracted from serum exosomes correlate with a lower survival rate of HCC patients. The authors imply that serum exosomal miR-638 affects liver carcinogenesis by inhibiting cancer cell proliferation. In contrast, the expression level of serum-derived exosomal miR-638 did not correlate with viral hepatitis B or C, tumour grade, or liver cirrhosis.
More complex studies were conducted to correlate the type of therapy and development of HCC with the expression of different exosomal miRNAs, based on the putative association of exosomal miRNA species with hepatitis and cirrhosis in HCC patients. The relative expression of exosomal miR-122 (calculated as relative expression of miR-122 after/before TACE) is significantly decreased after TACE [67]. One study was conducted on 57 patients with liver cirrhosis and 18 chronic hepatitis patients with HCC; miR-122 relative expression was higher in patients with liver cirrhosis and longer disease-specific survival. The authors point toward the applicability of miR-122 in the therapeutic guidance of TACE-treated patients.
Other studies suggest that exosomal miR-21 is a potential biomarker for the diagnosis of CHB (chronic hepatitis B) patients. Different reference genes (miRNAs and small RNAs) were used to normalize expression levels of exosomal serum-derived miRs—including miR-221, let-7a, miR-191, miR-26a, and miR-181a—in CHB patients, HCC patients, and healthy individuals. The expression level of exosomal miR-21 was significantly increased in the CHB group compared to the other two groups [78].
Similarly, Murakami et al. analysed miRNA expression profiles from the serum of 64 CHC (chronic hepatitis C) patients and 24 controls with normal liver (NL). The specific expression patterns of exosomal miRNAs expressed in chronic liver disease and inflammation correlated with the types and grades of liver disease. The expression patterns of nine miRNA species (miR-1225-5p, miR-1275, miR-638, miR-762, miR-320c, miR-451, miR-1974, miR-1207-5p, and miR-1246) identified CHC and NL with 96.59% accuracy [79].
Exosomes isolated from HCV J6/JFH-1-infected HuH-7.5 cells, and from the sera of chronic HCV-infected patients, contain the replication enhancers VHC-Ago2, miR-122, and HSP90. Recent studies have consistently shown that miR-122, Ago2, and HSP90 enhance HCV replication. Moreover, miR-122 is a host factor used by HCV for replication, and is present in exosomes isolated from HCV patients [80].
To date, we have identified many miRNA species isolated from exosomes, considering their potential roles as specific biomarkers for HCC. Therefore, scientists have used a more insightful approach of late: studies of miRNA profiling panels that match specific pathological conditions. Panels of miRNAs readily applicable as biomarkers or prognostic tools are yet to be discovered, but studies consider them to be sufficiently accurate to guide therapy in patients with advanced HCC disease. Table 3 reviews in detail these exosomal miRNA panels and their clinical relevance. The exosomes’ sources are indicated.
Table 3. Exosome panels and their clinical relevance.
| No. | miR Analysed in Panels | Expression Profile in HCC (Up- or Downregulated) | Exosome Isolation Methods | Source of Exosomes | Sample Size and Groups | Quantification Methods | Normalized (Reference miRs/Internal Control) | Clinical Relevance | References |
|---|---|---|---|---|---|---|---|---|---|
| 1 | miR-122, miR-125b, miR-145, miR-192, miR-194, miR-29a, miR-17-5p, and miR-106a. | up | Total Exosome Isolation Kit (GenePharma) | Human serum | HCC (n = 80) and healthy controls (n = 30) | qRT-PCR | miR-16 | Distinguish HCC patients from healthy controls. | [72] |
| 2 | miR-18a, miR-221, miR-222, miR-224 | up | ExoQuick Exosome Precipitation Solution | HCC (n = 20), CHB (n = 20), and LC (n = 20) | miR-16 | Distinguish HCC from LC and CHB. | [81] | ||
| miR-101, miR-106b, miR-122, miR-195 | down | ||||||||
| 3 | miR-26a, miR-29c, miR-21 | down | ExoQuick reagent | Human serum | HCC (n = 20), CHB (n = 20), and LC (n = 20) | let-7a | Lower levels in HCC patients than in cirrhotic and HBV patients. | [82] | |
| 4 | miR-122, miR-148a and miR-1246 | up | Polyethylene glycol (PEG) 6000 (Sigma-Aldrich, St Louis, MO) of 8% concentration | Discovery group: HCC (n = 5) with LC (n = 5) for deep sequencing. Validation group: NC (n = 64), CHB(n = 50), LC (n = 53), and HCC(n = 68) for further study 85 HCC patients | cel-miR-39 | Distinguish early-stage HCC from liver cirrhosis. | [83] | ||
| 5 | miR-21, miR-10b | up | Ultracentrifugation | Rat serum | Male fisher 344 rats (n = 108) (HCC models) | miRNA-484 | Biomarkers for early-stage HCC combined with exosomal miRNAs and AFP. | [84] | |
| miR-122, miR-200a | down | ||||||||
| 6 | miR-10b-5p, miR-18a-5p, miR-215-5p, and miR-940 | up | SeraMir Exosome RNA Amplification Kit (System Biosciences) | Human serum | HCC (n = 90), CHB (n = 27), LC (n = 33), and healthy controls (n = 28) | qRT-PCR and miR sequencing for discovery set | miR-1228-3p | Biomarker for early-stage HCC and poor disease-free survival | [73] |
| 7 | miR-519d, miR-21, miR-221 and miR-1228 | up | Ultracentrifugation | 30 patients (10 patients with liver cirrhosis without nodular liver lesions, 13 patients with early HCC, and 7 patients with advanced HCC) and HCC-derived cell lines (n = 7). HepG2, Hep3B, Huh-7, SNU449, SNU398, SNU182, and SNU475 | qRT-PCR | cel-miR-39 | More efficient diagnostic role than AFP for HCC patients. | [74] | |
| 8 | miR-140-3p, miR-30d-5p, miR-29b-3p, miR-130b-3p and miR-330-5p | up | 3D Medicine exosome isolation kit (CFDA license no. Hu min xie bei 20170019) | 10 fast-migrated and 10 slow-migrated PDC cultures from 36 HCC samples | Small RNA library construction and sequencing | Normalization | Migratory abilities of tumour cells | [85] | |
| miR-296-3p | down |
3.2. Effects of Exosomal microRNAs on Cell Survival, Proliferation, and Angiogenesis
HCC cells can release exosomes to promote tumour cell proliferation, angiogenesis, migration, or metastasis (Figure 2).
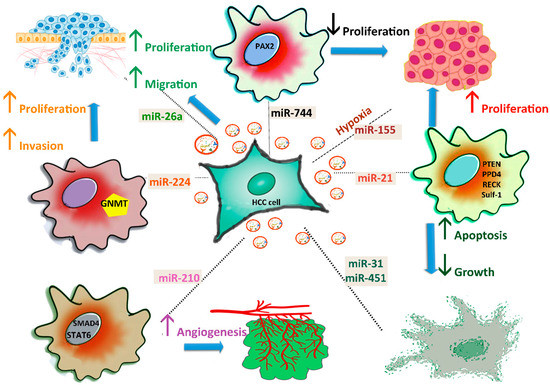
Figure 2. Roles of exosomal miRNAs in HCC progression. HCC cells secrete exosomes and affect proliferation by transferring miR-744, miR-21, and miR-26a into neighbouring cells. Angiogenesis is stimulated by miR-210, which blocks the SMAD4 and STAT6 pathways, while apoptosis is activated by miR-31 and miR-451. Inhibition of GNMT by miR-224 promotes the proliferation and invasion of HCC cells. miR-155 secretion increases under hypoxic conditions.
The two-way signalling pathways impact the drug resistance of HCC. For instance, high levels of miR-744 promote proliferation, and induce resistance of HepG2 cells to sorafenib, targeting paired box gene 2 (PAX2), which is overexpressed in HCC tissue. Following treatment with exosomes, miR-744 significantly decreases HCC cell proliferation and resistance to therapeutic drugs [71]. miR-21 induces cell proliferation and metastasis by inhibiting the expression of PTEN [86], PDD4, RECK, and SULF-1 (human sulphatase-1) [87], and confers resistance to chemotherapeutic drugs [88].
In vitro studies indicate that miR-224 decreases the expression of GNMT (glycine N-methyltransferase) by directly targeting the 3’-UTR mRNA and promoting the proliferation and invasion of HCC cells [58].
Exosomal miR-210 derived from HCC cells is internalized by endothelial cells, and promotes tumour angiogenesis through direct inhibition of the SMAD4 (SMAD family member 4) and STAT6 (signal transducer and activator of transcription 6) genes [63]. Matsuura et al. showed that both cellular and exosomal miR-155 expression levels were significantly increased under hypoxic conditions in HCC cells [64]. Of a group of six miRNAs, only miR-155 was upregulated under hypoxic conditions. Another study demonstrated that miR-31 and miR-451 species in exosomes derived from adult human liver stem cells inhibit HCC growth and stimulate apoptosis [89].
The release of miR-26a species in HepG2 cells treated with exosomes led to ectopic overexpression of miR-26a and decreased migration and tumour proliferation. Exosomes loaded with miR-26a blocked cell growth, pointing to the possibility of using engineered exosomes as therapeutic agents [90]. The next subsection of this review notes some results deciphering the mechanism of communication through exosomal miRNAs between HCC cells and cells from the tumour microenvironment.
3.3. Role of Exosomal microRNAs in Intra- and Intercellular Communication and Therapies
Exosomes secreted by HCC cells provide autocrine and paracrine signals to surrounding cells but also deliver products to distant cells, fine-tuning diverse biological responses. The process is two-way: blood cells or other cells from the tumoural microenvironment secrete exosomes, which mediate carcinogenesis. All of these processes have been intensively analysed, and many communications pathways have been depicted in recent years.
Of note, even during biogenesis, two miR species (miR-27b-3p and miR-92a-3p) inhibit VPS4A expression in HCC tissues [20]. In turn, dysregulation of VPS4A facilitates the secretion of oncogenic miRNAs in exosomes. The expression of VPS4A in HCC tissue is linked to tumour growth and metastasis.
Exosomal miR-21 derived from HCC cell lines converts normal hepatic stellate cells (HSCs) into cancer-associated fibroblasts (CAFs) through directly targeting the PTEN gene, which activates the PDK1/AKT signalling pathway. Finally, angiogenic cytokines (VEGF, MMP-2, MMP-9, FGF2, and TGF) are secreted, which promote cancer progression. These results concur with the clinical data: patients with HCC have high levels of serum exosomal miR-21—correlated with CAF activation, higher vessel density, and lower survival rates [91].
A newly described mechanism shows that miRNAs secreted by HCC cells inhibit T cell function, escaping the immune system. Next-generation sequencing analysis (NGS) reveals that miR-23a-3p has a high expression level in exosomes derived from tunicamycin-treated HCC cells (Exo-TM). Co-cultivation of T cells with macrophages treated with Exo-TM leads to decreased cell ratio of CD8 T and decreased interleukin-2 expression, as a result of the activation of the PTEN/AKT pathway [92].
Drug-resistant HCC cells (Bel/5-FU) secrete high levels of exosomal miR-32-5p, and low levels of the tumour suppressor gene PTEN. This discrepancy results from miR-32-5p overexpression, which inhibits PTEN and, subsequently, activates the PI3K/AKT signalling pathway. Inhibited PTEN induces drug resistance, and promotes angiogenesis and EMT. Clinically, miR-32-5p overexpression and low PTEN expression are positively associated with poor prognosis in HCC patients [93].
Exosome-derived miRNAs are mediators of the carcinogenesis induced by environmental chemicals. Exosomes derived from arsenite-transformed L-02 cells transfer miR-155 to normal L-02 and THLE-3 cells (grown in co-culture) and induce pro-inflammatory activity in normal liver cells. Moreover, exosome-derived miR-155 from serum was abundant in the patient group exposed to arsenite [94].
In hepatocytes, miR-155 binds to Toll receptor ligands, triggering Toll receptor-mediated inflammation and promoting liver injury and inflammation [95].
One study identifies new pathways of functional miRNA transfer from circulating tumour cells (CTCs) to the tumoural cells of origin, with the potential to inhibit tumour recurrence and metastatic spread. Exosomal miR-25-5p derived from HCC contributes to “tumour self-seeding”, defined as colonization by CTCs. This mechanism was linked to the expression of the LRRC7 (Leucine-rich repeat-containing 7) gene involved in CTC reattachment and the reformation of tumour cell seeding. The uptake of exosomes secreted by CTC cells greatly improved the migratory and invasive abilities of HCC cells, with miR-25-5p declared as the main target of LRRC7. Inhibition of exosomal miR-25-5p reversed these effects [96].
Exosomal miR-103 produced by hepatic cells and delivered to endothelial cells (HUVECs, LSECs) and normal liver sinusoidal endothelial cells (Sk-Hep-1) inhibited protein junctions, zonula occludens-1, VE cadherin, and p120-catenin. MiR-103 reduced the integrity of endothelial junctions, increased the grade of vascular permeability and, finally, facilitated tumour metastasis [97]. To analyse metastasis and proliferation of target cells, in vitro co-cultivation studies used the transfer of long non-coding RNA (lncRNA FAL1 (Focally amplified lncRNA on chromosome 1)) molecules to HepG2 and HuH-7 cell cultures. The competitive binding to miR-1236 in target cells leads to the overexpression of ZEB1 and AFP, and the subsequent promotion of metastasis and proliferation of the targeted cells [98].
The development of HCC is mainly based on LC disorders, as the tumour microenvironment is enriched with activated fibroblasts. Highly metastatic HCC cells release exosomal miR-1247-3p to convert fibroblasts into CAFs by targeting the B4GALT3 (β-1,4-galactosyltransferase 3) gene, which activates the β1-integrin/NF-κB signalling pathway. Consequently, the activated pathway decreases the secretion of the IL-6 and IL-8 cytokines. The same study showed that increased expression levels of exosomal miR-1247-3p significantly correlate with lung metastases in patients with HCC [61]. CAFs induce tumourigenic processes, tumour niches, EMT (epithelial–mesenchymal transition), and chemoresistance. All of these processes are important in the development of the inflammatory microenvironment, and promote lung metastases derived from liver cancer. A different group performed sequencing analysis of CAFs and corresponding para-cancer fibroblasts (PAFs), and found a significant reduction in exosomal miR-320a levels in CAF-derived exosomes. The transfer of stromal-cell-derived miR-320a inhibits the tumour progression of HCC cells by binding to its direct target PBX3. These results qualify the method of miRNA transfer as a potential treatment option for HCC progression [99].
Human macrophages transfer miRNA species to HCC cells in a manner that requires cell contact through gap junctions. Two specific miRNAs—miR-142 and miR-223—are efficiently transferred; they block the stathmin-1 and IGFR-1 receptors, and finally HCC cell proliferation [100].
Exosomal miR-21, miR-192, and miR-221 derived from colorectal cancer cell lines and transferred to HepG2 and A549 cells promoted the invasion and metastasis of recipient cells [101]. Hep3B-derived exosomes containing specific miRNAs—such as miR-584, miR-517c, miR-378, miR-520f, miR-142-5p, miR-451, miR-518d, miR-215, miR-376a, miR-133b, and miR-367—were internalized in another HCC cell line, HepG2. Consecutively, these miRNAs modulated the TAK1 (TGF-β-activated kinase 1) inhibition process. Loss of TAK1 has been associated with hepatocarcinogenesis [102].
Exosomes released from umbilical mesenchymal stem cells (UMSCs) inhibit HCV infection, especially viral replication. Qian et al. found that the anti-hepatitis C virus infection effect was mediated by various miRNAs (let-7f, miR-145, miR-199a, and miR-221) specifically transported by exosomes derived from UMSCs [103].
The E2 envelope glycoprotein of the hepatitis C virus (HCV-E2) stimulated mast cells to secrete exosomes with high levels of miR-490. The exosome shuttled into HCC cells inhibited the ERK1/2 signalling pathway and, in the end, suppressed HCC cells’ metastasis [104].
Using exosomes to mediate the transfer of miRNAs between different cells encourages clinical application, but for the moment, studies are limited to experiments using cell culture models. Exosomes operate on hepatocytes even when they derive from the cells of other tissues or organs, and contribute to the physiological or pathological processes of the liver [105][106][107][108].
Adipose tissue-derived mesenchymal stem cells (AMSCs) transfected with a plasmid-encoding miR-122 (122-Exo) effectively loaded miR-122 in secreted exosomes. Then, 122-Exo was transported to HCC cells, and altered the expression profiles of cyclin G1 (CCNG1), ADAM10 (a disintegrin and metalloprotease domain-containing protein 10), and IGFR-1 (insulin-like growth factor receptor 1). These changes occurred after HepG2 cells were exposed to 122-Exo for 24 h [109]. AMSCs infected with pre-miR-199a-3p (LV-199a) plasmid abundantly expressed miR-199, and transferred the species via exosomes after puromycin selection. This process increased the sensitivity of HCC cells to doxorubicin by inhibiting the mTOR signalling pathway. Intravenous injection of an orthotopic HCC murine model with the AMSC-Exo-199a produces a rapid increase in the efficiency of doxorubicin therapy in HCC tumours [110].
Other groups report the use of exosomes as nanotransporters in therapies, paving the way for new applications in HCC treatment. Exosomes transfected with antineoplastic and antifibrotic miR-335-5p were administered in vitro and in vivo to mice with developed HCC tumours; this treatment inhibited cell proliferation, invasion, and tumour growth. Previous reports demonstrated that HSC-derived exosomes can encapsulate and transfer miR-335-5p to HCC cells in vitro or in vivo, inhibiting tumour growth [111]. In a recent study, exosomes were used as vehicles for the transport and targeted internalization of small RNA molecules in HCC cells. HepG2 cells were conditioned to bind (through the Apo-A1 receptor) and internalize exosomes designed by genetic engineering [90].
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424.
- Bellentani, S. Epidemiology of hepatocellular carcinoma in metabolic liver disease. Hepatoma Res. 2020, 6, 23.
- Welzel, T.M.; Graubard, B.I.; Quraishi, S.; Zeuzem, S.; Davila, J.A.; El-Serag, H.B.; McGlynn, K.A. Population-attributable fractions of risk factors for hepatocellular carcinoma in the United States. Am. J. Gastroenterol. 2013, 108, 1314–1321.
- Younossi, Z.M.; Otgonsuren, M.; Henry, L.; Venkatesan, C.; Mishra, A.; Erario, M.; Hunt, S. Association of non-alcoholic fatty liver disease (NAFLD) with hepatocellular carcinoma (HCC) in the United States from 2004 to 2009. Hepatology 2015, 62, 1723–1730.
- Mittal, S.; El-Serag, H.B.; Sada, Y.H.; Kanwal, F.; Duan, Z.; Temple, S.; May, S.B.; Kramer, J.R.; Richardson, P.A.; Davila, J.A. Hepatocellular Carcinoma in the Absence of Cirrhosis in United States Veterans Is Associated with Nonalcoholic Fatty Liver Disease. Clin. Gastroenterol. Hepatol. 2016, 14, 124–131.e1.
- Varghese, J.; Kedarisetty, C.K.; Venkataraman, J.; Srinivasan, V.; Deepashree, T.; Uthappa, M.C.; Ilankumaran, K.; Govil, S.; Reddy, M.S.; Rela, M. Combination of TACE and Sorafenib Improves Outcomes in BCLC Stages B/C of Hepatocellular Carcinoma: A Single Centre Experience. Ann. Hepatol. 2017, 16, 247–254.
- Kuczynski, E.A.; Lee, C.R.; Man, S.; Chen, E.; Kerbel, R.S. Effects of Sorafenib Dose on Acquired Reversible Resistance and Toxicity in Hepatocellular Carcinoma. Cancer Res. 2015, 75, 2510–2519.
- Shu, H.; Li, W.; Shang, S.; Qin, X.; Zhang, S.; Liu, Y. Diagnosis of AFP-negative early-stage hepatocellular carcinoma using Fuc-PON1. Discov. Med. 2017, 23, 163–168.
- Schraml, C.; Kaufmann, S.; Rempp, H.; Syha, R.; Ketelsen, D.; Notohamiprodjo, M.; Nikolaou, K. Imaging of HCC—Current State of the Art. Diagnostics 2015, 5, 513–545.
- Forner, A.; Reig, M.; Bruix, J. Hepatocellular carcinoma. Lancet 2018, 391, 1301–1314.
- Wu, Q.; Zhou, L.; Lv, D.; Zhu, X.; Tang, H. Exosome-mediated communication in the tumour microenvironment contributes to hepatocellular carcinoma development and progression. J. Hematol. Oncol. 2019, 12, 53.
- Wang, H.; Lu, Z.; Zhao, X. Tumorigenesis, diagnosis, and therapeutic potential of exosomes in liver cancer. J. Hematol. Oncol. 2019, 12, 133.
- Kosaka, N. Decoding the Secret of Cancer by Means of Extracellular Vesicles. J. Clin. Med. 2016, 5, 22.
- Wong, C.-M.; Tsang, F.H.-C.; Ng, I.O.-L. Non-coding RNAs in hepatocellular carcinoma: Molecular functions and pathological implications. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 137–151.
- Li, S.; Yao, J.; Xie, M.; Liu, Y.; Zheng, M. Exosomal miRNAs in hepatocellular carcinoma development and clinical responses. J. Hematol. Oncol. 2018, 11, 54.
- György, B.; Szabó, T.G.; Pásztói, M.; Pál, Z.; Misják, P.; Aradi, B.; László, V.; Pállinger, É.; Pap, E.; Kittel, Á.; et al. Membrane vesicles, current state-of-the-art: Emerging role of extracellular vesicles. Cell. Mol. Life Sci. 2011, 68, 2667–2688.
- Heijnen, H.F.; Schiel, A.E.; Fijnheer, R.; Geuze, H.J.; Sixma, J.J. Activated platelets release two types of mem-brane vesicles: Microvesicles by surface shedding and exosomes derived from exocytosis of multivesicular bodies and alpha-granules. Blood 1999, 94, 3791–3799.
- Yanez-Mo, M.; Siljander, P.R.; Andreu, Z.; Zavec, A.B.; Borras, F.E.; Buzas, E.I.; Buzas, K.; Casal, E.; Cappello, F.; Carvalho, J.; et al. Biological properties of extracellular vesi-cles and their physiological functions. J. Extracell. Vesicles 2015, 4, 27066.
- Adell, M.A.Y.; Migliano, S.M.; Teis, D. ESCRT-III and Vps4: A dynamic multipurpose tool for membrane budding and scission. FEBS J. 2016, 283, 3288–3302.
- Wei, J.X.; Lv, L.H.; Wan, Y.L.; Cao, Y.; Li, G.L.; Lin, H.M.; Zhou, R.; Shang, C.Z.; Cao, J.; He, H.; et al. Vps4A functions as a tumour suppressor by regulating the secretion and uptake of exo-somal microRNAs in human hepatoma cells. Hepatology 2015, 61, 1284–1294.
- Kowal, J.; Tkach, M.; Théry, C. Biogenesis and secretion of exosomes. Curr. Opin. Cell Biol. 2014, 29, 116–125.
- Stoorvogel, W. Resolving sorting mechanisms into exosomes. Cell Res. 2015, 25, 531–532.
- Sinha, S.; Hoshino, D.; Hong, N.H.; Kirkbride, K.C.; Grega-Larson, N.E.; Seiki, M.; Tyska, M.J.; Weaver, A.M. Cortactin promotes exosome secretion by controlling branched actin dynamics. J. Cell Biol. 2016, 214, 197–213.
- Zila, V.; Difato, F.; Klimova, L.; Huerfano, S.; Forstova, J. Involvement of Microtubular Network and Its Motors in Productive Endocytic Trafficking of Mouse Polyomavirus. PLoS ONE 2014, 9, e96922.
- Ghossoub, R.; Lembo, F.; Rubio, A.; Gaillard, C.B.; Bouchet, J.; Vitale, N.; Slavík, J.; Machala, M.; Zimmermann, P. Syntenin-ALIX exosome biogenesis and budding into multivesicular bodies are controlled by ARF6 and PLD2. Nat. Commun. 2014, 5, 3477.
- Villarroya-Beltri, C.; Gutierrez-Vazquez, C.; Sanchez-Cabo, F.; Perez-Hernandez, D.; Vazquez, J.; Martin-Cofreces, N.; Martinez-Herrera, D.J.; Pascual-Montano, A.; Mittelbrunn, M.; Sanchez-Madrid, F. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 2013, 4, 2980.
- Karim, M.A.; Samyn, D.R.; Brett, C.L. A Cell-Free Content Mixing Assay for SNARE-Mediated Multivesicular Body-Vacuole Membrane Fusion. Adv. Struct. Saf. Stud. 2018, 1860, 289–301.
- Koles, K.; Nunnari, J.; Korkut, C.; Barria, R.; Brewer, C.; Li, Y.; Leszyk, J.; Zhang, B.; Budnik, V. Mechanism of Evenness Interrupted (Evi)-Exosome Release at Synaptic Boutons. J. Biol. Chem. 2012, 287, 16820–16834.
- Koles, K.; Budnik, V. Exosomes go with the Wnt. Cell. Logist. 2012, 2, 169–173.
- Savina, A.; Furlan, M.; Vidal, M.; Colombo, M.I. Exosome release is regulated by a calcium-dependent mechanism in K562 cells. J. Biol. Chem. 2003, 278, 20083–20090.
- Schroeder, B.; Schulze, R.J.; Weller, S.G.; Sletten, A.C.; Casey, C.A.; McNiven, M.A. The small GTPase Rab7 as a central regulator of hepatocellular lipophagy. Hepatology 2015, 61, 1896–1907.
- Song, L.; Tang, S.; Han, X.; Jiang, Z.; Dong, L.; Liu, C.; Liang, X.; Dong, J.; Qiu, C.; Wang, Y.; et al. KIBRA controls exosome secretion via inhibiting the proteasomal degradation of Rab27a. Nat. Commun. 2019, 10, 1639.
- Yang, L.; Peng, X.; Li, Y.; Zhang, X.; Ma, Y.; Wu, C.; Fan, Q.; Wei, S.; Li, H.; Liu, J. Long non-coding RNA HO-TAIR promotes exosome secretion by regulating RAB35 and SNAP23 in hepatocellular carcinoma. Mol. Cancer 2019, 18, 78.
- Yue, B.; Yang, H.; Wang, J.; Ru, W.; Wu, J.; Huang, Y.; Lan, X.; Lei, C.; Chen, H. Exosome biogenesis, secretion and function of exosomal miRNAs in skeletal muscle myogenesis. Cell Prolif. 2020, 53, 12857.
- Hessvik, N.P.; Llorente, A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 2018, 75, 193–208.
- Mathivanan, S.; Fahner, C.J.; Reid, G.E.; Simpson, R.J. ExoCarta 2012: Database of exosomal proteins, RNA and lipids. Nucleic Acids Res. 2011, 40, D1241–D1244.
- Kalra, H.; Simpson, R.J.; Ji, H.; Aikawa, E.; Altevogt, P.; Askenase, P.; Bond, V.C.; Borras, F.E.; Breakefield, X.; Budnik, V.; et al. Vesi-clepedia: A compendium for extracellular vesicles with continuous community annotation. PLoS Biol. 2012, 10, e1001450.
- Kim, D.K.; Kang, B.; Kim, O.Y.; Choi, D.S.; Lee, J.; Kim, S.R.; Go, G.; Yoon, Y.J.; Kim, J.H.; Jang, S.C.; et al. EVpedia: An integrated da-tabase of high-throughput data for systemic analyses of extracellular vesicles. J. Extracell. Vesicles 2013, 2, 20384.
- Subra, C.; Laulagnier, K.; Perret, B.; Record, M. Exosome lipidomics unravels lipid sorting at the level of mul-tivesicular bodies. Biochimie 2007, 89, 205–212.
- Parolini, I.; Federici, C.; Raggi, C.; Lugini, L.; Palleschi, S.; De Milito, A.; Coscia, C.; Iessi, E.; Logozzi, M.; Moli-nari, A.; et al. Microenvironmental pH is a key factor for exosome traffic in tumour cells. J. Biol. Chem. 2009, 284, 34211–34222.
- Théry, C.; Duban, L.; Segura, E.; Véron, P.; Lantz, O.; Amigorena, S. Indirect activation of naïve CD4+ T cells by dendritic cell–derived exosomes. Nat. Immunol. 2002, 3, 1156–1162.
- Kalani, A.; Tyagi, A.; Tyagi, N. Exosomes: Mediators of Neurodegeneration, Neuroprotection and Therapeutics. Mol. Neurobiol. 2013, 49, 590–600.
- Ha, D.; Yang, N.; Nadithe, V. Exosomes as therapeutic drug carriers and delivery vehicles across biological membranes: Current perspectives and future challenges. Acta Pharm. Sin. B 2016, 6, 287–296.
- van Balkom, B.W.M.; Eisele, A.S.; Pegtel, D.M.; Bervoets, S.; Verhaar, M.C. Quantitative and qualitative analysis of small RNAs in human endothelial cells and exosomes provides insights into localized RNA processing, degradation and sorting. J. Extracell. Vesicles 2015, 4, 26760.
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659.
- Pegtel, D.M.; Cosmopoulos, K.; Thorley-Lawson, D.A.; Van Eijndhoven, M.A.J.; Hopmans, E.S.; Lindenberg, J.L.; De Gruijl, T.D.; Würdinger, T.; Middeldorp, J.M. Functional delivery of viral miRNAs via exosomes. Proc. Natl. Acad. Sci. USA 2010, 107, 6328–6333.
- Filipenko, N.R.; MacLeod, T.J.; Yoon, C.-S.; Waisman, D.M. Annexin A2 Is a Novel RNA-binding Protein. J. Biol. Chem. 2004, 279, 8723–8731.
- Hollas, H.; Aukrust, I.; Grimmer, S.; Strand, E.; Flatmark, T.; Vedeler, A. Annexin A2 recognises a specific region in the 3’-UTR of its cognate messenger RNA. Biochim. Biophys. Acta 2006, 1763, 1325–1334.
- Mickleburgh, I.; Burtle, B.; Hollas, H.; Campbell, G.; Chrzanowska-Lightowlers, Z.; Vedeler, A.; Hesketh, J. An-nexin A2 binds to the localization signal in the 3’ untranslated region of c-myc mRNA. FEBS J. 2005, 272, 413–421.
- Denzer, K.; Kleijmeer, M.J.; Heijnen, H.F.; Stoorvogel, W.; Geuze, H.J. Exosome: From internal vesicle of the multivesicular body to intercellular signaling device. J. Cell Sci. 2000, 113, 3365–3374.
- Lamparski, H.G.; Metha-Damani, A.; Yao, J.-Y.; Patel, S.; Hsu, D.-H.; Ruegg, C.; Le Pecq, J.-B. Production and characterization of clinical grade exosomes derived from dendritic cells. J. Immunol. Methods 2002, 270, 211–226.
- Lai, R.C.; Yeo, R.W.; Tan, K.H.; Lim, S.K. Exosomes for drug delivery—A novel application for the mesenchymal stem cell. Biotechnol. Adv. 2013, 31, 543–551.
- Vlassov, A.V.; Magdaleno, S.; Setterquist, R.; Conrad, R. Exosomes: Current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim. Biophys. Acta Gen. Subj. 2012, 1820, 940–948.
- Qin, J.; Xu, Q. Functions and application of exosomes. Acta Pol. Pharm. Drug Res. 2014, 71, 537–543.
- Batrakova, E.V.; Kim, M.S. Using exosomes, naturally-equipped nanocarriers, for drug delivery. J. Control Release 2015, 219, 396–405.
- Torre, L.A.; Siegel, R.L.; Ward, E.M.; Jemal, A. Global Cancer Incidence and Mortality Rates and Trends—An Update. Cancer Epidemiol. Biomark. Prev. 2015, 25, 16–27.
- O’Brien, K.; Breyne, K.; Ughetto, S.; Laurent, L.C.; Breakefield, X.O. RNA delivery by extracellular vesicles in mammalian cells and its applications. Nat. Rev. Mol. Cell Biol. 2020, 21, 585–606.
- Cui, Y.; Xu, H.-F.; Liu, M.-Y.; Xu, Y.-J.; He, J.-C.; Zhou, Y.; Cang, S.-D. Mechanism of exosomal microRNA-224 in development of hepatocellular carcinoma and its diagnostic and prognostic value. World J. Gastroenterol. 2019, 25, 1890–1898.
- Wang, H.; Hou, L.; Li, A.; Duan, Y.; Gao, H.; Song, X. Expression of Serum Exosomal MicroRNA-21 in Human Hepatocellular Carcinoma. BioMed Res. Int. 2014, 2014, 864894.
- Xue, X.; Wang, X.; Zhao, Y.; Hu, R.; Qin, L. Exosomal miR-93 promotes proliferation and invasion in hepato-cellular carcinoma by directly inhibiting TIMP2/TP53INP1/CDKN1A. Biochem. Biophys. Res. Commun. 2018, 502, 515–521.
- Fang, T.; Lv, H.; Lv, G.; Li, T.; Wang, C.; Han, Q.; Yu, L.; Su, B.; Guo, L.; Huang, S.; et al. Tumour-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat. Commun. 2018, 9, 191.
- Nakano, T.; Chen, I.; Wang, C.; Chen, P.; Tseng, H.; Huang, K.; Hu, T.; Li, L.; Goto, S.; Cheng, Y.; et al. Circulating exosomal miR-92b: Its role for cancer immunoediting and clinical value for prediction of posttransplant hepatocellular carcinoma recurrence. Arab. Archaeol. Epigr. 2019, 19, 3250–3262.
- Lin, X.-J.; Fang, J.-H.; Yang, X.-J.; Zhang, C.; Yuan, Y.; Zheng, L.; Zhuang, S. Hepatocellular carcinoma cell-secreted exosomal microRNA-210 promotes angiogenesis in vitro and in vivo. Mol. Ther. Nucleic Acids 2018, 11, 243–252.
- Matsuura, Y.; Wada, H.; Eguchi, H.; Gotoh, K.; Kobayashi, S.; Kinoshita, M.; Kubo, M.; Hayashi, K.; Iwagami, Y.; Yamada, D.; et al. Exosomal miR-155 Derived from Hepatocellular Carcinoma Cells Under Hypoxia Promotes Angiogenesis in Endothelial Cells. Dig. Dis. Sci. 2018, 64, 792–802.
- Qu, Z.; Wu, J.; Wu, J.; Ji, A.; Qiang, G.; Jiang, Y.; Jiang, C.; Ding, Y. Exosomal miR-665 as a novel minimally invasive biomarker for hepatocellular carcinoma diagnosis and prognosis. Oncotarget 2017, 8, 80666–80678.
- Sugimachi, K.; Matsumura, T.; Hirata, H.; Uchi, R.; Ueda, M.; Ueo, H.; Shinden, Y.; Iguchi, T.; Eguchi, H.; Shirabe, K.; et al. Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. Br. J. Cancer 2015, 112, 532–538.
- Suehiro, T.; Miyaaki, H.; Kanda, Y.; Shibata, H.; Honda, T.; Ozawa, E.; Miuma, S.; Taura, N.; Nakao, K. Serum exosomal microRNA-122 and microRNA-21 as predictive biomarkers in transarterial chemoembolization-treated hepatocellular carcinoma patients. Oncol. Lett. 2018, 16, 3267–3273.
- Shi, M.; Jiang, Y.; Yang, L.; Yan, S.; Wang, Y.; Lu, X. Decreased levels of serum exosomal miR-638 predict poor prognosis in hepatocellular carcinoma. J. Cell. Biochem. 2018, 119, 4711–4716.
- Liu, W.; Hu, J.; Zhou, K.; Chen, F.; Wang, Z.; Liao, B.; Dai, Z.; Cao, Y.; Fan, J.; Zhou, J. Serum exosomal miR-125b is a novel prognostic marker for hepatocellular carcinoma. Oncotargets Ther. 2017, 10, 3843–3851.
- Tang, J.; Li, Y.; Liu, K.; Zhu, Q.; Yang, W.-H.; Xiong, L.-K.; Guo, D.-L. Exosomal miR-9-3p suppresses HBGF-5 expression and is a functional biomarker in hepatocellular carcinoma. Minerva Med. 2017, 109, 15–23.
- Wang, G.; Zhao, W.; Wang, H.; Qiu, G.; Jiang, Z.; Wei, G.; Li, X. Exosomal MiR-744 Inhibits Proliferation and Sorafenib Chemoresistance in Hepatocellular Carcinoma by Targeting PAX2. Med. Sci. Monit. 2019, 25, 7209–7217.
- Xue, X.; Zhao, Y.; Wang, X.; Qin, L.; Hu, R. Development and validation of serum exosomal microRNAs as di-agnostic and prognostic biomarkers for hepatocellular carcinoma. J. Cell. Biochem. 2019, 120, 135–142.
- Sohn, W.; Kim, J.; Kang, S.H.; Yang, S.R.; Cho, J.Y.; Cho, H.C.; Shim, S.G.; Paik, Y.H. Serum exosomal microRNAs as novel biomarkers for hepatocellular carcinoma. Exp. Mol. Med. 2015, 47, e184.
- Lin, H.; Zhang, Z. Diagnostic value of a microRNA signature panel in exosomes for patients with hepatocellular carcinoma. Int. J. Clin. Exp. Pathol. 2019, 12, 1478–1487.
- Wang, Y.; Zhang, C.; Zhang, P.; Guo, G.; Jiang, T.; Zhao, X.; Jiang, J.; Huang, X.; Tong, H.; Tian, Y. Serum exosomal microRNAs combined with alpha-fetoprotein as diagnostic markers of hepatocellular carcinoma. Cancer Med. 2018, 7, 1670–1679.
- Liu, W.H.; Ren, L.N.; Wang, X.; Wang, T.; Zhang, N.; Gao, Y.; Luo, H.; Navarro-Alvarez, N.; Tang, L.J. Combination of exosomes and circulating microRNAs may serve as a promising tumour marker complementary to alpha-fetoprotein for early-stage hepatocellular carcinoma diagnosis in rats. J. Cancer Res. Clin. Oncol. 2015, 141, 1767–1778.
- Cho, H.J.; Eun, J.W.; Baek, G.O.; Seo, C.W.; Ahn, H.R.; Kim, S.S.; Cho, S.W.; Cheong, J.Y. Serum exosomal microRNA, miR-10b-5p, as a potential diagnostic biomarker for early-stage hepatocellular carcinoma. J. Clin. Med. 2020, 9, 281.
- Fornari, F.; Ferracin, M.; Trerè, D.; Milazzo, M.; Marinelli, S.; Galassi, M.; Venerandi, L.; Pollutri, D.; Patrizi, C.; Borghi, A.; et al. Circulating microRNAs, miR-939, miR-595, miR-519d and miR-494, Identify Cirrhotic Patients with HCC. PLoS ONE 2015, 10, e0141448.
- Ning, S.; Liu, H.; Gao, B.; Wei, W.; Yang, A.; Li, J.; Zhang, L. miR-155, miR-96 and miR-99a as potential diag-nostic and prognostic tools for the clinical management of hepatocellular carcinoma. Oncol. Lett. 2019, 18, 3381–3387.
- Mohamed, A.A.; Omar, A.A.A.; El-Awady, R.R.; Hassan, S.M.A.; Eitah, W.M.S.; Ahmed, R.; Khater, A.; Tantawi, O.M.S. MiR-155 and MiR-665 Role as Potential Non-invasive Biomarkers for Hepatocellular Carcinoma in Egyptian Patients with Chronic Hepatitis C Virus Infection. J. Transl. Intern. Med. 2020, 8, 32–40.
- Guan, C.; Yang, F.; He, X.; Li, T.; Yang, Q.; He, H.; Xu, M. Clinical significance of microRNA-155 expression in hepatocellular carcinoma. Oncol. Lett. 2016, 11, 1574–1580.
- Li, Y.; Zhang, L.; Liu, F.; Xiang, G.; Jiang, D.; Pu, X. Identification of Endogenous Controls for Analyzing Serum Exosomal miRNA in Patients with Hepatitis B or Hepatocellular Carcinoma. Dis. Markers 2015, 2015, 1–12.
- Murakami, Y.; Toyoda, H.; Tanahashi, T.; Tanaka, J.; Kumada, T.; Yoshioka, Y.; Kosaka, N.; Ochiya, T.; Taguchi, Y.-H. Comprehensive miRNA Expression Analysis in Peripheral Blood Can Diagnose Liver Disease. PLoS ONE 2012, 7, e48366.
- Bukong, T.N.; Momen-Heravi, F.; Kodys, K.; Bala, S.; Szabo, G. Exosomes from Hepatitis C Infected Patients Transmit HCV Infection and Contain Replication Competent Viral RNA in Complex with Ago2-miR122-HSP90. PLoS Pathog. 2014, 10, e1004424.
- Yu, L.X.; Zhang, B.L.; Yang, Y.; Wang, M.C.; Lei, G.L.; Gao, Y.; Liu, H.; Xiao, C.H.; Xu, J.J.; Qin, H.; et al. Exosomal microRNAs as potential biomarkers for can-cer cell migration and prognosis in hepatocellular carcinoma patient-derived cell models. Oncol. Rep. 2019, 41, 257–269.
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. MicroRNA-21 regulates expression of the PTEN tumour suppressor gene in human hepatocellular cancer. Gastroenterology 2007, 133, 647–658.
- Bao, L.; Yan, Y.; Xu, C.; Ji, W.; Shen, S.; Xu, G.; Zeng, Y.; Sun, B.; Qian, H.; Chen, L.; et al. Mi-croRNA-21 suppresses PTEN and hSulf-1 expression and promotes hepatocellular carcinoma progression through AKT/ERK pathways. Cancer Lett. 2013, 337, 226–236.
- He, X.; Li, J.; Guo, W.; Liu, W.; Yu, J.; Song, W.; Dong, L.; Wang, F.; Yu, S.; Zheng, Y.; et al. Targeting the microRNA-21/AP1 axis by 5-fluorouracil and pirarubicin in human hepatocellular carcinoma. Oncotarget 2014, 6, 2302–2314.
- Fonsato, V.; Collino, F.; Herrera, M.B.; Cavallari, C.; Deregibus, M.C.; Cisterna, B.; Bruno, S.; Romagnoli, R.; Salizzoni, M.; Tetta, C.; et al. Human liver stem cell-derived microvesicles inhibit hepatoma growth in SCID mice by delivering antitumour microRNAs. Stem Cells 2012, 30, 1985–1998.
- Liang, G.; Kan, S.; Zhu, Y.; Feng, S.; Feng, W.; Gao, S. Engineered exosome-mediated delivery of functionally active miR-26a and its enhanced suppression effect in HepG2 cells. Int. J. Nanomed. 2018, 13, 585–599.
- Zhou, Y.; Ren, H.; Dai, B.; Li, J.; Shang, L.; Huang, J.; Shi, X. Hepatocellular carcinoma-derived exosomal miR-NA-21 contributes to tumour progression by converting hepatocyte stellate cells to cancer-associated fibro-blasts. J. Exp. Clin. Cancer Res. 2018, 37, 324.
- Liu, J.; Fan, L.; Yu, H.; Zhang, J.; He, Y.; Feng, D.; Wang, F.; Li, X.; Liu, Q.; Li, Y.; et al. Endoplasmic Reticulum Stress Causes Liver Cancer Cells to Release Exosomal miR-23a-3p and Up-regulate Programmed Death Ligand 1 Expression in Macrophages. Hepatology 2019, 70, 241–258.
- Fu, X.; Liu, M.; Qu, S.; Ma, J.; Zhang, Y.; Shi, T.; Wen, H.; Yang, Y.; Wang, S.; Wang, J.; et al. Exosomal microRNA-32-5p induces multidrug resistance in hepatocellular carcinoma via the PI3K/Akt path-way. J. Exp. Clin. Cancer Res. 2018, 37, 52.
- Chen, C.; Luo, F.; Liu, X.; Lu, L.; Xu, H.; Yang, Q.; Xue, J.; Shi, L.; Li, J.; Zhang, A.; et al. NF-kB-regulated exo-somal miR-155 promotes the inflammation associated with arsenite carcinogenesis. Cancer Lett. 2017, 388, 21–33.
- Bala, S.; Petrasek, J.; Mundkur, S.; Catalano, D.; Levin, I.; Ward, J.; Alao, H.; Kodys, K.; Szabo, G. Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology 2012, 56, 1946–1957.
- Liu, H.; Chen, W.; Zhi, X.; Chen, E.J.; Wei, T.; Zhang, J.; Shen, J.; Hu, L.Q.; Zhao, B.; Feng, X.H.; et al. Tumour-derived exosomes promote tumour self-seeding in hepatocellular carcinoma by transferring miRNA-25-5p to enhance cell motility. Oncogene 2018, 37, 4964–4978.
- Fang, J.H.; Zhang, Z.J.; Shang, L.R.; Luo, Y.W.; Lin, Y.F.; Yuan, Y.; Zhuang, S.M. Hepatoma cell-secreted ex-osomal microRNA-103 increases vascular permeability and promotes metastasis by targeting junction proteins. Hepatology 2018, 68, 1459–1475.
- Li, B.; Mao, R.; Liu, C.; Zhang, W.; Tang, Y.; Guo, Z. LncRNA FAL1 promotes cell proliferation and migration by acting as a CeRNA of miR-1236 in hepatocellular carcinoma cells. Life Sci. 2018, 197, 122–129.
- Zhang, Z.; Li, X.; Sun, W.; Yue, S.; Yang, J.; Li, J.; Ma, B.; Wang, J.; Yang, X.; Pu, M.; et al. Loss of exosomal miR-320a from cancer-associated fibroblasts contributes to HCC proliferation and metastasis. Cancer Lett. 2017, 397, 33–42.
- Aucher, A.; Rudnicka, D.; Davis, D.M. MicroRNAs Transfer from Human Macrophages to Hepato-Carcinoma Cells and Inhibit Proliferation. J. Immunol. 2013, 191, 6250–6260.
- Chiba, M.; Kimura, M.; Asari, S. Exosomes secreted from human colorectal cancer cell lines contain mRNAs, microRNAs and natural antisense RNAs, that can transfer into the human hepatoma HepG2 and lung cancer A549 cell lines. Oncol. Rep. 2012, 28, 1551–1558.
- Kogure, T.; Lin, W.-L.; Yan, I.K.; Braconi, C.; Patel, T. Intercellular nanovesicle-mediated microRNA transfer: A mechanism of environmental modulation of hepatocellular cancer cell growth. Hepatology 2011, 54, 1237–1248.
- Qian, X.; Xu, C.; Fang, S.; Zhao, P.; Wang, Y.; Liu, H.; Yuan, W.; Qi, Z. Exosomal MicroRNAs Derived from Umbilical Mesenchymal Stem Cells Inhibit Hepatitis C Virus Infection. Stem Cells Transl. Med. 2016, 5, 1190–1203.
- Xiong, L.; Zhen, S.; Yu, Q.; Gong, Z. HCV-E2 inhibits hepatocellular carcinoma metastasis by stimulating mast cells to secrete exosomal shuttle microRNAs. Oncol. Lett. 2017, 14, 2141–2146.
- Deng, Z.B.; Zhuang, X.; Ju, S.; Xiang, X.; Mu, J.; Liu, Y.; Jiang, H.; Zhang, L.; Mobley, J.; McClain, C.; et al. Exosome-like nanoparticles from intestinal mu-cosal cells carry prostaglandin E2 and suppress activation of liver NKT cells. J. Immunol. 2013, 190, 3579–3589.
- Li, T.; Yan, Y.; Wang, B.; Qian, H.; Zhang, X.; Shen, L.; Wang, M.; Zhou, Y.; Zhu, W.; Li, W.; et al. Exosomes Derived from Human Umbilical Cord Mesenchymal Stem Cells Alleviate Liver Fibrosis. Stem Cells Dev. 2013, 22, 845–854.
- Tan, C.Y.; Lai, R.C.; Wong, W.; Dan, Y.Y.; Lim, S.-K.; Ho, H.K. Mesenchymal stem cell-derived exosomes promote hepatic regeneration in drug-induced liver injury models. Stem Cell Res. Ther. 2014, 5, 76.
- Wang, S.; Wang, J.-Q.; Lv, X.-W. Exosomal miRNAs as biomarkers in the diagnosis of liver disease. Biomark. Med. 2017, 11, 491–501.
- Lou, G.; Song, X.; Yang, F.; Wu, S.; Wang, J.; Chen, Z.; Liu, Y. Exosomes derived from miR-122-modified adipose tissue-derived MSCs increase chemosensitivity of hepatocellular carcinoma. J. Hematol. Oncol. 2015, 8, 122.
- Lou, G.; Chen, L.; Xia, C.; Wang, W.; Qi, J.; Li, A.; Zhao, L.; Chen, Z.; Zheng, M.; Liu, Y. MiR-199a-modified exosomes from adipose tissue-derived mesenchymal stem cells improve hepatocellular carcinoma chemosensitivity through mTOR pathway. J. Exp. Clin. Cancer Res. 2020, 39, 4.
- Wang, F.; Li, L.; Piontek, K.; Sakaguchi, M.; Selaru, F.M. Exosome miR-335 as a novel therapeutic strategy in hepatocellular carcinoma. Hepatology 2018, 67, 940–954.
More
Information
Subjects:
Biochemistry & Molecular Biology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
736
Revisions:
2 times
(View History)
Update Date:
26 May 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





