| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Arianna Di Stadio | + 3380 word(s) | 3380 | 2020-12-15 09:44:23 | | | |
| 2 | Rita Xu | -1668 word(s) | 1712 | 2020-12-16 05:29:50 | | |
Video Upload Options
Coronavirus disease 2019 (COVID-19) is caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and is characterized by variable clinical presentation that ranges from asymptomatic to fatal multi-organ damage. The site of entry and the response of the host to the infection aect the outcomes. The role of the upper airways and the nasal barrier in the prevention of infection is increasingly being recognized. Besides the epithelial lining and the local immune system, the upper airways harbor a community of microorganisms, or microbiota, that takes an active part in mucosal homeostasis and in resistance to infection. However, the role of the upper airway microbiota in COVID-19 is not yet completely understood and likely goes beyond protection from viral entry to include the regulation of the immune response to the infection. Herein, we discuss the hypothesis that restoring endogenous barriers and anti-inflammatory pathways that are defective in COVID-19 patients might represent a valid strategy to reduce infectivity and ameliorate clinical symptomatology.
1. Introduction
Coronavirus disease 2019 (COVID-19), similarly to the flu virus, is transmitted through micro drops produced by sneezing, coughing, or simply speaking. The virus penetrates in the host through the upper airways, and the nasal barrier is the first defensive line to avoid infection [1]. Once the virus has entered target cells in the respiratory tract, the replication, maturation, and release of the virus occur while the host cell undergoes pyroptosis with the release of inflammatory molecules [2]. These initial events initiate an immune response that may lead to the resolution of the infection or, if dysfunctional, trigger a cytokine storm that exacerbates lung inflammation and increases the susceptibility to secondary bacterial or fungal infections [2]. In addition, the cytokine storm may cause potentially fatal multi-organ damage [2].
Host characteristics can influence the infectivity, severity, and outcomes of SARS-CoV-2 infection [3], and the local and systemic immune activities play a key role in the response to the virus aggression. Age ([4] and references therein) and gender [5][6] both impact on local and systemic immunity, but while the modulation of systemic immune responses can be quite easy and rapid to obtain, improving the local upper respiratory tract (URT) immune competence is slightly complex. In fact, age and gender determine substantial variation in the URT. Aging reduces the immune response capacity in the nasal mucosa and decreases muco-nasal clearance, a fundamental process to prevent virus infection. Furthermore, Di Stadio et al. [5] have shown that estrogen stimulation (hormone/gender effect) on the URT mucosa could reduce virus virulence thanks to the improvement of both nasal clearance and the local immune response. The authors speculated that this phenomenon could explain why women, despite affected as well as men, present less severe forms of infection with relative better outcomes (i.e., lower mortality).
Gender and age also affect the constituents of gut [7][8] and skin [9] microbiota, and we speculate that they can similarly affect the normal nasal flora [10]. Normal and healthy nasal flora is important to maintain a good balance in the upper airway tract [10][11] and can be helpful to contain and contrast viral infection [12]. Thus, therapies that affect the URT immune reactivity and microbiome composition and diversity may show promise in the treatment of a variety of human diseases, including respiratory infections. Up to date, despite the benefit shown on URT infection [13][14], the fine mechanisms through which local therapies with corticosteroids, antibiotics, and a variety of immunomodulators and nutraceuticals affect the local immune reactivity and microbial composition are largely unknown.
While alterations of the gut microbiota [15][16] and the lower respiratory tract [17] have been reported in COVID-19 patients, with potential diagnostic and therapeutic implications [18][19][20][21], the relationship between COVID-19 and the nasal microbiota has received little attention. Based on a previous hypothesis that emphasized the importance of the nasal barrier to fight COVID-19 [22].
2. The Nose Ecosystem at Work: The Microbiota
The nose likely represents the major site of entry and infection by SARS-CoV-2. Indeed, most of inhaled air enters the body through the nose, and the nasal epithelium show the highest expression of the ACE2 coronavirus receptor [23][24]. The nose might also regulate the subsequent immune response and disease severity. For instance, stimulation of the nasal innate immune response by low doses of mouse hepatitis virus type 1 was able to prime lung immunity for protection against subsequent lethal SARS-CoV or influenza A virus infection [25]. The barrier function of the nasal mucosa as well as the regulation of the local and distal immune response are likely modulated by the microbiota, i.e., the communities of microorganisms that colonize all of the surfaces of the human body exposed to the external environment. The microbiota may act either locally or distally to modulate host physiological and pathological processes. For instance, in the so-called gut-lung axis, the microbiota of the gastrointestinal tract can cross-talk with the lung microbiota and both can influence the immune system to gauge the susceptibility of the host to respiratory infections [25]. Similar to other mucosal sites of the body, the nasal cavity also harbors a community of commensal microorganisms that likely represent an important player in mucosal homeostasis and protection against infection.
Several studies have analyzed the composition of the nasal microbiome in healthy adult subjects. The major phyla belong to Actinobacteria and Firmicutes, followed by Proteobacteria and Bacteroidetes [11][26][27][28][29][30][31]. At the genus level, the nares are mainly colonized by lipophilic bacteria, such as Corynebacterium, Propionibacterium (Cutibacterium), and Staphylococcus, followed by other genera such as Moraxella, Streptococcus, and Dolosigranulum [28][29][30][31][32][33][34]. It is believed that commensal bacteria in the nasal cavity protect from opportunistic pathogens by competing for space and nutrients, and also by producing toxic molecules [35]. In addition, it has been recently shown that Staphylococcus epidermidis, which increases during nasal microbiome maturation in humans, stimulates the production of antimicrobial peptides by the nasal epithelium, which efficiently reduce pathogen colonization [36]. Moreover, S. epidermidis can also promote the production of interferon λ-dependent innate immunity by normal nasal epithelial cells to protect against the influenza virus [12], although interferon λ can in turn change the composition of the nasal microbiome and favor Staphylococcus aureus superinfection [37]. Although the interactions between the host, pathogens, and commensal bacteria are complex and still hard to decipher, it is becoming increasingly clear that dysbiosis in the nasal cavity, i.e., changes in the composition of the microbial communities, modulates the susceptibility of the host to pathological conditions, including, among others, acute respiratory tract infections [35].
Different microorganisms causing respiratory tract infections have been associated with changes in the nasal microbiome and include nonviral and viral pathogens. The former includes not only bacteria, such as Streptococcus pneumoniae, where it has been shown that IL-17 modulates the composition of the nasal microbiome to influence pneumococcal colonization [38], but also fungi. Indeed, we have recently characterized the microbiome of the URT of a large population of hematological patients and identified microbial signatures associated with invasive fungal infections (manuscript submitted), a major threat in this category of patients [39]. The associations between the nasal microbiome and viral infections have received more attention, although a causal relationship could not always be established. A recent study has evaluated the nasal and throat microbiomes in patients with influenza A and B infection and household contacts, and identified microbial signatures that could predict the risk of infection [40]. Similarly, it was shown that the nasal and throat microbiomes influenced the susceptibility to influenza infection [41] and were associated with influenza symptoms and duration of shedding in a household transmission study [42]. Importantly, the nasal microbiome cannot only influence the susceptibility to infection, but may also affect the efficacy of a live attenuated influenza vaccine [43], thus modulating its therapeutic efficacy. Similar associations with the nasal microbiome have been also demonstrated for respiratory syncytial virus bronchiolitis, with changes in the bacterial composition, alone [44] or in combination with host immune response [45], associated with the severity of symptoms, and the presence of specific taxa with the risk of recurrent wheezing after severe bronchiolitis [46]. Therefore, there may be shifts in the composition of the nasal microbiota that may result in pro- or anti-inflammatory patterns with effects on the susceptibility and course of infection. This paradigm has been elegantly demonstrated for the lung microbiome, in which the acellular bronchoalveolar lavage sample was distinguished in two patterns or pneumotypes, supraglottic predominant taxa (SPT) and background predominant taxa (BPT), based on the enrichment with taxa from the upper respiratory tract or the environment, respectively [47]. Notably, the pneumotype SPT was associated with increased lung inflammation and Th17 activation, and a blunted TLR4 response [47]. The presence of microbial patterns associated with distinct inflammatory responses has been also observed in the nasal microbiota. Healthy young adults were evaluated for their nasal microbiota before and at different times upon challenge with rhinovirus type 39 [48]. Upon grouping of the nasal microbiota into six clusters based on the predominant genera, it was demonstrated that the baseline nasal microbiota was associated with distinct nasal inflammatory responses, viral load, and symptom severity upon infection [48].
While we are just beginning to understand the role of the nasal microbiota in bacterial, fungal, and viral infections, its involvement in SARS-CoV-2 infection has remained largely unexplored and may reveal peculiar properties that reflect the specific etiopathogenesis of COVID-19. A study comparing the nasopharyngeal microbiota of patients positive or negative for COVID-19 was recently published [49]. No differences were revealed in diversity or composition [49], although the small number of samples and the presence of potential confounders may have hindered the identification of distinct signatures. Nevertheless, accumulating evidence points to a role of dysbiosis in barrier impairment with increased susceptibility to infection and dysregulated immune response. In this regard, potential approaches aimed at restoring the mucosal homeostasis with direct activity on microbiome composition and/or modulation of the immune response may represent valuable options for the prevention of COVID-19, and the use of treatments not associated with side effects would be desirable to increase the efficacy/safety profile (Figure 1).
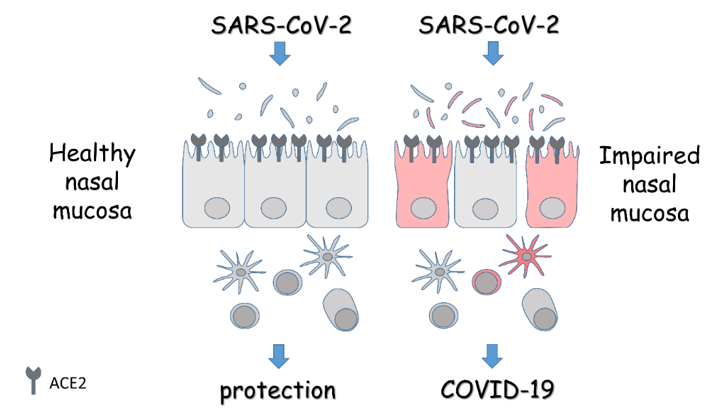
Figure 1. Schematic overview of the role of the nasal mucosa in COVID-19. The panel depicts the epithelial lining of the nasal mucosa expressing the entry receptors for SARS-CoV-2, and the associated microbiota and immune cells. In the presence of a healthy mucosa, the response to the virus is protective. On the contrary, in the presence of an impaired mucosa, the homeostasis is lost and alterations in microbiota, barrier integrity, and immune response turn the viral infection into overt disease. Details are described in the text.
References
- Baron, ; Fons, M.; Albrecht, T. Viral Pathogenesis. In Medical Microbiology; 4th ed., University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996.
- Tay, Z.; Poh, C.M.; Rénia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of COVID-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, doi:10.1038/s41577-020-0311-8.
- Di Stadio, A.; Ricci, G.; Greco, A.; de Vincentiis, M.; Ralli, M. Mortality rate and gender differences in COVID-19 patients dying in Italy: A comparison with other Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 4066–4067, doi:10.26355/eurrev_202004_20980.
- Mallapaty, S. The coronavirus is most deadly if you are older and male-new data reveal the Nature 2020, 585, 16–17, doi:10.1038/d41586-020-02483-2.
- Di Stadio, A.; Della Volpe, A.; Ralli, M.; Ricci, G. Gender differences in COVID-19 The estrogen effect on upper and lower airways. Can it help to figure out a treatment? Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 5195–5196, doi:10.26355/eurrev_202005_21298.
- Scully, E.P.; Haverfield, J.; Ursin, R.L.; Tannenbaum, C.; Klein, S.L. Considering how biological sex impacts immune responses and COVID-19 Nat. Rev. Immunol. 2020, 20, 442–447, doi:10.1038/s41577-020-0348-8.
- Yurkovetskiy, L.; Burrows, M.; Khan, A.A.; Graham, L.; Volchkov, P.; Becker, L.; Antonopoulos, D.; Umesaki, Y.; Chervonsky, A.V. Gender bias in autoimmunity is influenced by Immunity 2013, 39, 400–412, doi:10.1016/j.immuni.2013.08.013.
- Markle, J.G.; Frank, D.N.; Mortin-Toth, S.; Robertson, C.E.; Feazel, L.M.; Rolle-Kampczyk, U.; von Bergen, M.; McCoy, K.D.; Macpherson, A.J.; Danska, J.S. Sex differences in the gut microbiome drive hormone-dependent regulation of Science 2013, 339, 1084–1088, doi:10.1126/science.1233521.
- Ying, S.; Zeng, D.N.; Chi, L.; Tan, Y.; Galzote, C.; Cardona, C.; Lax, S.; Gilbert, J.; Quan, Z.X. The Influence of Age and Gender on Skin-Associated Microbial Communities in Urban and Rural Human PLoS ONE 2015, 10, e0141842, doi:10.1371/journal.pone.0141842.
- Proctor, D.M.; Relman, D.A. The Landscape Ecology and Microbiota of the Human Nose, Mouth, and Cell Host Microbe 2017, 21, 421–432, doi:10.1016/j.chom.2017.03.011.
- Kumpitsch, C.; Koskinen, K.; Schopf, V.; Moissl-Eichinger, C. The microbiome of the upper respiratory tract in health and BMC Biol. 2019, 17, 87, doi:10.1186/s12915-019-0703-z.
- Kim, H.J.; Jo, A.; Jeon, Y.J.; An, S.; Lee, K.M.; Yoon, S.S; Choi, J.Y. Nasal commensal Staphylococcus epidermidis enhances interferon-lambda-dependent immunity against influenza Microbiome 2019, 7, 80, doi:10.1186/s40168-019-0691-9.
- Della Volpe, A.; Ricci, G.; Ralli, M.; Gambacorta, V.; De Lucia, A.; Minni, A.; Pirozzi, C.; Paccone, M.; Pastore, V.; Di Stadio, A. The effects of oral supplements with Sambucus nigra, Zinc, Tyndallized Lactobacillus acidophilus (HA122), Arabinogalactans, vitamin D, vitamin E and vitamin C in otitis media with effusion in children: A randomized controlled Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 6360–6370, doi:10.26355/eurrev_201907_18460.
- Di Stadio, A.; Della Volpe, A.; Korsch, F.M.; De Lucia, A.; Ralli, M.; Martines, F.; Ricci G. Difensil Immuno Reduces Recurrence and Severity of Tonsillitis in Children: A Randomized Controlled Nutrients 2020, 126, doi:10.3390/nu12061637.
- Gu, S.; Chen, Y.; Wu, Z.; Chen, Y.; Gao, H.; Lv, L.; Guo, F.; Zhang, X.; Luo, R.; Huang, C.; et Alterations of the Gut Microbiota in Patients with COVID-19 or H1N1 Influenza. Clin. Infect. Dis. 2020, doi:10.1093/cid/ciaa709.
- Zuo, T.; Zhang, F.; Lui, G.C.Y.; Yeoh, Y.K.; Li, A.Y.L.; Zhan, H.; Wan, Y.; Chung, A.C.K.; Cheung, C.P.; Chen, N.; et Alterations in Gut Microbiota of Patients With COVID-19 During Time of Hospitalization. Gastroenterology 2020, 159, 944–955.E8, doi:10.1053/j.gastro.2020.05.048.
- Shen, Z.; Xiao, Y.; Kang, L.; Ma, W.; Shi, L.; Zhang, L.; Zhou, Z.; Yang, J.; Zhong, J.;, Yang, D.; et Genomic Diversity of Severe Acute Respiratory Syndrome-Coronavirus 2 in Patients with Coronavirus Disease 2019. Clin. Infect. Dis. 2020, 71, 713–720, doi:10.1093/cid/ciaa203.
- Akour, A. Probiotics and COVID-19: Is there any link? Appl. Microbiol. 2020, doi:10.1111/lam.13334.
- Iddir, M.; Brito, A.; Dingeo, G.; Fernandez Del Campo, S.S.; Samouda, H.; La Frano, M.R.; Bohn, T. Strengthening the Immune System and Reducing Inflammation and Oxidative Stress through Diet and Nutrition: Considerations during the COVID-19 Nutrients 2020, 126, doi:10.3390/nu12061562.
- Kalantar-Zadeh, K.; Ward, S.A.; Kalantar-Zadeh, K.; El-Omar, E.M. Considering the Effects of Microbiome and Diet on SARS-CoV-2 Infection: Nanotechnology ACS Nano 2020, 14, 5179–5182, doi:10.1021/acsnano.0c03402.
- Khatiwada, S.; Subedi, A. Lung microbiome and coronavirus disease 2019 (COVID-19): Possible link and Hum. Microb J. 2020, 17, 100073, doi:10.1016/j.humic.2020.100073.
- Ruggieri, A.; Anticoli, S.; D’Ambrosio, A.; Giordani, L.; Viora, M. The influence of sex and gender on immunity, infection and Annali Ist. Super. Sanita 2016, 52, 198–204, doi:10.4415/ANN_16_02_11.
- Hou, Y.J.; Okuda, K.; Edwards, C.E.; Martinez, D.R.; Asakura, T.; Dinnon, K.H., 3rd; Kato, T.; Lee, R.E.; Yount, B.L.; Mascenik, T.M.; et SARS-CoV-2 Reverse Genetics Reveals a Variable Infection Gradient in the Respiratory Tract. Cell 2020, 182, 429–446.e14, doi:10.1016/j.cell.2020.05.042.
- Sungnak, W.; Huang, N.; Becavin, C.; Berg, M.; Queen, R.; Litvinukova, M.; Talavera-López, C.; Maatz, H.; Reichart, D.; Sampaziotis, F.; et SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 2020, doi:10.1038/s41591-020-0868-6.
- Hua, X.; Vijay, R.; Channappanavar, R.; Athmer, J.; Meyerholz, D.K.; Pagedar, N.; Tilley, S.; Perlman, S. Nasal priming by a murine coronavirus provides protective immunity against lethal heterologous virus JCI Insight 2018, 311, doi:10.1172/jci.insight.99025.
- Bassis, C.M.; Tang, A.L.; Young, V.B.; Pynnonen, M.A. The nasal cavity microbiota of healthy Microbiome 2014, 2, 27, doi:10.1186/2049-2618-2-27.
- Brugger, S.D.; Bomar, L.; Lemon, K.P. Commensal-Pathogen Interactions along the Human Nasal PLoS Pathog. 2016, 12, e1005633, doi:10.1371/journal.ppat.1005633.
- Bomar, L.; Brugger, S.D.; Lemon, K.P. Bacterial microbiota of the nasal passages across the span of human Curr. Opin. Microbiol. 2018, 41, 8–14, doi:10.1016/j.mib.2017.10.023.
- Koskinen, K.; Reichert, J.L.; Hoier, S.; Schachenreiter, J.; Duller, S.; Moissl-Eichinger, C.; Schöpf, V. The nasal microbiome mirrors and potentially shapes olfactory Sci. Rep. 2018, 8, 1296, doi:10.1038/s41598-018-19438-3.
- Rawls, M.; Ellis, A.K. The microbiome of the Ann. Allergy Asthma Immunol. 2019, 122, 17-24, doi:10.1016/j.anai.2018.05.009.
- Hardy, B.L.; Merrell, D.S. Friend or Foe: Interbacterial Competition in the Nasal J. Bacteriol. 2020, doi:10.1128/JB.00480-20.
- Man, W.H.; de Steenhuijsen Piters, W.A.; Bogaert, D. The microbiota of the respiratory tract: Gatekeeper to respiratory Nat. Rev. Microbiol. 2017, 15, 259–270, doi:10.1038/nrmicro.2017.14.
- Escapa, I.F.; Chen, T.; Huang, Y.; Gajare, P.; Dewhirst, F.E.; Lemon, K.P. New Insights into Human Nostril Microbiome from the Expanded Human Oral Microbiome Database (eHOMD): A Resource for the Microbiome of the Human Aerodigestive mSystems 2018, 36, doi:10.1128/mSystems.00187-18.
- De Boeck, I.; van den Broek, M.F.L.; Allonsius, C.N.; Spacova, I.; Wittouck, S.; Martens, K.; Wuyts, S; Cauwenberghs, E.; Jokicevic, K.; Vandenheuvel, D.; et Lactobacilli Have a Niche in the Human Nose. Cell Rep. 2020, 31, 107674, doi:10.1016/j.celrep.2020.107674.
- Dimitri-Pinheiro, S.; Soares, R.; Barata, P. The Microbiome of the Nose-Friend or Foe? Allergy Rhinol. 2020, 11, 2152656720911605, doi:10.1177/2152656720911605.
- Liu, Q.; Liu, Q.; Meng, H.; Lv, H.; Liu, Y.; Liu, J.; Wang, H.; He, L.; Qin, J.; Wang, Y.; et Staphylococcus epidermidis Contributes to Healthy Maturation of the Nasal Microbiome by Stimulating Antimicrobial Peptide Production. Cell Host Microbe 2020, 27, 68–78.e5, doi:10.1016/j.chom.2019.11.003.
- Planet, P.J.; Parker, D.; Cohen, T.S.; Smith, H.; Leon, J.D.; Ryan, C.; Hammer, T.J.; Fierer, N.; Chen, E.I.; Prince, A.S. Lambda Interferon Restructures the Nasal Microbiome and Increases Susceptibility to Staphylococcus aureus mBio 2016, 7, e01939-15, doi:10.1128/mBio.01939-15.
- Ritchie, N.D.; Ijaz, U.Z.; Evans, T.J. IL-17 signalling restructures the nasal microbiome and drives dynamic changes following Streptococcus pneumoniae BMC Genom. 2017, 18, 807, doi:10.1186/s12864-017-4215-3.
- Pagano, L.; Busca, A.; Candoni, A.; Cattaneo, C.; Cesaro, S.; Fanci, R.; Nadali, G.; Potenza, L.; Russo, D.; Tumbarello, M.; et Risk stratification for invasive fungal infections in patients with hematological malignancies: SEIFEM recommendations. Blood Rev. 2017, 31, 17–29, doi:10.1016/j.blre.2016.09.002.
- Tsang, T.K.; Lee, K.H.; Foxman, B.; Balmaseda, A.; Gresh, L.; Sanchez, N.; Ojeda, S.; Lopez, R.; Yang, Y.; Kuan, G; et Association between the respiratory microbiome and susceptibility to influenza virus infection. Clin. Infect. Dis. 2019, doi:10.1093/cid/ciz968.
- Lee, K.H.; Gordon, A.; Shedden, K.; Kuan, G.; Ng, S.; Balmaseda, A.; Foxman, B. The respiratory microbiome and susceptibility to influenza virus PLoS ONE 2019, 14, e0207898, doi:10.1371/journal.pone.0207898.
- Lee, K.H.; Foxman, B.; Kuan, G.; Lopez, R.; Shedden, K.; Ng, S.; Balmaseda, A.; Gordon, A. The respiratory microbiota: Associations with influenza symptomatology and viral Ann. Epidemiol. 2019, 37, 51–56.e6, doi:10.1016/j.annepidem.2019.07.013.
- Salk, H.M.; Simon, W.L.; Lambert, N.D.; Kennedy, R.B.; Grill, D.E.; Kabat, B.F.; Poland, G.A. Taxa of the Nasal Microbiome Are Associated with Influenza-Specific IgA Response to Live Attenuated Influenza PLoS ONE 2016, 11, e0162803, doi:10.1371/journal.pone.0162803.
- Schippa, S.; Frassanito, A.; Marazzato, M.; Nenna, R.; Petrarca, L.; Neroni, B.; Bonfiglio, G.; Guerrieri, F.; Frasca, F.; Oliveto, G.; et Nasal Microbiota in RSV Bronchiolitis. Microorganisms 2020, 85, doi:10.3390/microorganisms8050731.
- Sonawane, A.R.; Tian, L.; Chu, C.Y.; Qiu, X.; Wang, L.; Holden-Wiltse, J.; Grier, A.; Gill, S.R.; Caserta, M.T.; Falsey, A.R.; et Microbiome-Transcriptome Interactions Related to Severity of Respiratory Syncytial Virus Infection. Sci. Rep. 2019, 9, 13824, doi:10.1038/s41598-019-50217-w.
- Mansbach, J.M.; Luna, P.N.; Shaw, C.A.; Hasegawa, K.; Petrosino, J.F.; Piedra, P.A; Sullivan, A.F.; Espinola, J.A.; Stewart, C.J.; Camargo, C.A. Jr. Increased Moraxella and Streptococcus species abundance after severe bronchiolitis is associated with recurrent J. Allergy Clin. Immunol. 2020, 145, 518–527.e8, doi:10.1016/j.jaci.2019.10.034.
- Segal, L.N.; Clemente, J.C.; Tsay, J.C.; Koralov, S.B.; Keller, B.C.; Wu, B.G.; Li, Y.; Shen, N.; Ghedin, E.; Morris, A.; et Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat. Microbiol. 2016, 1, 16031, doi:10.1038/nmicrobiol.2016.31.
- Lehtinen, M.J.; Hibberd, A.A.; Mannikko, S.; Yeung, N.; Kauko, T.; Forssten, S.; Lehtoranta, L.; Lahtinen, S.J.; Stahl, B.; Lyra, A.; et al. Nasal microbiota clusters associate with inflammatory response, viral load, and symptom severity in experimental rhinovirus Sci. Rep. 2018, 8, 11411, doi:10.1038/s41598-018-29793-w.
- De Maio, F.; Posteraro, B.; Ponziani, F.R.; Cattani, P.; Gasbarrini, A.; Sanguinetti, M. Nasopharyngeal Microbiota Profiling of SARS-CoV-2 Infected Biol. Proced. Online 2020, 22, 18, doi:10.1186/s12575-020-00131-7.