1000/1000
Hot
Most Recent

Metformin is a metabolic modulator widely used to treat type II diabetes and metabolic syndrome patients. It shows a safe profile and its use in additional indications, such as cancer, is an important matter of investigation.
Metformin (N′,N′-dimethylbiguanide) represents the most frequently administered drug to treat patients with metabolic syndrome and type 2 diabetes, worldwide. Its use has spanned over 60 years and is partially due to a very positive risk–benefit profile [1][2][3][4]. Systemically, metformin therapy lowers blood glucose in type 2 diabetes patients by targeting hepatic gluconeogenesis and by increasing glucose uptake in the peripheral tissues, mainly muscles, and indirectly reduces the insulin blood levels by counteracting insulin resistance[3][5]. Before digging into the relationships between metformin and cancer, we believe it may be useful to quickly review the main mechanism of action (MoA) of metformin in diabetic, non-cancer patients.
Undisputedly, the finding that metformin indirectly activates the adenosine 5′-monophosphate protein kinase (AMPK)[6][7][8], represented an important turn in the still unfinished quest for its mechanism of action (MoA)[9] (Figure 1). The “secret recipe” in metformin’s MoA consists of its hydrophilic nature, cationic behavior, Fe and Cu-binding properties and a pKa within the physiological pH range[10]. Metformin accumulates in the mitochondria of intact cells by virtue of its positive charge[11](Figure 1), causing inhibition of complex 1 of the respiratory chain [12]. In detail, metformin interferes with the coupling of redox and proton transfer domains in complex 1, resulting in altered redox status at the mitochondria and cytosol [10][13] and reactive oxygen species (ROS) accumulation [11][14]. The reduced phosphorylation of adenine nucleotides and accumulation of AMP allosterically determines the liver kinase B1-STE20-related pseudokinase-calcium binding protein-39 (LKB1-STRAD-CAB39)-mediated activation of AMPK[15]. Furthermore, “non-canonical” activation of AMPK takes place at lysosomes and is triggered either by changes in fructose-1,6-bisphosphate (F1, 6P2) which affects LKB1-mediated phosphorylation [16], or by galectin-9-promoted transforming growth factor-β-activated kinase-1 (TAK1)-mediated phosphorylation [17], the latter being linked to induction of autophagy[18][19]. Interestingly, activation of AMPK seems to be spatially and temporally regulated: a mild increase in AMP may activate the cytoplasmic and lysosomal pool while a more sustained increase in AMP may promote phosphorylation of AMPK by the LKB1 complex in mitochondria[20]. This may have a functional consequence in light of the topographic constraints and gradients existing within both normal tissues and in the tumor mass.
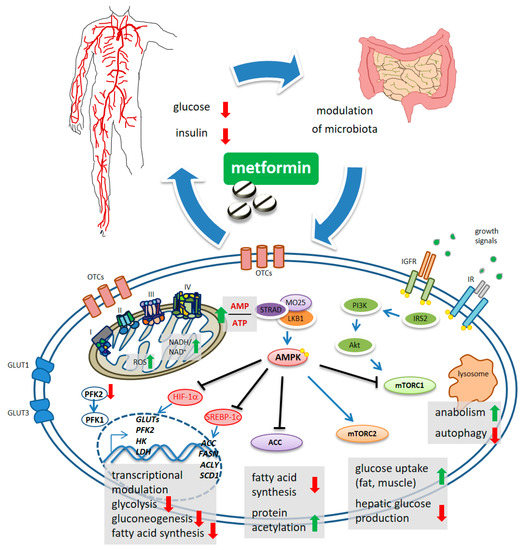
Figure 1. Schematic model of metformin action. Systemically, metformin lowers the glucose levels and, indirectly, reduces insulin levels. Metformin crosses the plasma membrane at least partially through organic cation transporters (OCTs and related) and enters mitochondria where it affects complex 1 coupling and causes altered redox status and increased AMP/ATP ratio. This latter activates, allosterically, the kinase LKB1 which phosphorylates AMPK. mTORC1 functions as an environmental sensor and is activated by insulin signaling and growth factors to promote anabolic metabolism and to inhibit autophagy. Metformin-stimulated AMPK reverses mTORC1 actions. mTORC2 is activated by AMPK and stimulates increased glucose entry into muscles and reduced glucose production in the liver. ACC inhibition reduces fatty acid synthesis and may increase collective protein acetylation (increased acetyl-CoA) thus exerting transcriptional modulation. Metformin-stimulated-AMPK controls transcriptionally hepatic gluconeogenesis. Metformin interferes with the increase in HIF-1α and the control of glycolytic genes and GLUT transporters, including the expression of PFK2 which increases the levels of the metabolite fructose-2,6-P2, allosterically activating PFK-1. Downregulated expression of ACC, FASN, ACLY and SCD1 was promoted by metformin, through interference with SREBP-1c and TR4.
Regarding the inhibition of gluconeogenesis, recent evidence shows that the early, acute downregulation of gluconeogenesis is AMPK independent and is possibly linked to compromised functioning of complex 1 that causes unbalanced NADH/NAD in mitochondria and an altered cytosolic redox state[20][21] and/or by inhibition of the mitochondrial glycerol-phosphate dehydrogenase (mGPD)[22].
The late inhibition of gluconeogenesis happens transcriptionally and is AMPK dependent. In this latter process, the small heterodimer partner (SHP), a transcriptional co-repressor, and phosphorylation of CREB binding protein (CBP) by AMPK-PKCι/λ (protein kinase C) were shown to play critical roles[23][24][25][26] (Figure 1). Additionally, even with some context specificity, AMPK targets additional metabolic enzymes such as HK2, glycogen synthase (GS), and hydroxy-methyl-glutaryl-CoA reductase (HMGCR)[27][28]. Metformin also downregulated the expression of glucose transporters (GLUT1, GLUT3) and of glycolytic enzymes such as hexokinase 2 (HK2), 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4 (PFKFB4), pyruvate kinase (PKM) and lactate dehydrogenase A(LDH)[29] . In hepatocarcinoma cells (HCC), metformin inhibited phosphofructokinase-1 (PFK1) activity by suppressing the expression of PFK2, thereby reducing the allosteric activation of PFK1 by fructose-2,6-bisphosphate[30] (Figure 1). These effects were achieved through metformin-induced inhibition of HIF-1α activity and its binding to the hypoxia-responsive elements (HRE) within the promoter region of these genes[31]. Collectively, the effect of metformin-activated AMPK accounts for increased catabolism and decreased anabolism by modulating protein synthesis, lipid homeostasis, glycolysis—and mitochondrial homeostasis, in addition to transcriptionally modulating gluconeogenesis (Figure 1).
Mammalian target of rapamycin (mTOR) is the core of two, functionally distinct, multiprotein complexes, mTOR complex 1 (mTORC1) and mTORC2[32], oppositely modulated by metformin via AMPK[33][34]. mTORC1 activation exerts anabolic effects (through increased ribosome biogenesis, lipid, nucleotide and protein synthesis) and suppresses autophagy. The mTORC1 is activated by insulin and growth factors via phosphatidyl-inositol 3-kinase (PI3K)/AKT[34]. Activated AMPK directly phosphorylates tuberous sclerosis complex 2 (TSC2), thereby inhibiting mTORC1 [[33]. mTORC1 signaling can also be inhibited by a metformin-sensitive Ras-related GTPase, as shown in mouse embryo fibroblasts[35]. On the other hand, metformin-stimulated AMPK activates mTORC2, thereby promoting cell survival and systemically reducing hepatic glucose production[36].
Another direct AMPK target protein highly relevant for cellular energy consumption, is the acetyl-CoA carboxylase (ACC), deactivated by AMPK via phosphorylation[37]. The inhibition of ACC activity decreased fatty acid synthesis consequent to a reduced conversion of acetyl-CoA to malonyl-CoA[38] (Figure 1). ACC inhibition increases collective protein acetylation and thus, may exert transcriptional modulation[37][38] (Figure 1). Additionally, phosphorylation by metformin-activated AMPK inhibited the proteasome-mediated degradation of insulin-induced gene 1 (Insig-1), which in turn reduced the activating cleavage of the transcription factor sterol regulatory element-binding protein-1c (SREBP-1c) and consequently, reduced lipogenic gene expression[39](Figure 1). In a mouse hepatoma model, metformin decreased de novo fatty acid synthesis by reducing, transcriptionally, the expression of acetyl CoA carboxylase, fatty acid synthase (FASN) and ATP citrate lyase (ACLY)[40]. Additionally, metformin impaired—AMPK-dependently—the binding and transactivation of the nuclear receptor TR4 to its responsive elements in the stearoyl-CoA desaturase-1 (SCD1) promoter, in hepatocytes[41] (Figure 1).
Remodeling the gut microbiota mediates the therapeutic effects of metformin and is responsible for its known gastrointestinal side effects[42][43]. In fact, there is long-known evidence showing that the full glucose-lowering effect of metformin is bound to its oral administration[44] and that antibiotics may blunt the effect of metformin in animal models[45]. Additionally, the concentration of metformin in the jejunum was estimated to be from ten to a few hundred times higher than in plasma[46]. The changes in microbiota elicited by metformin in diabetic patients can be relevant for its anticancer ac[47]tion, given the involvement of gut microbiota in the pathogenesis of colorectal cancer (CRC) and other solid tumors[47]. For instance, an effect of metformin on the abundance of intestinal Akkermansia muciniphila has been reproducibly reported[45][[48]. A. Muciniphila was shown to increase the abundance of gut-targeted CD4+ T cells, providing an adjuvant effect to the action of anti- programmed cell death -1 (PD-1) agents, in animal models of melanoma and non-small cell lung cancer (NSCLC)[49] and in a model of microsatellite-stable (MSS) colorectal cancer[50].