You're using an outdated browser. Please upgrade to a modern browser for the best experience.
Encyclopedia:
From Scholars for Scholars
Trending Entry
Top Viewed Entry
News and Events
More >>
Announcement
21 Apr 2025
We are thrilled to announce the Encyclopedia 2024 Outstanding Contributor Award winners. The award has been granted to the following researchers:
Name: Dr. Michele Filippo FontefrancescoAffiliations: 1 Catholic University of Sacred Hearth, Italy; 2 Durham University, UK
Name: Dr. Ioannis PassasAffiliation: Hellenic Mediterranean University, Greece
As awardees, they will each receive CHF 300 (Swiss francs), a certificate, and an opportunity to publish one paper free of charge in the Encyclopedia journal before 31 December 2025, following peer review.
Testimonials from the winners:
Dr. Michele Filippo Fontefrancesco: “I am honored to receive the Outstanding Contribution Award from Encyclopedia. This recognition holds special significance for me, both personally and professionally, as it expresses the recognition of the contribution given to the growth of a young international journal.Publishing with Encyclopedia has offered me a unique opportunity to communicate complex anthropological and social science topics in accessible ways and reach audiences far beyond my normal space in academia. As a researcher committed to bridging knowledge and action, I deeply appreciate the journal’s commitment to high-quality, open access publishing and its support in shaping content that resonates with scholars, practitioners, and policymakers alike.In my experience, the professionalism of the Encyclopedia’s editorial team, from peer review to publication and video production, is outstanding. As an Editorial Board Member and contributor, I see Encyclopedia as a platform where scientific rigor meets public engagement—an essential frontier for researchers today.This award strengthens my motivation to continue contributing, mentoring, and fostering a culture of inclusive and innovative science communication. Thank you for this meaningful acknowledgment.”
Dr. Ioannis Passas: “Receiving the Encyclopedia Outstanding Contributor Award 2024 is a true honor. What makes this recognition particularly meaningful is the journal’s wide-reaching, interdisciplinary nature. Few academic platforms today manage to bring together so many diverse fields under one roof while maintaining high standards and clear editorial direction. Being part of such an ecosystem—where knowledge moves across disciplines and formats—is both intellectually fulfilling and professionally valuable.This award acknowledges more than a single contribution; it reflects a collaborative process involving editors, reviewers, and technical staff who treat each submission with care and respect. I’m especially grateful for the chance to share ideas in both written and audiovisual forms, as this has expanded the impact of my work far beyond traditional academic circles.I appreciate this recognition deeply and look forward to continuing my involvement with Encyclopedia, supporting its mission to foster open, informed, and inclusive scholarly dialogue.”
We extend our sincere gratitude to all applicants for their outstanding contributions. The evaluation process and subsequent decision proved to be challenging. On behalf of the award evaluation committee, we congratulate the winners on their remarkable achievements.
Encyclopedia Editorial Office
Ongoing
07 Apr 2025
Encyclopedia MDPI is thrilled to announce significant enhancements to its Academic Video Service, which aim to improve its quality, accessibility, and functionality. Since its launch, our video service has enabled numerous scholars to present their research in a dynamic and visually engaging format, greatly enhancing its visibility and impact.
Due to the overwhelmingly positive reception this service has received, we have reached a point where the number of orders we are receiving exceeds our current capacity. In order to maintain the quality of these videos and continue optimizing the service, we have made the decision to introduce a fee. However, to ensure that this service is still a cost-effective option, we have set our prices significantly below the market average.
Highlights of the Upgrades to the Service
Although the service will now be fee-based, we are committed to providing even more professional and comprehensive support, including the following:
One-on-one video production guidance
Personalized assistance to ensure your needs are fully met.
Scriptwriting and English editing
Expertly crafted narratives and professional English editing to ensure your research is presented clearly, accurately, and with impact.
High-quality animations
Visually engaging animations are created to simplify complex research and captivate your audience.
Whiteboard Animations: Clean and minimalist, using hand-drawn illustrations to explain ideas step-by-step.
Motion Graphics (MG) Animations:
Cartoon Style: Bright, colorful, and approachable, ideal for making technical or scientific content more accessible and engaging.
Hand-Drawn Style: Unique and artistic, adding a personal touch to your research while maintaining clarity and professionalism.
Customized infographics (optional)
We can also create tailored infographics to visually summarize key data or findings, enhancing the clarity and appeal of your video.
Native voiceover
Native speakers provide voiceovers to enhance the accessibility and reach of your research.
Multiple rounds of revision
To ensure your video accurately represents your work.
Social media promotion
Expanding your research's visibility and impact.
Why Choose Us?
The Proven Impact of Video Abstracts
Research shows that a well-crafted video abstract can significantly enhance the visibility and impact of your work. It has been shown to do the following:
Increase paper views by 120% (Source: 10.1007/ s11192-019-03108-w)
Boost citations by 20% (Source: Wiley Online Library)
Improve journal rankings by 33% (Source: Research Square)
Raise Altmetrics scores by 140% (Source: Research Square)
Our Expertise in Academic Research
Backed by MDPI, our experienced production team combines deep academic knowledge with creative excellence. We understand the nuances of scholarly communication and ensure that every frame accurately conveys the value of your research, meeting the highest standards of quality and precision.
Collaborations with SCI Journals
We have partnered with many SCI journals to create exclusive video series, enhancing the dissemination and impact of published research. For example, our collaborations with Entropy, Remote Sensing, Nanomaterials , Animals , Nutrients, Foods , Sustainability, Encyclopedia, Cancers, etc., have helped authors achieve greater visibility and recognition for their work.
Global visibility
The videos are linked to your paper's DOI for maximum exposure.
Available Video Services and Their Pricing
Video Abstract (up to 5 minutes long): Summarizes the key findings, methodology, and significance of your research paper.
Regular price: CHF 600
Discounted Price: CHF 400
Short Take (up to 2 minutes long): Uses original animations to explain the specific aspects of your research.
Regular price: CHF 500
Discounted Price: CHF 300
Scholar Interview: A face-to-face discussion offering deeper insights into your publication.
Regular price: CHF 400
Discounted Price: CHF 200
Special Offer
To express our gratitude for your support, we are pleased to offer a buy one get one free promotion. This offer is valid for two weeks, and the complimentary service provided can be utilized within one year.
Video Production Service
If you want to see some examples of our videos, please visit https://encyclopedia.pub/video.
If you would like to apply for the video service, please click https://encyclopedia.pub/video_service.
Others
If you have any other questions, please contact office@encyclopedia.pub.
Announcement
25 Apr 2025
Every year on April 25, the world observes World Malaria Day—a reminder that while progress has been made, malaria remains a global threat.
In 2022, malaria infected 247 million people and claimed over 619,000 lives—mostly children under 5 in sub-Saharan Africa. It's a disease we can prevent, treat, and eliminate, but it still kills a child every minute.
1. What Is Malaria?
Malaria is caused by Plasmodium parasites, spread by the bite of an infected Anopheles mosquito. Its symptoms—fever, chills, vomiting—can quickly turn deadly without treatment.
But we're not powerless. Over the past two decades, thanks to tools like:
Insecticide-treated netsRapid diagnostic testsArtemisinin-based therapies (ACTs)The RTS,S malaria vaccine
…millions of lives have been saved, and some countries have eliminated the disease entirely.
Source: https://encyclopedia.pub/image/3486
2. The Game-Changer: Tu Youyou
One of the biggest breakthroughs in malaria treatment came from Tu Youyou, a Chinese scientist who turned to traditional medicine for answers.
In the 1970s, during China's Project 523, Tu discovered artemisinin from the sweet wormwood plant (Artemisia annua). This became the foundation for ACTs, now the world's most effective antimalarial drugs.
In 2015, Tu became the first Chinese woman to win a Nobel Prize in Physiology or Medicine.
Her discovery has saved millions of lives—a shining example of how ancient knowledge and modern science can work together.
3. What We're Still Up Against
Despite progress, the fight isn't over. We face:
Drug and insecticide resistance
Climate change expanding mosquito habitats
Critical funding gaps
And yet, the goal of ending malaria remains within reach—with continued innovation, investment, and global collaboration.
For more information about Tu youyou, you can view our online video "Science Never Ends: Tu youyou".
Announcement
24 Apr 2025
The Sarcidano Horse, an autochthonous breed from Sardinia, is gaining attention in conservation efforts due to its semi-feral lifestyle and genetic isolation. This breed offers important insights into the genetics of equine populations that have remained largely untouched by modern breeding practices.
1. Study Overview
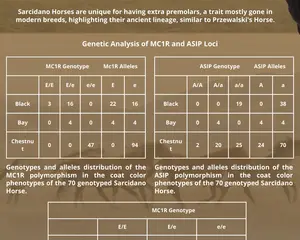
A recent study aimed to deepen the understanding of the Sarcidano Horse, specifically its coat color genetics. The focus was on two genes, MC1R and ASIP, which are responsible for determining the basic coat colors: bay, black, and chestnut.
70 horses were sampled for DNA analysis, and the study found the following:
The chestnut coat color was the most prevalent.
A smaller number of horses had black coats, with very few exhibiting bay coats.
There was no evidence of color dilution or spotting, suggesting that the population has remained genetically isolated, with little to no crossbreeding with domestic breeds.
Source: https://encyclopedia.pub/image/3410
2. Implications for Conservation
The lack of coat variations commonly associated with domesticated horses further supports the hypothesis that the Sarcidano Horse has not been affected by human selection. The breed’s genetic isolation and lack of crossbreeding are vital for its conservation, as they help preserve its unique genetic makeup.
Given the breed’s ancient origins and current genetic isolation, experts recommend a regional conservation plan. This plan would focus on:
Regular health and genetic monitoring of the population.
Collaboration with research centers and universities.
The collection and management of DNA, sperm, and embryos to support long-term conservation efforts.
3. Conclusion
The Sarcidano Horse represents an important genetic resource, offering valuable insights into the history and evolution of equine species. Continued efforts to preserve this breed, through both genetic research and conservation initiatives, are crucial to maintaining its genetic integrity and safeguarding it for future generations.
For more information about the research, you can view the online video entitled "Unveiling the Genes Behind the Sarcidano Horse's Coat Colors".
Announcement
21 Apr 2025
We are thrilled to announce the Encyclopedia 2024 Outstanding Contributor Award winners. The award has been granted to the following researchers:
Name: Dr. Michele Filippo FontefrancescoAffiliations: 1 Catholic University of Sacred Hearth, Italy; 2 Durham University, UK
Name: Dr. Ioannis PassasAffiliation: Hellenic Mediterranean University, Greece
As awardees, they will each receive CHF 300 (Swiss francs), a certificate, and an opportunity to publish one paper free of charge in the Encyclopedia journal before 31 December 2025, following peer review.
Testimonials from the winners:
Dr. Michele Filippo Fontefrancesco: “I am honored to receive the Outstanding Contribution Award from Encyclopedia. This recognition holds special significance for me, both personally and professionally, as it expresses the recognition of the contribution given to the growth of a young international journal.Publishing with Encyclopedia has offered me a unique opportunity to communicate complex anthropological and social science topics in accessible ways and reach audiences far beyond my normal space in academia. As a researcher committed to bridging knowledge and action, I deeply appreciate the journal’s commitment to high-quality, open access publishing and its support in shaping content that resonates with scholars, practitioners, and policymakers alike.In my experience, the professionalism of the Encyclopedia’s editorial team, from peer review to publication and video production, is outstanding. As an Editorial Board Member and contributor, I see Encyclopedia as a platform where scientific rigor meets public engagement—an essential frontier for researchers today.This award strengthens my motivation to continue contributing, mentoring, and fostering a culture of inclusive and innovative science communication. Thank you for this meaningful acknowledgment.”
Dr. Ioannis Passas: “Receiving the Encyclopedia Outstanding Contributor Award 2024 is a true honor. What makes this recognition particularly meaningful is the journal’s wide-reaching, interdisciplinary nature. Few academic platforms today manage to bring together so many diverse fields under one roof while maintaining high standards and clear editorial direction. Being part of such an ecosystem—where knowledge moves across disciplines and formats—is both intellectually fulfilling and professionally valuable.This award acknowledges more than a single contribution; it reflects a collaborative process involving editors, reviewers, and technical staff who treat each submission with care and respect. I’m especially grateful for the chance to share ideas in both written and audiovisual forms, as this has expanded the impact of my work far beyond traditional academic circles.I appreciate this recognition deeply and look forward to continuing my involvement with Encyclopedia, supporting its mission to foster open, informed, and inclusive scholarly dialogue.”
We extend our sincere gratitude to all applicants for their outstanding contributions. The evaluation process and subsequent decision proved to be challenging. On behalf of the award evaluation committee, we congratulate the winners on their remarkable achievements.
Encyclopedia Editorial Office
Ongoing
07 Apr 2025
Encyclopedia MDPI is thrilled to announce significant enhancements to its Academic Video Service, which aim to improve its quality, accessibility, and functionality. Since its launch, our video service has enabled numerous scholars to present their research in a dynamic and visually engaging format, greatly enhancing its visibility and impact.
Due to the overwhelmingly positive reception this service has received, we have reached a point where the number of orders we are receiving exceeds our current capacity. In order to maintain the quality of these videos and continue optimizing the service, we have made the decision to introduce a fee. However, to ensure that this service is still a cost-effective option, we have set our prices significantly below the market average.
Highlights of the Upgrades to the Service
Although the service will now be fee-based, we are committed to providing even more professional and comprehensive support, including the following:
One-on-one video production guidance
Personalized assistance to ensure your needs are fully met.
Scriptwriting and English editing
Expertly crafted narratives and professional English editing to ensure your research is presented clearly, accurately, and with impact.
High-quality animations
Visually engaging animations are created to simplify complex research and captivate your audience.
Whiteboard Animations: Clean and minimalist, using hand-drawn illustrations to explain ideas step-by-step.
Motion Graphics (MG) Animations:
Cartoon Style: Bright, colorful, and approachable, ideal for making technical or scientific content more accessible and engaging.
Hand-Drawn Style: Unique and artistic, adding a personal touch to your research while maintaining clarity and professionalism.
Customized infographics (optional)
We can also create tailored infographics to visually summarize key data or findings, enhancing the clarity and appeal of your video.
Native voiceover
Native speakers provide voiceovers to enhance the accessibility and reach of your research.
Multiple rounds of revision
To ensure your video accurately represents your work.
Social media promotion
Expanding your research's visibility and impact.
Why Choose Us?
The Proven Impact of Video Abstracts
Research shows that a well-crafted video abstract can significantly enhance the visibility and impact of your work. It has been shown to do the following:
Increase paper views by 120% (Source: 10.1007/ s11192-019-03108-w)
Boost citations by 20% (Source: Wiley Online Library)
Improve journal rankings by 33% (Source: Research Square)
Raise Altmetrics scores by 140% (Source: Research Square)
Our Expertise in Academic Research
Backed by MDPI, our experienced production team combines deep academic knowledge with creative excellence. We understand the nuances of scholarly communication and ensure that every frame accurately conveys the value of your research, meeting the highest standards of quality and precision.
Collaborations with SCI Journals
We have partnered with many SCI journals to create exclusive video series, enhancing the dissemination and impact of published research. For example, our collaborations with Entropy, Remote Sensing, Nanomaterials , Animals , Nutrients, Foods , Sustainability, Encyclopedia, Cancers, etc., have helped authors achieve greater visibility and recognition for their work.
Global visibility
The videos are linked to your paper's DOI for maximum exposure.
Available Video Services and Their Pricing
Video Abstract (up to 5 minutes long): Summarizes the key findings, methodology, and significance of your research paper.
Regular price: CHF 600
Discounted Price: CHF 400
Short Take (up to 2 minutes long): Uses original animations to explain the specific aspects of your research.
Regular price: CHF 500
Discounted Price: CHF 300
Scholar Interview: A face-to-face discussion offering deeper insights into your publication.
Regular price: CHF 400
Discounted Price: CHF 200
Special Offer
To express our gratitude for your support, we are pleased to offer a buy one get one free promotion. This offer is valid for two weeks, and the complimentary service provided can be utilized within one year.
Video Production Service
If you want to see some examples of our videos, please visit https://encyclopedia.pub/video.
If you would like to apply for the video service, please click https://encyclopedia.pub/video_service.
Others
If you have any other questions, please contact office@encyclopedia.pub.
Announcement
25 Apr 2025
Every year on April 25, the world observes World Malaria Day—a reminder that while progress has been made, malaria remains a global threat.
In 2022, malaria infected 247 million people and claimed over 619,000 lives—mostly children under 5 in sub-Saharan Africa. It's a disease we can prevent, treat, and eliminate, but it still kills a child every minute.
1. What Is Malaria?
Malaria is caused by Plasmodium parasites, spread by the bite of an infected Anopheles mosquito. Its symptoms—fever, chills, vomiting—can quickly turn deadly without treatment.
But we're not powerless. Over the past two decades, thanks to tools like:
Insecticide-treated netsRapid diagnostic testsArtemisinin-based therapies (ACTs)The RTS,S malaria vaccine
…millions of lives have been saved, and some countries have eliminated the disease entirely.
Source: https://encyclopedia.pub/image/3486
2. The Game-Changer: Tu Youyou
One of the biggest breakthroughs in malaria treatment came from Tu Youyou, a Chinese scientist who turned to traditional medicine for answers.
In the 1970s, during China's Project 523, Tu discovered artemisinin from the sweet wormwood plant (Artemisia annua). This became the foundation for ACTs, now the world's most effective antimalarial drugs.
In 2015, Tu became the first Chinese woman to win a Nobel Prize in Physiology or Medicine.
Her discovery has saved millions of lives—a shining example of how ancient knowledge and modern science can work together.
3. What We're Still Up Against
Despite progress, the fight isn't over. We face:
Drug and insecticide resistance
Climate change expanding mosquito habitats
Critical funding gaps
And yet, the goal of ending malaria remains within reach—with continued innovation, investment, and global collaboration.
For more information about Tu youyou, you can view our online video "Science Never Ends: Tu youyou".
Announcement
24 Apr 2025
The Sarcidano Horse, an autochthonous breed from Sardinia, is gaining attention in conservation efforts due to its semi-feral lifestyle and genetic isolation. This breed offers important insights into the genetics of equine populations that have remained largely untouched by modern breeding practices.
1. Study Overview
A recent study aimed to deepen the understanding of the Sarcidano Horse, specifically its coat color genetics. The focus was on two genes, MC1R and ASIP, which are responsible for determining the basic coat colors: bay, black, and chestnut.
70 horses were sampled for DNA analysis, and the study found the following:
The chestnut coat color was the most prevalent.
A smaller number of horses had black coats, with very few exhibiting bay coats.
There was no evidence of color dilution or spotting, suggesting that the population has remained genetically isolated, with little to no crossbreeding with domestic breeds.
Source: https://encyclopedia.pub/image/3410
2. Implications for Conservation
The lack of coat variations commonly associated with domesticated horses further supports the hypothesis that the Sarcidano Horse has not been affected by human selection. The breed’s genetic isolation and lack of crossbreeding are vital for its conservation, as they help preserve its unique genetic makeup.
Given the breed’s ancient origins and current genetic isolation, experts recommend a regional conservation plan. This plan would focus on:
Regular health and genetic monitoring of the population.
Collaboration with research centers and universities.
The collection and management of DNA, sperm, and embryos to support long-term conservation efforts.
3. Conclusion
The Sarcidano Horse represents an important genetic resource, offering valuable insights into the history and evolution of equine species. Continued efforts to preserve this breed, through both genetic research and conservation initiatives, are crucial to maintaining its genetic integrity and safeguarding it for future generations.
For more information about the research, you can view the online video entitled "Unveiling the Genes Behind the Sarcidano Horse's Coat Colors".
Announcement
21 Apr 2025
We are thrilled to announce the Encyclopedia 2024 Outstanding Contributor Award winners. The award has been granted to the following researchers:
Name: Dr. Michele Filippo FontefrancescoAffiliations: 1 Catholic University of Sacred Hearth, Italy; 2 Durham University, UK
Name: Dr. Ioannis PassasAffiliation: Hellenic Mediterranean University, Greece
As awardees, they will each receive CHF 300 (Swiss francs), a certificate, and an opportunity to publish one paper free of charge in the Encyclopedia journal before 31 December 2025, following peer review.
Testimonials from the winners:
Dr. Michele Filippo Fontefrancesco: “I am honored to receive the Outstanding Contribution Award from Encyclopedia. This recognition holds special significance for me, both personally and professionally, as it expresses the recognition of the contribution given to the growth of a young international journal.Publishing with Encyclopedia has offered me a unique opportunity to communicate complex anthropological and social science topics in accessible ways and reach audiences far beyond my normal space in academia. As a researcher committed to bridging knowledge and action, I deeply appreciate the journal’s commitment to high-quality, open access publishing and its support in shaping content that resonates with scholars, practitioners, and policymakers alike.In my experience, the professionalism of the Encyclopedia’s editorial team, from peer review to publication and video production, is outstanding. As an Editorial Board Member and contributor, I see Encyclopedia as a platform where scientific rigor meets public engagement—an essential frontier for researchers today.This award strengthens my motivation to continue contributing, mentoring, and fostering a culture of inclusive and innovative science communication. Thank you for this meaningful acknowledgment.”
Dr. Ioannis Passas: “Receiving the Encyclopedia Outstanding Contributor Award 2024 is a true honor. What makes this recognition particularly meaningful is the journal’s wide-reaching, interdisciplinary nature. Few academic platforms today manage to bring together so many diverse fields under one roof while maintaining high standards and clear editorial direction. Being part of such an ecosystem—where knowledge moves across disciplines and formats—is both intellectually fulfilling and professionally valuable.This award acknowledges more than a single contribution; it reflects a collaborative process involving editors, reviewers, and technical staff who treat each submission with care and respect. I’m especially grateful for the chance to share ideas in both written and audiovisual forms, as this has expanded the impact of my work far beyond traditional academic circles.I appreciate this recognition deeply and look forward to continuing my involvement with Encyclopedia, supporting its mission to foster open, informed, and inclusive scholarly dialogue.”
We extend our sincere gratitude to all applicants for their outstanding contributions. The evaluation process and subsequent decision proved to be challenging. On behalf of the award evaluation committee, we congratulate the winners on their remarkable achievements.
Encyclopedia Editorial Office
Featured Images
More >>
Encyclopedia Editorial Office
- 15 Apr 2025
Journal Encyclopedia - Peer-Reviewed Content
More >>
Peer Reviewed
Encyclopedia 2025, 5(2), 45; https://doi.org/10.3390/encyclopedia5020045
Encyclopedia 2025, 5(2), 43; https://doi.org/10.3390/encyclopedia5020043
Peer Reviewed
Encyclopedia 2025, 5(2), 54; https://doi.org/10.3390/encyclopedia5020054
Peer Reviewed
Encyclopedia 2025, 5(2), 45; https://doi.org/10.3390/encyclopedia5020045
Encyclopedia 2025, 5(2), 43; https://doi.org/10.3390/encyclopedia5020043
Peer Reviewed
Encyclopedia 2025, 5(2), 54; https://doi.org/10.3390/encyclopedia5020054
Peer Reviewed
Encyclopedia 2025, 5(2), 45; https://doi.org/10.3390/encyclopedia5020045
Encyclopedia 2025, 5(2), 43; https://doi.org/10.3390/encyclopedia5020043
See what people are saying about us
Shlomi Agmon
Encyclopedia Video provides potential readers with a tool to quickly understand what the work is about. That is important for casualreaders, whose time is thus spared, and for investedreaders, for whom it makes the decision to say "yes, I want to read the paper" much simpler.
School of Computer Science and Engineering, The Hebrew University of Jerusalem, Jerusalem 9190401, Israel
Ignacio Cea
For the video abstracts, the papers and authors could gain more visibility and increase citations. Also, it means a more diverse and interesting way of communicating research, which is something valuable in itself.
Center for Research, Innovation and Creation, and Faculty of Religious Sciences and Philosophy, Temuco Catholic University
Melvin R. Pete Hayden
Thank the video production crew for making such a wonderful video. The narrations have been significantly added to the video! Congratulations on such an outstanding job of Encyclopedia Video team.
University of Missouri School of Medicine, United States