
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Eva Turley | + 1598 word(s) | 1598 | 2021-10-26 08:21:00 | | | |
| 2 | Lindsay Dong | Meta information modification | 1598 | 2021-11-29 08:50:28 | | | | |
| 3 | Lindsay Dong | Meta information modification | 1598 | 2021-11-29 08:50:49 | | |
Video Upload Options
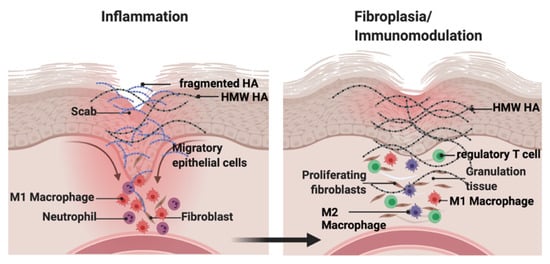
Hyaluronan (HA) is a major component of the ECM that consists of repeated disaccharide units of N-acetylglucosamine and glucuronic acid. HA is synthesized as a high molecular weight molecule but is degraded into heterogeneous fragments by hyaluronidases and reactive oxygen or nitrogen species. The biological activity of HA depends on its molecular weight. HA fragments stimulate tissue inflammation and fibrosis. In contrast, high molecular weight HA suppresses these processes and maintains tissue homeostasis. This functional duality is particularly important during wound repair where HA sequentially promotes then suppresses inflammation and fibrosis. In contrast, in tumors, HA effects are often co-opted to increase growth and invasion.
1. Background
2. The Hyaluronome
2.1. Hyaluronan
2.2. Hyaluronan Synthases
2.3. Hyaluronidases
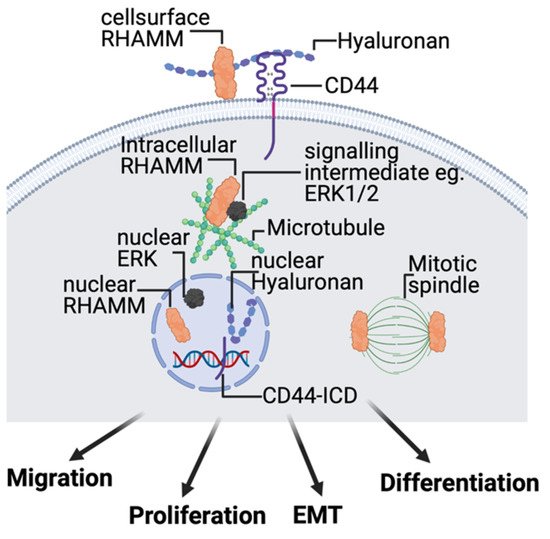
2.4. Hyaluronan Receptors, CD44, and RHAMM
References
- Dvorak, H.F. Tumors: Wounds That Do Not Heal–A Historical Perspective with a Focus on the Fundamental Roles of Increased Vascular Permeability and Clotting. Semin. Thromb. Hemost. 2019, 45, 576–592.
- Bissell, M.J.; Hines, W.C. Why don’t we get more cancer? A proposed role of the microenvironment in restraining cancer progression. Nat. Med. 2011, 17, 320–329.
- Allen, M.D.; Jones, L.J. The role of inflammation in progression of breast cancer: Friend or foe? Int. J. Oncol. 2015, 47, 797–805.
- Nakamura, N. A hypothesis: Radiation carcinogenesis may result from tissue injuries and subsequent recovery processes which can act as tumor promoters and lead to an earlier onset of cancer. Br. J. Radiol. 2020, 93, 20190843.
- Liu, M.; Tolg, C.; Turley, E. Dissecting the Dual Nature of Hyaluronan in the Tumor Microenvironment. Front. Immunol. 2019, 10, 947.
- Abatangelo, G.; Vindigni, V.; Avruscio, G.; Pandis, L.; Brun, P. Hyaluronic Acid: Redefining Its Role. Cells 2020, 9, 1743.
- Muto, J.; Sayama, K.; Gallo, R.L.; Kimata, K. Emerging evidence for the essential role of hyaluronan in cutaneous biology. J. Derm. Sci. 2019, 94, 190–195.
- Tolg, C.; McCarthy, J.B.; Yazdani, A.; Turley, E.A. Hyaluronan and RHAMM in wound repair and the “cancerization” of stromal tissues. Biomed. Res. Int. 2014, 2014, 103923.
- Turley, E.A.; Wood, D.K.; McCarthy, J.B. Carcinoma Cell Hyaluronan as a “Portable” Cancerized Prometastatic Microenvironment. Cancer Res. 2016, 76, 2507–2512.
- Kavasi, R.M.; Berdiaki, A.; Spyridaki, I.; Corsini, E.; Tsatsakis, A.; Tzanakakis, G.; Nikitovic, D. HA metabolism in skin homeostasis and inflammatory disease. Food Chem. Toxicol. 2017, 101, 128–138.
- Litwiniuk, M.; Krejner, A.; Speyrer, M.S.; Gauto, A.R.; Grzela, T. Hyaluronic Acid in Inflammation and Tissue Regeneration. Wounds 2016, 28, 78–88.
- Balazs, E.A. Hyaluronan as an ophthalmic viscoelastic device. Curr. Pharm. Biotechnol. 2008, 9, 236–238.
- Balazs, E.A.; Högberg, B.; Laurent, T.C. The Biological Activity of Hyaluron Sulfuric Acid. Acta Physiol. Scand. 1951, 23, 168–178.
- Snetkov, P.; Zakharova, K.; Morozkina, S.; Olekhnovich, R.; Uspenskaya, M. Hyaluronic Acid: The Influence of Molecular Weight on Structural, Physical, Physico–Chemical, and Degradable Properties of Biopolymer. Polymers 2020, 12, 1800.
- Valachova, K.; Soltes, L. Hyaluronan as a Prominent Biomolecule with Numerous Applications in Medicine. Int. J. Mol. Sci. 2021, 22, 7077.
- Ruiz Martinez, M.A.; Peralta Galisteo, S.; Castan, H.; Morales Hernandez, M.E. Role of proteoglycans on skin ageing: A review. Int. J. Cosmet. Sci. 2020, 42, 529–535.
- Turley, E.A. Hyaluronic acid stimulates protein kinase activity in intact cells and in an isolated protein complex. J. Biol. Chem. 1989, 264, 8951–8955.
- Aruffo, A.; Stamenkovic, I.; Melnick, M.; Underhill, C.B.; Seed, B. CD44 is the principal cell surface receptor for hyaluronate. Cell 1990, 61, 1303–1313.
- Hardwick, C.; Hoare, K.; Owens, R.; Hohn, H.P.; Hook, M.; Moore, D.; Cripps, V.; Austen, L.; Nance, D.M.; Turley, E.A. Molecular cloning of a novel hyaluronan receptor that mediates tumor cell motility. J. Cell Biol. 1992, 117, 1343–1350.
- Turley, E.A.; Noble, P.W.; Bourguignon, L.Y. Signaling properties of hyaluronan receptors. J. Biol. Chem. 2002, 277, 4589–4592.
- Schwertfeger, K.L.; Cowman, M.K.; Telmer, P.G.; Turley, E.A.; McCarthy, J.B. Hyaluronan, Inflammation, and Breast Cancer Progression. Front. Immunol. 2015, 6, 236.
- Kaul, A.; Short, W.D.; Keswani, S.G.; Wang, X. Immunologic Roles of Hyaluronan in Dermal Wound Healing. Biomolecules 2021, 11, 1234.
- Li, N.; Geng, C.; Hou, S.; Fan, H.; Gong, Y. Damage–Associated Molecular Patterns and Their Signaling Pathways in Primary Blast Lung Injury: New Research Progress and Future Directions. Int. J. Mol. Sci. 2020, 21, 6303.
- Roedig, H.; Damiescu, R.; Zeng–Brouwers, J.; Kutija, I.; Trebicka, J.; Wygrecka, M.; Schaefer, L. Danger matrix molecules orchestrate CD14/CD44 signaling in cancer development. Semin. Cancer Biol. 2020, 62, 31–47.
- Garantziotis, S.; Savani, R.C. Hyaluronan biology: A complex balancing act of structure, function, location and context. Matrix Biol. 2019, 78, 1–10.
- Savani, R.C. Modulators of inflammation in Bronchopulmonary Dysplasia. Semin. Perinatol. 2018, 42, 459–470.
- Camenisch, T.D.; Spicer, A.P.; Brehm–Gibson, T.; Biesterfeldt, J.; Augustine, M.L.; Calabro, A., Jr.; Kubalak, S.; Klewer, S.E.; McDonald, J.A. Disruption of hyaluronan synthase–2 abrogates normal cardiac morphogenesis and hyaluronan–mediated transformation of epithelium to mesenchyme. J. Clin. Investig. 2000, 106, 349–360.
- Bai, K.J.; Spicer, A.P.; Mascarenhas, M.M.; Yu, L.; Ochoa, C.D.; Garg, H.G.; Quinn, D.A. The role of hyaluronan synthase 3 in ventilator–induced lung injury. Am. J. Respir. Crit. Care Med. 2005, 172, 92–98.
- Kobayashi, N.; Miyoshi, S.; Mikami, T.; Koyama, H.; Kitazawa, M.; Takeoka, M.; Sano, K.; Amano, J.; Isogai, Z.; Niida, S.; et al. Hyaluronan deficiency in tumor stroma impairs macrophage trafficking and tumor neovascularization. Cancer Res. 2010, 70, 7073–7083.
- Mack, J.A.; Feldman, R.J.; Itano, N.; Kimata, K.; Lauer, M.; Hascall, V.C.; Maytin, E.V. Enhanced inflammation and accelerated wound closure following tetraphorbol ester application or full–thickness wounding in mice lacking hyaluronan synthases Has1 and Has3. J. Investig. Dermatol. 2012, 132, 198–207.
- Sindelar, M.; Jilkova, J.; Kubala, L.; Velebny, V.; Turkova, K. Hyaluronidases and hyaluronate lyases: From humans to bacteriophages. Colloids Surf. B Biointerfaces 2021, 208, 112095.
- Triggs–Raine, B.; Natowicz, M.R. Biology of hyaluronan: Insights from genetic disorders of hyaluronan metabolism. World J. Biol. Chem. 2015, 6, 110–120.
- Pibuel, M.A.; Poodts, D.; Diaz, M.; Hajos, S.E.; Lompardia, S.L. The scrambled story between hyaluronan and glioblastoma. J. Biol. Chem. 2021, 296, 100549.
- Piperigkou, Z.; Kyriakopoulou, K.; Koutsakis, C.; Mastronikolis, S.; Karamanos, N.K. Key Matrix Remodeling Enzymes: Functions and Targeting in Cancer. Cancers 2021, 13, 1441.
- Irie, F.; Tobisawa, Y.; Murao, A.; Yamamoto, H.; Ohyama, C.; Yamaguchi, Y. The cell surface hyaluronidase TMEM2 regulates cell adhesion and migration via degradation of hyaluronan at focal adhesion sites. J. Biol. Chem. 2021, 296, 100481.
- Liu, J.; Yan, W.; Han, P.; Tian, D. The emerging role of KIAA1199 in cancer development and therapy. Biomed. Pharmacother. 2021, 138, 111507.
- Bono, P.; Rubin, K.; Higgins, J.M.; Hynes, R.O. Layilin, a novel integral membrane protein, is a hyaluronan receptor. Mol. Biol. Cell 2001, 12, 891–900.
- Soroosh, A.; Albeiroti, S.; West, G.A.; Willard, B.; Fiocchi, C.; de la Motte, C.A. Crohn’s Disease Fibroblasts Overproduce the Novel Protein KIAA1199 to Create Proinflammatory Hyaluronan Fragments. Cell. Mol. Gastroenterol. Hepatol. 2016, 2, 358–368.e4.
- Yoshida, H.; Nagaoka, A.; Kusaka–Kikushima, A.; Tobiishi, M.; Kawabata, K.; Sayo, T.; Sakai, S.; Sugiyama, Y.; Enomoto, H.; Okada, Y.; et al. KIAA1199, a deafness gene of unknown function, is a new hyaluronan binding protein involved in hyaluronan depolymerization. Proc. Natl. Acad. Sci. USA 2013, 110, 5612–5617.
- Yoshida, H.; Okada, Y. Role of HYBID (Hyaluronan Binding Protein Involved in Hyaluronan Depolymerization), Alias KIAA1199/CEMIP, in Hyaluronan Degradation in Normal and Photoaged Skin. Int. J. Mol. Sci. 2019, 20, 5804.
- Johnson, P.; Arif, A.A.; Lee–Sayer, S.S.M.; Dong, Y. Hyaluronan and Its Interactions with Immune Cells in the Healthy and Inflamed Lung. Front. Immunol. 2018, 9, 2787.
- Toole, B.P. The CD147–HYALURONAN Axis in Cancer. Anat. Rec. 2020, 303, 1573–1583.
- Jackson, D.G. Hyaluronan in the lymphatics: The key role of the hyaluronan receptor LYVE–1 in leucocyte trafficking. Matrix Biol. 2019, 78, 219–235.
- Garantziotis, S.; Matalon, S. Sugarcoating Lung Injury: A Novel Role for High–Molecular–Weight Hyaluronan in Pneumonia. Am. J. Respir. Crit. Care Med. 2019, 200, 1197–1198.
- Weigel, P.H. Planning, evaluating and vetting receptor signaling studies to assess hyaluronan size–dependence and specificity. Glycobiology 2017, 27, 796–799.
- Weigel, P.H.; Baggenstoss, B.A. What is special about 200 kDa hyaluronan that activates hyaluronan receptor signaling? Glycobiology 2017, 27, 868–877.
- Ziebell, M.R.; Prestwich, G.D. Interactions of peptide mimics of hyaluronic acid with the receptor for hyaluronan mediated motility (RHAMM). J. Comput. Aided. Mol. Des. 2004, 18, 597–614.
- Yang, B.; Yang, B.L.; Savani, R.C.; Turley, E.A. Identification of a common hyaluronan binding motif in the hyaluronan binding proteins RHAMM, CD44 and link protein. EMBO J. 1994, 13, 286–296.
- Anderegg, U.; Simon, J.C.; Averbeck, M. More than just a filler—The role of hyaluronan for skin homeostasis. Exp. Dermatol. 2014, 23, 295–303.
- Kleiser, S.; Nystrom, A. Interplay between Cell–Surface Receptors and Extracellular Matrix in Skin. Biomolecules 2020, 10, 1170.
- Banerji, S.; Wright, A.J.; Noble, M.; Mahoney, D.J.; Campbell, I.D.; Day, A.J.; Jackson, D.G. Structures of the CD44–hyaluronan complex provide insight into a fundamental carbohydrate–protein interaction. Nat. Struct. Mol. Biol. 2007, 14, 234–239.
- Senbanjo, L.T.; AlJohani, H.; AlQranei, M.; Majumdar, S.; Ma, T.; Chellaiah, M.A. Identification of sequence–specific interactions of the CD44–intracellular domain with RUNX2 in the transcription of matrix metalloprotease–9 in human prostate cancer cells. Cancer Drug. Resist. 2020, 3, 586–602.
- Al–Othman, N.; Alhendi, A.; Ihbaisha, M.; Barahmeh, M.; Alqaraleh, M.; Al–Momany, B.Z. Role of CD44 in breast cancer. Breast Dis. 2020, 39, 1–13.
- Ooki, T.; Murata–Kamiya, N.; Takahashi–Kanemitsu, A.; Wu, W.; Hatakeyama, M. High–Molecular–Weight Hyaluronan Is a Hippo Pathway Ligand Directing Cell Density–Dependent Growth Inhibition via PAR1b. Dev. Cell 2019, 49, 590–604.e9.
- Hauser–Kawaguchi, A.; Luyt, L.G.; Turley, E. Design of peptide mimetics to block pro–inflammatory functions of HA fragments. Matrix Biol. 2019, 78, 346–356.
- Gulati, K.; Jamsandekar, M.; Poluri, K.M. Mechanistic insights into molecular evolution of species–specific differential glycosaminoglycan binding surfaces in growth–related oncogene chemokines. R. Soc. Open Sci. 2017, 4, 171059.
- Boittier, E.D.; Gandhi, N.S.; Ferro, V.; Coombe, D.R. Cross–Species Analysis of Glycosaminoglycan Binding Proteins Reveals Some Animal Models Are ”More Equal“ than Others. Molecules 2019, 24, 924.
- Carvalho, A.M.; Soares da Costa, D.; Paulo, P.M.R.; Reis, R.L.; Pashkuleva, I. Co–localization and crosstalk between CD44 and RHAMM depend on hyaluronan presentation. Acta Biomater. 2021, 119, 114–124.




