Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Dongfang Wang | + 1432 word(s) | 1432 | 2021-11-12 10:33:21 | | | |
| 2 | Vivi Li | Meta information modification | 1432 | 2021-11-24 02:24:45 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Wang, D. CNPase. Encyclopedia. Available online: https://encyclopedia.pub/entry/16309 (accessed on 07 February 2026).
Wang D. CNPase. Encyclopedia. Available at: https://encyclopedia.pub/entry/16309. Accessed February 07, 2026.
Wang, Dongfang. "CNPase" Encyclopedia, https://encyclopedia.pub/entry/16309 (accessed February 07, 2026).
Wang, D. (2021, November 23). CNPase. In Encyclopedia. https://encyclopedia.pub/entry/16309
Wang, Dongfang. "CNPase." Encyclopedia. Web. 23 November, 2021.
Copy Citation
As an unconventional PDE, CNPase catalyzes 2′,3′-cyclic nucleotides as substrates [9], which are reported to impair mitochondrial integrity and accelerate the opening of the mitochondrial permeability transition pore (mPTP). As a mitochondrial protein, protein–protein interactions between the functional complexes I–V of the mitochondrial inner membrane and CNPase, leads to the assumption that CNPase might be involved in mitochondrial respiration and energy production.
CNPase
heart failure animal model
1. Introduction
Heart failure is the end-stage of cardiovascular disease and represents the top threat to public health worldwide. Despite accomplishments in therapeutics for heart failure, with a 5-year mortality rate of 75%, the prognostic results are far from satisfactory [1]. Cardiac pathological hypertrophy usually precedes the onset of heart failure and is characterized by enlarged cardiomyocytes, thickening, and stiff ventricular walls due to pressure overload, mutations of sarcomeric proteins, or infarction-induced cardiomyocyte loss [2]. The deficiency of ATP production plays a central role in the pathogenesis and progression of heart failure phenotypes [3]. At present, improving myocardial energy homeostasis with neuroendocrine systems and heart rate management has emerged as a core strategy for treating heart failure [4].
Genetic modification, pharmacological and surgical methods have been used in small animals to decipher the pathogenesis of heart failure and develop new therapies. Zebrafish are well characterized and widely used cardiovascular disease models based on genotypic and genotyping similarities. They are suitable for high-throughput chemical screening and gene manipulation due to their tiny size, relatively low maintenance cost, rapid development, accessibility for in vivo imaging, and the optical clarity of embryos [5]. Additionally, zebrafish embryos can survive without blood flow by oxygen diffusion, which makes them excellent models of heart failure [6]. Approximately 25% of mouse genes are embryonically lethal when knocked out [7], making it difficult to produce viable murine models for the study of heart failure.
Recently, the roles of 2′,3′-cyclic nucleotide-3′-phosphodiesterase (CNPase) in the pathophysiological processes of heart diseases have emerged, as indicated by researchers targeting CNPase enzymatic activity, which might represent a novel therapeutic strategy for heart failure [8]. As an unconventional PDE, CNPase catalyzes 2′,3′-cyclic nucleotides as substrates [9], which are reported to impair mitochondrial integrity and accelerate the opening of the mitochondrial permeability transition pore (mPTP) [10]. As a mitochondrial protein, protein–protein interactions between the functional complexes I–V of the mitochondrial inner membrane and CNPase, leads to the assumption that CNPase might be involved in mitochondrial respiration and energy production [11]. Increasing shreds of evidence suggest that CNPase plays a role during ageing through substrate hydrolysis activity and protein–protein interactions to protect mitochondria from mPTP, and the antiaging activity of CNPase in heart failure and brain cognition was suggested by two reports [12][13]. However, CNPase deficiency is potentially embryo lethal, and premature deaths are quite common among CNPase −/− litters [14]. Thus, suitable genetically modified animal models, which could be developed as powerful tools to provide direct evidence of CNPase function in heart failure, are lacking for studying CNPase in cardiovascular diseases.
2. A Zebrafish Larvae Model of Spontaneous Dilated Cardiomyopathy Induced by Morpholino-Mediated Knockdown of CNPase
To explore the role of CNPase in cardiomyopathy development, a translation-blocking cnpase morpholino (MO) was designed (Figure 1A) and injected onto one-cell stage embryos. Kaplan–Meier analysis of the survival rate showed that roughly >85% of the cnpase-blocked larvae were deceased within three days post-fertilization, suggesting that the cnpase loss-of-function might lead to embryonic or neonatal death (Figure 1B). Notably, the cnpase-blocked larval zebrafish developed pericardial edema, atrium and ventricle hypertrophy, bradycardia with a reduced ejection fraction, fractional shortening, and cardiac output, which was highly similar to the pathophysiological state of human heart failure (Figure 1C–H).
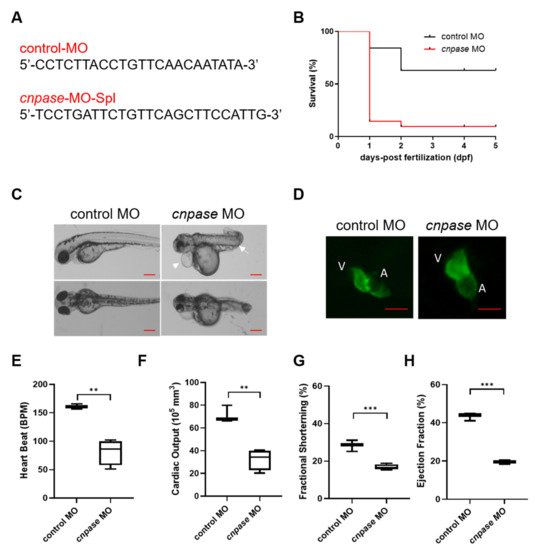
Figure 1. Morpholino-mediated cnpase knockdown results in a heart failure phenotype in zebrafish larvae. (A) The target sequences of control morpholino and morpholino targeting cnpase. (B) Effects of morpholino injection on the survival rate of zebrafish embryos over 120 hpf, n = 25~50. hpf, hours post-fertilization. (C) Phenotypes of the tg(myl7:gfp) zebrafish after cnpase-MO administration at 4 dpf. Arrows indicate the pericardial edema and malformation of body shape. dpf, days post-fertilization. Scale bar, 1000 µm. (D) cnpase-morpholino administration results in enlarged ventricular and atrium size. The images were captured by upright fluorescence microscope (Axio Imager A2, Zeiss). Scale bar, 1000 µm. (E–H) Cardiac function evaluation of zebrafish embryo after MO injection at 4 dpf, n = 25~50. Values are expressed as mean ± SD. Unpaired t-test, **, p < 0.005; ***, p < 0.001.
3. The Generation of a Zebrafish Cardiomyopathy Model by CRISPR-Cas9-Mediated CNPase Knockout
To establish a stable cardiomyopathy animal model, researchers herein generated a cnpase knockout zebrafish strain on a pure AB Tg(flk: egfp) background using the CRISPR-Cas9 gene-editing system. CRISPR-Cas9 founders (F0) were generated in which the first exon of the zebrafish CNPase ortholog was targeted simultaneously by a pair of sgRNAs (Figure 2A). Adult founders were inbred to give birth to the offspring, in which indels were detected in the PCR amplicons of the region around the target site (Figure 2B,C). Researchers found that frame-shift indels in the cnpase coding region could create mutant alleles in zebrafish in which it recapitulates MO phenotypes, excluding the possibility of MO-induced toxic and off-target effects (Figure 2C,D). Considering that homozygous mutations of CNPase cause embryo or neonatal lethality, the heterozygous cnpase+/− zebrafish strain was maintained for future study.
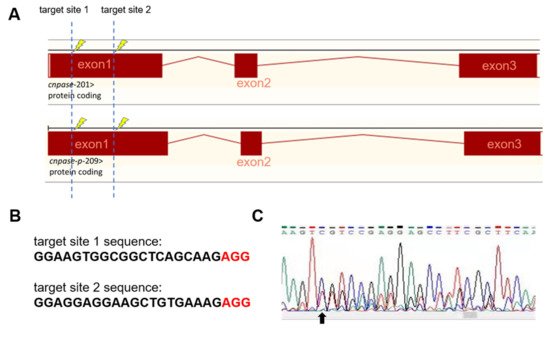
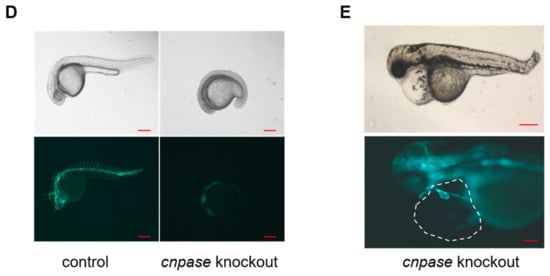
Figure 2. Generation of CNPase knockout zebrafish strain using the CRISPR-Cas9 editing system. (A) Schematic representation of the knockout strategy using CRISPR-Cas9 by targeting the first exon of cnpase. The cnpase-201, cnpase-209 encodes CNPase I (45 kDa) and CNPase II (47 kDa), respectively. (B) Sequences of the target site 1 and 2. “AGG” is the protospacer adjacent motif. (C) The genotyping of cnpase+/− zebrafish confirmed by tail-DNA PCR amplicons. The arrow indicates where the frame shift begins. (D,E) The cnpase deficient zebrafish exhibits developmental retardation, and cardiac failure phenotypes. Scale bar, 1000 µm.
4. AAV9-Mediated CNPase Myocardial Delivery Ameliorates Heart Dysfunction
Researchers reported that targeting CNPase enzymatic activity might give rise to novel therapies for heart failure [8]. Thus, the therapeutic potential of CNPase as a new target in a translational cardiac hypertrophy animal model was explored. Heart hypertrophy was surgically induced by abdominal aorta constriction in Sprague-Dawley rats. Two weeks later, CNPase was introduced into the left ventricular myocardium through ultrasound-guided AAV9 delivery. Four weeks after the AAV9 delivery, the echocardiographic analysis revealed that the hypertrophic response was less pronounced in the AAV9-CNPase delivered group, supported by the decrease of left ventricular internal diameter in the vesicle-delivered group (Figure 3A). Further, the heart dysfunction had recovered upon the AAV9-mediated gene therapy, wherein the cardiac output (CO), left ventricular ejection fraction (EF), as well as fraction shortening (FS) were enhanced (Figure 3C–E). No effects of CNPase expression on heart rate were observed (Figure 3B).
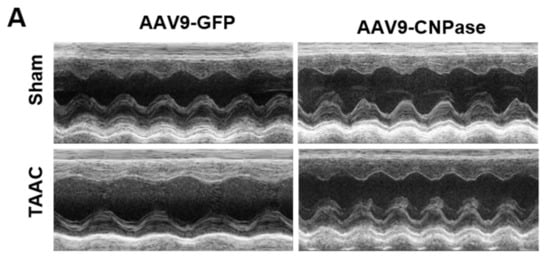
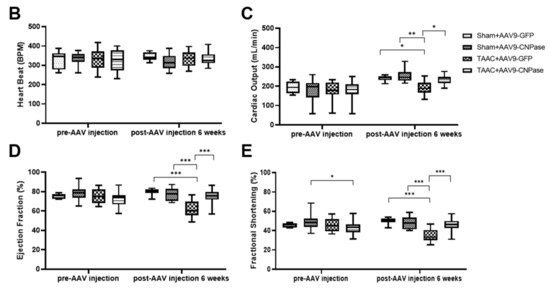
Figure 3. Myocardial CNPase delivery protects the heart from cardiac hypertrophy and dysfunction. (A) Representative echocardiography images of left ventricles. (B–E) The therapeutic effects of CNPase on heart rate, left ventricular ejection fraction, fraction shortening, cardiac output. The pressure overload-induced heart hypertrophy was carried out by transverse abdominal aorta constriction (TAAC) surgery. The AAV9-CNPase vectors (1 × 1012 genome copies) were delivered into the left ventricular myocardium guided by echocardiography. n = 8 in the sham group, n = 14–16 in the TAAC group. One-way ANOVA, Tukey’s multiple comparisons test, *, p < 0.05; **, p < 0.005; ***, p < 0.001.
5. CNPase Overexpression Improves Mitochondrial Respiration Function in Cardiac Hypertrophy
As the most energy-demanding organ, the heart relies on mitochondria as the powerhouse to produce ATP. The mitochondrial bioenergetics were profiled by the mitoStress assay using a Seahorse XFe96 cellular energy analyzer (Figure 4A). Current data demonstrated that the respiratory capacity was decreased upon AngII-induced hypertrophic stress. In contrast, the mitochondrial bioenergetics were well preserved in CNPase-overexpressed cells, as suggested by the basal, maximal, and reserved mitochondrial oxygen consumption rate (OCR) (Figure 4B–E). Notably, there was an increase of ATP-linked OCR in CNPase-overexpressed cells compared to the empty vector-transfected cells (Figure 4D).
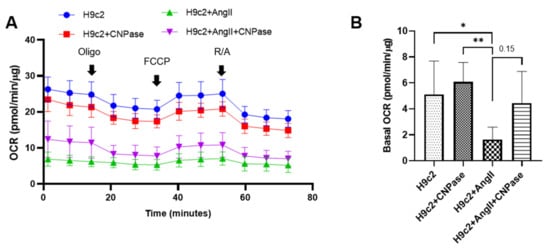
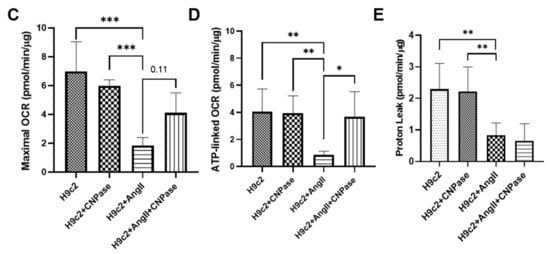
Figure 4. CNPase expression is essential for mitochondrial integrity, respiration capacity, and ATP production. (A) The general scheme of the mitochondrial stress test. The mitoStress assay was carried out on a Seahorse XFp extracellular flux analyzer (Agilent, Santa Clara, CA, USA) to directly measure the oxygen consumption rate (OCR) of cells. H9C2 cells were plated at a density of 2000 cells/well onto a 96-well seahorse microplate and transfected with pcDNA3.1-HA or pcDNA3.1-HA-CNPase vectors. For the mitoStress assay, programmable injection of selective inhibitors, oligomycin (2.5 µM), FCCP (1 µM), and rotenone/antimycin (2.5 µM) was performed. The Seahorse data shown are acquired from 8–12 independent replicates per experiment. Oligo, Oligomycin; FCCP, Potent uncoupler of mitochondrial oxidative phosphorylation; R/A, Rotenone/Antimycin. (B–E) Effects of CNPase overexpression on the basal, maximal and spare mitochondrial respiration, ATP production, and proton leak. One-way ANOVA, Tukey’s multiple comparisons test, *, p < 0.05; **, p < 0.005; ***, p < 0.001.
References
- Shah, K.S.; Xu, H.; Matsouaka, R.A.; Bhatt, D.L.; Heidenreich, P.A.; Hernandez, A.F.; Devore, A.D.; Yancy, C.W.; Fonarow, G.C. Heart Failure with Preserved, Borderline, and Reduced Ejection Fraction: 5-Year Outcomes. J. Am. Coll. Cardiol. 2017, 70, 2476–2486.
- Frey, N.; Katus, H.A.; Olson, E.N.; Hill, J.A. Hypertrophy of the Heart. Circulation 2004, 109, 1580–1589.
- Ingwall, J.S.; Weiss, R.G. Is the Failing Heart Energy Starved? Circ. Res. 2004, 95, 135–145.
- He, Y.; Huang, W.; Zhang, C.; Chen, L.; Xu, R.; Li, N.; Wang, F.; Han, L.; Yang, M.; Zhang, D. Energy metabolism disorders and potential therapeutic drugs in heart failure. Acta Pharm. Sin. B 2021, 11, 1098–1116.
- Narumanchi, S.; Wang, H.; Perttunen, S.; Tikkanen, I.; Lakkisto, P.; Paavola, J. Zebrafish Heart Failure Models. Front. Cell Dev. Biol. 2021, 9, 662583.
- Shi, X.; Chen, R.; Zhang, Y.; Yun, J.; Brand-Arzamendi, K.; Liu, X.; Wen, X.-Y. Zebrafish heart failure models: Opportunities and challenges. Amino Acids 2018, 50, 787–798.
- Wu, Y.; Zhang, J.; Peng, B.; Tian, D.; Zhang, D.; Li, Y.; Feng, X.; Liu, J.; Li, J.; Zhang, T.; et al. Generating viable mice with heritable embryonically lethal mutations using the CRISPR-Cas9 system in two-cell embryos. Nat. Commun. 2019, 10, 2883.
- Wang, D.; Tan, K.S.; Arias-Moreno, X.; Tan, W.; Cheng, G. Sirt3 increases CNPase enzymatic activity through deacetylation and facilitating substrate accessibility. Biochem. Biophys. Res. Commun. 2021, 571, 181–187.
- Heaton, P.A.; Eckstein, F. Diastereomeric specificity of 2′,3′-cyclic nucleotide 3′-phosphodiesterase. Nucleic Acids Res. 1996, 24, 850–853.
- Azarashvili, T.; Krestinina, O.; Galvita, A.; Grachev, D.; Baburina, Y.; Stricker, R.; Evtodienko, Y.; Reiser, G. Ca2+-dependent permeability transition regulation in rat brain mitochondria by 2′,3′-cyclic nucleotides and 2′,3′-cyclic nucleotide 3′-phosphodiesterase. Am. J. Physiol. Cell Physiol. 2009, 296, C1428–C1439.
- Baburina, Y.; Azarashvili, T.; Grachev, D.; Krestinina, O.; Galvita, A.; Stricker, R.; Reiser, G. Mitochondrial 2′, 3′-cyclic nucleotide 3′-phosphodiesterase (CNP) interacts with mPTP modulators and functional complexes (I–V) coupled with release of apoptotic factors. Neurochem. Int. 2015, 90, 46–55.
- Baburina, Y.; Odinokova, I.; Azarashvili, T.; Akatov, V.; Lemasters, J.J.; Krestinina, O. 2′,3′-Cyclic nucleotide 3′-phosphodiesterase as a messenger of protection of the mitochondrial function during melatonin treatment in aging. Biochim. et Biophys. Acta (BBA)—Biomembr. 2017, 1859, 94–103.
- Zhao, Y.Y.; Shi, X.Y.; Zhang, L.; Wu, H.; Chao, F.L.; Huang, C.X.; Gao, Y.; Qiu, X.; Chen, L.; Lu, W.; et al. Enriched environment induces higher CNPase positive cells in aged rat hippocampus. Neurosci. Lett. 2013, 555, 177–181.
- Lappe-Siefke, C.; Goebbels, S.; Gravel, M.; Nicksch, E.; Lee, J.; Braun, P.E.; Griffiths, I.R.; Nave, K.-A. Disruption of Cnp1 uncouples oligodendroglial functions in axonal support and myelination. Nature Genet. 2003, 33, 366–374.
More
Information
Subjects:
Physiology; Allergy
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.0K
Entry Collection:
Hypertension and Cardiovascular Diseases
Revisions:
2 times
(View History)
Update Date:
24 Nov 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




