Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Satoshi Nagayama | + 2678 word(s) | 2678 | 2021-09-27 11:51:55 | | | |
| 2 | Rita Xu | Meta information modification | 2678 | 2021-09-28 08:05:01 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Nagayama, S. Precision Medicine for Colorectal Cancer. Encyclopedia. Available online: https://encyclopedia.pub/entry/14655 (accessed on 07 February 2026).
Nagayama S. Precision Medicine for Colorectal Cancer. Encyclopedia. Available at: https://encyclopedia.pub/entry/14655. Accessed February 07, 2026.
Nagayama, Satoshi. "Precision Medicine for Colorectal Cancer" Encyclopedia, https://encyclopedia.pub/entry/14655 (accessed February 07, 2026).
Nagayama, S. (2021, September 28). Precision Medicine for Colorectal Cancer. In Encyclopedia. https://encyclopedia.pub/entry/14655
Nagayama, Satoshi. "Precision Medicine for Colorectal Cancer." Encyclopedia. Web. 28 September, 2021.
Copy Citation
In the field of colorectal cancer (CRC) treatment, diagnostic modalities and chemotherapy regimens have progressed remarkably in the last two decades. However, it is still difficult to identify minimal residual disease (MRD) necessary for early detection of recurrence/relapse of tumors and to select and provide appropriate drugs timely before a tumor becomes multi-drug-resistant and more aggressive.
liquid biopsy
minimal residual disease
cancer precision medicine
immune checkpoint inhibitor
neoantigen
1. Introduction
In spite of remarkable progress in diagnosis and treatment of colorectal cancer (CRC), there are still big challenges to improve the overall prognosis. Current diagnosis of recurrence/relapse is based on tumor biomarkers or imaging modalities including CT, MR, and PET examinations, which fail to detect minute lesions (micrometastases). It should be favorable to know the precise condition of the disease earlier than imaging diagnosis and initiate a proper treatment before clinically overt recurrences are identified. In addition, most patients who receive systemic chemotherapy become resistant during the treatment course and end up in the termination of the treatment with the standardized guidelines. A suitable drug should be selected and provided based on the molecular biological profiles of the tumors before the disease culminates in a far-advanced stage.
With the advance of genomic sequencing technologies typified by next-generation sequencing (NGS), it becomes much easier, more rapid, and less expensive to access comprehensive genomic information of the tumors. There is a high possibility that usage of the in-depth genomic profiles contributes to detection of recurrent/relapsed tumors and to proper choice of beneficial treatment options, including molecularly targeted therapies. In order to obtain the genomic profiles, blood-based tests, so called ‘liquid biopsies’, are considered to be a more useful method as compared to tumor biopsies, since liquid biopsies can be performed repeatedly and less invasively during a monitoring period and can provide the genomic information that is highly concordant with that of tumor biopsies. Hence, by leveraging the genomic profiles obtained by liquid biopsies, recurrence/relapse can be detected earlier than the current imaging diagnosis, and more suitable therapies can be provided without the delay in the time course of the treatment. Furthermore, analyzing the comprehensive genomic information, which also can be supplemented by liquid biopsies, will lead to a new development of effective immunotherapy related to mutations and/or neoantigens. In this review, we focus on potential roles of liquid biopsy in terms of clinical management of CRCs ranging from early detection of recurrence/relapse to acquisition of definitive clues leading to a promising and beneficial treatment, and touch on the potential of a promising immunotherapy in the treatment of CRCs (Figure 1).
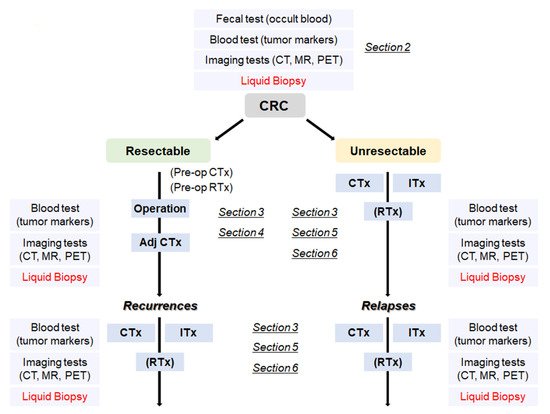
Figure 1. A flowchart from the diagnosis of CRC to treatment according to the disease progression. CRC: colorectal cancer, Adj CTx: adjuvant chemotherapy, CTx: chemotherapy, RTx: radiotherapy, Itx: immunotherapy.
2. Cancer Screening Using Liquid Biopsy
CRC is often diagnosed at a late stage owing to the lack of specific symptoms in early stages [1]. It is clinically important to develop an easy, cheap, and sensitive cancer screening method that detects cancer or precancerous lesions before clinical symptoms arise. If we can, we are able to begin treatment at earlier stages and increase probability of cure of the disease.
Colonoscopy screening remains the gold standard for early-stage diagnosis of CRC and has led to the reduction of CRC-related mortality [2][3]. Although colonoscopy is certainly effective, it is an invasive and relatively expensive procedure to screen CRC. Recent advancement of noninvasive screening approaches that were approved by the US Food and Drug Administration (FDA) includes stool-based tests and multitarget stool DNA tests (e.g., Cologuard) summarized in Table 1.
Table 1. Up-to-date stool- and blood-based screening method.
| Screening Method | Type of Sample | Description | Overall Performance |
|---|---|---|---|
| High sensitivity Guaiac-based fecal occult blood test (gFOBT) -FDA approved |
Stool |
|
In randomized controlled trials, screening with FOBT reduced CRC mortality rates by 15% to 33% [4][5][6] Specificity: 86.7% to 97.7%; Sensitivity: 33.3% to 79.4% [7][8] |
| Fecal Immunochemical test (FIT) -FDA approved |
Stool |
|
Randomized control: On-going Specificity: 81% to 96%; Sensitivity: 65% to 95% [9][10] |
| Multitarget stool DNA test (Cologuard) -FDA approved |
Stool |
|
Specificity: 86.6%; Sensitivity: 92.3% [11] |
| Epi proColon -FDA approved |
Blood |
|
Specificity: 80 to 82%; Sensitivity: 68 to 72% [12] |
| CancerSEEK | Blood |
|
All 8 cancer types including CRC Specificity > 99% Sensitivity: 33% to 70% [13] |
| GRAIL (CCGA and STRIVE) | Blood |
|
All 12 cancer types including CRC Specificity: 98.3% to 99.8% Stage I–III sensitivity: 60.7% to 73.3% [14] |
| DELFI | Blood |
|
Specificity: 95%; Sensitivity: 81% [15] |
A follow-up diagnostic colonoscopy is performed if abnormalities are indicated by these noninvasive tests. Even though the existing noninvasive stool-based tests for colon cancer have shown high sensitivity and specificity, adherence remains low. An observational study from Germany indicated the improved compliance to CRC screening using these tests. In this study, 97% of the subjects who had refused colonoscopy accepted an alternative noninvasive method; 83% of them chose the blood test, only 15% chose the stool test, and the remaining people refused to receive any screenings [16].
To improve the low adherence of colonoscopy and stool-based tests, liquid biopsy approaches, which have progressed substantially, are likely to be suitable to apply for the screening of CRC using blood samples. In 2016, the FDA approved the first blood-based screening test, Epi proColon, that possibly detects the promoter methylation status of the septin 9 (SEPT9) gene in cell-free DNAs (cfDNAs) for colon cancer using qPCR. Methylated SEPT9 level is increased in CRC and thus serves as a differential biomarker for early detection of CRC. The Epi proColon assay showed sensitivity and specificity values ranging from 68 to 72% and 80 to 82%, respectively (Table 1) [17][18]. Importantly, results from the multicenter randomized ADMIT trial indicated that adherence of Epi proColon blood-based screening was 99.5% compared with 88.1% for the FIT stool-based test, demonstrating a preferential acceptance of the blood test [12].
In addition to methylation signatures, Cohen and colleagues reported a multianalyte detection systems (CancerSEEK) by combining the detection of specific mutations of circulating tumor DNA (ctDNA) with conventional biomarkers for the detection of eight common surgically resectable cancer types. The CancerSEEK approach detected cancer with a sensitivity range from 69% to 98% and a specificity of 99% [13]. Specifically, 65% (252/388) of stage I–III resectable CRC were positive with CancerSEEK [13]. A subsequent prospective interventional study, DETECT-A (Detecting cancers Earlier Through Elective mutation-based blood Collection and Testing) evaluated 10,006 women with no prior history of cancer and followed-up for 12 months with the combination of the CancerSEEK study and imaging. Among those participants, 127 of 134 had a positive blood test underwent PET–CT imaging examination to evaluate the presence or absence of as well as the location of cancer. A total of 26 women were diagnosed to have a cancer, and 65% of them were found to be at a localized stage, potentially amenable to surgical resection [19].
On the other hand, Guardant Health has initiated the ECLIPSE study (https://clinicaltrials.gov/ct2/show/NCT04136002, accessed on 26 April 2021) for early detection of CRC with the LUNAR-2 blood test. The LUNAR-2 assay could detect somatic variants, methylation alterations, and other epigenomic changes and reported a high sensitivity in detecting CRC. This assay will be further tested on approximately 10,000 individuals aged 45–84 who are at average risk for CRC.
GRAIL started the Circulating Cell-free Genome Atlas Study (CCGA) as a discovery study and found that whole-genome bisulfite sequencing (WGBS) interrogating genome-wide methylation patterns outperformed whole-genome sequencing (WGS) and targeted sequencing approaches interrogating copy-number variants (CNVs) and single-nucleotide variants (SNVs)/small insertions and deletions, respectively [20][21]. GRAIL established a high-specificity (low false positive rate) targeted bisulfite sequencing, which focused on more than 100,000 methylation sites in our genome and assessed methylation patterns to evaluate the presence or absence of cancer with machine learning. The results of CCGA and the STRIVE study reported a sensitivity of 67.3% for 12 cancer types at stages I to III with an accuracy of 93% to predict tissue of origin [14].
Cristiano et al. [15] developed a method called DELFI (DNA evaluation of fragments for early interception) for early cancer detection. This method utilized the differences of genome-wide cfDNA fragmentation profiles as well as machine learning to distinguish cancer patients from healthy individuals. DELFI detected 152 of 208 patients with eight cancer types including breast, CRC, lung cancer, ovarian cancer, pancreatic cancer, gastric cancer, and cholangiocarcinoma. The overall sensitivity and specificity were 73% and 98%, respectively and 81% and 95% for CRC, respectively. Furthermore, among the 126 patients who were evaluated by both targeted sequencing and DELFI, the sensitivity of DELFI alone was 66% (83 of the 126 patients), but when combining both tests, the sensitivity improved to 82% (103 of the 126 cases) [15].
3. Genomic Analysis for Selection of Molecular-Targeted Drugs
In cases of systemic/distant recurrences after the curative resection of primary CRCs or in those with surgically unresectable stage IV CRCs, intensive systemic chemotherapy is provided to halt the progression of the disease. However, it is difficult to completely eradicate cancer cells using the current regimens of systemic chemotherapy, so novel therapies based on the genomic profiles of the tumors of individual patients should be developed. Some of targeting genetic mutations include KRAS, BRAF, HER2, and microsatellite instability (MSI), which are leveraged in the current clinical setting [22] (Table 2).
Table 2. Genomic biomarkers in CRCs.
| Gene | Biomarkers | Frequencies (%) | Anticancer Agents |
|---|---|---|---|
| KRAS | Wild type | 60 [23] | Cetuximab Panitumumab |
| KRAS | G12C | 8 [23] | Sotorasib (AMG510) Adagrasib (MRTX849) |
| BRAF | V600E | 10 [24] | Vemurafenib Dabrafenib Ecorafenib |
| HER2 | Amplification | 2–3 [25][26] | Pertuzumab Trastuzumab Lapatinib |
| MLH1 MSH2 MSH6 PMS2 |
MSI-H | 10–15 [27][28] | Nivolumab Pembrolizumab Ipilimumab |
As for application of the KRAS mutation status to select chemotherapy regimens, phase III clinical trials such as CRYATAL, OPUS, CO.17, and FIRE-3 have shown that the benefit of adding cetuximab (anti-epidermal growth factor (EGFR) antibody) to FOLFOX or FOLFIRI was confined to patients with CRCs not having KRAS mutations [29][30][31][32][33][34][35]. With respect to KRAS-mutant tumors, the complexity of the signaling network of the KRAS-mutant alleles has made it difficult to develop molecularly-targeted therapies against KRAS mutations. Mutant KRAS protein has thus been regarded as an undruggable target, so most therapeutic strategies have been designed to inhibit downstream effector pathways such as the ERK/MAPK cascade. However, the clinical efficacy of targeting downstream effectors has been marginal [36]. Two covalently-binding inhibitors, AMG510 (Sotorasib) and MRTX849 (Adagrasib), which specifically target the KRAS G12C mutation, have recently been developed [37][38][39][40][41][42], and their encouraging efficacy in solid tumors harboring the KRAS G12C mutation including non-small cell lung cancers (NSCLCs) and CRCs has been demonstrated in several clinical trials [43][44][45][46][47]. However, most NSCLC patients with the KRAS G12C mutant showed a favorable response to selective KRAS G12C inhibition, while CRC patients harboring the same mutation rarely revealed clinical benefits. This drug resistance is speculated to result from a possible mechanism where a novel mutation can appear [48][49] and/or the feedback reactivation of the RAS pathway following KRAS G12C inhibition may occur. To overcome the acquired resistance by the adaptive RAS pathway feedback reactivation in CRCs, combinatorial targeting of EGFR and KRAS G12C or, theoretically, concomitant inhibition of SHP2 and KRAS G12C is expected as a promising treatment strategy, since SHP2 mediates signaling from multiple receptor tyrosine kinases to RAS, and its inhibition can more comprehensively hamper the feedback reactivation [50][51].
Regarding the treatment for CRCs with the BRAF V600E mutation, the administration of a BRAF inhibitor (vemurafenib) alone showed only limited clinical efficacy compared to the favorable responses observed in melanoma patients [52][53]. As in the case of KRAS G12C inhibition, adaptive feedback reactivation of the RAS-signaling pathway is considered to be a major mechanism of therapeutic resistance or poor response. Specifically, BRAF inhibition in cancers with the BRAF V600E mutation led to loss of negative feedback signals through the MAPK pathway in CRCs, resulting in receptor tyrosine kinase-mediated reactivation of MAPK signaling by wild-type RAS and RAF [54][55][56][57][58]. The concomitant administration of dabrafenib and trametinib therefore has a substantial impact on clinical efficacy in a subset of patients with BRAF-V600E CRCs [59]. Furthermore, combined BRAF + EGFR + MEK inhibitions are tolerable and result in favorable clinical responses and a significantly longer overall survival compared to standard therapy in patients with BRAF-V600E CRCs [60][61].
Regarding HER2-positive CRCs, HER2-inhibiting antibodies and small molecules can suppress the activity of HER2-amplification or mutations [62]. According to several clinical trials including HERACLES, MyPathway, and DESTINY-CRC01, HER2-targeted therapies including anti-HER2 antibody conjugated with or without cytotoxic drugs showed promising and long-lasting outcome in HER2-positive CRCs that had been refractory to standard treatment [63][64][65][66][67]. Although HER2 amplification is identified in only 2–3% of CRCs, these results could provide hope to a substantial number of CRC patients who have experienced progression of the disease with the standardized guidelines.
In MSI-high CRCs due to mismatch-repair deficiency, immune checkpoint inhibitors such as pembrolizumab and nivolumab have shown a significant clinical benefit in clinical trials including CheckMate-142 and KEYNOTE-164 [68][69][70][71][72][73]. Furthermore, pembrolizumab monotherapy has led to clinically meaningful improvements in health-related quality of life compared with chemotherapy in MSI-high CRC patients (KEYNOTE-177) [74][75]. The administration of immune checkpoint inhibitors is thus regarded as a first-line treatment option for this population [76]. However, acquired resistance to anti-PD-1 immunotherapy was reported in a subset of cases where the expression of MHC and/or B2M was reduced or either of these genes were lost in tumor cells, leading to impaired antigen presentation and resulting in immune evasion [77][78][79]. Hence, therapies targeting CTLA4 or PD-1 still have some limitations in treatment efficacy. It is also an intriguing approach to upregulate MHC-I expression to enhance sensitivity to immunotherapy [80].
NGS-based targeted-gene panel tests have recently been used in a clinical setting to identify patients with actionable genetic alterations for enrollment in genotype-matched clinical trials. According to the mutational landscape of metastatic cancers of more than 10,000 patients with clinical sequencing using a comprehensive assay MSK-IMPACT, one or more potentially actionable genetic alterations was detected in 36.7% of the patients, and 11% could be enrolled to genome-guided clinical trials [81]. In the literature covering genomic testing of advanced cancers, a small proportion of patients (4–31%) had a chance to receive genetic-alteration-matched therapy [82][83][84][85][86][87][88][89][90][91][92][93][94]. Furthermore, an observational study involving more than 1000 patients showed that overall response rates, time-to-treatment failure, and overall survival were higher with matched targeted therapy than those observed without matching, suggesting that identifying specific genetic alterations and choosing therapy based on these alterations are associated with a better prognosis than standard systemic therapy [83]. This finding is consistent with a recent meta-analysis of phase I trials that showed a higher overall response rate (30.6% vs. 4.9%, p < 0.001) and median progression-free survival (5.7 months vs. 2.95 months, p < 0.001) for genotype-matched trials, compared with non-selected therapies [95].
With the advent of WGS and whole-exome sequencing (WES), we can share more comprehensive information on the genomic alterations of individual tumors than with targeted gene-panel sequencing [22]. A recent WGS analysis comprising 2520 samples in 22 types of metastatic tumors showed that 62% of these tumors harbored at least one actionable mutation [96]. With the advances in sequencing technologies, more actionable biomarkers and/or oncogenic mutations have been detected in individual cancers. While it might suggest that a high proportion of actionable alterations are detectable in cancer patients by WGS and WES, the clinical benefits for cancer patients are still very limited. The limited contribution of gene panel tests is attributable to a variety of reasons ranging from patient-dependent factors such as health deterioration in those with an advanced cancer and patient preferences to physician-dependent factors including strict inclusion criteria of clinical trials, the cost of off-label use of drugs, and limited supply of molecular-targeted drugs [84][85][88]. Since genomic profiles can alter clinical management in diverse scenarios, the combination of comprehensive molecular testing and better access to genome-guided trials can improve rates of clinical trial enrollment, thereby enabling precision cancer medicine on a large scale. To expand opportunities for genome-matched therapies, further development of novel molecular-targeted drugs and other treatment options is urgently required.
References
- Vega, P.; Valentin, F.; Cubiella, J. Colorectal cancer diagnosis: Pitfalls and opportunities. World J. Gastrointest. Oncol. 2015, 7, 422–433.
- Brenner, H.; Stock, C.; Hoffmeister, M. Effect of screening sigmoidoscopy and screening colonoscopy on colorectal cancer incidence and mortality: Systematic review and meta-analysis of randomised controlled trials and observational studies. BMJ 2014, 348, g2467.
- Elmunzer, B.J.; Singal, A.G.; Sussman, J.B.; Deshpande, A.R.; Sussman, D.A.; Conte, M.L.; Dwamena, B.A.; Rogers, M.A.; Schoenfeld, P.S.; Inadomi, J.M.; et al. Comparing the effectiveness of competing tests for reducing colorectal cancer mortality: A network meta-analysis. Gastrointest. Endosc. 2015, 81, 700–709.e3.
- Mandel, J.S.; Bond, J.H.; Church, T.R.; Snover, D.C.; Bradley, G.M.; Schuman, L.M.; Ederer, F. Reducing mortality from colorectal cancer by screening for fecal occult blood. Minnesota Colon Cancer Control Study. N. Engl. J. Med. 1993, 328, 1365–1371.
- Hardcastle, J.D.; Chamberlain, J.O.; Robinson, M.H.; Moss, S.M.; Amar, S.S.; Balfour, T.W.; James, P.D.; Mangham, C.M. Randomised controlled trial of faecal-occult-blood screening for colorectal cancer. Lancet 1996, 348, 1472–1477.
- Kronborg, O.; Fenger, C.; Olsen, J.; Jorgensen, O.D.; Sondergaard, O. Randomised study of screening for colorectal cancer with faecal-occult-blood test. Lancet 1996, 348, 1467–1471.
- Allison, J.E.; Tekawa, I.S.; Ransom, L.J.; Adrain, A.L. A comparison of fecal occult-blood tests for colorectal-cancer screening. N. Engl. J. Med. 1996, 334, 155–159.
- Brenner, H.; Tao, S. Superior diagnostic performance of faecal immunochemical tests for haemoglobin in a head-to-head comparison with guaiac based faecal occult blood test among 2235 participants of screening colonoscopy. Eur. J. Cancer 2013, 49, 3049–3054.
- Imperiale, T.F.; Gruber, R.N.; Stump, T.E.; Emmett, T.W.; Monahan, P.O. Performance Characteristics of Fecal Immunochemical Tests for Colorectal Cancer and Advanced Adenomatous Polyps: A Systematic Review and Meta-analysis. Ann. Intern. Med. 2019, 170, 319–329.
- Niedermaier, T.; Balavarca, Y.; Brenner, H. Stage-Specific Sensitivity of Fecal Immunochemical Tests for Detecting Colorectal Cancer: Systematic Review and Meta-Analysis. Am. J. Gastroenterol. 2020, 115, 56–69.
- Imperiale, T.F.; Ransohoff, D.F.; Itzkowitz, S.H.; Levin, T.R.; Lavin, P.; Lidgard, G.P.; Ahlquist, D.A.; Berger, B.M. Multitarget stool DNA testing for colorectal-cancer screening. N. Engl. J. Med. 2014, 370, 1287–1297.
- Liles, E.G.; Coronado, G.D.; Perrin, N.; Harte, A.H.; Nungesser, R.; Quigley, N.; Potter, N.T.; Weiss, G.; Koenig, T.; deVos, T. Uptake of a colorectal cancer screening blood test is higher than of a fecal test offered in clinic: A randomized trial. Cancer Treat. Res. Commun. 2017, 10, 27–31.
- Cohen, J.D.; Li, L.; Wang, Y.; Thoburn, C.; Afsari, B.; Danilova, L.; Douville, C.; Javed, A.A.; Wong, F.; Mattox, A.; et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 2018, 359, 926–930.
- Liu, M.C.; Oxnard, G.R.; Klein, E.A.; Swanton, C.; Seiden, M.V.; Consortium, C. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann. Oncol. 2020, 31, 745–759.
- Cristiano, S.; Leal, A.; Phallen, J.; Fiksel, J.; Adleff, V.; Bruhm, D.C.; Jensen, S.O.; Medina, J.E.; Hruban, C.; White, J.R.; et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature 2019, 570, 385–389.
- Adler, A.; Geiger, S.; Keil, A.; Bias, H.; Schatz, P.; deVos, T.; Dhein, J.; Zimmermann, M.; Tauber, R.; Wiedenmann, B. Improving compliance to colorectal cancer screening using blood and stool based tests in patients refusing screening colonoscopy in Germany. BMC Gastroenterol. 2014, 14, 183.
- Potter, N.T.; Hurban, P.; White, M.N.; Whitlock, K.D.; Lofton-Day, C.E.; Tetzner, R.; Koenig, T.; Quigley, N.B.; Weiss, G. Validation of a real-time PCR-based qualitative assay for the detection of methylated SEPT9 DNA in human plasma. Clin. Chem. 2014, 60, 1183–1191.
- Johnson, D.A.; Barclay, R.L.; Mergener, K.; Weiss, G.; Konig, T.; Beck, J.; Potter, N.T. Plasma Septin9 versus fecal immunochemical testing for colorectal cancer screening: A prospective multicenter study. PLoS ONE 2014, 9, e98238.
- Lennon, A.M.; Buchanan, A.H.; Kinde, I.; Warren, A.; Honushefsky, A.; Cohain, A.T.; Ledbetter, D.H.; Sanfilippo, F.; Sheridan, K.; Rosica, D.; et al. Feasibility of blood testing combined with PET-CT to screen for cancer and guide intervention. Science 2020, 369, 6499.
- Oxnard, G.R.; Klein, E.A.; Seiden, M.; Hubbell, E.; Venn, O.; Jamshidi, A.; Zhang, N.; Beausang, J.F.; Gross, S.; Kurtzman, K.N.; et al. Simultaneous multi-cancer detection and tissue of origin (TOO) localization using targeted bisulfite sequencing of plasma cell-free DNA (cfDNA). J. Glob. Oncol. 2019, 5, 44.
- Liu, M.C.; Klein, E.; Hubbell, E.; Maddala, T.; Aravanis, A.M.; Beausang, J.F.; Filippova, D.; Gross, S.; Jamshidi, A.; Kurtzman, K.; et al. Plasma cell-free DNA (cfDNA) assays for early multi-cancer detection: The circulating cell-free genome atlas (CCGA) study. Ann. Oncol. 2018, 29, viii14.
- Lieu, C.H.; Corcoran, R.B.; Overman, M.J. Integrating Biomarkers and Targeted Therapy Into Colorectal Cancer Management. Am. Soc. Clin. Oncol. Educ. Book. 2019, 39, 207–215.
- Neumann, J.; Zeindl-Eberhart, E.; Kirchner, T.; Jung, A. Frequency and type of KRAS mutations in routine diagnostic analysis of metastatic colorectal cancer. Pathol. Res. Pract. 2009, 205, 858–862.
- Di Nicolantonio, F.; Martini, M.; Molinari, F.; Sartore-Bianchi, A.; Arena, S.; Saletti, P.; De Dosso, S.; Mazzucchelli, L.; Frattini, M.; Siena, S.; et al. Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J. Clin. Oncol. 2008, 26, 5705–5712.
- Valtorta, E.; Martino, C.; Sartore-Bianchi, A.; Penaullt-Llorca, F.; Viale, G.; Risio, M.; Rugge, M.; Grigioni, W.; Bencardino, K.; Lonardi, S.; et al. Assessment of a HER2 scoring system for colorectal cancer: Results from a validation study. Mod. Pathol. 2015, 28, 1481–1491.
- Siena, S.; Sartore-Bianchi, A.; Marsoni, S.; Hurwitz, H.I.; McCall, S.J.; Penault-Llorca, F.; Srock, S.; Bardelli, A.; Trusolino, L. Targeting the human epidermal growth factor receptor 2 (HER2) oncogene in colorectal cancer. Ann. Oncol. 2018, 29, 1108–1119.
- Goldstein, J.; Tran, B.; Ensor, J.; Gibbs, P.; Wong, H.L.; Wong, S.F.; Vilar, E.; Tie, J.; Broaddus, R.; Kopetz, S.; et al. Multicenter retrospective analysis of metastatic colorectal cancer (CRC) with high-level microsatellite instability (MSI-H). Ann. Oncol. 2014, 25, 1032–1038.
- Koopman, M.; Kortman, G.A.; Mekenkamp, L.; Ligtenberg, M.J.; Hoogerbrugge, N.; Antonini, N.F.; Punt, C.J.; van Krieken, J.H. Deficient mismatch repair system in patients with sporadic advanced colorectal cancer. Br. J. Cancer 2009, 100, 266–273.
- Van Cutsem, E.; Kohne, C.H.; Hitre, E.; Zaluski, J.; Chang Chien, C.R.; Makhson, A.; D’Haens, G.; Pinter, T.; Lim, R.; Bodoky, G.; et al. Cetuximab and chemotherapy as initial treatment for metastatic colorectal cancer. N. Engl. J. Med. 2009, 360, 1408–1417.
- Bokemeyer, C.; Bondarenko, I.; Makhson, A.; Hartmann, J.T.; Aparicio, J.; de Braud, F.; Donea, S.; Ludwig, H.; Schuch, G.; Stroh, C.; et al. Fluorouracil, leucovorin, and oxaliplatin with and without cetuximab in the first-line treatment of metastatic colorectal cancer. J. Clin. Oncol. 2009, 27, 663–671.
- Karapetis, C.S.; Khambata-Ford, S.; Jonker, D.J.; O’Callaghan, C.J.; Tu, D.; Tebbutt, N.C.; Simes, R.J.; Chalchal, H.; Shapiro, J.D.; Robitaille, S.; et al. K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N. Engl. J. Med. 2008, 359, 1757–1765.
- Heinemann, V.; von Weikersthal, L.F.; Decker, T.; Kiani, A.; Vehling-Kaiser, U.; Al-Batran, S.E.; Heintges, T.; Lerchenmuller, C.; Kahl, C.; Seipelt, G.; et al. FOLFIRI plus cetuximab versus FOLFIRI plus bevacizumab as first-line treatment for patients with metastatic colorectal cancer (FIRE-3): A randomised, open-label, phase 3 trial. Lancet Oncol. 2014, 15, 1065–1075.
- Van Cutsem, E.; Kohne, C.H.; Lang, I.; Folprecht, G.; Nowacki, M.P.; Cascinu, S.; Shchepotin, I.; Maurel, J.; Cunningham, D.; Tejpar, S.; et al. Cetuximab plus irinotecan, fluorouracil, and leucovorin as first-line treatment for metastatic colorectal cancer: Updated analysis of overall survival according to tumor KRAS and BRAF mutation status. J. Clin. Oncol. 2011, 29, 2011–2019.
- Van Cutsem, E.; Lenz, H.J.; Kohne, C.H.; Heinemann, V.; Tejpar, S.; Melezinek, I.; Beier, F.; Stroh, C.; Rougier, P.; van Krieken, J.H.; et al. Fluorouracil, leucovorin, and irinotecan plus cetuximab treatment and RAS mutations in colorectal cancer. J. Clin. Oncol. 2015, 33, 692–700.
- Bokemeyer, C.; Kohne, C.H.; Ciardiello, F.; Lenz, H.J.; Heinemann, V.; Klinkhardt, U.; Beier, F.; Duecker, K.; van Krieken, J.H.; Tejpar, S. FOLFOX4 plus cetuximab treatment and RAS mutations in colorectal cancer. Eur. J. Cancer 2015, 51, 1243–1252.
- Ryan, M.B.; Corcoran, R.B. Therapeutic strategies to target RAS-mutant cancers. Nat. Rev. Clin. Oncol. 2018, 15, 709–720.
- Patricelli, M.P.; Janes, M.R.; Li, L.S.; Hansen, R.; Peters, U.; Kessler, L.V.; Chen, Y.; Kucharski, J.M.; Feng, J.; Ely, T.; et al. Selective Inhibition of Oncogenic KRAS Output with Small Molecules Targeting the Inactive State. Cancer Discov. 2016, 6, 316–329.
- Janes, M.R.; Zhang, J.; Li, L.S.; Hansen, R.; Peters, U.; Guo, X.; Chen, Y.; Babbar, A.; Firdaus, S.J.; Darjania, L.; et al. Targeting KRAS Mutant Cancers with a Covalent G12C-Specific Inhibitor. Cell 2018, 172, 578–589.e17.
- Fell, J.B.; Fischer, J.P.; Baer, B.R.; Ballard, J.; Blake, J.F.; Bouhana, K.; Brandhuber, B.J.; Briere, D.M.; Burgess, L.E.; Burkard, M.R.; et al. Discovery of Tetrahydropyridopyrimidines as Irreversible Covalent Inhibitors of KRAS-G12C with In Vivo Activity. ACS Med. Chem. Lett. 2018, 9, 1230–1234.
- Ostrem, J.M.; Shokat, K.M. Direct small-molecule inhibitors of KRAS: From structural insights to mechanism-based design. Nat. Rev. Drug Discov. 2016, 15, 771–785.
- Lito, P.; Solomon, M.; Li, L.S.; Hansen, R.; Rosen, N. Allele-specific inhibitors inactivate mutant KRAS G12C by a trapping mechanism. Science 2016, 351, 604–608.
- Zuberi, M.; Khan, I.; O’Bryan, J.P. Inhibition of RAS: Proven and potential vulnerabilities. Biochem. Soc. Trans. 2020, 48, 1831–1841.
- Canon, J.; Rex, K.; Saiki, A.Y.; Mohr, C.; Cooke, K.; Bagal, D.; Gaida, K.; Holt, T.; Knutson, C.G.; Koppada, N.; et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 2019, 575, 217–223.
- Lanman, B.A.; Allen, J.R.; Allen, J.G.; Amegadzie, A.K.; Ashton, K.S.; Booker, S.K.; Chen, J.J.; Chen, N.; Frohn, M.J.; Goodman, G.; et al. Discovery of a Covalent Inhibitor of KRAS(G12C) (AMG 510) for the Treatment of Solid Tumors. J. Med. Chem. 2020, 63, 52–65.
- Hallin, J.; Engstrom, L.D.; Hargis, L.; Calinisan, A.; Aranda, R.; Briere, D.M.; Sudhakar, N.; Bowcut, V.; Baer, B.R.; Ballard, J.A.; et al. The KRAS(G12C) Inhibitor MRTX849 Provides Insight toward Therapeutic Susceptibility of KRAS-Mutant Cancers in Mouse Models and Patients. Cancer Discov. 2020, 10, 54–71.
- Fell, J.B.; Fischer, J.P.; Baer, B.R.; Blake, J.F.; Bouhana, K.; Briere, D.M.; Brown, K.D.; Burgess, L.E.; Burns, A.C.; Burkard, M.R.; et al. Identification of the Clinical Development Candidate MRTX849, a Covalent KRAS(G12C) Inhibitor for the Treatment of Cancer. J. Med. Chem. 2020, 63, 6679–6693.
- Hong, D.S.; Fakih, M.G.; Strickler, J.H.; Desai, J.; Durm, G.A.; Shapiro, G.I.; Falchook, G.S.; Price, T.J.; Sacher, A.; Denlinger, C.S.; et al. KRAS(G12C) Inhibition with Sotorasib in Advanced Solid Tumors. N. Engl. J. Med. 2020, 383, 1207–1217.
- Tanaka, N.; Lin, J.J.; Li, C.; Ryan, M.B.; Zhang, J.; Kiedrowski, L.A.; Michel, A.G.; Syed, M.U.; Fella, K.A.; Sakhi, M.; et al. Clinical acquired resistance to KRASG12C inhibition through a novel KRAS switch-II pocket mutation and polyclonal alterations converging on RAS-MAPK reactivation. Cancer Discov. 2021.
- Awad, M.M.; Liu, S.; Rybkin, I.I.; Arbour, K.C.; Dilly, J.; Zhu, V.W.; Johnson, M.L.; Heist, R.S.; Patil, T.; Riely, G.J.; et al. Acquired Resistance to KRASG12C Inhibition in Cancer. N. Engl. J. Med. 2021, 384, 2382–2393.
- Amodio, V.; Yaeger, R.; Arcella, P.; Cancelliere, C.; Lamba, S.; Lorenzato, A.; Arena, S.; Montone, M.; Mussolin, B.; Bian, Y.; et al. EGFR Blockade Reverts Resistance to KRAS(G12C) Inhibition in Colorectal Cancer. Cancer Discov. 2020, 10, 1129–1139.
- Ryan, M.B.; Fece de la Cruz, F.; Phat, S.; Myers, D.T.; Wong, E.; Shahzade, H.A.; Hong, C.B.; Corcoran, R.B. Vertical Pathway Inhibition Overcomes Adaptive Feedback Resistance to KRAS(G12C) Inhibition. Clin. Cancer Res. 2020, 26, 1633–1643.
- Hyman, D.M.; Puzanov, I.; Subbiah, V.; Faris, J.E.; Chau, I.; Blay, J.Y.; Wolf, J.; Raje, N.S.; Diamond, E.L.; Hollebecque, A.; et al. Vemurafenib in Multiple Nonmelanoma Cancers with BRAF V600 Mutations. N. Engl. J. Med. 2015, 373, 726–736.
- Kopetz, S.; Desai, J.; Chan, E.; Hecht, J.R.; O’Dwyer, P.J.; Maru, D.; Morris, V.; Janku, F.; Dasari, A.; Chung, W.; et al. Phase II Pilot Study of Vemurafenib in Patients With Metastatic BRAF-Mutated Colorectal Cancer. J. Clin. Oncol. 2015, 33, 4032–4038.
- Paraiso, K.H.; Fedorenko, I.V.; Cantini, L.P.; Munko, A.C.; Hall, M.; Sondak, V.K.; Messina, J.L.; Flaherty, K.T.; Smalley, K.S. Recovery of phospho-ERK activity allows melanoma cells to escape from BRAF inhibitor therapy. Br. J. Cancer 2010, 102, 1724–1730.
- Montero-Conde, C.; Ruiz-Llorente, S.; Dominguez, J.M.; Knauf, J.A.; Viale, A.; Sherman, E.J.; Ryder, M.; Ghossein, R.A.; Rosen, N.; Fagin, J.A. Relief of feedback inhibition of HER3 transcription by RAF and MEK inhibitors attenuates their antitumor effects in BRAF-mutant thyroid carcinomas. Cancer Discov. 2013, 3, 520–533.
- Corcoran, R.B.; Ebi, H.; Turke, A.B.; Coffee, E.M.; Nishino, M.; Cogdill, A.P.; Brown, R.D.; Della Pelle, P.; Dias-Santagata, D.; Hung, K.E.; et al. EGFR-mediated re-activation of MAPK signaling contributes to insensitivity of BRAF mutant colorectal cancers to RAF inhibition with vemurafenib. Cancer Discov. 2012, 2, 227–235.
- Prahallad, A.; Sun, C.; Huang, S.; Di Nicolantonio, F.; Salazar, R.; Zecchin, D.; Beijersbergen, R.L.; Bardelli, A.; Bernards, R. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature 2012, 483, 100–103.
- Ahronian, L.G.; Sennott, E.M.; Van Allen, E.M.; Wagle, N.; Kwak, E.L.; Faris, J.E.; Godfrey, J.T.; Nishimura, K.; Lynch, K.D.; Mermel, C.H.; et al. Clinical Acquired Resistance to RAF Inhibitor Combinations in BRAF-Mutant Colorectal Cancer through MAPK Pathway Alterations. Cancer Discov. 2015, 5, 358–367.
- Corcoran, R.B.; Atreya, C.E.; Falchook, G.S.; Kwak, E.L.; Ryan, D.P.; Bendell, J.C.; Hamid, O.; Messersmith, W.A.; Daud, A.; Kurzrock, R.; et al. Combined BRAF and MEK Inhibition With Dabrafenib and Trametinib in BRAF V600-Mutant Colorectal Cancer. J. Clin. Oncol. 2015, 33, 4023–4031.
- Corcoran, R.B.; Andre, T.; Atreya, C.E.; Schellens, J.H.M.; Yoshino, T.; Bendell, J.C.; Hollebecque, A.; McRee, A.J.; Siena, S.; Middleton, G.; et al. Combined BRAF, EGFR, and MEK Inhibition in Patients with BRAF(V600E)-Mutant Colorectal Cancer. Cancer Discov. 2018, 8, 428–443.
- Kopetz, S.; Grothey, A.; Yaeger, R.; Van Cutsem, E.; Desai, J.; Yoshino, T.; Wasan, H.; Ciardiello, F.; Loupakis, F.; Hong, Y.S.; et al. Encorafenib, Binimetinib, and Cetuximab in BRAF V600E-Mutated Colorectal Cancer. N. Engl. J. Med. 2019, 381, 1632–1643.
- Pahuja, K.B.; Nguyen, T.T.; Jaiswal, B.S.; Prabhash, K.; Thaker, T.M.; Senger, K.; Chaudhuri, S.; Kljavin, N.M.; Antony, A.; Phalke, S.; et al. Actionable Activating Oncogenic ERBB2/HER2 Transmembrane and Juxtamembrane Domain Mutations. Cancer Cell 2018, 34, 792–806.e5.
- Sartore-Bianchi, A.; Trusolino, L.; Martino, C.; Bencardino, K.; Lonardi, S.; Bergamo, F.; Zagonel, V.; Leone, F.; Depetris, I.; Martinelli, E.; et al. Dual-targeted therapy with trastuzumab and lapatinib in treatment-refractory, KRAS codon 12/13 wild-type, HER2-positive metastatic colorectal cancer (HERACLES): A proof-of-concept, multicentre, open-label, phase 2 trial. Lancet Oncol. 2016, 17, 738–746.
- Sartore-Bianchi, A.; Lonardi, S.; Martino, C.; Fenocchio, E.; Tosi, F.; Ghezzi, S.; Leone, F.; Bergamo, F.; Zagonel, V.; Ciardiello, F.; et al. Pertuzumab and trastuzumab emtansine in patients with HER2-amplified metastatic colorectal cancer: The phase II HERACLES-B trial. ESMO Open 2020, 5, e000911.
- Hainsworth, J.D.; Meric-Bernstam, F.; Swanton, C.; Hurwitz, H.; Spigel, D.R.; Sweeney, C.; Burris, H.; Bose, R.; Yoo, B.; Stein, A.; et al. Targeted Therapy for Advanced Solid Tumors on the Basis of Molecular Profiles: Results From MyPathway, an Open-Label, Phase IIa Multiple Basket Study. J. Clin. Oncol. 2018, 36, 536–542.
- Meric-Bernstam, F.; Hurwitz, H.; Raghav, K.P.S.; McWilliams, R.R.; Fakih, M.; VanderWalde, A.; Swanton, C.; Kurzrock, R.; Burris, H.; Sweeney, C.; et al. Pertuzumab plus trastuzumab for HER2-amplified metastatic colorectal cancer (MyPathway): An updated report from a multicentre, open-label, phase 2a, multiple basket study. Lancet Oncol. 2019, 20, 518–530.
- Siena, S.; Di Bartolomeo, M.; Raghav, K.; Masuishi, T.; Loupakis, F.; Kawakami, H.; Yamaguchi, K.; Nishina, T.; Fakih, M.; Elez, E.; et al. Trastuzumab deruxtecan (DS-8201) in patients with HER2-expressing metastatic colorectal cancer (DESTINY-CRC01): A multicentre, open-label, phase 2 trial. Lancet Oncol. 2021, 22, 779–789.
- Le, D.T.; Uram, J.N.; Wang, H.; Bartlett, B.R.; Kemberling, H.; Eyring, A.D.; Skora, A.D.; Luber, B.S.; Azad, N.S.; Laheru, D.; et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N. Engl. J. Med. 2015, 372, 2509–2520.
- Dudley, J.C.; Lin, M.T.; Le, D.T.; Eshleman, J.R. Microsatellite Instability as a Biomarker for PD-1 Blockade. Clin. Cancer Res. 2016, 22, 813–820.
- Le, D.T.; Durham, J.N.; Smith, K.N.; Wang, H.; Bartlett, B.R.; Aulakh, L.K.; Lu, S.; Kemberling, H.; Wilt, C.; Luber, B.S.; et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 2017, 357, 409–413.
- Overman, M.J.; McDermott, R.; Leach, J.L.; Lonardi, S.; Lenz, H.J.; Morse, M.A.; Desai, J.; Hill, A.; Axelson, M.; Moss, R.A.; et al. Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142): An open-label, multicentre, phase 2 study. Lancet Oncol. 2017, 18, 1182–1191.
- Le, D.T.; Kim, T.W.; Van Cutsem, E.; Geva, R.; Jager, D.; Hara, H.; Burge, M.; O’Neil, B.; Kavan, P.; Yoshino, T.; et al. Phase II Open-Label Study of Pembrolizumab in Treatment-Refractory, Microsatellite Instability-High/Mismatch Repair-Deficient Metastatic Colorectal Cancer: KEYNOTE-164. J. Clin. Oncol. 2020, 38, 11–19.
- Hargadon, K.M.; Johnson, C.E.; Williams, C.J. Immune checkpoint blockade therapy for cancer: An overview of FDA-approved immune checkpoint inhibitors. Int. Immunopharmacol. 2018, 62, 29–39.
- Andre, T.; Shiu, K.K.; Kim, T.W.; Jensen, B.V.; Jensen, L.H.; Punt, C.; Smith, D.; Garcia-Carbonero, R.; Benavides, M.; Gibbs, P.; et al. Pembrolizumab in Microsatellite-Instability-High Advanced Colorectal Cancer. N. Engl. J. Med. 2020, 383, 2207–2218.
- Andre, T.; Amonkar, M.; Norquist, J.M.; Shiu, K.K.; Kim, T.W.; Jensen, B.V.; Jensen, L.H.; Punt, C.J.A.; Smith, D.; Garcia-Carbonero, R.; et al. Health-related quality of life in patients with microsatellite instability-high or mismatch repair deficient metastatic colorectal cancer treated with first-line pembrolizumab versus chemotherapy (KEYNOTE-177): An open-label, randomised, phase 3 trial. Lancet Oncol. 2021, 22, 665–677.
- Trullas, A.; Delgado, J.; Genazzani, A.; Mueller-Berghaus, J.; Migali, C.; Muller-Egert, S.; Zander, H.; Enzmann, H.; Pignatti, F. The EMA assessment of pembrolizumab as monotherapy for the first-line treatment of adult patients with metastatic microsatellite instability-high or mismatch repair deficient colorectal cancer. ESMO Open 2021, 6, 100145.
- Rodig, S.J.; Gusenleitner, D.; Jackson, D.G.; Gjini, E.; Giobbie-Hurder, A.; Jin, C.; Chang, H.; Lovitch, S.B.; Horak, C.; Weber, J.S.; et al. MHC proteins confer differential sensitivity to CTLA-4 and PD-1 blockade in untreated metastatic melanoma. Sci. Transl. Med. 2018, 10.
- Zaretsky, J.M.; Garcia-Diaz, A.; Shin, D.S.; Escuin-Ordinas, H.; Hugo, W.; Hu-Lieskovan, S.; Torrejon, D.Y.; Abril-Rodriguez, G.; Sandoval, S.; Barthly, L.; et al. Mutations Associated with Acquired Resistance to PD-1 Blockade in Melanoma. N. Engl. J. Med. 2016, 375, 819–829.
- Sade-Feldman, M.; Jiao, Y.J.; Chen, J.H.; Rooney, M.S.; Barzily-Rokni, M.; Eliane, J.P.; Bjorgaard, S.L.; Hammond, M.R.; Vitzthum, H.; Blackmon, S.M.; et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 2017, 8, 1136.
- Gu, S.S.; Zhang, W.; Wang, X.; Jiang, P.; Traugh, N.; Li, Z.; Meyer, C.; Stewig, B.; Xie, Y.; Bu, X.; et al. Therapeutically Increasing MHC-I Expression Potentiates Immune Checkpoint Blockade. Cancer Discov. 2021, 11, 1524–1541.
- Zehir, A.; Benayed, R.; Shah, R.H.; Syed, A.; Middha, S.; Kim, H.R.; Srinivasan, P.; Gao, J.; Chakravarty, D.; Devlin, S.M.; et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 2017, 23, 703–713.
- Johnson, D.B.; Dahlman, K.H.; Knol, J.; Gilbert, J.; Puzanov, I.; Means-Powell, J.; Balko, J.M.; Lovly, C.M.; Murphy, B.A.; Goff, L.W.; et al. Enabling a genetically informed approach to cancer medicine: A retrospective evaluation of the impact of comprehensive tumor profiling using a targeted next-generation sequencing panel. Oncologist 2014, 19, 616–622.
- Tsimberidou, A.M.; Iskander, N.G.; Hong, D.S.; Wheler, J.J.; Falchook, G.S.; Fu, S.; Piha-Paul, S.; Naing, A.; Janku, F.; Luthra, R.; et al. Personalized medicine in a phase I clinical trials program: The MD Anderson Cancer Center initiative. Clin. Cancer Res. 2012, 18, 6373–6383.
- Stockley, T.L.; Oza, A.M.; Berman, H.K.; Leighl, N.B.; Knox, J.J.; Shepherd, F.A.; Chen, E.X.; Krzyzanowska, M.K.; Dhani, N.; Joshua, A.M.; et al. Molecular profiling of advanced solid tumors and patient outcomes with genotype-matched clinical trials: The Princess Margaret IMPACT/COMPACT trial. Genome Med. 2016, 8, 109.
- Meric-Bernstam, F.; Brusco, L.; Shaw, K.; Horombe, C.; Kopetz, S.; Davies, M.A.; Routbort, M.; Piha-Paul, S.A.; Janku, F.; Ueno, N.; et al. Feasibility of Large-Scale Genomic Testing to Facilitate Enrollment Onto Genomically Matched Clinical Trials. J. Clin. Oncol. 2015, 33, 2753–2762.
- Von Hoff, D.D.; Stephenson, J.J., Jr.; Rosen, P.; Loesch, D.M.; Borad, M.J.; Anthony, S.; Jameson, G.; Brown, S.; Cantafio, N.; Richards, D.A.; et al. Pilot study using molecular profiling of patients’ tumors to find potential targets and select treatments for their refractory cancers. J. Clin. Oncol. 2010, 28, 4877–4883.
- Laskin, J.; Jones, S.; Aparicio, S.; Chia, S.; Ch’ng, C.; Deyell, R.; Eirew, P.; Fok, A.; Gelmon, K.; Ho, C.; et al. Lessons learned from the application of whole-genome analysis to the treatment of patients with advanced cancers. Cold Spring Harb. Mol. Case Stud. 2015, 1, a000570.
- Sholl, L.M.; Do, K.; Shivdasani, P.; Cerami, E.; Dubuc, A.M.; Kuo, F.C.; Garcia, E.P.; Jia, Y.; Davineni, P.; Abo, R.P.; et al. Institutional implementation of clinical tumor profiling on an unselected cancer population. JCI Insight 2016, 1, e87062.
- Kikuchi, J.; Ohhara, Y.; Takada, K.; Tanabe, H.; Hatanaka, K.; Amano, T.; Hatanaka, K.C.; Hatanaka, Y.; Mitamura, T.; Kato, M.; et al. Clinical significance of comprehensive genomic profiling tests covered by public insurance in patients with advanced solid cancers in Hokkaido, Japan. Jpn. J. Clin. Oncol. 2021, 51, 753–761.
- Takeda, M.; Takahama, T.; Sakai, K.; Shimizu, S.; Watanabe, S.; Kawakami, H.; Tanaka, K.; Sato, C.; Hayashi, H.; Nonagase, Y.; et al. Clinical Application of the FoundationOne CDx Assay to Therapeutic Decision-Making for Patients with Advanced Solid Tumors. Oncologist 2021, 26, e588–e596.
- Kondo, T.; Matsubara, J.; Quy, P.N.; Fukuyama, K.; Nomura, M.; Funakoshi, T.; Doi, K.; Sakamori, Y.; Yoshioka, M.; Yokoyama, A.; et al. Comprehensive genomic profiling for patients with chemotherapy-naive advanced cancer. Cancer Sci. 2021, 112, 296–304.
- Schwaederle, M.; Daniels, G.A.; Piccioni, D.E.; Fanta, P.T.; Schwab, R.B.; Shimabukuro, K.A.; Parker, B.A.; Kurzrock, R. On the Road to Precision Cancer Medicine: Analysis of Genomic Biomarker Actionability in 439 Patients. Mol. Cancer Ther. 2015, 14, 1488–1494.
- Hirshfield, K.M.; Tolkunov, D.; Zhong, H.; Ali, S.M.; Stein, M.N.; Murphy, S.; Vig, H.; Vazquez, A.; Glod, J.; Moss, R.A.; et al. Clinical Actionability of Comprehensive Genomic Profiling for Management of Rare or Refractory Cancers. Oncologist 2016, 21, 1315–1325.
- Schwaederle, M.; Parker, B.A.; Schwab, R.B.; Daniels, G.A.; Piccioni, D.E.; Kesari, S.; Helsten, T.L.; Bazhenova, L.A.; Romero, J.; Fanta, P.T.; et al. Precision Oncology: The UC San Diego Moores Cancer Center PREDICT Experience. Mol. Cancer Ther. 2016, 15, 743–752.
- Schwaederle, M.; Zhao, M.; Lee, J.J.; Lazar, V.; Leyland-Jones, B.; Schilsky, R.L.; Mendelsohn, J.; Kurzrock, R. Association of Biomarker-Based Treatment Strategies With Response Rates and Progression-Free Survival in Refractory Malignant Neoplasms: A Meta-analysis. JAMA Oncol. 2016, 2, 1452–1459.
- Priestley, P.; Baber, J.; Lolkema, M.P.; Steeghs, N.; de Bruijn, E.; Shale, C.; Duyvesteyn, K.; Haidari, S.; van Hoeck, A.; Onstenk, W.; et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 2019, 575, 210–216.
More
Information
Subjects:
Oncology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
817
Revisions:
2 times
(View History)
Update Date:
28 Sep 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




