
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Ramon Alvarez-Puebla | + 2387 word(s) | 2387 | 2021-09-02 04:34:33 | | | |
| 2 | Vivi Li | Meta information modification | 2387 | 2021-09-15 04:33:38 | | |
Video Upload Options
As medicine continues to advance our understanding of and knowledge about the complex and multifactorial nature of cancer, new major technological challenges have emerged in the design of analytical methods capable of characterizing and assessing the dynamic heterogeneity of cancer for diagnosis, prognosis and monitoring, as required by precision medicine. With this aim, novel nanotechnological approaches have been pursued and developed for overcoming intrinsic and current limitations of conventional methods in terms of rapidity, sensitivity, multiplicity, non-invasive procedures and cost. Eminently, a special focus has been put on their implementation in liquid biopsy analysis. Among optical nanosensors, those based on surface-enhanced Raman scattering (SERS) have been attracting tremendous attention due to the combination of the intrinsic prerogatives of the technique (e.g., sensitivity and structural specificity) and the high degree of refinement in nano-manufacturing, which translate into reliable and robust real-life applications.
1. Introduction
2. SERS Imaging of Cancer Tissues and Single Cells
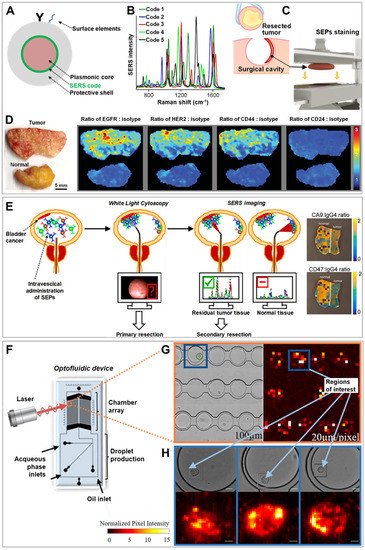
References
- Sant, G.R.; Knopf, K.B.; Albala, D.M. Live-single-cell phenotypic cancer biomarkers-future role in precision oncology? NPJ Precis. Oncol. 2017, 1, 21.
- Borrebaeck, C.A.K. Precision diagnostics: moving towards protein biomarker signatures of clinical utility in cancer. Nat. Rev. Cancer 2017, 17, 199–204.
- Cohen, J.D.; Javed, A.A.; Thoburn, C.; Wong, F.; Tie, J.; Gibbs, P.; Schmidt, C.M.; Yip-Schneider, M.T.; Allen, P.J.; Schattner, M.; et al. Combined circulating tumor DNA and protein biomarker-based liquid biopsy for the earlier detection of pancreatic cancers. Proc. Natl. Acad. Sci. USA 2017, 114, 10202–10207.
- Shi, J.; Kantoff, P.W.; Wooster, R.; Farokhzad, O.C. Cancer nanomedicine: Progress, challenges and opportunities. Nat. Rev. Cancer 2017, 17, 20–37.
- Barreto, J.A.; O’Malley, W.; Kubeil, M.; Graham, B.; Stephan, H.; Spiccia, L. Nanomaterials: Applications in Cancer Imaging and Therapy. Adv. Mater. 2011, 23, H18–H40.
- Laing, S.; Jamieson, L.E.; Faulds, K.; Graham, D. Surface-enhanced Raman spectroscopy for in vivo biosensing. Nat. Rev. Chem. 2017, 1, e0060.
- Wang, Z.Y.; Zong, S.F.; Wu, L.; Zhu, D.; Cui, Y.P. SERS-Activated Platforms for Immunoassay: Probes, Encoding Methods, and Applications. Chem. Rev. 2017, 117, 7910–7963.
- Zong, C.; Xu, M.X.; Xu, L.J.; Wei, T.; Ma, X.; Zheng, X.S.; Hu, R.; Ren, B. Surface-Enhanced Raman Spectroscopy for Bioanalysis: Reliability and Challenges. Chem. Rev. 2018, 118, 4946–4980.
- Guerrini, L.; Pazos-Perez, N.; Garcia-Rico, E.; Alvarez-Puebla, R. Cancer characterization and diagnosis with SERS-encoded particles. Cancer Nanotechnol. 2017, 8, e5.
- Schlücker, S. Surface-Enhanced Raman Spectroscopy: Concepts and Chemical Applications. Angew. Chem. Int. Edit. 2014, 53, 4756–4795.
- Pedrol, E.; Garcia-Algar, M.; Massons, J.; Nazarenus, M.; Guerrini, L.; Martínez, J.; Rodenas, A.; Fernandez-Carrascal, A.; Aguiló, M.; Estevez, L.G.; et al. Optofluidic device for the quantification of circulating tumor cells in breast cancer. Sci. Rep. 2017, 7, e3677.
- Pedrol, E.; Martínez, J.; Aguiló, M.; Garcia-Algar, M.; Nazarenus, M.; Guerrini, L.; Garcia-Rico, E.; Álvarez-Puebla, R.A.; Díaz, F.; Massons, J. Microfluidic device with dual-channel fluorescence acquisition for quantification/identification of cancer cells. Microfluid. Nanofluid. 2017, 21, e181.
- Fernandez-Carrascal, A.; Garcia-Algar, M.; Nazarenus, M.; Torres-Nuñez, A.; Guerrini, L.; Feliu, N.; Parak, W.J.; Garcia-Rico, E.; Alvarez-Puebla, R.A. Metabolic pathway for the universal fluorescent recognition of tumor cells. Oncotarget 2017, 8, 76108–76115.
- Jiang, R.; Li, B.; Fang, C.; Wang, J. Metal/Semiconductor Hybrid Nanostructures for Plasmon-Enhanced Applications. Adv. Mater. 2014, 26, 5274–5309.
- Wang, Y.Q.; Yan, B.; Chen, L.X. SERS Tags: Novel Optical Nanoprobes for Bioanalysis. Chem. Rev. 2013, 113, 1391–1428.
- Lane, L.A.; Qian, X.M.; Nie, S.M. SERS Nanoparticles in Medicine: From Label-Free Detection to Spectroscopic Tagging. Chem. Rev. 2015, 115, 10489–10529.
- Fabris, L. SERS Tags: The Next Promising Tool for Personalized Cancer Detection? ChemNanoMat 2016, 2, 249–258.
- Wang, Y.W.; Doerksen, J.D.; Kang, S.Y.; Walsh, D.; Yang, Q.; Hong, D.; Liu, J.T.C. Multiplexed Molecular Imaging of Fresh Tissue Surfaces Enabled by Convection-Enhanced Topical Staining with SERS-Coded Nanoparticles. Small 2016, 12, 5612–5621.
- Davis, R.M.; Kiss, B.; Trivedi, D.R.; Metzner, T.J.; Liao, J.C.; Gambhir, S.S. Surface-Enhanced Raman Scattering Nanoparticles for Multiplexed Imaging of Bladder Cancer Tissue Permeability and Molecular Phenotype. ACS Nano 2018, 12, 9669–9679.
- Willner, M.R.; McMillan, K.S.; Graham, D.; Vikesland, P.J.; Zagnoni, M. Surface-Enhanced Raman Scattering Based Microfluidics for Single-Cell Analysis. Anal. Chem. 2018, 90, 12004–12010.
- Willets, K.A.; Van Duyne, R.P. Localized surface plasmon resonance spectroscopy and sensing. Annu. Rev. Phys. Chem. 2007, 58, 267–297.
- Le Ru, E.C.; Etchegoin, P.G. Principles of Surface-Enhanced Raman Spectroscopy; Elsevier: Amsterdam, The Netherlands, 2009.
- Lim, E.-K.; Kim, T.; Paik, S.; Haam, S.; Huh, Y.-M.; Lee, K. Nanomaterials for Theranostics: Recent Advances and Future Challenges. Chem. Rev. 2015, 115, 327–394.
- Cassano, D.; Santi, M.; D’Autilia, F.; Mapanao, A.K.; Luin, S.; Voliani, V. Photothermal effect by NIR-responsive excretable ultrasmall-in-nano architectures. Mater. Horiz. 2019, 6, 531–537.
- Wang, Y.; Kang, S.; Khan, A.; Ruttner, G.; Leigh, S.Y.; Murray, M.; Abeytunge, S.; Peterson, G.; Rajadhyaksha, M.; Dintzis, S.; et al. Quantitative molecular phenotyping with topically applied SERS nanoparticles for intraoperative guidance of breast cancer lumpectomy. Sci. Rep. 2016, 6, 21242.
- Mallia, R.J.; McVeigh, P.Z.; Fisher, C.J.; Veilleux, I.; Wilson, B.C. Wide-field multiplexed imaging of EGFR-targeted cancers using topical application of NIR SERS nanoprobes. Nanomedicine 2015, 10, 89–101.
- Zavaleta, C.L.; Garai, E.; Liu, J.T.C.; Sensarn, S.; Mandella, M.J.; Van de Sompel, D.; Friedland, S.; Van Dam, J.; Contag, C.H.; Gambhir, S.S. A Raman-based endoscopic strategy for multiplexed molecular imaging. Proc. Natl. Acad. Sci. USA 2013, 110, E2288–E2297.
- Oseledchyk, A.; Andreou, C.; Wall, M.A.; Kircher, M.F. Folate-Targeted Surface-Enhanced Resonance Raman Scattering Nanoprobe Ratiometry for Detection of Microscopic Ovarian Cancer. ACS Nano 2017, 11, 1488–1497.
- Zavagno, G.; Goldin, E.; Mencarelli, R.; Capitanio, G.; Bianco, P.D.; Marconato, R.; Mocellin, S.; Marconato, G.; Belardinelli, V.; Marcon, F.; et al. Role of resection margins in patients treated with breast conservation surgery. Cancer 2008, 112, 1923–1931.
- Kim, Y.I.; Jeong, S.; Jun, B.H.; Lee, Y.S.; Lee, Y.S.; Jeong, D.H.; Lee, D.S. Endoscopic imaging using surface-enhanced Raman scattering. Eur. J. Nanomed. 2017, 9, 14.
- Garai, E.; Sensarn, S.; Zavaleta, C.L.; Loewke, N.O.; Rogalla, S.; Mandella, M.J.; Felt, S.A.; Friedland, S.; Liu, J.T.C.; Gambhir, S.S.; et al. A real-time clinical endoscopic system for intraluminal, multiplexed imaging of surface-enhanced Raman scattering nanoparticles. PLoS ONE 2015, 10, e0123185.
- Zavaleta, C.L.; Smith, B.R.; Walton, I.; Doering, W.; Davis, G.; Shojaei, B.; Natan, M.J.; Gambhir, S.S. Multiplexed imaging of surface enhanced Raman scattering nanotags in living mice using noninvasive Raman spectroscopy. Proc. Natl. Acad. Sci. USA 2009, 106, 13511–13516.
- Garcia-Algar, M.; Fernandez-Carrascal, A.; Olano-Daza, A.; Guerrini, L.; Feliu, N.; Parak, W.J.; Guimera, R.; Garcia-Rico, E.; Alvarez-Puebla, R.A. Adaptive metabolic pattern biomarker for disease monitoring and staging of lung cancer with liquid biopsy. NPJ Precis. Oncol. 2018, 2, 16.
- Zeng, Y.; Koo, K.M.; Trau, M.; Shen, A.-G.; Hu, J.-M. Watching SERS glow for multiplex biomolecular analysis in the clinic: A review. Appl. Mater. Today 2019, 15, 431–444.
- Mir-Simon, B.; Reche-Perez, I.; Guerrini, L.; Pazos-Perez, N.; Alvarez-Puebla, R.A. Universal One-Pot and Scalable Synthesis of SERS Encoded Nanoparticles. Chem. Mat. 2015, 27, 950–958.
- Pazos-Perez, N.; Fitzgerald, J.M.; Giannini, V.; Guerrini, L.; Alvarez-Puebla, R.A. Modular assembly of plasmonic core–satellite structures as highly brilliant SERS-encoded nanoparticles. Nanoscale Adv. 2019, 1, 122–131.
- Bohndiek, S.E.; Wagadarikar, A.; Zavaleta, C.L.; Van de Sompel, D.; Garai, E.; Jokerst, J.V.; Yazdanfar, S.; Gambhir, S.S. A small animal Raman instrument for rapid, wide-area, spectroscopic imaging. Proc. Natl. Acad. Sci. USA 2013, 110, 12408–12413.
- Alvarez-Puebla, R.A.; Pazos-Perez, N.; Guerrini, L. SERS-fluorescent encoded particles as dual-mode optical probes. Appl. Mater. Today 2018, 13, 1–14.
- Jeong, S.; Kim, Y.-i.; Kang, H.; Kim, G.; Cha, M.G.; Chang, H.; Jung, K.O.; Kim, Y.-H.; Jun, B.-H.; Hwang, D.W.; et al. Fluorescence-Raman Dual Modal Endoscopic System for Multiplexed Molecular Diagnostics. Sci. Rep. 2015, 5, 9455.
- Aroca, R.F. Plasmon enhanced spectroscopy. Phys. Chem. Chem. Phys. 2013, 15, 5355–5363.
- Li, J.-F.; Li, C.-Y.; Aroca, R.F. Plasmon-enhanced fluorescence spectroscopy. Chem. Soc. Rev. 2017, 46, 3962–3979.
- Bodelon, G.; Montes-García, V.; Fernández-Lõpez, C.; Pastoriza-Santos, I.; Pérez-Juste, J.; Liz-Marzán, L.M. [email protected] SERRS Tags for Multiplex Immunophenotyping Cellular Receptors and Imaging Tumor Cells. Small 2015, 11, 4149–4157.
- Jahn, I.J.; Zukovskaja, O.; Zheng, X.S.; Weber, K.; Bocklitz, T.W.; Cialla-May, D.; Popp, J. Surface-enhanced Raman spectroscopy and microfluidic platforms: challenges, solutions and potential applications. Analyst 2017, 142, 1022–1047.
- Johnson-Chavarria, E.M.; Agrawal, U.; Tanyeri, M.; Kuhlman, T.E.; Schroeder, C.M. Automated single cell microbioreactor for monitoring intracellular dynamics and cell growth in free solution. Lab Chip 2014, 14, 2688–2697.




