Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Runlong Zhang | + 2987 word(s) | 2987 | 2021-08-12 11:16:35 | | | |
| 2 | Ron Wang | + 308 word(s) | 3295 | 2021-08-31 09:51:35 | | | | |
| 3 | Ron Wang | + 1019 word(s) | 4006 | 2021-08-31 09:52:21 | | | | |
| 4 | Ron Wang | + 1019 word(s) | 4006 | 2021-08-31 09:52:50 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Zhang, R. Chilling Requirement Validation. Encyclopedia. Available online: https://encyclopedia.pub/entry/13732 (accessed on 07 February 2026).
Zhang R. Chilling Requirement Validation. Encyclopedia. Available at: https://encyclopedia.pub/entry/13732. Accessed February 07, 2026.
Zhang, Runlong. "Chilling Requirement Validation" Encyclopedia, https://encyclopedia.pub/entry/13732 (accessed February 07, 2026).
Zhang, R. (2021, August 31). Chilling Requirement Validation. In Encyclopedia. https://encyclopedia.pub/entry/13732
Zhang, Runlong. "Chilling Requirement Validation." Encyclopedia. Web. 31 August, 2021.
Copy Citation
The introduction of herbaceous peony (Paeonia lactiflora Pall.) in low-latitude areas is of great significance to expand the landscape application of this world-famous ornamental. With the hazards of climate warming, warm winters occurs frequently, which makes many excellent northern herbaceous peony cultivars unable to meet their chilling requirements (CR) and leads to their poor growth and flowering in southern China. Exploring the endodormancy release mechanism of underground buds is crucial for improving low-CR cultivar screening and breeding.
abscisic acid
chilling requirement (CR)
endodormancy release
herbaceous peony
Paeonia lactiflora Pall
reactive oxygen species
underground bud dormancy
1. Introduction
Bud dormancy is an important survival strategy of temperate perennial plants in response to detrimental environments, such as cold in winter. With global climate change, bud dormancy has gradually become a popular topic in plant growth and development research fields in recent decades [1][2]. The complete process of bud dormancy includes three contiguous stages: paradormancy, endodormancy and ecodormancy [3]. Paradormancy is well accepted as the cessation of bud growth after shoot elongation stops. As the daylength shortens and the temperature decreases, perennials defoliate, and their buds generally transfer from paradormancy to endodormancy [4]. In autumn, with a decrease in temperature or day length, buds enter endodormancy, at which stage they are unable to sprout even under favorable conditions until the indispensable chilling accumulation has been completed [5][6]. Ecodormancy is the dormancy caused by adverse environmental factors such as low temperature and drought stress [3]. Once the environmental conditions are suitable, dormancy is released. However, endodormancy cannot be released before accomplishing indispensable chilling accumulation. Due to the global warming trend, many plants with winter dormancy traits suffer from hot autumns and warm winters, which disrupt normal endodormancy release because of insufficient chilling accumulation and then negatively affect their subsequent growth and development. Sufficient daylength and low temperature are the two most important environmental factors affecting the bud dormancy process. Short daylength could induce Populus L. and Morus alba L. to enter dormancy, and a low-temperature environment could induce Pyrus spp. and Malus pumila Mill. to enter dormancy [7][8][9]. Shortening daylength alone does not induce these fruit trees to enter bud dormancy without decreasing temperature. The dormancy of Vitis vinifera L. and Armeniaca vulgaris Lam. was mainly controlled by sunshine duration and low temperature [10][11].
The internal factors controlling dormancy processes are complicated and involve carbohydrate conversion, hormone signaling, antioxidant metabolism, water transport, gene regulation, and DNA methylation and demethylation [12][13][14]. During the paradormancy stage, sugar metabolic activity is continuously vigorous, and the large amount of accumulated soluble total sugar and sucrose not only helps to improve cell osmotic potential to resist the cold in winter but also acts as a regulatory substance to inhibit the growth of buds in perennials [15][16][17]. In addition, the content and balance of endogenous hormones are currently widely accepted to control the occurrence and release of bud endodormancy. A previous study showed that the formation and maintenance of abscisic acid (ABA) and the status transition of bud dormancy were closely interrelated in potato. The ABA content increased with the deepening of bud endodormancy but decreased regularly with the endodormancy release [18]. Generally, the ABA content continuously increased in paradormancy, reached the highest level in endodormancy and finally decreased to the lowest level in ecodormancy [18][19][20]. Gibberellin (GA) was confirmed to promote the dissolution of dormancy. The accumulation of low temperature in winter resulted in an increase in GA content and then promoted the degradation of callose in the sieve tube, which was conducive to promoting the release of endodormancy [21]. Additionally, reactive oxygen species (ROS) function as signal transduction factors and are also involved in many regulatory processes related to plant growth and development. The activities of superoxide dismutase (SOD) and peroxidase (POD) gradually increased, while catalase (CAT) activity showed the opposite trend after the end of endodormancy in pear [22].
In addition to the control of physiological factors, bud endodormancy induction, transition and release are also regulated by various genes and associated pathways. Hormone signaling and metabolism-related genes, such as 9-cis-epoxycarotenoid dioxygenase (NCED), ABA-insensitive 5 (ABI5), protein phosphatase 2ca (PP2C), and Arabidopsis thaliana gibberellin 2-oxidase (GA2ox), are popular research topics in plant bud endodormancy, while sucrose synthase 3 (SUS3), Arabidopsis thaliana cell wall invertase 1 (CWINV), cyclin D3 (CYCD) and cyclin-dependent kinase b2;2 (CDKB22) are also frequently involved in carbohydrate and ROS metabolism. These genes promote or inhibit plant bud endodormancy and may play an important role in breeding plant germplasm with long or short endodormancy periods through gene overexpression and silencing. In addition, dormancy-associated MADS-box (DAM) genes belong to the MADS-box gene family and are crucial for bud endodormancy regulation [23]. DAM genes were isolated and identified in a study on bud endodormancy in peach with rare evergreen traits [24]. Since then, researchers have identified related genes in many other species (pear, plum, kiwifruit) and further confirmed their association with the endodormancy process [25][26][27]. These genes form a complex dormancy regulatory network with other MADS-box genes and genes related to carbohydrate, phytohormone and antioxidant metabolism [28].
The bud endodormancy of Paeonia suffruticosa belonging to the same genus as Paeonia lactiflora has been studied for decades. During the process of herbaceous peony flower bud endodormancy release, accompanied by the hydrolysis of soluble sugar, starch, and the increase of antioxidant enzyme activity [29]. Hormone regulation of herbaceous peony flower bud endodormancy is mainly dependent on the ratio of endodormancy release promoter to inhibitor, especially GA and ABA, the ratio between the two is closely related to the endodormancy process of herbaceous peony flower bud [18][30][31]. In addition, in-depth research at the molecular level has mainly focused on methylation and gene regulation [32][33]. GA and low temperature both have the effect of reducing DNA methylation. The gene GA2ox8 related to GA synthesis is also one of the important genes regulating endodormancy [34].
Herbaceous peony (Paeonia lactiflora Pall.) is a world-famous ornamental plant that originates from China and has a long cultivation history worldwide [35]. It is widely used in cut flowers, potted plants and gardens. This species is mainly distributed in temperate and cool regions of the Northern Hemisphere, such as Europe, North America and Asia [36]. To survive in cold winters, the aboveground parts of herbaceous peony wither in autumn, and the underground bud undergoes endodormancy. Only after accumulating enough of a chilling requirement (CR) in winter can it grow well in the coming year. Therefore, chilling is an essential factor in the endodormancy-growth cycle of herbaceous peony [37]. In this dormant period, the underground buds of herbaceous peony undergo a series of physiological changes, such as carbohydrate conversion, hormone metabolism and ROS signal transduction [38]. The introduction of herbaceous peony from high latitudes to low latitudes is beneficial to expand its distribution range and increase its application in landscaping. However, the winter temperature at low latitudes is too high to meet the CR, which poses a great obstacle to the release of bud endodormancy in the coming spring [39][40].
2. Current Insights
2.1. CR Evaluation Is an Important Method to Popularize Herbaceous Peony at Low Latitudes
Compared with long-term breeding, CR evaluation and screening of cultivars with low cold demand is more direct and can help select target cultivars in a shorter time. Typical materials with high cold demand and low cold demand can be selected after evaluation and can be used as parents for scientific research and long-term breeding [41]. For herbaceous peonies, it will take a long time to create new cultivars with the low-CR trait by a traditional crossbreeding strategy. Therefore, the introduction of herbaceous peonies from high-latitude areas and CR evaluation are needed to screen for low-CR cultivars [37]. In this study, the CR of ‘Meiju’ was calculated as 677.5 CUs by the UT model, and thus ‘Meiju’ was verified as a low-CR cultivar compared with other northern cultivars (‘Zhuguang’: 1182 CUs, ‘Qiaoling’: 1017 CUs, ‘Qihua Lushuang’: 685.5 CUs, ‘Fen Yunu’: 685.5 CUs [37]). In our previous study, the bud endodormancy of ‘Meiju’ was released on 9 January 2017 and the CR was correspondingly calculated as 1017.0 CUs based on the UT model [37]. As a matter of fact, we analyzed the morphological data more carefully afterwards, and deduced that the bud endodormancy of ‘Meiju’ had probably been released on 26 December 2016. Therefore, the true CR value of ‘Meiju’ should be 685.5 CUs during the winter of 2016 to 2017, which is very close to 677.5 CUs verified in this study. This northern cultivar grew vigorously and bloomed exuberantly in Hangzhou based on multiyear observations; thus, it could be used as a pioneer cultivar to popularize herbaceous peony in Hangzhou and other low-latitude areas [37][42].
2.2. Carbohydrate Metabolism Participates in the Regulation of Endodormancy Release
The conversion of sucrose, total soluble sugar and starch is an important factor in regulating bud endodormancy in perennials [15][16][17][43][44]. Generally, in the stage of endodormancy, starch tends to decrease, while sucrose and total soluble sugar increase accordingly. Starch is a polysaccharide that cannot be directly used by plants, and it can only be used when it is converted into monosaccharides. Therefore, the increase in sucrose and total soluble sugar content in herbaceous peony buds before 9 January may come from the decomposition of starch, which can provide energy for the life activities of herbaceous peony buds and promote the gradual release of endodormancy in herbaceous peony buds [15][16]. However, this is different from the results from a study in spinach, which shows that different species have different regulatory mechanisms [45].
On the other hand, the increase in sucrose and total soluble sugar content may also help to reduce the osmotic pressure of cells and ensure that the cells can fully absorb water, thus increasing the cold resistance of herbaceous peony buds and helping them survive the lowest temperature period in Hangzhou in January (Figure 2) [46].
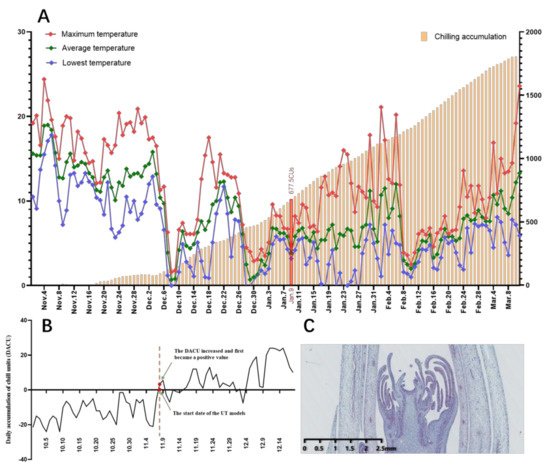
Figure 2. (A): Temperature change and chilling accumulation of underground buds from ‘Meiju’ during dormancy (using the UT model to calculate the chilling accumulation). The line chart represents the daytime temperature, and the bar chart represents the chilling accumulation value. (B): Trends in the daily chill-unit accumulation and start dates for CR evaluations via the UT model. (C): A slice of the underground bud closest to the start date.
2.3. Relationship between Hormone Metabolism and Endodormancy Release
Plant endogenous hormones play an important role in the induction, transformation and release of bud endodormancy [4][47][48]. Plant endogenous hormones mainly include two categories: promoting the endodormancy release of buds (such as GA, ZR, BR, JA, and IAA) and inhibiting the release (maintaining endodormancy, such as ABA) [12][49][50]. The endogenous hormone content in plants is an important factor regulating the induction and release of endodormancy, and the dynamic balance between hormone contents is often highly correlated with the process of dormancy [4][48].
Figure 10 shows that among the genes associated with ABA, only the expression pattern of ABI5 was completely consistent with the change in ABA level from endodormancy to ecological dormancy. ABI5 is a key transcription factor in the ABA signaling pathway that helps to promote ABA signal transduction [41]. The rapid downregulation of ABI5 around 9 January may be one of the reasons for the decrease in ABA content, which is the key hormone-related gene to accelerate the release of endodormancy [51].
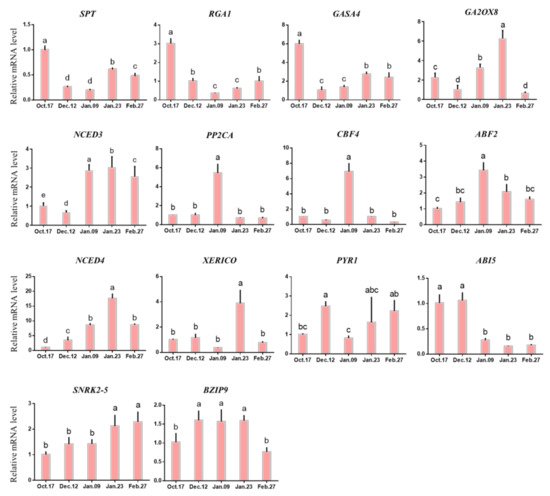
Figure 10. Expression of hormone metabolism-related genes in different dormancy stages. Each bar represents the mean ± SEM (n = 3). Different letters indicate a significant difference among different dates (p < 0.05). The picture was drawn by GraphPad Prism 8.0 and the data were processed by IBM SPSS Statistics 26.
As an ABA binding factor, ABF2 often combines with AREB1 to form the AREB1/ABF2 transcription factor, which is the key factor downstream of the ABA signaling pathway [52][53]. In the process of bud dormancy, the expression of ABF2 is theoretically conducive to ABA signal transduction, thus maintaining bud endodormancy [6][54]. In this study, ABF2 showed a unimodal curve, which reached its peak on 9 January (Figure 10), suggesting that its high expression promoted the release of endodormancy in buds. However, the trend of ABF2 expression during the whole endodormancy period was opposite that of ABA (Figure 6 and Figure 10), suggesting that its expression was upregulated but that the ABA content was decreased.
However, the expression of other genes is only in line with expectations at some stages. For example, the expression of upstream genes NCED3 and NCED4 in the ABA synthesis pathway increases during endodormancy (Figure 10, 17 October–9 January); this gene has been proven to be an important gene for promoting ABA synthesis, which is consistent with the results of ABA content determination (Figure 6) [31][55][56]. One of the possible reasons why ‘Meiju’ is a low CR herbaceous peony is that the NCED gene is upregulated in early winter. However, in the stage of ecodormancy in which the content of ABA decreased, NCED3 and NCED4 continued to be upregulated and maintained at a relatively high expression level (9 January–27 February). Similar results are also reflected in pear, which shows that the genes in the NCED gene family regulate different biological processes, and these genes may also be involved in the resistance of herbaceous peony buds to low temperature in winter during ecodormancy [4].
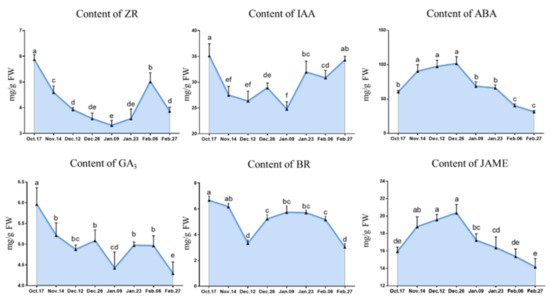
Figure 6. The changes in hormone contents during the underground bud dormancy period of ‘Meiju’. Each bar represents the mean ± SEM (n = 3). Different letters indicate a significant difference among different dates (p < 0.05). The experimental time was from 2018 to 2019, and the data were processed by IBM SPSS Statistics 26.
GA3 is one of the important hormones regulating bud endodormancy [57]. In this study, the content of GA3 was gradually decreased before the endodormancy release, and the decrease might be related to the antagonism of ABA. In the stage when endodormancy release (before 9 January), the ABA content decreased rapidly and the ratio of GA/ABA began to increase, which promoted the release of endodormancy. In terms of gene expression, GA2ox has been proved to have a negative regulatory effect on GA synthesis in Arabidopsis thaliana and poplar [58][59]. After the endodormancy released, the rapid increase in GA3 content might be related to the up-regulation of GASA4 gene expression, which is related to gibberellin synthesis.
In summary, the chilling requirement of ‘Meiju’ is shorter, and the endodormancy release period is earlier, which is closely related to the early decrease in ABA content and the regulation of related genes such as ABI5, NCED3, and PP2C [60][56][61][62]. More gene function verification experiments are needed to confirm this hypothesis. In addition to ABA, hormones that promote the release of bud endodormancy, such as GA3, IAA, JA, ZR and BR, which act in opposition to ABA, can also be involved in determining the characteristics of low chilling requirements [63][64][65][66].
2.4. Antioxidant Metabolism Involved in Endodormancy Release Regulation
Many studies have shown that antioxidant metabolism is also involved in the regulation of bud endodormancy [22][67]. Similar to a study in pears, the activity of SOD decreased before the release of ecodormancy but began to increase after this release, and the activity of CAT gradually increased before the termination of endodormancy and began to decline after termination [22]. The activity of POD was upregulated rapidly before the release of endodormancy and then decreased rapidly. These findings are completely different from those in pears [22]. The enzyme activity related to the antioxidant metabolic pathway responded rapidly during paradormancy and coincided with the degree of dormancy. This suggests that ROS may act as a signal transduction substance to sense environmental changes and participate in the regulation of paradormancy induction and release. However, the related regulatory mechanisms may be different in different species [22][68].
At the level of gene expression, the expression of most genes related to antioxidant metabolism (EXPA6, CDKB22, CYCD3, BG3, actin-related protein 6 (ARP6), PER52, histone h1-3 (HIS1-3)) decreased gradually before the release of endodormancy and then increased again (Figure 9). The expression of these genes changed dramatically, decreased rapidly when entering endodormancy, and then increased rapidly after endodormancy was released.
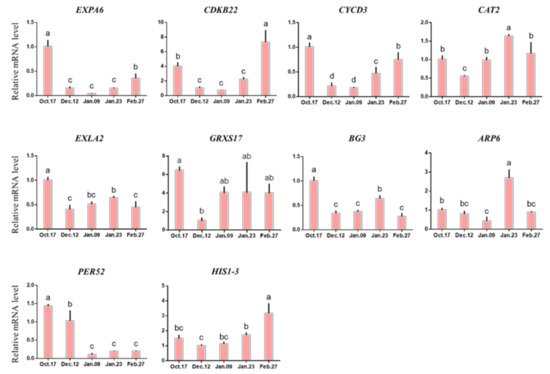
Figure 9. Change in the expression levels of antioxidant metabolism-related genes. Each bar represents the mean ± SEM (n = 3). Different letters indicate a significant difference among different dates (p < 0.05). The picture was drawn by GraphPad Prism 8.0 and the data were processed by IBM SPSS Statistics 26.
In other metabolic pathways, the expression trend of genes related to ROS signal transduction was opposite those related to sucrose content (Figure 5), total soluble sugar content (Figure 5) and ABA content (Figure 6). ROS may play a role in sensing and transmitting signals in the process of dormancy release in buds of ‘Meiju’. This shows that the efficient signal transmission of ROS and the timely reflection of other physiological activities may be one of the reasons for the low cooling requirement of ‘Meiju’.

Figure 5. The changes in sugar and carbohydrate contents during the bud dormancy period of ‘Meiju’. Each bar represents the mean ± SEM (n = 3). The experimental time was from 2018 to 2019, and the data were processed by IBM SPSS Statistics 26. Different letters indicate a significant difference among different dates (p < 0.05).
2.5. Judgment of Endodormancy Release Should Be Based on Comprehensive Research
According to the experiment of moving outdoor potted herbaceous peony to a greenhouse, several key indices (BPF, ANS, APW, APH, ADS, WFS) were used as the most intuitive evidence to judge that the endodormancy of herbaceous peony was released around 9 January.
Additionally, the state of cells in leaf primordia (LP), growing point (GP), vascular strand primordia (VS), bud primordia (BP) and other parts is also quite different from that in the previous period, with an obvious tight arrangement of cells and a deeper color of nuclei (Figure 4). The continuous accumulation of these soluble carbohydrates possibly helped to increase the osmotic potential of underground bud cells and enhanced the cold resistance of ‘Meiju’ (Figure 5). High concentrations of sugar, together with ROS signaling, may lead to a decrease in the contents of hormones such as ABA, JAME, and IAA (Figure 12). The expression of many important genes related to endodormancy release, such as CBF4, PP2C, and NCEDs, reached the highest level around 9 January (Figure 10) [42][45][46][48].
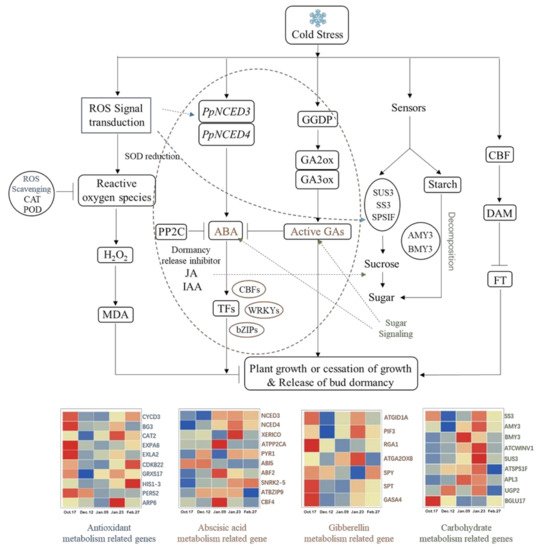
Figure 12. Underground bud endodormancy regulatory network. The picture was drawn by TBtools and Microsoft Office 2019. The data were processed by IBM SPSS Statistics 26. Full name and function of the genes are shown in Table 2.
Table 2. Selection of genes related to carbohydrate metabolism, hormone metabolism and dormancy regulation.
| Abbreviations | Full Names | Gene | Physiological Metabolic Pathways |
|---|---|---|---|
| SS3 | starch synthase 3 | AT1G11720 | Carbohydrate transport and metabolism |
| AMY3 | α-amylase-like 3 | AT1G69830 | Carbohydrate transport and metabolism |
| BMY3 | β-amylase 3 | AT5G18670 | putative beta-amylase |
| ATCWINV1 | Arabidopsis thalianacell wall invertase 1 | AT3G13790 | Carbohydrate transport and metabolism |
| SUS3 | sucrose synthase 3 | AT4G02280 | Cell all/membrane/envelope biogenesis |
| ATSPS1F | sucrose phosphate synthase 1f | AT5G20280 | Cell all/membrane/envelope biogenesis |
| PFK3 | phosphofructokinase 3 | AT4G26270 | Carbohydrate transport and metabolism |
| APL3 | / | AT4G39210 | Carbohydrate transport and metabolism |
| UGP2 | UDP-glucose pyrophosphorylase 2 | AT5G17310 | Carbohydrate transport and metabolism |
| BGLU17 | β-glucosidase 17 | AT2G44480 | Carbohydrate transport and metabolism |
| NCED3 | 9-cis-epoxycarotenoid dioxygenase 3 | AT3G14440 | Secondary metabolites biosynthesis, transport and catabolism |
| NCED4 | 9-cis-epoxycarotenoid dioxygenase 4 | AT4G19170 | Secondary metabolites biosynthesis, transport and catabolism |
| XERICO | AT2G04240 | Involved in ABA metabolism | |
| ATPP2CA | protein phosphatase 2CA | AT3G11410 | Signal transduction mechanisms |
| PYR1 | pyrabactin resistance 1 | AT4G17870 | PYR/PYL/RCAR family proteins function as abscisic acid sensors |
| ABI5 | ABA insensitive 5 | AT2G36270 | involved in ABA signalling during seed maturation and germination |
| ABF2 | abscisic acid responsive elements-binding factor 2 | AT1G45249 | Leucine zipper transcription factor that binds to the abscisic acid (ABA)–responsive element (ABRE) motif in the promoter region of ABA-inducible genes. |
| SNRK2-5 | SNF1-related protein kinase 2.5 | AT5G63650 | encodes a member of SNF1-related protein kinases (SnRK2) whose activity is activated by ionic (salt) and non-ionic (mannitol) osmotic stress. |
| ATBZIP9 | Arabidopsis thalianabasic leucine zipper 9 | AT5G24800 | |
| CYCD3 | cyclin d3 | AT4G34160 | Cell cycle control, cell division, chromosome partitioning |
| BG3 | β-1,3-glucanase 3 | AT3G57240 | encodes a member of glycosyl hydrolase family 17 |
| CAT2 | catalase 2 | AT4G35090 | Catalase domain-containing protein/Catalase-rel domain-containing protein [Cephalotus follicularis] |
| EXPA6 | Arabidopsis thalianatexpansin 6 | AT2G28950 | Encodes an expansin. Involved in the formation of nematode-induced syncytia in roots of Arabidopsis thaliana. |
| EXLA2 | expansin-like a2 | AT4G38400 | member of EXPANSIN-LIKE. |
| CDKB22 | cyclin-dependent kinase b2;2 | AT1G20930 | Expressed in the shoot apical meristem. Involved in regulation of the G2/M transition of the mitotic cell cycle. |
| GRXS17 | Arabidopsis thalianamonothiol glutaredoxin 17 | AT4G04950 | Encodes a monothiol glutaredoxin that is a critical component involved in ROS accumulation, auxin signaling, and temperature-dependent postembryonic growth in plants. |
| HIS1-3 | histone H1-3 | AT2G18050 | encodes a structurally divergent linker histone whose gene expression is induced by dehydration and ABA. The mRNA is cell-to-cell mobile. |
| PER52 | peroxidase 52 | AT5G05340 | Encodes a protein with sequence similarity to peroxidases that is involved in lignin biosynthesis. Loss of function mutations show abnormal development of xylem fibers and reduced levels of lignin biosynthetic enxymes. |
| ARP6 | actin-related protein 6 | AT3G33520 | Encodes ACTIN-RELATED PROTEIN6 (ARP6), a putative component of a chromatin-remodeling complex. |
| GID1A | GA insensitive dwarf1a | AT3G05120 | Encodes a gibberellin (GA) receptor ortholog of the rice GA receptor gene (OsGID1). |
| PIF3 | phytochrome interacting factor 3 | AT1G09530 | Transcription factor interacting with photoreceptors phyA and phyB. |
| RGA1 | repressor of GAL-3 1 | AT2G01570 | Putative transcriptional regulator repressing the gibberellin response and integration of phytohormone signalling. |
| ATGA2OX8 | Arabidopsis thalianagibberellin 2-oxidase 8 | AT4G21200 | Encodes a protein with gibberellin 2-oxidase activity which acts specifically on C-20 gibberellins. |
| SPY | spindly | AT3G11540 | Encodes a N-acetyl glucosamine transferase that may glycosylate other molecules involved in GA signaling. |
| SPT | spatula | AT4G36930 | Encodes a transcription factor of the bHLH protein family. Mutants have abnormal, unfused carpels and reduced seed dormancy. |
| GASA4 | GAST1 protein homolog 4 | AT5G15230 | Encodes gibberellin-regulated protein GASA4. Promotes GA responses and exhibits redox activity. |
| CBF4 | C-repeat-binding factor 4 | AT5G51990 | This gene is involved in response to drought stress and abscisic acid treatment, but not to low temperature. |
| SPL9 | squamosa promoter binding protein-like 9 | AT2G42200 | Encodes a putative transcriptional regulator that is involved in the vegetative to reproductive phase transition. |
| SVP | short vegetative phase | AT2G22540 | Encodes a nuclear protein that acts as a floral repressor and that functions within the thermosensory pathway. |
| AGL20 | agamous-like 20 | AT2G45660 | Controls flowering and is required for CO to promote flowering. |
| AP1 | apetala1 | AT1G69120 | Floral homeotic gene encoding a MADS domain protein homologous to SRF transcription factors. |
| GRF2 | general regulatory factor 2, | AT1G78300 | G-box binding factor GF14 omega encoding a 14-3-3 protein The mRNA is cell-to-cell mobile. |
Note: Full names and notes of genes in the table are from “TAIR—Home Page (arabidopsis.org)”. “/” indicates the information could not be found.
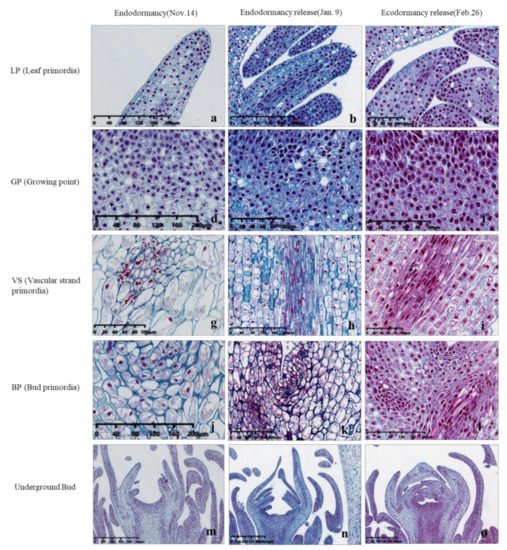
Figure 4. Cytological observation of ‘Meiju’ underground buds from endodormancy to ecodormancy. The dyeing method in the picture is as follows: Safranin O-Fast Green staining. (a–c): Leaf primordia; (d–f): Growing point; (g–i): Vascular strand primordia; (j–l): Bud primordia; m-o: Underground bud. The scale bars are in the lower left corner of the picture.
These indices may be used as potential judgment indices of CR satisfaction and the endodormancy release period to provide some new judgment indices for the general method of endodormancy release when the germination rate reaches 50%, but they need to be further studied and verified [69].
2.6. Possible Regulatory Networks for Dormancy Release of Underground Buds in Herbaceous Peony
Temperature is closely related to the induction, maintenance and release of perennial bud endodormancy [43][70]. It has been proven that there is a relationship between cold signal transduction and molecular networks related to bud endodormancy in some temperature-sensitive plants [71]. There is a great similarity between these plants and ‘Meiju’. In the underground bud of ‘Meiju’, the change in external temperature causes ‘Meiju’ to produce a large amount of MDA, causing damage to the bud of “Meiju”.ROS signal transduction responded quickly and transmitted signals to other related metabolic processes in time, among which glucose metabolism and hormone metabolism are the most important. The ABA content increased rapidly through the regulation of the NCED3 and NCED4 genes. In the later stage of endodormancy, GA2ox8 and other genes regulated gibberellin accumulation and inhibited the effect of ABA. Under the joint regulation of the DAM gene, the endodormancy of ‘Meiju’ underground buds was released (Figure 12).
A large number of antioxidant metabolism-related genes and some carbohydrate metabolism-related genes (SS3, BGLU) responded quickly (Figure 12). Which made the sugar and hormone metabolism extremely active, allowing earlier completion of the accumulation of soluble sugars and the synthesis and metabolism of endodormancy release inhibitor (ABA). Therefore, the ABA/GA ratio decreased rapidly in the earlier period (9 January), which may be an important reason for the short endodormancy period and low cooling capacity of ‘Meiju’.
References
- Faust, M.; Erez, A.; Rowland, L.; Wang, S.; Norman, H. Bud Dormancy in Perennial Fruit Trees: Physiological Basis for Dormancy Induction, Maintenance, and Release. HortScience 1997, 32, 623–629.
- Chang, C.Y.; Bräutigam, K.; Hüner, N.P.A.; Ensminger, I. Champions of winter survival: Cold acclimation and molecular regulation of cold hardiness in evergreen conifers. New Phytol. 2021, 229, 675–691.
- Lang, G.A.; Early, J.D.; Martin, G.C.; Darnell, R.L. Endodormancy, paradormancy, and ecodormancy-physiological terminology and classification for dormancy research. Hortscience 1987, 22, 371–377.
- Li, J.; Xu, Y.; Niu, Q.; He, L.; Teng, Y.; Bai, S. Abscisic Acid (ABA ) Promotes the Induction and Maintenance of Pear (Pyrus pyrifolia White Pear Group) Flower Bud Endodormancy. Int. J. Mol. Sci. 2018, 19, 310.
- Yang, Q.; Gao, Y.; Wu, X.; Moriguchi, T.; Bai, S.; Teng, Y. Bud endodormancy in deciduous fruit trees: Advances and prospects. Hortic. Res. 2021, 8, 139.
- Yang, Q.; Yang, B.; Li, J.; Wang, Y.; Tao, R.; Yang, F.; Wu, X.; Yan, X.; Ahmad, M.; Shen, J.; et al. ABA-responsive ABRE-BINDING FACTOR3 activates DAM3 expression to promote bud dormancy in Asian pear. Plant, Cell Environ. 2020, 43, 1360–1375.
- Jian, L.-C.; Li, P.H.; Sun, L.-H.; Chen, T.H. Alterations in ultrastructure and subcellular localization of Ca2+in poplar apical bud cells during the induction of dormancy. J. Exp. Bot. 1997, 48, 1195–1207.
- Heide, O.M.; Prestrud, A.K. Low temperature, but not photoperiod, controls growth cessation and dormancy induction and release in apple and pear. Tree Physiol. 2005, 25, 109–114.
- Ghelardini, L.; Santini, A.; Black-Samuelsson, S.; Myking, T.; Falusi, M. Bud dormancy release in elm (Ulmus spp.) clones—A case study of photoperiod and temperature responses. Tree Physiol. 2010, 30, 264–274.
- Campoy, J.; Ruiz, D.; Egea, J. Dormancy in temperate fruit trees in a global warming context: A review. Sci. Hortic. 2011, 130, 357–372.
- Perez, F.J.; Kuhn, N.; Vergara, R. Expression analysis of phytochromes A, B and floral integrator genes during the entry and exit of grapevine-buds from endodormancy. J. Plant Physiol. 2011, 168, 1659–1666.
- Horvath, D.P.; Anderson, J.; Chao, W.S.; Foley, M.E. Knowing when to grow: Signals regulating bud dormancy. Trends Plant Sci. 2003, 8, 534–540.
- E Hedley, P.; Russell, J.R.; Jorgensen, L.; Gordon, S.; A Morris, J.; A Hackett, C.; Cardle, L.; Brennan, R. Candidate genes associated with bud dormancy release in blackcurrant (Ribes nigrum L.). BMC Plant Biol. 2010, 10, 202.
- Sasaki, R.; Yamane, H.; Ooka, T.; Jotatsu, H.; Kitamura, Y.; Akagi, T.; Tao, R. Functional and Expressional Analyses of PmDAM Genes Associated with Endodormancy in Japanese Apricot. Plant Physiol. 2011, 157, 485–497.
- Arora, R.; Rowland, L.J.; Tanino, K. Induction and Release of Bud Dormancy in Woody Perennials: A Science Comes of Age. HortScience 2003, 38, 911–921.
- Horvath, D.P. Role of mature leaves in inhibition of root bud growth inEuphorbia esulaL. Weed Sci. 1999, 47, 544–550.
- Horvath, D.; Chao, W.S.; Anderson, J. Molecular Analysis of Signals Controlling Dormancy and Growth in Underground Adventitious Buds of Leafy Spurge. Plant Physiol. 2002, 128, 1439–1446.
- Destefano-Beltrán, L.; Knauber, D.; Huckle, L.; Suttle, J.C. Effects of postharvest storage and dormancy status on ABA content, metabolism, and expression of genes involved in ABA biosynthesis and metabolism in potato tuber tissues. Plant Mol. Biol. 2006, 61, 687–697.
- Zheng, C.; Halaly, T.; Acheampong, A.K.; Takebayashi, Y.; Jikumaru, Y.; Kamiya, Y.; Or, E. Abscisic acid (ABA) regulates grape bud dormancy, and dormancy release stimuli may act through modification of ABA metabolism. J. Exp. Bot. 2015, 66, 1527–1542.
- Tuan, P.A.; Bai, S.; Saito, T.; Ito, A.; Moriguchi, T. Dormancy-Associated MADS-Box (DAM) and the Abscisic Acid Pathway Regulate Pear Endodormancy Through a Feedback Mechanism. Plant Cell Physiol. 2017, 58, 1378–1390.
- Rinne, P.L.; Welling, A.; Vahala, J.; Ripel, L.; Ruonala, R.; Kangasjärvi, J.; van der Schoot, C. Chilling of Dormant Buds Hyperinduces FLOWERING LOCUS T and Recruits GA-Inducible 1,3-β-Glucanases to Reopen Signal Conduits and Release Dormancy in Populus. Plant Cell 2011, 23, 130–146.
- Hussain, S.; Liu, G.; Liu, D.; Ahmed, M.; Hussain, N.; Teng, Y. Study on the expression of dehydrin genes and activities of antioxidativeenzymes in floral buds of two sand pear (Pyrus pyrifolia Nakai) cultivarsrequiring different chilling hours for bud break. Turk. J. Agric. For. 2015, 39, 930–939.
- Niu, Q.; Li, J.; Cai, D.; Qian, M.; Jia, H.; Bai, S.; Hussain, S.; Liu, G.; Teng, Y.; Zheng, X. Dormancy-associated MADS-box genes and microRNAs jointly control dormancy transition in pear (Pyrus pyrifoliawhite pear group) flower bud. J. Exp. Bot. 2016, 67, 239–257.
- Bielenberg, D.G.; Wang, Y.; Li, Z.; Zhebentyayeva, T.; Fan, S.; Reighard, G.L.; Scorza, R.; Abbott, A.G. Sequencing and annotation of the evergrowing locus in peach [Prunus persica (L.) Batsch] reveals a cluster of six MADS-box transcription factors as candidate genes for regulation of terminal bud formation. Tree Genet. Genomes 2008, 4, 495–507.
- Ubi, B.; Sakamoto, D.; Ban, Y.; Shimada, T.; Ito, A.; Nakajima, I.; Takemura, Y.; Tamura, F.; Saito, T.; Moriguchi, T. Molecular Cloning of Dormancy-associated MADS-box Gene Homologs and Their Characterization during Seasonal Endodormancy Transitional Phases of Japanese Pear. J. Am. Soc. Hortic. Sci. 2010, 135, 174–182.
- Yamane, H.; Kashiwa, Y.; Ooka, T.; Tao, R.; Yonemori, K. Suppression Subtractive Hybridization and Differential Screening Reveals Endodormancy-associated Expression of an SVP/AGL24-type MADS-box Gene in Lateral Vegetative Buds of Japanese Apricot. J. Am. Soc. Hortic. Sci. 2008, 133, 708–716.
- Wu, R.-M.; Walton, E.F.; Richardson, A.; Wood, M.; Hellens, R.; Varkonyi-Gasic, E. Conservation and divergence of four kiwifruit SVP-like MADS-box genes suggest distinct roles in kiwifruit bud dormancy and flowering. J. Exp. Bot. 2011, 63, 797–807.
- Liu, B.; Zhao, S.; Tan, F.; Zhao, H.; Wang, D.; Si, H.; Chen, Q. Changes in ROS production and antioxidant capacity during tuber sprouting in potato. Food Chem. 2017, 237, 205–213.
- Zhang, J.; Zhang, X.; Wang, C.; Gai, S.; Mu, P. Correlation Analysis Between C/N and the Amount of Chilling Treatment During the Release of Dormant Floral Buds in Paeonia suffruticosa. Chin. Agric. Sci. Bull. 2012, 28, 231–235.
- Liu, B.; Zheng, G.-S.; Yan, Z.-P.; Wang, Z.-Z. Effect of different chilling treatments on forcing culture of tree peony in winter and changes of nutrient substance. Xibei Zhiwu Xuebao 2004, 24, 1635–1639.
- Destefano-Beltrán, L.; Knauber, D.; Huckle, L.; Suttle, J. Chemically forced dormancy termination mimics natural dormancy progression in potato tuber meristems by reducing ABA content and modifying expression of genes involved in regulating ABA synthesis and metabolism. J. Exp. Bot. 2006, 57, 2879–2886.
- Zhang, T.; Si, F.; Zhang, Y.; Gai, S. Effect of Exogenous Gibberellin on DNA Methylation Level and Expression of Related Enzyme Genes in Tree Peony Floral Buds. Sci. Agric. Sin. 2018, 51, 3561–3569.
- Xin, H.; Zhang, Y.; Wang, X.; Liu, C.; Feng, W.; Gai, S. Morphological, anatomical and DNA methylation changes of tree peony buds during chilling induced dormancy release. Plant Physiol. Biochem. 2019, 144, 64–72.
- Tong, H.; Xiao, Y.; Liu, D.; Gao, S.; Liu, L.; Yin, Y.; Jin, Y.; Qian, Q.; Chu, C. Brassinosteroid Regulates Cell Elongation by Modulating Gibberellin Metabolism in Rice. Plant Cell 2014, 26, 4376–4393.
- Zhao, D.; Tang, Y.; Xia, X.; Sun, J.; Meng, J.; Shang, J.; Tao, J. Integration of Transcriptome, Proteome, and Metabolome Provides Insights into How Calcium Enhances the Mechanical Strength of Herbaceous Peony Inflorescence Stems. Cells 2019, 8, 102.
- Hall, A.; Catley, J.; Walton, E. The effect of forcing temperature on peony shoot and flower development. Sci. Hortic. 2007, 113, 188–195.
- Wang, X.; Li, D.; Zhang, D.; Shi, X.; Wu, Y.; Qi, Z.; Ding, H.; Zhu, K.; Xia, Y.; Zhang, J. Improving crucial details and selecting the optimal model for evaluating the chilling requirement of Paeonia lactiflora Pall. at low latitudes during four winters. Sci. Hortic. 2020, 265, 109175.
- Yuxi, Z.; Dan, Y.; Chunying, L.; Shupeng, G. Dynamic of carbohydrate metabolism and the related genes highlights PPP pathway activation during chilling induced bud dormancy release in tree peony (Paeonia suffruticosa). Sci. Hortic. 2018, 242, 36–43.
- Luedeling, E. Climate change impacts on winter chill for temperate fruit and nut production: A review. Sci. Hortic. 2012, 144, 218–229.
- Zhang, J.; Li, D.; Shi, X.; Zhang, D.; Qiu, S.; Wei, J.; Zhang, J.; Zhou, J.; Zhu, K.; Xia, Y. Mining and expression analysis of candidate genes involved in regulating the chilling requirement fulfillment of Paeonia lactiflora ‘Hang Baishao’. BMC Plant Biol. 2017, 17, 262.
- Hao, X.; Tang, H.; Wang, B.; Yue, C.; Wang, L.; Zeng, J.; Yang, Y.; Wang, X. Integrative transcriptional and metabolic analyses provide insights into cold spell response mechanisms in young shoots of the tea plant. Tree Physiol. 2018, 38, 1655–1671.
- Zhang, J.; Wang, X.; Zhang, N.; Qiu, S.; Wei, J.; Guo, J.; Li, D.; Xia, Y. Evaluating the Comprehensive Performance of Herbaceous Peonies at low latitudes by the Integration of Long-running Quantitative Observation and Multi-Criteria Decision Making Approach. Sci. Rep. 2019, 9, 15079.
- Anderson, J.V.; Horvath, D.P.; Chao, W.S.; Foley, M.E. Bud Dormancy in Perennial Plants: A Mechanism for Survival. In Dormancy and Resistance in Harsh Environments; Lubzens, E., Cerda, J., Clark, M., Eds.; Topics in Current Genetics: Weed Biology Research Unit; USDA-Agricultural Research Service: Fargo, ND, USA, 2010; Volume 21, pp. 69–90.
- Philip, M.M.; FangYun, C.; HongYan, L. Chronological changes in plant hormone and sugar contents in cv. Ao-Shuang autumn flowering tree peony. Hortic. Sci. 2011, 38, 104–112.
- Guy, C.L.; Huber, J.L.A.; Huber, S.C. Sucrose Phosphate Synthase and Sucrose Accumulation at Low Temperature. Plant Physiol. 1992, 100, 502–508.
- Sasaki, H.; Ichimura, K.; Oda, M. Changes in Sugar Content during Cold Acclimation and Deacclimation of Cabbage Seedlings. Ann. Bot. 1996, 78, 365–369.
- Cooke, J.E.K.; Eriksson, M.E.; Junttila, O. The dynamic nature of bud dormancy in trees: Environmental control and molecular mechanisms. Plant, Cell Environ. 2012, 35, 1707–1728.
- Sonnewald, S.; Sonnewald, U. Regulation of potato tuber sprouting. Planta 2013, 239, 27–38.
- Lee, S.; Cheng, H.; King, K.E.; Wang, W.; He, Y.; Hussain, A.; Lo, J.; Harberd, N.P.; Peng, J. Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev. 2002, 16, 646–658.
- Bouquin, T.; Meier, C.; Foster, R.; Nielsen, M.E.; Mundy, J. Control of Specific Gene Expression by Gibberellin and Brassinosteroid. Plant Physiol. 2001, 127, 450–458.
- Martinez-Andujar, C.; Ordiz, M.I.; Huang, Z.; Nonogaki, M.; Beachy, R.N. Induction of 9-cis-epoxycarotenoid dioxygenase in Arabidopsis thaliana seeds enhances seed dormancy. Proc. Natl. Acad. Sci. USA 2011, 108, 17225–17229.
- Gao, S.; Gao, J.; Zhu, X.; Song, Y.; Li, Z.; Ren, G.; Zhou, X.; Kuai, B. ABF2, ABF3, and ABF4 Promote ABA-Mediated Chlorophyll Degradation and Leaf Senescence by Transcriptional Activation of Chlorophyll Catabolic Genes and Senescence-Associated Genes in Arabidopsis. Mol. Plant 2016, 9, 1272–1285.
- Lee, S.-J.; Kang, J.-Y.; Park, H.-J.; Kim, M.D.; Bae, M.S.; Choi, H.-I.; Kim, S.Y. DREB2C Interacts with ABF2, a bZIP Protein Regulating Abscisic Acid-Responsive Gene Expression, and Its Overexpression Affects Abscisic Acid Sensitivity. Plant Physiol. 2010, 153, 716–727.
- Wu, J.; Jin, Y.; Liu, C.; Vonapartis, E.; Liang, J.; Wu, W.; Gazzarrini, S.; He, J.; Yi, M. GhNAC83 inhibits corm dormancy release by regulating ABA signaling and cytokinin biosynthesis in Gladiolus hybridus. J. Exp. Bot. 2019, 70, 1221–1237.
- Cutler, A.J.; E Krochko, J. Formation and breakdown of ABA. Trends Plant Sci. 1999, 4, 472–478.
- Lefebvre, V.; North, H.; Frey, A.; Sotta, B.; Seo, M.; Okamoto, M.; Nambara, E.; Marion-Poll, A. Functional analysis of ArabidopsisNCED6andNCED9genes indicates that ABA synthesized in the endosperm is involved in the induction of seed dormancy. Plant J. 2006, 45, 309–319.
- Shu, K.; Zhou, W.; Yang, W. APETALA 2-domain-containing transcription factors: Focusing on abscisic acid and gibberellins antagonism. New Phytol. 2018, 217, 977–983.
- An, F.; Zhang, X.; Zhu, Z.; Ji, Y.; He, W.; Jiang, Z.; Li, M.; Guo, H. Coordinated regulation of apical hook development by gibberellins and ethylene in etiolated Arabidopsis seedlings. Cell Res. 2012, 22, 915–927.
- Zawaski, C.; Kadmiel, M.; Pickens, J.; Ma, C.; Strauss, S.; Busov, V. Repression of gibberellin biosynthesis or signaling produces striking alterations in poplar growth, morphology, and flowering. Planta 2011, 234, 1285–1298.
- Frey, A.; Effroy, D.; Lefebvre, V.; Seo, M.; Perreau, F.; Berger, A.; Sechet, J.; To, A.; North, H.M.; Marion-Poll, A. Epoxycarotenoid cleavage by NCED5 fine-tunes ABA accumulation and affects seed dormancy and drought tolerance with other NCED family members. Plant J. 2012, 70, 501–512.
- Rodríguez, M.V.; Mendiondo, G.M.; Maskin, L.; Gudesblat, G.E.; Iusem, N.D.; Benech-Arnold, R.L. Expression of ABA signalling genes and ABI5 protein levels in imbibed Sorghum bicolor caryopses with contrasting dormancy and at different developmental stages. Ann. Bot. 2009, 104, 975–985.
- Née, G.; Kramer, K.; Nakabayashi, K.; Yuan, B.; Xiang, Y.; Miatton, E.; Finkemeier, I.; Soppe, W.J.J. DELAY OF GERMINATION1 requires PP2C phosphatases of the ABA signalling pathway to control seed dormancy. Nat. Commun. 2017, 8, 72.
- Steber, C.M.; McCourt, P. A Role for Brassinosteroids in Germination in Arabidopsis. Plant Physiol. 2001, 125, 763–769.
- Liu, X.; Zhang, H.; Zhao, Y.; Feng, Z.; Li, Q.; Yang, H.-Q.; Luan, S.; Li, J.; He, Z.-H. Auxin controls seed dormancy through stimulation of abscisic acid signaling by inducing ARF-mediated ABI3 activation in Arabidopsis. Proc. Natl. Acad. Sci. USA 2013, 110, 15485–15490.
- Xu, Q.; Truong, T.T.; Barrero, J.M.; Jacobsen, J.V.; Hocart, C.H.; Gubler, F. A role for jasmonates in the release of dormancy by cold stratification in wheat. J. Exp. Bot. 2016, 67, 3497–3508.
- Suttle, J.C.; Banowetz, G.M. Changes in cis -zeatin and cis -zeatin riboside levels and biological activity during potato tuber dormancy. Physiol. Plant. 2000, 109, 68–74.
- Noriega, X.; Burgos, B.; Pérez, F.J. Short day-photoperiod triggers and low temperatures increase expression of peroxidase RNA transcripts and basic peroxidase isoenzyme activity in grapevine buds. Phytochemistry 2007, 68, 1376–1383.
- Mazzitelli, L.; Hancock, R.D.; Haupt, S.; Walker, P.G.; Pont, S.D.A.; McNicol, J.; Cardle, L.; Morris, J.; Viola, R.; Brennan, R.; et al. Co-ordinated gene expression during phases of dormancy release in raspberry (Rubus idaeus L.) buds. J. Exp. Bot. 2007, 58, 1035–1045.
- Liang, W.; Xu, C.; Wang, Z.; Zhang, C.; Lin, M. Physiological changes of three jujube cultivars with different cooling requirements during dormancy. J. Arid. Land Resour. Environ. 2020, 34, 187–193.
- Atkinson, C.; Brennan, R.; Jones, H. Declining chilling and its impact on temperate perennial crops. Environ. Exp. Bot. 2013, 91, 48–62.
- Welling, A.; Palva, E.T. Molecular control of cold acclimation in trees. Physiol. Plant. 2006, 127, 167–181.
More
Information
Subjects:
Genetics & Heredity
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
685
Revisions:
4 times
(View History)
Update Date:
31 Aug 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




