
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Alexandra Peregrina | + 4219 word(s) | 4219 | 2021-08-25 08:10:42 | | | |
| 2 | Lindsay Dong | -122 word(s) | 4097 | 2021-08-30 04:25:45 | | |
Video Upload Options
The large production of non-degradable petrol-based plastics has become a major global issue due to its environmental pollution. Biopolymers produced by microorganisms such as polyhydroxyalkanoates (PHAs) are gaining potential as a sustainable alternative, but the high cost associated to their industrial production has been a limiting factor. Post-transcriptional regulation is a key step to control gene expression in changing environments and has been reported to play a major role in numerous cellular processes. However, limited reports are available concerning the regulation of PHA accumulation in bacteria, and many essential regulatory factors still need to be identified. Here, we review studies where the synthesis of PHA has been reported to be regulated at the post-transcriptional level, and we analyze the RNA-mediated networks involved. Finally, we discuss the forthcoming research on riboregulation, synthetic and metabolic engineering which could lead to improved strategies for PHAs synthesis in industrial production, thereby reducing the costs currently associated with this procedure.
1. Introduction
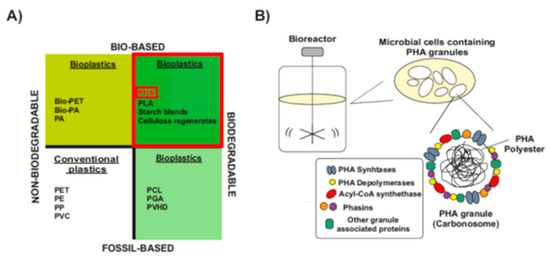
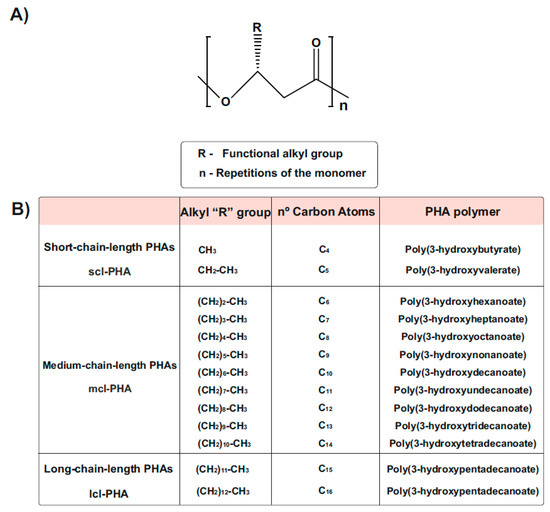
Types and Chemical Structure of PHAs Polymers
Natural PHA Producers and Engineering of Non-PHA Producers
PHA Composition and Preferred Carbon Source
2. RNA-Mediated Control in Native Synthesis of PHAs
2.1. The Expanding RNA World: Non-Coding Bacterial RNome
For years, it was considered that the expression of the bacterial genome resulted in three large groups of RNA molecules: mRNA, which contains open reading frames that translate into proteins, and two more types of RNA, the ribosomal rRNA and transfer tRNA, which are essential for protein biosynthesis carried out by ribosomes. Therefore, regulation of gene expression was exclusively associated with the activity of protein regulators [36][37]. However, post-genomic research is revealing an unprecedented high abundance and diversity of untranslated small RNA molecules (50–350 nt of average length) called sRNAs or non-coding RNAs, expanding the total of RNA species that together constitute the bacterial “RNome” [38][39]. These sRNAs are commonly encoded by single transcriptional units between open reading frames (ORFs) and although do not translate to protein, play very important roles in the gene regulation of diverse physiological processes at the post-transcriptional level [40][38][41].
These riboregulators are deeply conserved in prokaryotes and adjust gene expression in response to specific environmental or physiological signals, facilitating adaptation to diverse environmental stimuli. This is especially important to allow the cell to profit from transiently available nutrients [38][42].
Depending on their genomic location relative to the mRNA targets that they regulate, sRNAs are classified as cis- or trans-encoded. The latter constitute a majority group and are expressed from intergenic regions (IGRs), generally far from the target messenger counterparts [43][44][45].
2.1.1. RNA-Binding Proteins and Regulatory Networks
Some regulatory RBPs can act as chaperones by facilitating the intermolecular base pairing between sRNAs and mRNAs [46][47]. One of the best characterized RBP is Hfq that also exerts a central role in post-transcriptional gene regulation, as evidenced by the pleiotropic effect of the inactivation of the hfq gene in many Gram-negative bacteria [48][49][50][51].
As most sRNAs have typical bacterial Rho-independent terminators that usually contain a poly-U 3′-terminus, Hfq can interact with the terminators and influence sRNA stability [52]. The distal face has strong affinity for A-rich sequences of mRNAs [53][54].
Hfq also offers a scaffold for the interaction with several other proteins [51], e.g., Crc, which is involved in catabolite repression control in some Pseudomonas sp. [46][51][55][56].
2.1.2. Post-Transcriptional Regulation by Ribonucleases
2.2. Post-Transcriptional Regulation of sRNAs and Their Implications for Microbial PHAs Synthesis in Different Microorganisms
2.2.1. MmgR sRNA Is a Negative Regulator of PHB Accumulation in Sinorhizobium meliloti
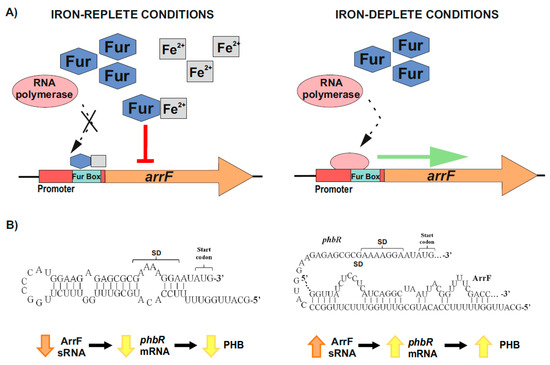
2.2.2. Post-Transcriptional Control of PhbR as Key Step during PHB Production in Azotobacter vinelandii
The main regulatory mechanism leading to the accumulation of PHB in this organism involves the phbBAC operon, which encodes for key enzymes of the PHB biosynthesis pathway. This operon is in turn controlled by the transcriptional activator PhbR and the sigma factor RpoS [76][77]. Interestingly, PhbR expression has been reported to be post-transcriptionally controlled by the two-component GacS–GacA global regulator [78]. This system (global antibiotic and cyanide control) belongs to the Gac–Rsm cascade and is involved in the regulation of many cellular processes in numerous bacterial organisms, as reviewed by Lapouge et al. [79].
2.2.3. Global Post-Transcriptional Regulatory Protein Crc as Main Target of sRNAs CrcZ and CrcY in Pseudomonas putida
The catabolite repression control protein (Crc) plays a key role in the CCR process, impeding the expression of genes involved in the synthesis of catabolic enzymes for the use of non-preferred carbon sources in pseudomonads [89][90][91][92]. This global regulator recognizes AANAANAA sequences in the genome called catabolite activity (CA) motif, located near the Shine–Dalgarno sequence of target mRNAs, and with a function in translation inhibition [93][55][89][94][92][95]. This process is in cooperation with the distal face of the protein Hfq, which is required by Crc to bind the mRNA motif through the formation of a stable ribonucleoprotein complex at the targets [96][97][95][98][99]. In Figure 9, Moreno et al. [89] summarizes the procedure by which the action of this complex is modulated and antagonized by two small RNAs (CrcZ and CrcY) in P. putida. The levels of both sRNAs significantly increase when bacteria grow with a non-preferred carbon source or have reached stationary growth phase. These sRNAs sequester one or both of the Crc/Hfq proteins, therefore decreasing the CCR, and allowing translation of the target mRNAs with A-rich motifs involved in the transport and/or assimilation of compounds [100][89][101][96][97][95][98][99]. During the formation of this multilayered and complex Hfq/Crc/CrcZ-CrcY regulatory system, the Crc–Hfq complex protects the sRNAs from ribonucleases by increasing their stability [48][97][102]. These sRNAs are mainly transcribed from σ54/RpoN-dependent promoters (PcrcZ and PcrcY) regulated by the two-component sensor-regulator system CbrA–CbrB (mainly CrcZ) together with other protein factors [93][100][89][101][96][97][92]. In this regulatory complex, each component affects either the transcription or the stability of the other components, e.g., the activity of the sRNA promoters relies on the type of carbon source and carbon/nitrogen (C/N) ratio. In turn, this promotes that the cellular metabolism adopts distinct pathways that allow the cell to adapt its requirements for energy and molecular biosynthesis [93][89][101][97][92].
2.2.4. Post-Transcriptional Control of phaC1 Synthase as a Key Aspect along PHA Synthesis in P. putida CA-3
3. Conclusions and Perspectives
3.1. Role of Post-Transcriptional Regulation during the Native Synthesis of PHAs
3.2. Controlling PHAs Production in Bacteria via Synthetic Small Non-Coding RNAs
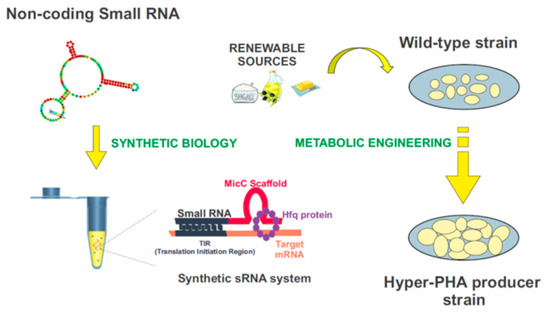
References
- Rujnic-Sokele, M.; Pilipovic, A. Challenges and opportunities of biodegradable plastics: A mini review. Waste Manag. Res. 2017, 35, 132–140.
- European Bioplastics Conference, Berlin. Available online: https://www.european-bioplastics.org/903/ (accessed on 29 November 2016).
- Razza, F.; Innocentii, F. Bioplastics from renewable resources: The benefits of biodegradability. Asia Pac. J. Chem. Eng. 2012, 7, S301–S309.
- Iwata, T. Biodegradable and Bio-Based Polymers: Future Prospects of Eco-Friendly Plastics. Angew. Chem. 2015, 54.
- Rehm, B.H. Polyester synthases: Natural catalysts for plastics. Biochem. J. 2003, 376, 15–33.
- Możejko, J.; Ciesielski, S. Pulsed Feeding Strategy Is More Favorable to Medium-Chain-Length Polyhydroxyalkanoates Production from Waste Rapeseed Oil. Biotechnol. Prog. 2014, 30, 1243–1246.
- Bresan, S.; Sznajder, A.; Hauf, W.; Forchhammer, K.; Pfeiffer, D.; Jendrossek, D. Polyhydroxyalkanoate (PHA) Granules Have No Phospholipids. Sci. Rep. 2016, 6, 26612.
- de Eugenio, L.I.; Escapa, I.F.; Morales, V.; Dinjaski, N.; Galán, B.; García, J.L.; Prieto, M.A. The Turnover of Medium-Chain-Length Polyhydroxyalkanoates in Pseudomonas putida KT2442 and the Fundamental Role of PhaZ Depolymerase for the Metabolic Balance. Environ. Microbiol. 2010, 12, 207–221.
- Luengo, J.M.; Garcia, B.; Sandoval, A.; Naharro, G.; Olivera, E.R. Bioplastics from microorganisms. Curr. Opin. Microbiol. 2003, 6, 251–260.
- Chen, G.Q. A microbial polyhydroxyalkanoates (PHA) based bio- and materials industry. Chem. Soc. Rev. 2009, 38, 2434–2446.
- Lukasiewicz, B.; Basnett, P.; Nigmatullin, R.; Matharu, R.; Knowles, J.C.; Roy, I. Binary Polyhydroxyalkanoate Systems for Soft Tissue Engineering. Acta Biomater. 2018, 71, 225–234.
- Elbahloul, Y.; Steinbüchel, A. Large-scale Production of poly(3-hydroxyoctanoic Acid) by Pseudomonas putida GPo1 and a Simplified Downstream Process. Appl. Environ. Microbiol. 2009, 75, 643–651.
- Martínez, V.; García, P.; García, J.L.; Prieto, M.A. Controlled Autolysis Facilitates the Polyhydroxyalkanoate Recovery in Pseudomonas putida KT2440. Microb. Biotechnol. 2011, 4, 533–547.
- Rebocho, A.; Pereira, J.; Freitas, F.; Neves, L.; Alves, V.; Sevrin, C.; Grandfils, C.; Reis, M. Production of Medium-Chain Length Polyhydroxyalkanoates by Pseudomonas citronellolis Grown in Apple Pulp Waste. Appl. Food Biotechnol. 2019, 6, 71–82.
- Cruz, M.V.; Freitas, F.; Paiva, A.; Mano, F.; Dionísio, M.; Ramos, A.M.; Reis, M.A. Valorization of fatty acids-containing wastes and byproducts into short- and medium-chain length polyhydroxyalkanoates. New Biotechnol. 2016, 33, 206–215.
- Koller, M. Biodegradable and Biocompatible Polyhydroxy-alkanoates (PHA): Auspicious Microbial Macromolecules for Pharmaceutical and Therapeutic Applications. Molecules 2018, 23, 362.
- Khanna, S.; Srivastava, A.K. Recent advances in microbial polyhydroxyalkanoates. Process Biochem. 2005, 40, 607–619.
- Tan, G.-Y.A.; Chen, C.-L.; Liya, L.L.G.; Wang, L.; Razaad, I.M.N.; Li, Y.; Zhao, L.; Mo, Y.; Wang, J.-Y. Start a Research on Biopolymer Polyhydroxyalkanoate (PHA): A Review. Polymers 2014, 6, 706–754.
- Domínguez-Díaz, M.; Romo-Uribe, A. Viscoelastic behavior of biodegradable polyhydroxyalkanoates. Bioinspired Biomim. Nanobiomaterials 2012, 1, 214–220.
- Mozejko-Ciesielska, J.; Mostek, A. Time-Course Proteomic Analysis of Pseudomonas putida KT2440 during Mcl-Polyhydroxyalkanoate Synthesis under Nitrogen Deficiency. Polymers 2019, 11, 748.
- Kourmentza, C.; Plácido, J.; Venetsaneas, N.; Burniol-Figols, A.; Varrone, C.; Gavala, H.N.; Reis, M.A.M. Recent Advances and Challenges Towards Sustainable Polyhydroxyalkanoate (PHA) Production. Bioengineering 2017, 4, 55.
- Madison, L.L.; Huisman, G.W. Metabolic engineering of poly(3-hydroxyalkanoates): From DNA to plastic. Microbiol. Mol. Biol. Rev. 1999, 63, 21–53.
- Jendrossek, D.; Handrick, R. Microbial Degradation of Polyhydroxyalkanoates. Annu. Rev. Microbiol. 2002, 56, 403–432.
- Cruz, M.V.; Gouveia, A.R.; Dionísio, M.; Freitas, F.; Reis, M.A.M. A Process Engineering Approach to Improve Production of P(3HB) by Cupriavidus necator from Used Cooking Oil. Int. J. Polym. Sci. 2019, 2019, 191650.
- Castro-Mayorga, J.L.; Freitas, F.; Reis, M.A.M.; Prieto, M.A.; Lagaron, J.M. Biosynthesis of Silver Nanoparticles and Polyhydroxybutyrate Nanocomposites of Interest in Antimicrobial Applications. Int. J. Biol. Macromol. 2018, 108, 426–435.
- Escapa, I.F.; del Cerro, C.; García, J.L.; Prieto, M.A. The Role of GlpR Repressor in Pseudomonas putida KT2440 Growth and PHA Production from Glycerol. Environ. Microbiol. 2013, 15, 93–110.
- Escapa, I.; Morales, V.; Martino, V.; Pollet, E.; Avérous, L.; García, J.; Prieto, M. Disruption of β-oxidation pathway in Pseudomonas putida KT2442 to produce new functionalized PHAs with thioester groups. Appl. Microbiol. Biotechnol. 2011, 89, 1583–1598.
- Khosravi-Darani, K.; Mokhtari, Z.B.; Amai, T.; Tanaka, K. Microbial Production of Poly(hydroxybutyrate) From C₁ Carbon Sources. Appl. Microbiol. Biotechnol. 2013, 97, 1407–1424.
- Anderson, A.J.; Dawes, E.A. Occurrence, Metabolism, Metabolic Role, and Industrial Uses of Bacterial Polyhydroxyalkanoates. Microbiol. Rev. 1990, 54, 450–472.
- Steinbüchel, A.; Hein, S. Biochemical and Molecular Basis of Microbial Synthesis of Polyhydroxyalkanoates in Microorganisms. Adv. Biochem. Eng./Biotechnol. 2001, 71, 81–123.
- Mezzina, M.P.; Manoli, M.T.; Prieto, M.A.; Nikel, P.I. Engineering Native and Synthetic Pathways in Pseudomonas putida for the Production of Tailored Polyhydroxyalkanoates. Biotechnol. J. 2020, 16, e2000165.
- Prieto, M.A.; de Eugenio, L.I.; Galán, B.; Luengo, J.M.; Witholt, B.J.M. Synthesis and Degradation of Polyhydroxyalkanoates In Pseudomonas: A Model System in Biology; Springerlink: Berlin, Germany, 2007; Volume V, pp. 397–428.
- Poblete-Castro, I.; Becker, J.; Dohnt, K.; dos Santos, V.M.; Wittmann, C. Industrial biotechnology of Pseudomonas putida and related species. Appl. Microbiol. Biotechnol. 2012, 93, 2279–2290.
- Borrero-de Acuña, J.; Bielecka, A.; Häussler, S.; Schobert, M.; Jahn, M.; Wittmann, C.; Jahn, D.; Poblete-Castro, I. Production of medium chain length polyhydroxyalkanoate in metabolic flux optimized Pseudomonas putida. Microb. Cell Factories 2014, 13, 1–15.
- Mozejko-Ciesielska, J.; Dabrowska, D.; Szalewska-Palasz, A.; Ciesielski, S. Medium-chain-length polyhydroxyalkanoates synthesis by Pseudomonas putida KT2440 relA/spoT mutant: Bioprocess characterization and transcriptome analysis. AMB Express 2017, 7, 92.
- Crick, F. Central dogma of molecular biology. Nature 1970, 227, 561–563.
- Temin, H.M. Reverse transcription in the eukaryotic genome: Retroviruses, pararetroviruses, retrotransposons, and retrotranscripts. Mol. Biol. Evol. 1985, 2, 455–468.
- Wagner, E.G.H.; Romby, P. Small RNAs in Bacteria and Archaea: Who They Are, What They Do, and How They Do It. Adv. Genet. 2015, 90, 133–208.
- Jimenez-Zurdo, J.I.; Valverde, C.; Becker, A. Insights into the noncoding RNome of nitrogen-fixing endosymbiotic alpha-proteobacteria. Mol. Plant. Microbe Interact. 2013, 26, 160–167.
- Eddy, S.R. Non-coding RNA genes and the modern RNA world. Nat. Rev. Genet. 2001, 2, 919–929.
- Storz, G.; Vogel, J.; Wassarman, K.M. Regulation by small RNAs in bacteria: Expanding frontiers. Mol. Cell 2011, 43, 880–891.
- Gottesman, S.; McCullen, C.A.; Guillier, M.; Vanderpool, C.K.; Majdalani, N.; Benhammou, J.; Thompson, K.M.; FitzGerald, P.C.; Sowa, N.A.; FitzGerald, D.J. Small RNA Regulators and the Bacterial Response to Stress. Cold Spring Harb. Symp. Quant. Biol. 2006, 71, 1–11.
- Gottesman, S.; Storz, G. Bacterial small RNA regulators: Versatile roles and rapidly evolving variations. Cold Spring Harb. Perspect. Biol. 2011, 3, a003798.
- Storz, G.; Altuvia, S.; Wassarman, K.M. An abundance of RNA regulators. Annu. Rev. Biochem. 2005, 74, 199–217.
- Oliva, G.; Sahr, T.; Buchrieser, C. Small RNAs, 5’ UTR elements and RNA-binding proteins in intracellular bacteria: Impact on metabolism and virulence. FEMS Microbiol. Rev. 2015, 39, 331–349.
- Van Assche, E.; Van Puyvelde, S.; Vanderleyden, J.; Steenackers, H.P. RNA-binding proteins involved in post-transcriptional regulation in bacteria. Front. Microbiol. 2015, 6, 141.
- Herschlag, D. RNA Chaperones and the RNA Folding Problem. J. Biol. Chem. 1995, 270, 20871–20874.
- Saramago, M.; Barria, C.; Dos Santos, R.F.; Silva, I.J.; Pobre, V.; Domingues, S.; Andrade, J.M.; Viegas, S.C.; Arraiano, C.M. The role of RNases in the regulation of small RNAs. Curr. Opin. Microbiol. 2014, 18, 105–115.
- Torres-Quesada, O.; Reinkensmeier, J.; Schluter, J.P.; Robledo, M.; Peregrina, A.; Giegerich, R.; Toro, N.; Becker, A.; Jimenez-Zurdo, J.I. Genome-wide profiling of Hfq-binding RNAs uncovers extensive post-transcriptional rewiring of major stress response and symbiotic regulons in Sinorhizobium meliloti. RNA Biol. 2014, 11, 563–579.
- Torres-Quesada, O.; Oruezabal, R.I.; Peregrina, A.; Jofre, E.; Lloret, J.; Rivilla, R.; Toro, N.; Jimenez-Zurdo, J.I. The Sinorhizobium meliloti RNA chaperone Hfq influences central carbon metabolism and the symbiotic interaction with alfalfa. BMC Microbiol. 2010, 10, 71.
- Sobrero, P.; Valverde, C. The bacterial protein Hfq: Much more than a mere RNA-binding factor. Crit. Rev. Microbiol. 2012, 38, 276–299.
- Vogel, J.; Luisi, B.F. Hfq and its constellation of RNA. Nat. Rev. Microbiol. 2011, 9, 578–589.
- De Lay, N.; Schu, D.J.; Gottesman, S. Bacterial Small RNA-based Negative Regulation: Hfq and Its Accomplices. J. Biol. Chem. 2013, 288, 7996–8003.
- Link, T.M.; Valentin-Hansen, P.; Brennan, R.G. Structure of Escherichia Coli Hfq Bound to Polyriboadenylate RNA. Proc. Natl. Acad. Sci. USA 2009, 106, 19292–19297.
- Moreno, R.; Ruiz-Manzano, A.; Yuste, L.; Rojo, F. The Pseudomonas Putida Crc Global Regulator Is an RNA Binding Protein That Inhibits Translation of the AlkS Transcriptional Regulator. Mol. Microbiol. 2007, 64, 665–675.
- Malecka, E.M.; Bassani, F.; Dendooven, T.; Sonnleitner, E.; Rozner, M.; Albanese, T.G.; Resch, A.; Luisi, B.; Woodson, S.; Bläsi, U. Stabilization of Hfq-mediated translational repression by the co-repressor Crc in Pseudomonas aeruginosa. Nucleic Acids Res. 2021, 49, 7075–7087.
- Viegas, S.C.; Arraiano, C.M. Regulating the Regulators: How Ribonucleases Dictate the Rules in the Control of Small Non-Coding RNAs. RNA Biol. 2008, 5, 230–243.
- Arraiano, C.M.; Maquat, L.E. Post-transcriptional control of gene expression: Effectors of mRNA decay. Mol. Microbiol. 2003, 49, 267–276.
- Ow, M.C.; Perwez, T.; Kushner, S.R. RNase G of Escherichia coli exhibits only limited functional overlap with its essential homologue, RNase E. Mol. Microbiol. 2003, 49, 607–622.
- Morita, T.; Maki, K.; Aiba, H. RNase E-based ribonucleoprotein complexes: Mechanical basis of mRNA destabilization mediated by bacterial noncoding RNAs. Genes. Dev. 2005, 19, 2176–2186.
- Santos, J.M.; Drider, D.; Marujo, P.E.; Lopez, P.; Arraiano, C.M. Determinant role of E. coli RNase III in the decay of both specific and heterologous mRNAs. FEMS Microbiol. Lett. 1997, 157, 31–38.
- Arraiano, C.M.; Andrade, J.M.; Domingues, S.; Guinote, I.B.; Malecki, M.; Matos, R.G.; Moreira, R.N.; Pobre, V.; Reis, F.P.; Saramago, M.; et al. The critical role of RNA processing and degradation in the control of gene expression. FEMS Microbiol. Rev. 2010, 34, 883–923.
- Lamontagne, B.; Elela, S.A. Evaluation of the RNA determinants for bacterial and yeast RNase III binding and cleavage. J. Biol. Chem. 2004, 279, 2231–2241.
- Saramago, M.; Peregrina, A.; Robledo, M.; Matos, R.G.; Hilker, R.; Serrania, J.; Becker, A.; Arraiano, C.M.; Jimenez-Zurdo, J.I. Sinorhizobium meliloti YbeY is an endoribonuclease with unprecedented catalytic features, acting as silencing enzyme in riboregulation. Nucleic Acids Res. 2017, 45, 1371–1391.
- Schlüter, J.P.; Reinkensmeier, J.; Barnett, M.J.; Lang, C.; Krol, E.; Giegerich, R.; Long, S.R.; Becker, A. Global mapping of transcription start sites and promoter motifs in the symbiotic alpha-proteobacterium Sinorhizobium meliloti 1021. BMC Genom. 2013, 14, 156.
- Torres-Quesada, O.; Millan, V.; Nisa-Martinez, R.; Bardou, F.; Crespi, M.; Toro, N.; Jimenez-Zurdo, J.I. Independent activity of the homologous small regulatory RNAs AbcR1 and AbcR2 in the legume symbiont Sinorhizobium meliloti. PLoS ONE 2013, 8, e68147.
- Lagares, A., Jr.; Ceizel Borella, G.; Linne, U.; Becker, A.; Valverde, C. Regulation of polyhydroxybutyrate accumulation in Sinorhizobium meliloti by the trans-encoded small RNA MmgR. J. Bacteriol. 2017, 199, e00776-16.
- Baumgardt, K.; Smidova, K.; Rahn, H.; Lochnit, G.; Robledo, M.; Evguenieva-Hackenberg, E. The stress-related, rhizobial small RNA RcsR1 destabilizes the autoinducer synthase encoding mRNA sinI in Sinorhizobium meliloti. RNA Biol. 2016, 13, 486–499.
- Robledo, M.; Peregrina, A.; Millan, V.; Garcia-Tomsig, N.I.; Torres-Quesada, O.; Mateos, P.F.; Becker, A.; Jimenez-Zurdo, J.I. A conserved alpha-proteobacterial small RNA contributes to osmoadaptation and symbiotic efficiency of rhizobia on legume roots. Environ. Microbiol. 2017.
- Sobrero, P.; Valverde, C. Evidences of autoregulation of hfq expression in Sinorhizobium meliloti strain 2011. Arch. Microbiol. 2011, 193, 629–639.
- Valverde, C.; Livny, J.; Schluter, J.P.; Reinkensmeier, J.; Becker, A.; Parisi, G. Prediction of Sinorhizobium meliloti sRNA genes and experimental detection in strain 2011. BMC Genom. 2008, 9, 416.
- Lagares, A.; Roux, I.; Valverde, C. Phylogenetic distribution and evolutionary pattern of an α-proteobacterial small RNA gene that controls polyhydroxybutyrate accumulation in Sinorhizobium meliloti. Mol. Phylogenetics Evol. 2016, 99, 182–193.
- Ceizel-Borella, G.; Lagares, A., Jr.; Valverde, C. Expression of the small regulatory RNA gene mmgR is regulated negatively by AniA and positively by NtrC in Sinorhizobium meliloti 2011. Microbiology 2018, 164, 88–98.
- Pötter, M.; Steinbüchel, A. Poly(3-hydroxybutyrate) granule-associated proteins: Impacts on poly(3-hydroxybutyrate) synthesis and degradation. Biomacromolecules 2005, 6, 552–560.
- Ceizel-Borella, G.; Lagares, A., Jr.; Valverde, C. Expression of the Sinorhizobium meliloti small RNA gene mmgR is controlled by the nitrogen source. FEMS Microbiol. Lett. 2016, 363, fnw069.
- Castañeda, M.; Guzmán, J.; Moreno, S.; Espín, G. The GacS Sensor Kinase Regulates Alginate and Poly-β-Hydroxybutyrate Production in Azotobacter vinelandii. J. Bacteriol. 2000, 182, 2624–2628.
- Hernandez-Eligio, A.; Castellanos, M.; Moreno, S.; Espín, G. Transcriptional activation of the Azotobacter vinelandii polyhydroxybutyrate biosynthetic genes phbBAC by PhbR and RpoS. Microbiology 2011, 157, 3014–3023.
- Hernandez-Eligio, A.; Moreno, S.; Castellanos, M.; Castañeda, M.; Nuñez, C.; Muriel-Millan, L.F.; Espín, G. RsmA post-transcriptionally controls PhbR expression and polyhydroxybutyrate biosynthesis in Azotobacter vinelandii. Microbiology 2012, 158, 1953–1963.
- Lapouge, K.; Schubert, M.; Allain, F.H.; Haas, D. Gac/Rsm signal transduction pathway of gamma-proteobacteria: From RNA recognition to regulation of social behaviour. Mol. Microbiol. 2008, 67, 241–253.
- Velázquez-Sánchez, C.; Espín, G.; Peña, C.; Segura, D. The Modification of Regulatory Circuits Involved in the Control of Polyhydroxyalkanoates Metabolism to Improve Their Production. Front. Bioeng. Biotechnol. 2020, 8, 386.
- Bedoya-Pérez, L.P.; Muriel-Millán, L.F.; Moreno, S.; Quiroz-Rocha, E.; Rivera-Gómez, N.; Espín, G. The pyrophosphohydrolase RppH is involved in the control of RsmA/CsrA expression in Azotobacter vinelandii and Escherichia coli. Microbiol. Res. 2018, 214, 91–100.
- Muriel-Millán, L.F.; Castellanos, M.; Hernandez-Eligio, J.A.; Moreno, S.; Espín, G. Posttranscriptional Regulation of PhbR, the Transcriptional Activator of Polyhydroxybutyrate Synthesis, by Iron and the sRNA ArrF in Azotobacter Vinelandii. Appl. Microbiol. Biotechnol. 2014, 98, 2173–2182.
- Massé, E.; Salvail, H.; Desnoyers, G.; Arguin, M. Small RNAs controlling iron metabolism. Curr. Opin. Microbiol. 2007, 10, 140–145.
- Oglesby-Sherrouse, A.G.; Murphy, E.R. Iron-responsive bacterial small RNAs: Variations on a theme. Met. Integr. Biometal Sci. 2013, 5, 276–286.
- Massé, E.; Gottesman, S. A small RNA regulates the expression of genes involved in iron metabolism in Escherichia coli. Proc. Natl. Acad. Sci. USA 2002, 99, 4620–4625.
- Jung, Y.S.; Kwon, Y.M. Small RNA ArrF regulates the expression of sodB and feSII genes in Azotobacter vinelandii. Curr. Microbiol. 2008, 57, 593–597.
- Noar, J.D.; Bruno-Bárcena, J.M. Azotobacter vinelandii: The source of 100 years of discoveries and many more to come. Microbiology 2018, 164, 421–436.
- Pyla, R.; Kim, T.J.; Silva, J.L.; Jung, Y.S. Overproduction of poly-beta-hydroxybutyrate in the Azotobacter vinelandii mutant that does not express small RNA ArrF. Appl. Microbiol. Biotechnol. 2009, 84, 717–724.
- Moreno, R.; Fonseca, P.; Rojo, F. Two Small RNAs, CrcY and CrcZ, Act in Concert to Sequester the Crc Global Regulator in Pseudomonas putida, Modulating Catabolite Repression. Mol. Microbiol. 2012, 83, 24–40.
- Ruiz-Manzano, A.; Yuste, L.; Rojo, F. Levels and activity of the Pseudomonas putida global regulatory protein Crc vary according to growth conditions. J. Bacteriol. 2005, 187, 3678–3686.
- Moreno, R.; Rojo, F. The target for the Pseudomonas putida Crc global regulator in the benzoate degradation pathway is the BenR transcriptional regulator. J. Bacteriol. 2008, 190, 1539–1545.
- García-Mauriño, S.M.; Pérez-Martínez, I.; Amador, C.I.; Canosa, I.; Santero, E. Transcriptional activation of the CrcZ and CrcY regulatory RNAs by the CbrB response regulator in Pseudomonas putida. Mol. Microbiol. 2013, 89, 189–205.
- La Rosa, R.; de la Pena, F.; Prieto, M.A.; Rojo, F. The Crc protein inhibits the production of polyhydroxyalkanoates in Pseudomonas putida under balanced carbon/nitrogen growth conditions. Environ. Microbiol. 2014, 16, 278–290.
- Moreno, R.; Martínez-Gomariz, M.; Yuste, L.; Gil, C.; Rojo, F. The Pseudomonas putida Crc global regulator controls the hierarchical assimilation of amino acids in a complete medium: Evidence from proteomic and genomic analyses. Proteomics 2009, 9, 2910–2928.
- López, N.I.; Pettinari, M.J.; Nikel, P.I.; Méndez, B.S. Polyhydroxyalkanoates: Much More than Biodegradable Plastics. Adv. Appl. Microbiol. 2015, 93, 73–106.
- La Rosa, R.; Nogales, J.; Rojo, F. The Crc/CrcZ-CrcY Global Regulatory System Helps the Integration of Gluconeogenic and Glycolytic Metabolism in Pseudomonas putida. Environ. Microbiol. 2015, 17, 3362–3378.
- Hernandez-Arranz, S.; Sanchez-Hevia, D.; Rojo, F.; Moreno, R. Effect of Crc and Hfq proteins on the transcription, processing, and stability of the Pseudomonas putida CrcZ sRNA. RNA 2016, 22, 1902–1917.
- Sonnleitner, E.; Bläsi, U. Regulation of Hfq by the RNA CrcZ in Pseudomonas aeruginosa carbon catabolite repression. PLoS Genet. 2014, 10, e1004440.
- Moreno, R.; Hernández-Arranz, S.; La Rosa, R.; Yuste, L.; Madhushani, A.; Shingler, V.; Rojo, F. The Crc and Hfq proteins of Pseudomonas putida cooperate in catabolite repression and formation of ribonucleic acid complexes with specific target motifs. Environ. Microbiol. 2015, 17, 105–118.
- Marzi, S.; Romby, P. RNA Mimicry, a Decoy for Regulatory Proteins. Mol. Microbiol. 2012, 83, 1–6.
- Sonnleitner, E.; Abdou, L.; Haas, D. Small RNA as Global Regulator of Carbon Catabolite Repression in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2009, 106, 21866–21871.
- Moll, I.; Afonyushkin, T.; Vytvytska, O.; Kaberdin, V.R.; Blasi, U. Coincident Hfq binding and RNase E cleavage sites on mRNA and small regulatory RNAs. RNA 2003, 9, 1308–1314.
- Ryan, W.J.; O’Leary, N.D.; O’Mahony, M.; Dobson, A.D. GacS-dependent Regulation of Polyhydroxyalkanoate Synthesis in Pseudomonas putida CA-3. Appl. Environ. Microbiol. 2013, 79, 1795–1802.
- O’Leary, N.D.; O’Connor, K.E.; Ward, P.; Goff, M.; Dobson, A.D.W. Genetic Characterization of Accumulation of Polyhydroxyalkanoate from Styrene in Pseudomonas putida CA-3. Appl. Environ. Microbiol. 2005, 71, 4380–4387.
- Chen, G.Q.; Jian, X.R. Engineering Bacteria for Enhanced Polyhydroxyalkanoates (PHA) Biosynthesis. Synth. Syst. Biotechnol. 2017, 2, 192–197.
- Kalia, V.C.; Lal, S.; Cheema, S. Insight in to the phylogeny of polyhydroxyalkanoate biosynthesis: Horizontal gene transfer. Gene 2007, 389, 19–26.
- Quendera, A.P.; Seixas, A.F.; Dos Santos, R.F.; Santos, I.; Silva, J.P.N.; Arraiano, C.M.; Andrade, J.M. RNA-Binding Proteins Driving the Regulatory Activity of Small Non-coding RNAs in Bacteria. Front. Mol. Biosci. 2020, 7, 78.
- Apura, P.; Saramago, M.; Peregrina, A.; Viegas, S.C.; Carvalho, S.M.; Saraiva, L.M.; Arraiano, C.M.; Domingues, S. Tailor-made sRNAs: A plasmid tool to control the expression of target mRNAs in Pseudomonas putida. Plasmid 2020, 109, 102503.
- Freemont, P.S. Synthetic biology industry: Data-driven design is creating new opportunities in biotechnology. Emerg. Top. Life Sci. 2019, 3, 651–657.
- Foley, P.L.; Shuler, M.L. Considerations for the design and construction of a synthetic platform cell for biotechnological applications. Biotechnol. Bioeng. 2010, 105, 26–36.
- Nikel, P.I.; Martínez-García, E.; de Lorenzo, V. Biotechnological domestication of pseudomonads using synthetic biology. Nat. Rev. Microbiol. 2014, 12, 368–379.
- Copeland, M.F.; Politz, M.C.; Pfleger, B.F. Application of TALEs, CRISPR/Cas and sRNAs as trans-acting regulators in prokaryotes. Curr. Opin. Biotechnol. 2014, 29, 46–54.
- Chappell, J.; Watters, K.E.; Takahashi, M.K.; Lucks, J.B. A renaissance in RNA synthetic biology: New mechanisms, applications and tools for the future. Curr. Opin. Chem. Biol. 2015, 28, 47–56.
- Isaacs, F.J.; Dwyer, D.J.; Collins, J.J. RNA synthetic biology. Nat. Biotechnol. 2006, 24, 545–554.
- Na, D.; Yoo, S.M.; Chung, H.; Park, H.; Park, J.H.; Lee, S.Y. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat. Biotechnol. 2013, 31, 170–174.
- Yoo, S.M.; Na, D.; Lee, S.Y. Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nat. Protoc. 2013, 8, 1694–1707.
- Xie, W.H.; Deng, H.K.; Hou, J.; Wang, L.J. Synthetic small regulatory RNAs in microbial metabolic engineering. Appl. Microbiol. Biotechnol. 2021, 105, 1–12.
- Martínez-García, E.; Nikel, P.I.; Aparicio, T.; de Lorenzo, V. Pseudomonas 2.0: Genetic upgrading of P. putida KT2440 as an enhanced host for heterologous gene expression. Microb. Cell Factories 2014, 13, 159.
- Martínez-García, E.; Goñi-Moreno, A.; Bartley, B.; McLaughlin, J.; Sánchez-Sampedro, L.; Del Pozo, H.P.; Hernández, C.P.; Marletta, A.S.; De Lucrezia, D.; Sánchez-Fernández, G.; et al. SEVA 3.0: An update of the Standard European Vector Architecture for enabling portability of genetic constructs among diverse bacterial hosts. Nucleic Acids Res. 2020, 48, D1164–D1170.
- Li, M.; Ma, Y.; Zhang, X.; Zhang, L.; Chen, X.; Ye, J.W.; Chen, G.Q. Tailor-Made Polyhydroxyalkanoates by Reconstructing Pseudomonas entomophila. Adv. Mater. 2021, e2102766.





