
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Alejandro Romero | + 3309 word(s) | 3309 | 2021-08-13 05:50:14 | | | |
| 2 | Vivi Li | Meta information modification | 3309 | 2021-08-24 11:59:50 | | |
Video Upload Options
The wide variety of epigenetic controls available is rapidly expanding the knowledge of molecular biology even overflowing it. At the same time, it can illuminate unsuspected ways of understanding the etiology of cancer. New emerging therapeutic horizons, then, promise to overcome the current antitumor strategies need. The translational utility of this complexity is particularly welcome in oral cancer (OC), in which natural history is alarmingly disappointing due to the invasive and mutilating surgery, the high relapsing rate, the poor quality of life and the reduced survival after diagnosis. Melatonin activates protective receptor-dependent and receptor-independent processes that prevent tissue cancerisation and inhibit progressive tumor malignancy and metastasis.
1. Introduction
2. The Role of Melatonin in the Oral Cavity: Functionality and Alterations in Oral Cancer
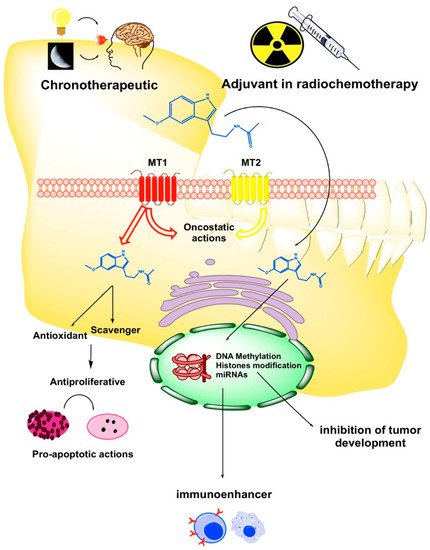
3. Epigenetic Regulation of Melatonin in Oral Cancer
3.1. Epigenetic Methylation of DNA
3.2. Epigenetic Modification of Chromatin Structure
3.3. Non-Coding micro-RNAs
| Epigenetic Control | Experimental Model | Melatonin Treatment | Main Findings | References |
|---|---|---|---|---|
| DNA methylation | OSCC cell lines | The loss by homozygous deletion or silencing by CpG hypermethylation of the MLT receptor 1A (MTNR1A) gene was associated with cancer status and tumor phenotype | [33] | |
| Histone modification | Patient-derived tumor xenografts models overexpressing LSD1. Mouse-based subcutaneous OC SCC25-xenograft model. OSCC cell lines. |
20 mg/kg daily, i.p., for 24 and 42 days. 0–20 mM for 24 h 2–4 mM for 24 and 48 h |
MLT demonstrated anti-OC activity through LSD1 down-regulation | [58] |
| HSC-3 and OECM-1 OC cell lines | 1 mM for 24 h | MLT inhibited migration of tumor cells through down-regulation of MMP-9 expression and activity by decreasing CREBBP/EP300-dependent H3 and H4 histone acetylation on MMP-9 promoter | [62] | |
| Promoter activity | HONE-1, NPC-39 and NPC-BM nasopharyngeal carcinoma cell lines. | 0.5–1 mM | MLT reduced MMP-9 promoter activity through inhibition of SP-1 transcription factor expression | [63] |
| SCC9, SCC25 and CAL27 OSCC cell lines. | 10 μg/mL for 72 h | MLT reduced miR-155 and increased miR-21. | [74] | |
| Non-coding micro-RNAs | 121 OC specimens and 66 normal counterparts for the study of miR-24 expression HCT 116 and MCF-7 cells. |
1 μM for 72 h | MLT decreased miR-24 expression, which pairs with the regulation of cell proliferation, DNA damage and oncogenic transformation genes. | [72] |
References
- Warnakulasuriya, S. Global epidemiology of oral and oropharyngeal cancer. Oral Oncol. 2009, 45, 309–316.
- Chin, D.; Boyle, G.M.; Porceddu, S.; Theile, D.R.; Parsons, P.G.; Coman, W.B. Head and neck cancer: Past, present and future. Expert Rev. Anticancer Ther. 2006, 6, 1111–1118.
- Montero, P.H.; Patel, S.G. Cancer of the oral cavity. Surg. Oncol. Clin. N. Am. 2015, 24, 491–508.
- Van der Waal, I.; de Bree, R.; Brakenhoff, R.; Coebergh, J.W. Early diagnosis in primary oral cancer: Is it possible? Med. Oral Patol. Oral Cir. Bucal 2011, 16, e300–e305.
- Cancer Genome Atlas, N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 2015, 517, 576–582.
- Razzouk, S. Translational genomics and head and neck cancer: Toward precision medicine. Clin. Genet. 2014, 86, 412–421.
- Viet, C.T.; Schmidt, B.L. Understanding oral cancer in the genome era. Head Neck 2010, 32, 1246–1268.
- Li, C.C.; Shen, Z.; Bavarian, R.; Yang, F.; Bhattacharya, A. Oral cancer: Genetics and the role of precision medicine. Dent. Clin. N. Am. 2018, 62, 29–46.
- Lo, W.L.; Kao, S.Y.; Chi, L.Y.; Wong, Y.K.; Chang, R.C. Outcomes of oral squamous cell carcinoma in Taiwan after surgical therapy: Factors affecting survival. J. Oral Maxillofac. Surg. 2003, 61, 751–758.
- Zhang, X.; Li, L.; Wei, D.; Yap, Y.; Chen, F. Moving cancer diagnostics from bench to bedside. Trends Biotechnol. 2007, 25, 166–173.
- Cutando, A.; Gomez-Moreno, G.; Arana, C.; Acuna-Castroviejo, D.; Reiter, R.J. Melatonin: Potential functions in the oral cavity. J. Periodontol. 2007, 78, 1094–1102.
- Laakso, M.L.; Porkka-Heiskanen, T.; Alila, A.; Stenberg, D.; Johansson, G. Correlation between salivary and serum melatonin: Dependence on serum melatonin levels. J. Pineal Res. 1990, 9, 39–50.
- Praninskiene, R.; Dumalakiene, I.; Kemezys, R.; Mauricas, M.; Jucaite, A. Diurnal melatonin patterns in children: Ready to apply in clinical practice? Pediatr. Neurol. 2012, 46, 70–76.
- Russo, D.; Merolla, F.; Varricchio, S.; Salzano, G.; Zarrilli, G.; Mascolo, M.; Strazzullo, V.; Di Crescenzo, R.M.; Celetti, A.; Ilardi, G. Epigenetics of oral and oropharyngeal cancers. Biomed. Rep. 2018, 9, 275–283.
- Kulkarni, V.; Uttamani, J.R.; Naqvi, A.R.; Nares, S. microRNAs: Emerging players in oral cancers and inflammatory disorders. Tumour Biol. 2017, 39, 1010428317698379.
- Shen, Y.Q.; Guerra-Librero, A.; Fernández-Gil, B.I.; Florido, J.; Garcia-Lopez, S.; Martínez-Ruiz, L.; Mendivil-Perez, M.; Soto-Mercado, V.; Acuna-Castroviejo, D.; Ortega-Arellano, H.; et al. Combination of melatonin and rapamycin for head and neck cancer therapy: Suppression of AKT/mTOR pathway activation, and activation of mitophagy and apoptosis via mitochondrial function regulation. J. Pineal Res. 2018, 64, e12461.
- Fernández-Gil, B.I.; Guerra-Librero, A.; Shen, Y.Q.; Florido, J.; Martínez-Ruiz, L.; García-López, S.; Adan, C.; Rodríguez-Santana, C.; Acuna-Castroviejo, D.; Quinones-Hinojosa, A.; et al. Melatonin enhances cisplatin and radiation cytotoxicity in head and neck squamous cell carcinoma by stimulating mitochondrial ROS generation, apoptosis, and autophagy. Oxid. Med. Cell Longev. 2019, 2019, 7187128.
- Bondy, S.C.; Campbell, A. Mechanisms underlying tumor suppressive properties of melatonin. Int. J. Mol. Sci. 2018, 19, 2205.
- Reiter, R.J.; Rosales-Corral, S.A.; Liu, X.Y.; Acuna-Castroviejo, D.; Escames, G.; Tan, D.X. Melatonin in the oral cavity: Physiological and pathological implications. J. Periodontal Res. 2015, 50, 9–17.
- Li, Y.; Li, S.; Zhou, Y.; Meng, X.; Zhang, J.J.; Xu, D.P.; Li, H.B. Melatonin for the prevention and treatment of cancer. Oncotarget 2017, 8, 39896–39921.
- Gil-Martin, E.; Egea, J.; Reiter, R.J.; Romero, A. The emergence of melatonin in oncology: Focus on colorectal cancer. Med. Res. Rev. 2019.
- Zare, H.; Shafabakhsh, R.; Reiter, R.J.; Asemi, Z. Melatonin is a potential inhibitor of ovarian cancer: Molecular aspects. J. Ovarian Res. 2019, 12, 26.
- Liu, R.; Wang, H.L.; Deng, M.J.; Wen, X.J.; Mo, Y.Y.; Chen, F.M.; Zou, C.L.; Duan, W.F.; Li, L.; Nie, X. Melatonin inhibits reactive oxygen species-driven proliferation, epithelial-mesenchymal transition, and vasculogenic mimicry in oral cancer. Oxid. Med. Cell Longev. 2018, 2018, 3510970.
- Gonçalves Ndo, N.; Rodrigues, R.V.; Jardim-Perassi, B.V.; Moschetta, M.G.; Lopes, J.R.; Colombo, J.; Zuccari, D.A. Molecular markers of angiogenesis and metastasis in lines of oral carcinoma after treatment with melatonin. Anticancer Agents Med. Chem. 2014, 14, 1302–1311.
- Cutando, A.; López-Valverde, A.; De Vicente, J.; Gimenez, J.L.; CarcÍa, I.A.; De Diego, R.G. Action of melatonin on squamous cell carcinoma and other tumors of the oral cavity (Review). Oncol. Lett. 2014, 7, 923–926.
- Lv, J.W.; Zheng, Z.Q.; Wang, Z.X.; Zhou, G.Q.; Chen, L.; Mao, Y.P.; Lin, A.H.; Reiter, R.J.; Ma, J.; Chen, Y.P.; et al. Pan-cancer genomic analyses reveal prognostic and immunogenic features of the tumor melatonergic microenvironment across 14 solid cancer types. J. Pineal Res. 2019, 66, e12557.
- Mehta, A.; Kaur, G. Potential role of melatonin in prevention and treatment of oral carcinoma. Indian J. Dent. 2014, 5, 86–91.
- D’Souza, S.; Addepalli, V. Preventive measures in oral cancer: An overview. Biomed. Pharmacother. 2018, 107, 72–80.
- Ha, P.K.; Califano, J.A. Promoter methylation and inactivation of tumour-suppressor genes in oral squamous-cell carcinoma. Lancet Oncol. 2006, 7, 77–82.
- Potaczek, D.P.; Harb, H.; Michel, S.; Alhamwe, B.A.; Renz, H.; Tost, J. Epigenetics and allergy: From basic mechanisms to clinical applications. Epigenetics 2017, 9, 539–571.
- Lee, J.K.; Kim, M.J.; Hong, S.P.; Hong, S.D. Inactivation patterns of p16/INK4A in oral squamous cell carcinomas. Exp. Mol. Med. 2004, 36, 165–171.
- Saito, K.; Uzawa, K.; Endo, Y.; Kato, Y.; Nakashima, D.; Ogawara, K.; Shiba, M.; Bukawa, H.; Yokoe, H.; Tanzawa, H. Plasma membrane Ca2+ ATPase isoform 1 down-regulated in human oral cancer. Oncol. Rep. 2006, 15, 49–55.
- Nakamura, E.; Kozaki, K.; Tsuda, H.; Suzuki, E.; Pimkhaokham, A.; Yamamoto, G.; Irie, T.; Tachikawa, T.; Amagasa, T.; Inazawa, J.; et al. Frequent silencing of a putative tumor suppressor gene melatonin receptor 1 A (MTNR1A) in oral squamous-cell carcinoma. Cancer Sci. 2008, 99, 1390–1400.
- Shaw, R. The epigenetics of oral cancer. Int. J. Oral Maxillofac. Surg. 2006, 35, 101–108.
- Smith, I.M.; Mydlarz, W.K.; Mithani, S.K.; Califano, J.A. DNA global hypomethylation in squamous cell head and neck cancer associated with smoking, alcohol consumption and stage. Int. J. Cancer 2007, 121, 1724–1728.
- Ghantous, Y.; Schussel, J.L.; Brait, M. Tobacco and alcohol-induced epigenetic changes in oral carcinoma. Curr. Opin. Oncol. 2018, 30, 152–158.
- Shaw, R.J.; Liloglou, T.; Rogers, S.N.; Brown, J.S.; Vaughan, E.D.; Lowe, D.; Field, J.K.; Risk, J.M. Promoter methylation of P16, RARbeta, E-cadherin, cyclin A1 and cytoglobin in oral cancer: Quantitative evaluation using pyrosequencing. Br. J. Cancer 2006, 94, 561–568.
- Shaw, R.J.; Hall, G.L.; Lowe, D.; Bowers, N.L.; Liloglou, T.; Field, J.K.; Woolgar, J.A.; Risk, J.M. CpG island methylation phenotype (CIMP) in oral cancer: Associated with a marked inflammatory response and less aggressive tumour biology. Oral Oncol. 2007, 43, 878–886.
- Zhong, L.; Liu, Y.; Wang, K.; He, Z.; Gong, Z.; Zhao, Z.; Yang, Y.; Gao, X.; Li, F.; Wu, H.; et al. Biomarkers: Paving stones on the road towards the personalized precision medicine for oral squamous cell carcinoma. BMC Cancer 2018, 18, 911.
- Morandi, L.; Gissi, D.; Tarsitano, A.; Asioli, S.; Gabusi, A.; Marchetti, C.; Montebugnoli, L.; Foschini, M.P. CpG location and methylation level are crucial factors for the early detection of oral squamous cell carcinoma in brushing samples using bisulfite sequencing of a 13-gene panel. Clin. Epigenet. 2017, 9, 85.
- Bai, G.; Song, J.; Yuan, Y.; Chen, Z.; Tian, Y.; Yin, X.; Niu, Y.; Liu, J. Systematic analysis of differentially methylated expressed genes and site-speci fi c methylation as potential prognostic markers in head and neck cancer. J. Cell Physiol. 2019, 234, 22687–22702.
- Zhou, C.; Ye, M.; Ni, S.; Li, Q.; Ye, D.; Li, J.; Shen, Z.; Deng, H. DNA methylation biomarkers for head and neck squamous cell carcinoma. Epigenetics 2018, 13, 398–409.
- Arayataweegool, A.; Srisuttee, R.; Mahattanasakul, P.; Tangjaturonsasme, N.; Kerekhanjanarong, V.; Kitkumthorn, N.; Mutirangura, A. Head and neck squamous cell carcinoma drives long interspersed element-1 hypomethylation in the peripheral blood mononuclear cells. Oral Dis. 2019, 25, 64–72.
- Haim, A.; Zubidat, A.E. Artificial light at night: Melatonin as a mediator between the environment and epigenome. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370, 20140121.
- Cajochen, C.; Jud, C.; Munch, M.; Kobialka, S.; Wirz-Justice, A.; Albrecht, U. Evening exposure to blue light stimulates the expression of the clock gene PER2 in humans. Eur. J. Neurosci. 2006, 23, 1082–1086.
- Rana, S.; Mahmood, S. Circadian rhythm and its role in malignancy. J. Circadian Rhythm. 2010, 8, 3.
- Hardeland, R. Melatonin and the pathologies of weakened or dysregulated circadian oscillators. J. Pineal Res. 2017, 62, e12377.
- Schwimmer, H.; Metzer, A.; Pilosof, Y.; Szyf, M.; Machnes, Z.M.; Fares, F.; Harel, O.; Haim, A. Light at night and melatonin have opposite effects on breast cancer tumors in mice assessed by growth rates and global DNA methylation. Chronobiol. Int. 2014, 31, 144–150.
- Zubidat, A.E.; Fares, B.; Fares, F.; Haim, A. Artificial light at night of different spectral compositions differentially affects tumor growth in mice: Interaction with melatonin and epigenetic pathways. Cancer Control. 2018, 25, 1073274818812908.
- Agbaria, S.; Haim, A.; Fares, F.; Zubidat, A.E. Epigenetic modification in 4T1 mouse breast cancer model by artificial light at night and melatonin—the role of DNA-methyltransferase. Chronobiol. Int. 2019, 36, 629–643.
- Smiraglia, D.J.; Smith, L.T.; Lang, J.C.; Rush, L.J.; Dai, Z.; Schuller, D.E.; Plass, C. Differential targets of CpG island hypermethylation in primary and metastatic head and neck squamous cell carcinoma (HNSCC). J. Med. Genet. 2003, 40, 25–33.
- Von Zeidler, S.V.; Miracca, E.C.; Nagai, M.A.; Birman, E.G. Hypermethylation of the p16 gene in normal oral mucosa of smokers. Int. J. Mol. Med. 2004, 14, 807–811.
- Kresty, L.A.; Mallery, S.R.; Knobloch, T.J.; Song, H.; Lloyd, M.; Casto, B.C.; Weghorst, C.M. Alterations of p16(INK4a) and p14(ARF) in patients with severe oral epithelial dysplasia. Cancer Res. 2002, 62, 5295–5300.
- Kulkarni, V.; Saranath, D. Concurrent hypermethylation of multiple regulatory genes in chewing tobacco associated oral squamous cell carcinomas and adjacent normal tissues. Oral Oncol. 2004, 40, 145–153.
- Eberharter, A.; Becker, P.B. Histone acetylation: A switch between repressive and permissive chromatin. Second in review series on chromatin dynamics. EMBO Rep. 2002, 3, 224–229.
- Webber, L.P.; Wagner, V.P.; Curra, M.; Vargas, P.A.; Meurer, L.; Carrard, V.C.; Squarize, C.H.; Castilho, R.M.; Martins, M.D. Hypoacetylation of acetyl-histone H3 (H3K9ac) as marker of poor prognosis in oral cancer. Histopathology 2017, 71, 278–286.
- Yuan, C.; Li, Z.; Qi, B.; Zhang, W.; Cheng, J.; Wang, Y. High expression of the histone demethylase LSD1 associates with cancer cell proliferation and unfavorable prognosis in tongue cancer. J. Oral Pathol. Med. 2015, 44, 159–165.
- Yang, C.Y.; Lin, C.K.; Tsao, C.H.; Hsieh, C.C.; Lin, G.J.; Ma, K.H.; Shieh, Y.S.; Sytwu, H.K.; Chen, Y.W. Melatonin exerts anti-oral cancer effect via suppressing LSD1 in patient-derived tumor xenograft models. Oncotarget 2017, 8, 33756–33769.
- Chung, Y.L.; Lee, M.Y.; Pui, N.N. Epigenetic therapy using the histone deacetylase inhibitor for increasing therapeutic gain in oral cancer: Prevention of radiation-induced oral mucositis and inhibition of chemical-induced oral carcinogenesis. Carcinogenesis 2009, 30, 1387–1397.
- Onseng, K.; Johns, N.P.; Khuayjarernpanishk, T.; Subongkot, S.; Priprem, A.; Hurst, C.; Johns, J. Beneficial effects of adjuvant melatonin in minimizing oral mucositis complications in head and neck cancer patients receiving concurrent chemoradiation. J. Altern. Complement. Med. 2017, 23, 957–963.
- Abdel Moneim, A.E.; Guerra-Librero, A.; Florido, J.; Shen, Y.Q.; Fernández-Gil, B.; Acuna-Castroviejo, D.; Escames, G. Oral mucositis: Melatonin gel an effective new treatment. Int. J. Mol. Sci. 2017, 18, 1003.
- Yeh, C.M.; Lin, C.W.; Yang, J.S.; Yang, W.E.; Su, S.C.; Yang, S.F. Melatonin inhibits TPA-induced oral cancer cell migration by suppressing matrix metalloproteinase-9 activation through the histone acetylation. Oncotarget 2016, 7, 21952–21967.
- Ho, H.Y.; Lin, C.W.; Chien, M.H.; Reiter, R.J.; Su, S.C.; Hsieh, Y.H.; Yang, S.F. Melatonin suppresses TPA-induced metastasis by downregulating matrix metalloproteinase-9 expression through JNK/SP-1 signaling in nasopharyngeal carcinoma. J. Pineal Res. 2016, 61, 479–492.
- Selbach, M.; Schwanhausser, B.; Thierfelder, N.; Fang, Z.; Khanin, R.; Rajewsky, N. Widespread changes in protein synthesis induced by microRNAs. Nature 2008, 455, 58–63.
- Sethi, N.; Wright, A.; Wood, H.; Rabbitts, P. MicroRNAs and head and neck cancer: Reviewing the first decade of research. Eur. J. Cancer 2014, 50, 2619–2635.
- Kolokythas, A.; Miloro, M.; Zhou, X. Review of microRNA proposed target genes in oral cancer. Part II. J. Oral Maxillofac. Res. 2011, 2, e2.
- Manasa, V.G.; Kannan, S. Impact of microRNA dynamics on cancer hallmarks: An oral cancer scenario. Tumour Biol. 2017, 39, 1010428317695920.
- Schneider, A.; Victoria, B.; Lopez, Y.N.; Suchorska, W.; Barczak, W.; Sobecka, A.; Golusinski, W.; Masternak, M.M.; Golusinski, P. Tissue and serum microRNA profile of oral squamous cell carcinoma patients. Sci. Rep. 2018, 8, 675.
- Min, A.; Zhu, C.; Peng, S.; Rajthala, S.; Costea, D.E.; Sapkota, D. MicroRNAs as important players and biomarkers in oral carcinogenesis. Biomed. Res. Int. 2015, 2015, 186904.
- Lin, S.C.; Liu, C.J.; Lin, J.A.; Chiang, W.F.; Hung, P.S.; Chang, K.W. miR-24 up-regulation in oral carcinoma: Positive association from clinical and in vitro analysis. Oral Oncol. 2010, 46, 204–208.
- Lee, S.E.; Kim, S.J.; Youn, J.P.; Hwang, S.Y.; Park, C.S.; Park, Y.S. MicroRNA and gene expression analysis of melatonin-exposed human breast cancer cell lines indicating involvement of the anticancer effect. J. Pineal Res. 2011, 51, 345–352.
- Mori, F.; Ferraiuolo, M.; Santoro, R.; Sacconi, A.; Goeman, F.; Pallocca, M.; Pulito, C.; Korita, E.; Fanciulli, M.; Muti, P.; et al. Multitargeting activity of miR-24 inhibits long-term melatonin anticancer effects. Oncotarget 2016, 7, 20532–20548.
- Lee, S.E.; Kim, S.J.; Yoon, H.J.; Yu, S.Y.; Yang, H.; Jeong, S.I.; Hwang, S.Y.; Park, C.S.; Park, Y.S. Genome-wide profiling in melatonin-exposed human breast cancer cell lines identifies differentially methylated genes involved in the anticancer effect of melatonin. J. Pineal Res. 2013, 54, 80–88.
- Hunsaker, M.; Barba, G.; Kingsley, K.; Howard, K.M. Differential microRNA expression of miR-21 and miR-155 within oral cancer extracellular vesicles in response to melatonin. Dent. J. 2019, 7, 48.
- Avissar, M.; Christensen, B.C.; Kelsey, K.T.; Marsit, C.J. MicroRNA expression ratio is predictive of head and neck squamous cell carcinoma. Clin. Cancer Res. 2009, 15, 2850–2855.
- Lamperska, K.M.; Kozlowski, P.; Kolenda, T.; Teresiak, A.; Blizniak, R.; Przybyla, W.; Masternak, M.M.; Golusinski, P.; Golusinski, W. Unpredictable changes of selected miRNA in expression profile of HNSCC. Cancer Biomark. 2016, 16, 55–64.
- Cervigne, N.K.; Reis, P.P.; Machado, J.; Sadikovic, B.; Bradley, G.; Galloni, N.N.; Pintilie, M.; Jurisica, I.; Perez-Ordonez, B.; Gilbert, R.; et al. Identification of a microRNA signature associated with progression of leukoplakia to oral carcinoma. Hum. Mol. Genet. 2009, 18, 4818–4829.
- He, Q.; Chen, Z.; Cabay, R.J.; Zhang, L.; Luan, X.; Chen, D.; Yu, T.; Wang, A.; Zhou, X. microRNA-21 and microRNA-375 from oral cytology as biomarkers for oral tongue cancer detection. Oral Oncol. 2016, 57, 15–20.
- Zeljic, K.; Jovanovic, I.; Jovanovic, J.; Magic, Z.; Stankovic, A.; Supic, G. MicroRNA meta-signature of oral cancer: Evidence from a meta-analysis. Upsala J. Med. Sci. 2018, 123, 43–49.
- Yan, Z.Y.; Luo, Z.Q.; Zhang, L.J.; Li, J.; Liu, J.Q. Integrated analysis and microRNA expression profiling identified seven miRNAs associated with progression of oral squamous cell carcinoma. J. Cell. Physiol. 2017, 232, 2178–2185.
- Gissi, D.B.; Morandi, L.; Gabusi, A.; Tarsitano, A.; Marchetti, C.; Cura, F.; Palmieri, A.; Montebugnoli, L.; Asioli, S.; Foschini, M.P.; et al. A noninvasive test for microRNA expression in oral squamous cell carcinoma. Int. J. Mol. Sci. 2018, 19, 1789.
- Shah, S.; Jadhav, K.; Shah, V.; Gupta, N.; Dagrus, K. miRNA 21: Diagnostic prognostic and therapeutic marker for oral cancer. Microrna 2016, 5, 175–179.
- Yap, T.; Koo, K.; Cheng, L.; Vella, L.J.; Hill, A.F.; Reynolds, E.; Nastri, A.; Cirillo, N.; Seers, C.; McCullough, M. Predicting the presence of oral squamous cell carcinoma using commonly dysregulated MicroRNA in oral swirls. Cancer Prev. Res. 2018, 11, 491–502.




