
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Shanta Hinton | + 2197 word(s) | 2197 | 2021-07-13 05:42:08 | | | |
| 2 | Dean Liu | Meta information modification | 2197 | 2021-07-14 04:41:52 | | |
Video Upload Options
Pseudophosphatases are important regulators, and their proper regulation is important for maintaining homeostasis and preventing diseases.
1. Introduction
Over the past decade, the relevance of pseudoenzymes has been elevated and solidified as important signaling regulators. These modes of action and their roles in various cellular processes serve as an idea link for the roles of pseudoenzymes in pathologies and diseases. Misregulation of pseudoenzymes has been implicated in the etiology of various diseases such as cancer, obesity, and neurological disorders [1][2][3][4][5]. This review highlights the signaling pseudophosphatases and their implications in diseases, while demonstrating some of their essential roles in preventing or causing such diseases.
Pseudophosphatases of PTPs are widely accepted as having mutations within their signature active site motif (HCX5R) They maintain the three-dimensional fold and the ability to bind phosphorylated proteins [1][6][7]. Recent reports have advanced the field, complicating the mutated sequence definition [6][8] and the “grab and hold on” perspective, where pseudophosphatases stably interact with phosphorylated residues, thereby competitively inhibiting active phosphatases or kinases [6][7]. For example, the histidine domain containing protein tyrosine phosphatase (HD-PTP) is reported as a pseudophosphatase [1][2][9][10][11][12][13] and an active phosphatase [1][2][14][15][16].
This dual and contradictory functionality of HD-PTP highlights the complexities of investigating pseudophosphatases. Wishart and Dixon defined pseudophosphatases as STYX domains, and demonstrated a loss of function of dephosphorylating phosphorylated substrates [17]; thus, they defined the prototypical STYX domain beyond pure bioinformatics [17]. The analysis of pseudophosphatases through a combination of sequencing and functional experiments—dephosphorylation assays with appropriate substrates and contextual situations (localization, substrates/interactors, or abundance of interactor or pseudphosphatase—will provide more of an insight into the full spectrum [1][2][3]. Nevertheless, the roles of pseudophosphatases as regulators in many cellular processes such as spermatogenesis, stress response, neuronal differentiation, cell fate, migration, ubiquitylation, demyelination, oocyte-to-zygote transition, transcription, and apoptosis [1][18][3][7][11][13][19][20] have led to interest into their apparent roles in diseases.
The increased interest in pseudophosphatases resulted in the discovery of their relevance in many diseases. provided a useful framework for pseudophosphatases’ linkage to diseases [3]. Mutations of these molecules or their misregulation leads to diseases such as leukemia, breast cancer, colorectal cancer, hepatocarcinoma, glioblastoma, other cancers, Charcot-Marie-Tooth (CMT) disease (abnormal nerve myelination), obesity, diabetes, chronic obstructive pulmonary disease (COPD), and nephrotic syndrome [2][3][21][22][23][24][25][26][27]. A comprehensive list of the known functions of pseudophosphatases and their implications in diseases is provided in Table 1.
| Pseudophosphatase Name | Alternative Names | Normal Function | Role in Disease |
|---|---|---|---|
| MTMR5 (Myotubularin-related protein 5) |
SBF1 |
|
|
| MTMR9 (Myotubularin-related protein 9) |
|||
| MTMR10 (Myotubularin-related protein 10) |
- |
|
|
| MTMR11 (Myotubularin-related protein 11) |
- | ||
| MTMR12 (Myotubularin-related protein 12) |
3-PAP | - | |
| MTMR13 (Myotubularin-related protein 13) |
SBF2 |
|
|
| STYX (Serine/threonine/tyrosine -interacting protein) |
|||
| MK-STYX (Mitogen-activated protein kinase phosphoserine/ threonine/tyrosine-binding protein) |
STYXL1, DUSP-24 |
|
|
| STYXL2 (Serine/threonine/tyrosine-interacting-like protein 2) |
DUSP-27 (duplicated) |
|
|
| TAB1 (TGF-beta-activated kinase 1 and MAP3K7-binding protein 1) |
MAP3K7IP1 | ||
| Tensin 1 | TNS1 |
The proteins here all contain alterations in their catalytic active site motifs that are associated with loss of phosphatase function.
Furthermore, those that have not been linked to diseases have functions that may allude to a possible role in diseases. The diversity of these diseases, such as various types of cancer, COPD, CMT, obesity, diabetes, etc. Furthermore, their diverse range, while validating the intricate roles of pseudophosphatases in signaling cascades, also makes understanding their many functions in diseases more complicated. Considering this special edition focused on the roles of protein tyrosine phosphatases in signaling, with an emphasis on therapeutic strategies, the pseudophosphatases’ signaling mechanisms linked to diseases, myotubularins, tensins, and STYX pseudophosphatases, was discussed in more detail.
2. Myotubularin in Diseases
Six of these genes produce pseudophosphatases, which form complexes with their active homologs [21][64][65] (Figure 1A). This dimerization is context dependent, resulting in complex stability, enhancement of catalytic function (increased phosphatase activity), and regulation of the subcellular localization of the active phosphatase [66][67]. The coupling of active and inactive MTMs to form heterodimers is common; however, self-association to form homodimers also occurs among both active and inactive MTMs (Figure 1A) It is of interest that some MTMs have not been reported to interact directly as dimers [65].
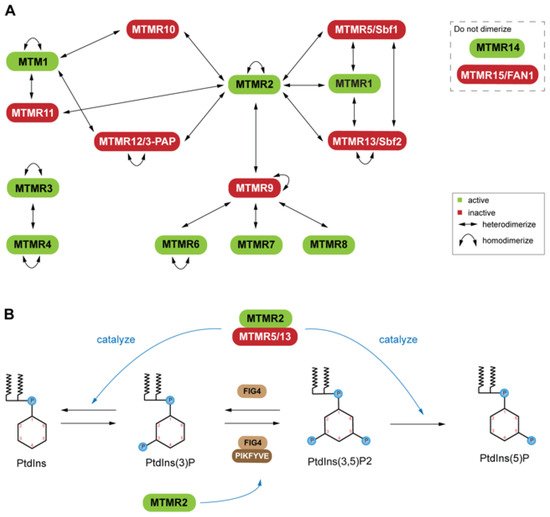
For example, mutations of MTMR2 and MTMR13 leads to misregulation of the AKT signaling in Type 4B Charcot-Marie-Tooth (CMT4B) disease [68]. Overexpression of MTMR2 prevents sustained activation of epidermal growth factor, which leads to sustained AKT activation in transgenic mice [68]. PtdIns(3)P and PtdIns(3,5)P2are important molecules for endosomal-lysosomal membrane trafficking [69], which is tightly regulated by PtdIns kinase and phosphatase activity. MTMs have also been linked to X-linked centronuclear myopathy (XLCNM)
(phosphatase) forms heterodimers with MTMR13 (pseudophosphatase) or MTMR5 (pseudophosphatase), and heterodimers with MTMR13 (pseudophosphatase) or MTMR5 (pseudophosphatase), and Genetic studies in mice revealed that MTMR12 interacts with another phosphatase, polyphosphoinositde phosphatase (FIG4) MTM pseudophosphatase: phosphatase heterodimer signaling complexes are required for cellular processes such as differentiation, membrane trafficking, endocytosis, and survival [70][21][71][69].
3. STYX Pseudophosphatases in Disease
The term “STYX” (phosphoserine/threonine/tyrosine-interacting protein) was coined to designate the phosphotyrosine binding domain that has no catalytic activity, or “dead” phosphatases or pseudophosphatase [17]. These “dead” or STYX domain phosphatases allude to the Greek mythological STYX river of the dead [6][7]. Usage of a point mutation to “restore” catalytic activity in the STYX domain has proved to be a helpful tool to initiate the process of investigating molecules that contain STYX domains. STYX domain phosphatases are pseudophosphatases; therefore, the authors have referred to them as STYX pseudophosphatases, as previously reported [6].
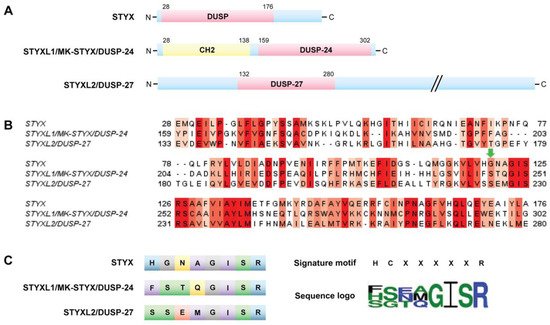
They are all members of the dual specificity protein tyrosine phosphatase (DUSP) family; however, they belong to different subfamilies [2]. These domains vary both from the active site signature motif and each other (Figure 2B,C). Of note, all these STYX domain proteins have glycine, isoleucine, serine, and arginine in the active site motif (Figure 2B,C). Furthermore, early reports classified glycine as a conserved residue of the active site motif, HCxxGxxR [70][16].
STYX and MK-STYX have been shown to be important signaling regulators in cellular processes such as cell cycle, spermatogenesis, cell-fate decisions, cell migration, ubiquitylation and protein degradation, apoptosis, and neuronal differentiation [19][20][46][47][49][72][73], alluding to their roles in the etiology of various diseases. Furthermore, these STYX pseudophosphatases have roles in cancer such as colorectal, Ewing sarcoma, and hepatocarcinoma [4][22][27]. In addition, STYX promotes oncogenesis in colorectal cancer by inhibiting FBXW7, blocking the degradation of cyclin E and c-Jun Therefore, free (unbound) cyclin E and c-Jun are able to promote proliferation in colorectal cancer.
MK-STYX lacks the critical cysteine in the active site signature motif (HCX5R) An increase in MK-STYX expression also promotes hepatocellular carcinoma (Figure 3 (Bii)) mRNA expression results in upregulation of PI3K (phosphatidyl 3-kinase)/AKT pathway proteins and an enhancement of proliferation, while reducing apoptosis in hepatocellular carcinoma STYXL2 is a downstream target of the Janus activated kinase/Signal transducers and activators of transcription signaling pathway [74], suggesting that STYXL2 has important roles in signaling cascades.
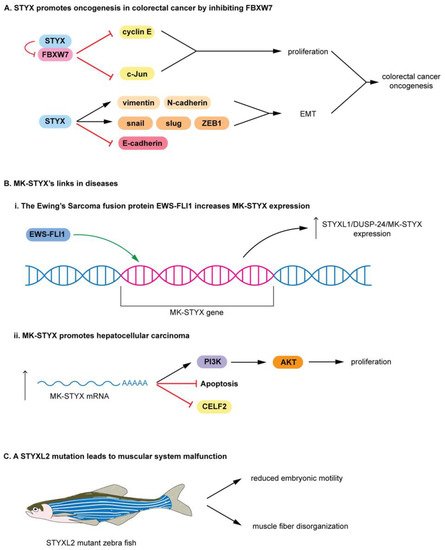
In addition, a transgene integration intoSTYXL2reduces STYXL2 expression in zebrafish, resulting in muscular system malfunction (Figure 3C) [2][74]. STYXL2 mutants have reduced embryonic motility (low paralysis), displaying low spontaneous coiling movements and severely reduced touch response (Figure 3C) [74]. Furthermore, STYXL2 mutants have a major disruptions in the contractile apparatus of their muscle fibers–disorganized muscle fiber formation (Figure 3C) [74].
4. Atypical Pseudophosphatases
This review focused on the less contradictory pseudophosphatases’ signaling in diseases, with DUSP27/STYXL2 as the exception; Reiterer et al. encompasses such contradictory pseudophosphatases [2]. In addition, the D2 domains, which are catalytically inactive and recognize substrate [70], of the PTP receptor molecules were excluded. Recently, the protein tyrosine phosphatase receptor U (PTPRU) was classified as pseudophosphatase [75], both its DUSP domains, D1 and D2, were considered as pseudophosphatase domains [75]. In addition, tensin 2 is thought to have possible activity by dephosphorylating insulin receptor substrate-1 [76].
Some pseudophosphatases were reported as having catalytic activity [1][2][3]. Phosphatase of regenerating liver-3 (PRL3) phosphatase is an example of the concept of a pseudo-pseudophosphatase, which recognizes the catalytic activity and the importance of the noncatalysis in signaling cascades [2][77][78]. In studies with PRL3 mutants specifically defective in either binding the substrate of PRL3 substrate, CBS domain divalent metal cations transport mediators (CNNM) or phosphatase activity [78] demonstrated that the phosphatase activity is dispensable [78]. Moreover, phosphatase activity prevents PRL3-CNNM interaction, which is necessary and sufficient for tumor metastasis [78].
References
- Hinton, S.D. The role of pseudophosphatases as signaling regulators. Biochim. Biophys. Acta Mol. Cell Res. 2019, 1866, 167–174.
- Reiterer, V.; Pawlowski, K.; Desrochers, G.; Pause, A.; Sharpe, H.J.; Farhan, H. The dead phosphatases society: A review of the emerging roles of pseudophosphatases. FEBS J. 2020, 287, 4198–4220.
- Reiterer, V.; Eyers, P.A.; Farhan, H. Day of the dead: Pseudokinases and pseudophosphatases in physiology and disease. Trends Cell Biol. 2014, 24, 489–505.
- Isrie, M.; Zamani Esteki, M.; Peeters, H.; Voet, T.; Van Houdt, J.; Van Paesschen, W.; Van Esch, H. Homozygous missense mutation in STYXL1 associated with moderate intellectual disability, epilepsy and behavioural complexities. Eur. J. Med. Genet. 2015, 58, 205–210.
- Siligan, C.; Ban, J.; Bachmaier, R.; Spahn, L.; Kreppel, M.; Schaefer, K.L.; Poremba, C.; Aryee, D.N.; Kovar, H. EWS-FLI1 target genes recovered from Ewing’s sarcoma chromatin. Oncogene 2005, 24, 2512–2524.
- Wishart, M.J.; Dixon, J.E. Gathering STYX: Phosphatase-like form predicts functions for unique protein-interaction domains. Trends Biochem. Sci. 1998, 23, 301–306.
- Tonks, N.K. Pseudophosphatases: Grab and hold on. Cell 2009, 139, 464–465.
- Chen, M.J.; Dixon, J.E.; Manning, G. Genomics and evolution of protein phosphatases. Sci. Signal. 2017, 10.
- Gingras, M.C.; Zhang, Y.L.; Kharitidi, D.; Barr, A.J.; Knapp, S.; Tremblay, M.L.; Pause, A. HD-PTP is a catalytically inactive tyrosine phosphatase due to a conserved divergence in its phosphatase domain. PLoS ONE 2009, 4, e5105.
- Manteghi, S.; Gingras, M.C.; Kharitidi, D.; Galarneau, L.; Marques, M.; Yan, M.; Cencic, R.; Robert, F.; Paquet, M.; Witcher, M.; et al. Haploinsufficiency of the ESCRT Component HD-PTP Predisposes to Cancer. Cell Rep. 2016, 15, 1893–1900.
- Gahloth, D.; Heaven, G.; Jowitt, T.A.; Mould, A.P.; Bella, J.; Baldock, C.; Woodman, P.; Tabernero, L. The open architecture of HD-PTP phosphatase provides new insights into the mechanism of regulation of ESCRT function. Sci. Rep. 2017, 7, 9151.
- Gahloth, D.; Levy, C.; Heaven, G.; Stefani, F.; Wunderley, L.; Mould, P.; Cliff, M.J.; Bella, J.; Fielding, A.J.; Woodman, P.; et al. Structural Basis for Selective Interaction between the ESCRT Regulator HD-PTP and UBAP1. Structure 2016, 24, 2115–2126.
- Bulgari, D.; Jha, A.; Deitcher, D.L.; Levitan, E.S. Myopic (HD-PTP, PTPN23) selectively regulates synaptic neuropeptide release. Proc. Natl. Acad. Sci. USA 2018, 115, 1617–1622.
- Zhang, S.; Fan, G.; Hao, Y.; Hammell, M.; Wilkinson, J.E.; Tonks, N.K. Suppression of protein tyrosine phosphatase N23 predisposes to breast tumorigenesis via activation of FYN kinase. Genes Dev. 2017, 31, 1939–1957.
- Lin, G.; Aranda, V.; Muthuswamy, S.K.; Tonks, N.K. Identification of PTPN23 as a novel regulator of cell invasion in mammary epithelial cells from a loss-of-function screen of the ‘PTP-ome’. Genes Dev. 2011, 25, 1412–1425.
- Flint, A.J.; Tiganis, T.; Barford, D.; Tonks, N.K. Development of “substrate-trapping” mutants to identify physiological substrates of protein tyrosine phosphatases. Proc. Natl. Acad. Sci. USA 1997, 94, 1680–1685.
- Wishart, M.J.; Denu, J.M.; Williams, J.A.; Dixon, J.E. A single mutation converts a novel phosphotyrosine binding domain into a dual-specificity phosphatase. J. Biol. Chem. 1995, 270, 26782–26785.
- Hinton, S.D. Pseudophosphatase MK-STYX: The atypical member of the MAP kinase phosphatases. FEBS J. 2020, 287, 4221–4231.
- Parry, J.M.; Velarde, N.V.; Lefkovith, A.J.; Zegarek, M.H.; Hang, J.S.; Ohm, J.; Klancer, R.; Maruyama, R.; Druzhinina, M.K.; Grant, B.D.; et al. EGG-4 and EGG-5 Link Events of the Oocyte-to-Embryo Transition with Meiotic Progression in C. elegans. Curr. Biol. 2009, 19, 1752–1757.
- Cheng, K.C.; Klancer, R.; Singson, A.; Seydoux, G. Regulation of MBK-2/DYRK by CDK-1 and the pseudophosphatases EGG-4 and EGG-5 during the oocyte-to-embryo transition. Cell 2009, 139, 560–572.
- Robinson, F.L.; Dixon, J.E. Myotubularin phosphatases: Policing 3-phosphoinositides. Trends Cell Biol. 2006, 16, 403–412.
- He, D.; Ma, Z.; Fang, C.; Ding, J.; Yang, W.; Chen, P.; Huang, L.; Wang, C.; Yu, Y.; Yang, L.; et al. Pseudophosphatase STYX promotes tumor growth and metastasis by inhibiting FBXW7 function in colorectal cancer. Cancer Lett. 2019, 454, 53–65.
- Gu, J.; Zhang, Z.; Lang, T.; Ma, X.; Yang, L.; Xu, J.; Tian, C.; Han, K.; Qiu, J. PTPRU, As A Tumor Suppressor, Inhibits Cancer Stemness By Attenuating Hippo/YAP Signaling Pathway. OncoTargets Ther. 2019, 12, 8095–8104.
- Robinson, F.L.; Dixon, J.E. The phosphoinositide-3-phosphatase MTMR2 associates with MTMR13, a membrane-associated pseudophosphatase also mutated in type 4B Charcot-Marie-Tooth disease. J. Biol. Chem. 2005, 280, 31699–31707.
- Cho, A.R.; Uchio-Yamada, K.; Torigai, T.; Miyamoto, T.; Miyoshi, I.; Matsuda, J.; Kurosawa, T.; Kon, Y.; Asano, A.; Sasaki, N.; et al. Deficiency of the tensin2 gene in the ICGN mouse: An animal model for congenital nephrotic syndrome. Mamm. Genome 2006, 17, 407–416.
- Stylianou, P.; Clark, K.; Gooptu, B.; Smallwood, D.; Brightling, C.E.; Amrani, Y.; Roach, K.M.; Bradding, P. Tensin1 expression and function in chronic obstructive pulmonary disease. Sci. Rep. 2019, 9, 18942.
- Wu, J.Z.; Jiang, N.; Lin, J.M.; Liu, X. STYXL1 promotes malignant progression of hepatocellular carcinoma via downregulating CELF2 through the PI3K/Akt pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 2977–2985.
- Kim, S.A.; Vacratsis, P.O.; Firestein, R.; Cleary, M.L.; Dixon, J.E. Regulation of myotubularin-related (MTMR)2 phosphatidylinositol phosphatase by MTMR5, a catalytically inactive phosphatase. Proc. Natl. Acad. Sci. USA 2003, 100, 4492–4497.
- Pareyson, D.; Stojkovic, T.; Reilly, M.M.; Leonard-Louis, S.; Laura, M.; Blake, J.; Parman, Y.; Battaloglu, E.; Tazir, M.; Bellatache, M.; et al. A multicenter retrospective study of charcot-marie-tooth disease type 4B (CMT4B) associated with mutations in myotubularin-related proteins (MTMRs). Ann. Neurol. 2019, 86, 55–67.
- Firestein, R.; Nagy, P.L.; Daly, M.; Huie, P.; Conti, M.; Cleary, M.L. Male infertility, impaired spermatogenesis, and azoospermia in mice deficient for the pseudophosphatase Sbf1. J. Clin. Investig. 2002, 109, 1165–1172.
- Zhou, M.; Wu, L.; Liang, J.; Shen, C.; Lin, J. Expression analysis and functional characterization of a novel cold-responsive gene CbCOR15a from Capsella bursa-pastoris. Mol. Biol. Rep. 2012, 39, 5169–5179.
- Hotta, K.; Kitamoto, T.; Kitamoto, A.; Mizusawa, S.; Matsuo, T.; Nakata, Y.; Kamohara, S.; Miyatake, N.; Kotani, K.; Komatsu, R.; et al. Association of variations in the FTO, SCG3 and MTMR9 genes with metabolic syndrome in a Japanese population. J. Hum. Genet. 2011, 56, 647–651.
- Baulac, S.; Gourfinkel-An, I.; Couarch, P.; Depienne, C.; Kaminska, A.; Dulac, O.; Baulac, M.; LeGuern, E.; Nabbout, R. A novel locus for generalized epilepsy with febrile seizures plus in French families. Arch. Neurol. 2008, 65, 943–951.
- Shoji, H.; Isomoto, H.; Yoshida, A.; Ikeda, H.; Minami, H.; Kanda, T.; Urabe, S.; Matsushima, K.; Takeshima, F.; Nakao, K.; et al. MicroRNA-130a is highly expressed in the esophageal mucosa of achalasia patients. Exp. Ther. Med. 2017, 14, 898–904.
- Kabir, N.N.; Ronnstrand, L.; Kazi, J.U. Deregulation of protein phosphatase expression in acute myeloid leukemia. Med. Oncol. 2013, 30, 517.
- Lucci, M.A.; Orlandi, R.; Triulzi, T.; Tagliabue, E.; Balsari, A.; Villa-Moruzzi, E. Expression profile of tyrosine phosphatases in HER2 breast cancer cells and tumors. Anal. Cell. Pathol. 2010, 32, 361–372.
- Gupta, V.A.; Hnia, K.; Smith, L.L.; Gundry, S.R.; McIntire, J.E.; Shimazu, J.; Bass, J.R.; Talbot, E.A.; Amoasii, L.; Goldman, N.E.; et al. Loss of catalytically inactive lipid phosphatase myotubularin-related protein 12 impairs myotubularin stability and promotes centronuclear myopathy in zebrafish. PLoS Genet. 2013, 9, e1003583.
- Nandurkar, H.H.; Layton, M.; Laporte, J.; Selan, C.; Corcoran, L.; Caldwell, K.K.; Mochizuki, Y.; Majerus, P.W.; Mitchell, C.A. Identification of myotubularin as the lipid phosphatase catalytic subunit associated with the 3-phosphatase adapter protein, 3-PAP. Proc. Natl. Acad. Sci. USA 2003, 100, 8660–8665.
- Lassuthova, P.; Vill, K.; Erdem-Ozdamar, S.; Schroder, J.M.; Topaloglu, H.; Horvath, R.; Muller-Felber, W.; Bansagi, B.; Schlotter-Weigel, B.; Glaser, D.; et al. Novel SBF2 mutations and clinical spectrum of Charcot-Marie-Tooth neuropathy type 4B2. Clin. Genet. 2018, 94, 467–472.
- Reiterer, V.; Fey, D.; Kolch, W.; Kholodenko, B.N.; Farhan, H. Pseudophosphatase STYX modulates cell-fate decisions and cell migration by spatiotemporal regulation of ERK1/2. Proc. Natl. Acad. Sci. USA 2013, 110, E2934–E2943.
- Reiterer, V.; Figueras-Puig, C.; Le Guerroue, F.; Confalonieri, S.; Vecchi, M.; Jalapothu, D.; Kanse, S.M.; Deshaies, R.J.; Di Fiore, P.P.; Behrends, C.; et al. The pseudophosphatase STYX targets the F-box of FBXW7 and inhibits SCFFBXW7 function. EMBO J. 2017, 36, 260–273.
- Buchanan, N.S.; Zhao, J.; Zhu, K.; Patwa, T.H.; Miller, F.R.; Lubman, D.M. Differential expression of acidic proteins with progression in the MCF10 model of human breast disease. Int. J. Oncol. 2007, 31, 941–949.
- Liu, L.; Jiang, H.; Wang, X.; Wang, X.; Zou, L. STYX/FBXW7 axis participates in the development of endometrial cancer cell via Notch-mTOR signaling pathway. Biosci. Rep. 2020, 40.
- Hinton, S.D.; Myers, M.P.; Roggero, V.R.; Allison, L.A.; Tonks, N.K. The pseudophosphatase MK-STYX interacts with G3BP and decreases stress granule formation. Biochem. J. 2010, 427, 349–357.
- Barr, J.E.; Munyikwa, M.R.; Frazier, E.A.; Hinton, S.D. The pseudophosphatase MK-STYX inhibits stress granule assembly independently of Ser149 phosphorylation of G3BP-1. FEBS J. 2013, 280, 273–284.
- Niemi, N.M.; Sacoman, J.L.; Westrate, L.M.; Gaither, L.A.; Lanning, N.J.; Martin, K.R.; MacKeigan, J.P. The pseudophosphatase MK-STYX physically and genetically interacts with the mitochondrial phosphatase PTPMT1. PLoS ONE 2014, 9, e93896.
- Banks, D.A.; Dahal, A.; McFarland, A.G.; Flowers, B.M.; Stephens, C.A.; Swack, B.; Gugssa, A.; Anderson, W.A.; Hinton, S.D. MK-STYX Alters the Morphology of Primary Neurons, and Outgrowths in MK-STYX Overexpressing PC-12 Cells Develop a Neuronal Phenotype. Front. Mol. Biosci. 2017, 4, 76.
- Flowers, B.M.; Rusnak, L.E.; Wong, K.E.; Banks, D.A.; Munyikwa, M.R.; McFarland, A.G.; Hinton, S.D. The pseudophosphatase MK-STYX induces neurite-like outgrowths in PC12 cells. PLoS ONE 2014, 9, e114535.
- Dahal, A.; Hinton, S.D. Antagonistic roles for STYX pseudophosphatases in neurite outgrowth. Biochem. Soc. Trans. 2017, 45, 381–387.
- Cao, Y.; Banks, D.A.; Mattei, A.M.; Riddick, A.T.; Reed, K.M.; Zhang, A.M.; Pickering, E.S.; Hinton, S.D. Pseudophosphatase MK-STYX Alters Histone Deacetylase 6 Cytoplasmic Localization, Decreases Its Phosphorylation, and Increases Detyrosination of Tubulin. Int. J. Mol. Sci. 2019, 20, 1455.
- Tomar, V.S.; Baral, T.K.; Nagavelu, K.; Somasundaram, K. Serine/threonine/tyrosine-interacting-like protein 1 (STYXL1), a pseudo phosphatase, promotes oncogenesis in glioma. Biochem. Biophys. Res. Commun. 2019, 515, 241–247.
- Kim, B.; Bae, H.; Lee, H.; Lee, S.; Park, J.C.; Kim, K.R.; Kim, S.J. Proton Beams Inhibit Proliferation of Breast Cancer Cells by Altering DNA Methylation Status. J. Cancer 2016, 7, 344–352.
- Winter, J.M.; Curry, N.L.; Gildea, D.M.; Williams, K.A.; Lee, M.; Hu, Y.; Crawford, N.P.S. Modifier locus mapping of a transgenic F2 mouse population identifies CCDC115 as a novel aggressive prostate cancer modifier gene in humans. BMC Genom. 2018, 19, 450.
- Fero, K.; Bergeron, S.A.; Horstick, E.J.; Codore, H.; Li, G.H.; Ono, F.; Dowling, J.J.; Burgess, H.A. Impaired embryonic motility in dusp27 mutants reveals a developmental defect in myofibril structure. Dis. Models Mech. 2014, 7, 289–298.
- Shibuya, H.; Yamaguchi, K.; Shirakabe, K.; Tonegawa, A.; Gotoh, Y.; Ueno, N.; Irie, K.; Nishida, E.; Matsumoto, K. TAB1: An activator of the TAK1 MAPKKK in TGF-beta signal transduction. Science 1996, 272, 1179–1182.
- Lu, G.; Kang, Y.J.; Han, J.; Herschman, H.R.; Stefani, E.; Wang, Y. TAB-1 modulates intracellular localization of p38 MAP kinase and downstream signaling. J. Biol. Chem. 2006, 281, 6087–6095.
- Zhu, Y.; Regunath, K.; Jacq, X.; Prives, C. Cisplatin causes cell death via TAB1 regulation of p53/MDM2/MDMX circuitry. Genes Dev. 2013, 27, 1739–1751.
- Lei, X.; Han, N.; Xiao, X.; Jin, Q.; He, B.; Wang, J. Enterovirus 71 3C inhibits cytokine expression through cleavage of the TAK1/TAB1/TAB2/TAB3 complex. J. Virol. 2014, 88, 9830–9841.
- Lo, S.H. Tensin. Int. J. Biochem. Cell Biol. 2004, 36, 31–34.
- Chen, H.; Duncan, I.C.; Bozorgchami, H.; Lo, S.H. Tensin1 and a previously undocumented family member, tensin2, positively regulate cell migration. Proc. Natl. Acad. Sci. USA 2002, 99, 733–738.
- Zhou, H.; Zhang, Y.; Wu, L.; Xie, W.; Li, L.; Yuan, Y.; Chen, Y.; Lin, Y.; He, X. Elevated transgelin/TNS1 expression is a potential biomarker in human colorectal cancer. Oncotarget 2018, 9, 1107–1113.
- Zhan, Y.; Liang, X.; Li, L.; Wang, B.; Ding, F.; Li, Y.; Wang, X.; Zhan, Q.; Liu, Z. MicroRNA-548j functions as a metastasis promoter in human breast cancer by targeting Tensin1. Mol. Oncol. 2016, 10, 838–849.
- Chang, K.C.; Diermeier, S.D.; Yu, A.T.; Brine, L.D.; Russo, S.; Bhatia, S.; Alsudani, H.; Kostroff, K.; Bhuiya, T.; Brogi, E.; et al. MaTAR25 lncRNA regulates the Tensin1 gene to impact breast cancer progression. Nat. Commun. 2020, 11, 6438.
- Ng, A.A.; Logan, A.M.; Schmidt, E.J.; Robinson, F.L. The CMT4B disease-causing phosphatases Mtmr2 and Mtmr13 localize to the Schwann cell cytoplasm and endomembrane compartments, where they depend upon each other to achieve wild-type levels of protein expression. Hum. Mol. Genet. 2013, 22, 1493–1506.
- St-Denis, N.; Gupta, G.D.; Lin, Z.Y.; Gonzalez-Badillo, B.; Pelletier, L.; Gingras, A.C. Myotubularin-related proteins 3 and 4 interact with polo-like kinase 1 and centrosomal protein of 55 kDa to ensure proper abscission. Mol. Cell. Proteomics MCP 2015, 14, 946–960.
- Raess, M.A.; Cowling, B.S.; Bertazzi, D.L.; Kretz, C.; Rinaldi, B.; Xuereb, J.M.; Kessler, P.; Romero, N.B.; Payrastre, B.; Friant, S.; et al. Expression of the neuropathy-associated MTMR2 gene rescues MTM1-associated myopathy. Hum. Mol. Genet. 2017, 26, 3736–3748.
- Raess, M.A.; Friant, S.; Cowling, B.S.; Laporte, J. WANTED—Dead or alive: Myotubularins, a large disease-associated protein family. Adv. Biol. Regul. 2017, 63, 49–58.
- Berger, P.; Tersar, K.; Ballmer-Hofer, K.; Suter, U. The CMT4B disease-causing proteins MTMR2 and MTMR13/SBF2 regulate AKT signalling. J. Cell. Mol. Med. 2011, 15, 307–315.
- Hnia, K.; Vaccari, I.; Bolino, A.; Laporte, J. Myotubularin phosphoinositide phosphatases: Cellular functions and disease pathophysiology. Trends Mol. Med. 2012, 18, 317–327.
- Tonks, N.K. Protein tyrosine phosphatases—From housekeeping enzymes to master regulators of signal transduction. FEBS J. 2013, 280, 346–378.
- Gal, J.; Kuang, L.; Barnett, K.R.; Zhu, B.Z.; Shissler, S.C.; Korotkov, K.V.; Hayward, L.J.; Kasarskis, E.J.; Zhu, H. ALS mutant SOD1 interacts with G3BP1 and affects stress granule dynamics. Acta Neuropathol. 2016, 132, 563–576.
- Wishart, M.J.; Dixon, J.E. The archetype STYX/dead-phosphatase complexes with a spermatid mRNA-binding protein and is essential for normal sperm production. Proc. Natl. Acad. Sci. USA 2002, 99, 2112–2117.
- Reiterer, V.; Pawlowski, K.; Farhan, H. STYX: A versatile pseudophosphatase. Biochem. Soc. Trans. 2017, 45, 449–456.
- Cooper, L.M.; Waddell, D.S. A tale of two DUSP27s: Proposed resolution for the naming of distinct dual-specificity phosphatases. Am. J. Physiol. Cell Physiol. 2020, 319, C148–C150.
- Hay, I.M.; Fearnley, G.W.; Rios, P.; Kohn, M.; Sharpe, H.J.; Deane, J.E. The receptor PTPRU is a redox sensitive pseudophosphatase. Nat. Commun. 2020, 11, 3219.
- Koh, A.; Lee, M.N.; Yang, Y.R.; Jeong, H.; Ghim, J.; Noh, J.; Kim, J.; Ryu, D.; Park, S.; Song, P.; et al. C1-Ten is a protein tyrosine phosphatase of insulin receptor substrate 1 (IRS-1), regulating IRS-1 stability and muscle atrophy. Mol. Cell. Biol. 2013, 33, 1608–1620.
- Gehring, K.; Miki, H. Phosphatase, pseudo-phosphatase, or both? Understanding PRL oncogenicity. Br. J. Cancer 2021, 124, 1035–1036.
- Kozlov, G.; Funato, Y.; Chen, Y.S.; Zhang, Z.; Illes, K.; Miki, H.; Gehring, K. PRL3 pseudophosphatase activity is necessary and sufficient to promote metastatic growth. J. Biol. Chem. 2020, 295, 11682–11692.




