
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jekaterina Erenpreisa | + 1857 word(s) | 1857 | 2021-06-30 10:31:45 | | | |
| 2 | Dean Liu | -5 word(s) | 1852 | 2021-07-05 05:03:51 | | | | |
| 3 | Jekaterina Erenpreisa | Meta information modification | 1852 | 2021-07-06 08:03:32 | | | | |
| 4 | Dean Liu | + 108 word(s) | 1960 | 2021-07-07 12:06:55 | | |
Video Upload Options
Recent data point to the leading role of heterochromatin in genome maintenance, highlighting the attractions between heterochromatic regions as being central to phase separation of the active and inactive chromatin domains. The three main properties of heterochromatin—the position silencing effect on transcription, stickiness, and flexible rigidity are important for its network formation.
1. Introduction
The DNA linear sequence code represents only the most basic principle of gene regulation. In 1944, Erwin Schrödinger (one of the founders of quantum mechanics and Physics Nobel Prize winner of 1933) published his famous lecture “What is life?” [1]. He proposed that DNA may possess the features of aperiodic crystal and use spatial organization as one of the regulatory elements. The physical features of heterochromatin composed of repeated short nucleotide sequences (satellite DNA) may confer the specificity to provide a gene with “a chromosomal address”[2]. In turn, Ilya Prigogine, another Nobel prize winner in chemistry 1977, formulated that any living dissipative system undergoing exchange of matter and energy with the environment is a subject of structured attenuated oscillations[3].Thus, the 3D nuclear organization experiences transcriptional oscillations coordinating its function and scaling many orders of magnitude, including cell–cell interaction[4][5][6]. At a very coarse grain, we can imagine such oscillations as a biphasic interchange of chromatin compartments between active euchromatin and inactive heterochromatin compartment[7]. Although the dominating view in the last decade was that the euchromatin and heterochromatin compartments behave as relatively independent self-organized crowded fluids, some recent data still point to the leading role of heterochromatin in genome maintenance, highlighting the attractions between heterochromatic regions as being central to phase separation of the active and inactive chromatin domains[8]. Recent experiments have shown that heterochromatin and bound HP1 protein undergo liquid-liquid phase separation, while euchromatin does not[9]. In[10] it was revealed that condensed chromatin exists in a solid-like state whose properties resist external forces and create an elastic gel providing a scaffold that supports liquid-liquid phase separation of chromatin binding proteins. The physical features of constitutive heterochromatin, namely the regular structure, condensation, and positioning of nucleosomes, are conformed by a set of associated chromosomal proteins, including subcompartmentation by CTCF and cohesin, [11][12] which also depend evolutionarily on condensin II.
The properties of herochromatin are consistent with the earlier ideas that its topology, in terms of the sum of its position-cis-silencing effects on transcription, can provide the 3D “genome active space” in addition to the DNA gene-coding function, “the morphogenic function”[13]—a source of positional information necessary for the whole genome regulation[14][15][16]. In the recent publication[18], we were able to link the timing of structural bursting of pericentromere-associated domains (PADs) with the large-scale activation of gene expression occurring by critical self-organization in cancer cells coinciding with biphasic activation of bivalent stress-response genes of the FOS family in the differentiation commitment model. Loss of the heterochromatin silencing threshold by splitting PAD clusters leading to the documented massive unfolding of euchromatin, was suggested as the first step for genome repatterning. The discovered basic features of PADs, converting the same amount of constitutive heterochromatin by fusing-splitting within the constraint of cell nucleus volume, suggests their involvement in the supra-chromosomal network as one of the most promising structural candidates supporting chromatin driving gradients and transcriptional pulsing by bursting synchronization[17].
The mesoscopic approach to the description of complex systems is guided by unveiling the presence of “organization principles”. Its main peculiarity is the independence from the detailed knowledge of the basic bricks of the system and the emergence of relevant features from the correlation structure of the studied phenomenon, where “all the entities can be considered as networks of interacting parts”. Here, we will follow this approach to individuate (besides a detailed molecular-level mechanistic approach) some basic pillars of the supra-chromosome wiring structure.
2. Flexible Heterochromatin, Transcription Regulation, and Related Mechanobiology of the Actomyosin Network
The mechanobiology of transcription and transcriptional pulsing itself. EM studies of speckles (also called interchromatin granule clusters) examined in very active cancer cells revealed their direct link to both perinucleolar and perinuclear heterochromatin. Suppression of rRNA synthesis by AcD provoked the radial-concentric organization of the main components involved in transcription machinery, with heterochromatin disclosing a spatial link between rRNA and mRNA synthesis and processing events. The situation with low AcD mimics a return phase of transcriptional pulsing.
An actomyosin-based cytoskeleton using ATP energy plays a role in heterochromatin segregation and the organization of specific heterochromatin compartments: β-actin deficient cells exhibit changes in the spatial organization of H3K9Me3/HP1α—positive heterochromatin[19]. At the same time, one of the primary functions of the elastic nuclear actomyosin network is to maintain a chromatin landscape compatible with transcription[20]. How are these two functions coupled through the putative nuclear heterochromatin network?
The involvement of fibrillar nuclear actin (F-actin) in the activation and process of transcription with all three RNA polymerases is well-established[20][21]. Upon serum stimulation of transcription, F-actin is seen accumulating in the nucleolus, transcription fabrics, and at the nuclear envelope outlining the route of the transcription conveyor. Nuclear actin and the motor of actin, nuclear myosin (NM1), are required for RNA polymerase I transcription of rRNA. Under AcD action, NM1 is transferred into the nucleolus, likely favouring the contraction of NORs.
Actin is part of pre-initiation complexes and is necessary for transcription by RNA polymerase II; it is involved both in the initiation and elongation of transcription, acting through ribonucleoprotein interactions and the chromatin remodelling, but also interacting with pure DNA[22]. NM1-F-actin polymerizing complex activating Pol II needs the binding of bipolar phosphatidylinositol 4,5-bisphosphate and adapting by its disordered protein domain, which also enables liquid phase separation of nuclear speckles [23]. The role of speckles in the amplification of gene expression was recently stressed. NM1 interacts with rRNA transcription initiation factor TIF-IA, which brings actin and NM1 in close proximity with each other. It thus provides feedback with the nucleolus sensing the cell environment, starting transcription pulsing aided by actomyosin. These studies show that the regulation of transcription is structured. At the same time, the process itself creates the assisting nucleoskeleton dynamic framework.
The involvement in RNA transcription of actomyosin activity of nuclear speckles, their phase separation, their anchoring to the perinucleolar and perinuclear heterochromatin, as well as their spring-like morphology, visualized by EM (Figure 1a,b), is apparent.
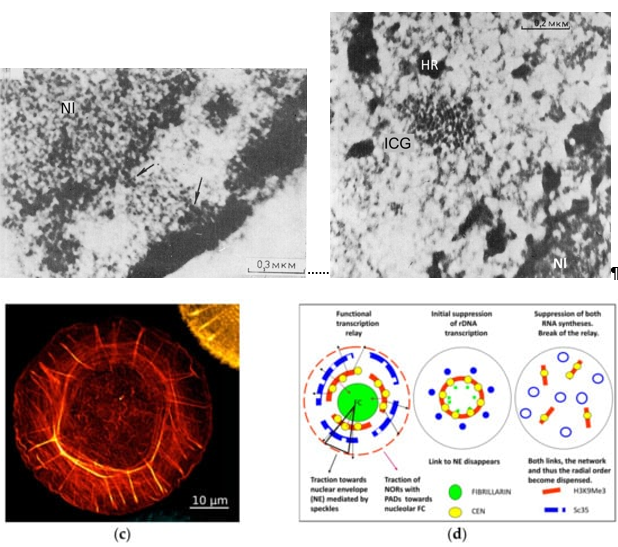
Figure 1. The speckles acting as radial spring pumps anchored between the nucleolar and perinuclear flexibly rigid heterochromatin shells and equipped by elastic actomyosin elements integrate the radial-concentric nuclear order for transcription pulsing. (a,b) EM thin sections of rat ascitic Zajdela hepatoma cells: (a) The links of the speckle (ICG) to the perinucleolar and perinuclear heterochromatin (arrows), Nl—nucleolus; (b) the view of the radial-concentric speckle substructure as revealed on shortly cold-PF-prefixed cells after short DNAse I treatment with 4.5 mM MgCl2 in phosphate buffer/sucrose before dehydration and epoxy-embedment (conventional contrasting). ICG—speckle, HR—heterochromatin, Nl—nucleolus. Republished from[24]. (c) The representative image of the circular-radial self-organization of F-actin filaments (stained by phalloidin) of human foreskin fibroblasts, structured soon after seeding on isotropic substrate; from[25]. (d) Schematic of the functional transcriptional relay in its relationship with speckles and the concentric rings of the nucleolar and perinuclear heterochromatin shells, deduced from experiments with suppression of RNA synthesis and the literature on the participation of the nuclear cytoskeleton in it. Modified from[26].
It presumes that radial nuclear order is much dependent on the speckle springy traction of the nucleolar and nuclear RNA synthesis conveyor between the perinucleolar and perinuclear heterochromatin stiffly bending concentric rings. Release of the connection to nuclear lamin contracts the system inside the cell nucleus and exposes its radial-concentric character. When nuclear synthesis has been fully suppressed, both anchors of speckles to the perinucleolar and perinuclear constraints are released and the radial-concentric order is dispensed. Thus, nuclear speckles, postulated as a structure integrating the whole process of transcription[27] seem indeed capable of executing it through transcription pulsing of the heterochromatin network united with the actomyosin network which, in its turn, also follows and supports its radial-concentric character.
3. Nucleolar Pulsing and Nucleolus-Associated Domains (NADs)
Balbiani (1864) was the first who discovered vacuoles and their discharge from the nucleolus in active cells and called the nucleolus the “heart of the cell”. Nucleolar pulsing was also described in plants, associated with transcriptional activity and in mouse oocytes with particularly high synthesis of rRNA. Pulsation of nucleolar rRNA synthesis was also revealed using confocal Raman microspectroscopy and pulsation and coalescence of human nucleoli were registered in living cells[28]. In[29] the authors suggest that observation of the nucleolus pulsing may reflect cycling in the assembly and disassembly of active chromatin structure in and/or around rDNA genes.
In this connection, the question arises whether the emergence of function in transcription alveoli may be associated with nucleolus-associated domains (NADs). rDNAs are the most abundant genes, and the stability of the rDNA cluster was considered a key element of genome maintenance.
NADs are defined as genomic regions enriched at the nucleolar periphery in interphase cells[30]. In addition to the nucleolar organizers (NORs) in five acrocentric chromosome pairs of the human genome, when using deep sequencing of chromatin associated with biochemically purified nucleoli in Hi-C and other labelling methods, it was revealed that specific chromatin domains from most human chromosomes may associate with nucleoli[31]. The NADs comprising 38% of the human genome contain rDNA sequences and H3K9me3-enriched heterochromatin associated with a-satellite DNA and centromeres[32]. The most recent data on NADs in mouse embryonic stem cells and fibroblasts, besides a large NADs subset overlapping with lamina-associated domains (LADs), revealed a subset of smaller NADs free from binding to nuclear lamina — corresponding more dynamic chromocenters with higher gene density, including developmental and differentiation genes[33][34]. Nucleoli in G1 cells are very dynamic—fusing and splitting; the reiterated ribosomal DNA sequences are also easily amplified and highly variable in size, flexibly adapting to cell needs, and are the first undergoing replication stress during genome instability and cell senescence.
In any case, the available molecular and morphological data indicate that most NADs are associated with PADs, and this link can be hypothesized to play a role in the formation of functional alveoli and scale-free transcriptional pulsing of their mutual network coordinating rRNA and mRNA synthesis in the radially directed relay. Therefore, we should shortly review some data on transcriptional pulsing.
4. Transcriptional Pulsing and Synchronization in a Cell Population
Transcriptional bursting, also known as transcriptional pulsing, is a fundamental property of genes, which have been observed in diverse organisms, from bacteria to mammals[35]. The burst frequency is primarily encoded in enhancers, and burst size in core promoters, while cell type-specific gene expression is primarily shaped by changes in burst frequencies[6][36]. Cultured cells display cell cycle-independent collective dynamics of gene transcription fluctuations with a period of 20–40 min[4]. The stress response of Zaidela ascites carcinoma cells (washed and incubated in PBS in a plugged vial), seen as a few ~90 min collective waves of mRNA synthesis increased up to two to three times, was registered along with a bi-stable switch for the increased superhelicity of DNA opposed by the emergence and subsequent repair of DNA single-strand-breaks; these oscillations further faded and led to cell death[5][37].
The sense of radial-circular nuclear organization for transcriptional pulsing is not only in the structural conveyor coordinating the production of pre-ribosomes with the synthesis and maturation of It means feedback from sensing the environment and cell community which, in case of life threat (and likely, in reprogramming), can become synchronised. In other words—it is an element of explorative adaptation requiring the system to act as a whole. But the question remains, how does this self-organising dynamicity fit with the robust regulation of gene expression in tissue differentiation[38] and, ultimately, how is the tissue-specific code established and read?
References
- Schroedinger, E. What is Life? The Physical Aspect of the Living Cell, Etc; Cambridge University Press: Cambridge, 1944.
- Singh, P.B.; Huskisson, N.S. Chromatin Complexes as Aperiodic Microcrystalline Arrays That Regulate Genome Organisation and Expression. Dev. Genet. 1998, 22, 85–99.
- The Nobel Prize in Chemistry 1977 Available online: https://www.nobelprize.org/prizes/chemistry/1977/prigogine/lecture/
- Tsuchiya, M.; Wong, S.T.; Yeo, Z.X.; Colosimo, A.; Palumbo, M.C.; Farina, L.; Crescenzi, M.; Mazzola, A.; Negri, R.; Bianchi, M.M.; et al. Gene Expression Waves. Cell Cycle Independent Collective Dynamics in Cultured Cells. FEBS J. 2007, 274, 2878–2886.
- Erenpreisa, J.; Budylin, A. Related Changes in RNA Synthesis and DNA Superhelicity during Starvation of Ehrlich Ascites Tumour Cells. Proc. Latv. Acad. Sci 1990, Ser B. 10, 90–94.
- Larsson, A.J.M.; Johnsson, P.; Hagemann-Jensen, M.; Hartmanis, L.; Faridani, O.R.; Reinius, B.; Segerstolpe, Å.; Rivera, C.M.; Ren, B.; Sandberg, R. Genomic Encoding of Transcriptional Burst Kinetics. Nature 2019, 565, 251–254.
- Lieberman-Aiden, E.; van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 2009, 326, 289–293.
- Falk, M.; Feodorova, Y.; Naumova, N.; Imakaev, M.; Lajoie, B.R.; Leonhardt, H.; Joffe, B.; Dekker, J.; Fudenberg, G.; Solovei, I.; et al. Heterochromatin Drives Compartmentalization of Inverted and Conventional Nuclei. Nature 2019, 570, 395–399.
- Gibson, B.A.; Doolittle, L.K.; Jensen, L.E.; Gamarra, N.; Redding, S.; Rosen, M.K. Organization and Regulation of Chromatin by Liquid-Liquid Phase Separation. bioRxiv 2019.
- Strickfaden, H.; Tolsma, T.; Sharma, A.; Alan Underhill, D.; Hansen, J.C.; Hendzel, M.J. Condensed Chromatin Behaves like a Solid on the Mesoscale in Vitro and in Living Cells. Int. J. Mol. Sci. 2019, 21, doi:10.3390/ijms21010240.
- Girard, M.; Olvera de la Cruz, M.; Marko, J.F.; Erbaş, A. Heterochromatin Flexibility Contributes to Chromosome Segregation in the Cell Nucleus. bioRxiv 2020.
- Thorn, G.J.; Clarkson, C.T.; Rademacher, A.; Mamayusupova, H.; Schotta, G.; Rippe, K.; Teif, V.B. DNA Sequence-Dependent Formation of Heterochromatin Nanodomains. BioRxiv 2020, doi:10.1101/2020.12.20.423673.
- Scherrer, K. A Unified Matrix Hypothesis of DNA-Directed Morphogenesis, Protodynamism and Growth Control. Biosci. Rep. 1989, 9, 157–188.
- Nagl, W. Condensed chromatin: Species-specificity, tissue-specificity, and cell cycle-specificity, as monitored by scanning cytometry. In Cell Growth; Springer New York: Boston, MA, 1982; pp. 171–218 ISBN 9781468440485.
- Ērenpreisa, J.; Zhukotsky, A. Interphase Genome as the Active Space: Chromatin Dynamics during Chick Embryo Chondrogenesis. Mechanisms of Ageing and Development 1993, 67, 21–32.
- Bártová, E.; Kozubek, S.; Jirsová, P.; Kozubek, M.; Gajová, H.; Lukášová, E.; Skalnı́ková, M.; Gaňová, A.; Koutná, I.; Hausmann, M. Nuclear Structure and Gene Activity in Human Differentiated Cells. J. Struct. Biol. 2002, 139, 76–89.
- Erenpreisa, J.; Giuliani, A. Resolution of Complex Issues in Genome Regulation and Cancer Requires Non-Linear and Network-Based Thermodynamics. Int. J. Mol. Sci. 2020, 21(1), 240.
- Krigerts, J.; Salmina, K.; Freivalds, T.; Zayakin, P.; Rumnieks, F.; Inashkina, I.; Giuliani, A.; Hausmann, M.; Erenpreisa, J. Differentiating Cancer Cells Reveal Early Large-Scale Genome Regulation by Pericentric Domains. Biophys. J. 2021, 120, 711–724.
- Venit, T.; Mahmood, S.R.; Endara-Coll, M.; Percipalle, P. Nuclear Actin and Myosin in Chromatin Regulation and Maintenance of Genome Integrity. Int. Rev. Cell Mol. Biol. 2020, 355, 67–108.
- de Lanerolle, P. Nuclear Actin and Myosins at a Glance. J. Cell Sci. 2012, 125, 4945–4949.
- Yamazaki, S.; Yamamoto, K.; Harata, M. Contribution of Nuclear Actin to Transcription Regulation. Genom Data 2015, 4, 127–129.
- Negishi, M.; Sakaue, T.; Takiguchi, K.; Yoshikawa, K. Cooperation between Giant DNA Molecules and Actin Filaments in a Microsphere. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2010, 81, 051921.
- Balaban, C.; Sztacho, M.; Blažíková, M.; Hozák, P. The F-Actin-Binding MPRIP Forms Phase-Separated Condensates and Associates with PI(4,5)P2 and Active RNA Polymerase II in the Cell Nucleus. Cells 2021, 10, doi:10.3390/cells10040848
- Erenpreisa, J. Organization of the Chromatin in Interphase Cell Nucleus; Zinatne Publishers: Riga, 1990.
- Jalal, S.; Shi, S.; Acharya, V.; Huang, R.Y.-J.; Viasnoff, V.; Bershadsky, A.D.; Tee, Y.H. Actin Cytoskeleton Self-Organization in Single Epithelial Cells and Fibroblasts under Isotropic Confinement. J. Cell Sci. 2019, 132, doi:10.1242/jcs.220780.
- Salmina, K.; Kurg, R.; Krigerts, J.; Erenpreisa, J. The Topological Relationship between Ribogenesis, mRNA Transcription/splicing and the Tension of Actin Cytoskeleton. Biopolym. Cell 2019, 35, 231–232.
- Galganski, L.; Urbanek, M.O.; Krzyzosiak, W.J. Nuclear Speckles: Molecular Organization, Biological Function and Role in Disease. Nucleic Acids Res. 2017, 45, 10350–10368.
- Caragine, C.M.; Haley, S.C.; Zidovska, A. Nucleolar Dynamics and Interactions with Nucleoplasm in Living Cells. Elife 2019, 8, doi:10.7554/eLife.47533
- Vankova Hausnerova, V.; Krizek, P.; Hagen, G.M.; Lanctot, C. Transcriptional Pulsing of a Nucleolar Transgene. bioRxiv 2016.
- Bizhanova, A.; Kaufman, P.D. Close to the Edge: Heterochromatin at the Nucleolar and Nuclear Peripheries. Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms 2021, 1864, 194666.
- van Koningsbruggen, S.; Gierlinski, M.; Schofield, P.; Martin, D.; Barton, G.J.; Ariyurek, Y.; den Dunnen, J.T.; Lamond, A.I. High-Resolution Whole-Genome Sequencing Reveals That Specific Chromatin Domains from Most Human Chromosomes Associate with Nucleoli. Mol. Biol. Cell 2010, 21, 3735–3748.
- Németh, A.; Conesa, A.; Santoyo-Lopez, J.; Medina, I.; Montaner, D.; Péterfia, B.; Solovei, I.; Cremer, T.; Dopazo, J.; Längst, G. Initial Genomics of the Human Nucleolus. PLoS Genet. 2010, 6, e1000889.
- Vertii, A.; Ou, J.; Yu, J.; Yan, A.; Pagès, H.; Liu, H.; Zhu, L.J.; Kaufman, P.D. Two Contrasting Classes of Nucleolus-Associated Domains in Mouse Fibroblast Heterochromatin. Genome Res. 2019, 29, 1235–1249.
- Bizhanova, A.; Yan, A.; Yu, J.; Zhu, L.J.; Kaufman, P.D. Distinct Features of Nucleolus-Associated Domains in Mouse Embryonic Stem Cells. Chromosoma 2020, 129, 121–139.
- Smirnov, E.; Hornáček, M.; Vacík, T.; Cmarko, D.; Raška, I. Discontinuous Transcription. Nucleus 2018, 9, 149–160.
- Beckman, W.F.; Jiménez, M.Á.L.; Moerland, P.D.; Westerhoff, H.V.; Verschure, P.J. 4sUDRB-Sequencing for Genome-Wide Transcription Bursting Quantification in Breast Cancer Cells. bioRxiv 2020.
- Budylin, A.; Erenpreisa, J.; Zhukotsky, A.; Zaleskaya, N. Fluctuations of the Reparable DNA Strand Breaks in Starving Tumour Cells. Proc.Latv.Acad.Sci. 1993, 5, 72–73.
- Tsuchiya, M.; Giuliani, A.; Hashimoto, M.; Erenpreisa, J.; Yoshikawa, K. Self-Organizing Global Gene Expression Regulated through Criticality: Mechanism of the Cell-Fate Change. PLoS One 2016, 11, e0167912.




