| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Byung-Hoon Lee | + 2555 word(s) | 2555 | 2021-06-16 05:52:26 | | | |
| 2 | Peter Tang | Meta information modification | 2555 | 2021-06-23 03:47:35 | | |
Video Upload Options
The 26S proteasome is the principal protease for regulated intracellular proteolysis. This multi-subunit complex is also pivotal for clearance of harmful proteins that are produced throughout the lifetime of eukaryotes. Recent structural and kinetic studies have revealed a multitude of conformational states of the proteasome in substrate-free and substrate-engaged forms. These conformational transitions demonstrate that proteasome is a highly dynamic machinery during substrate processing that can be also controlled by a number of proteasome-associated factors. Essentially, three distinct family of deubiquitinases–USP14, RPN11, and UCH37–are associated with the 19S regulatory particle of human proteasome. USP14 and UCH37 are capable of editing ubiquitin conjugates during the process of their dynamic engagement into the proteasome prior to the catalytic commitment. In contrast, RPN11-mediated deubiquitination is directly coupled to substrate degradation by sensing the proteasome’s conformational switch into the commitment steps.
1. Introduction
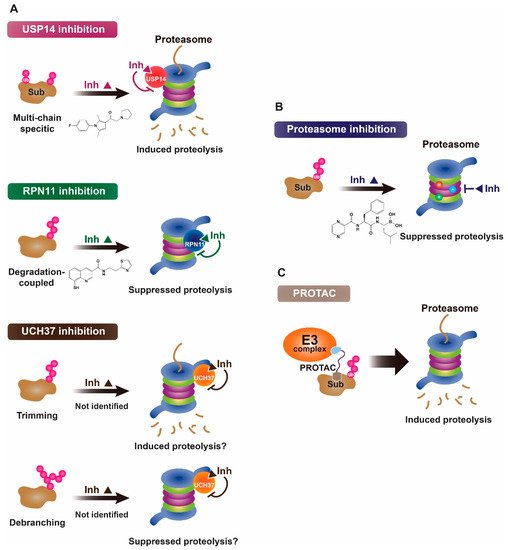
2. Proteasomal Deubiquitinases as Therapeutic Targets
3. Proteasomal Deubiquitinase Inhibitors
3.1. USP14 Inhibitors
|
Target |
Compound Name |
Structure |
Notes |
Reference |
|---|---|---|---|---|
|
USP14 |
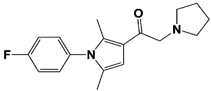
IU1 |
|
IC50: 4.7 μM (Ub-AMC) |
Lee et al., 2010 [16] |
|
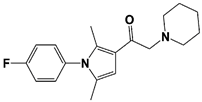
IU1-2 |
|
IC50: 1.7 μM (Ub-AMC) |
Boselli et al., 2017 [62] |
|
|
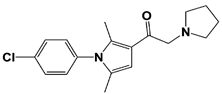
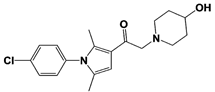
IU1-33 |
|
IC50: 1.1 μM (Ub-AMC) |
||
|
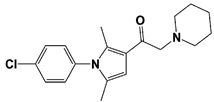
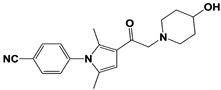
IU1-47 |
|
IC50: 0.6 μM (Ub-AMC) |
||
|
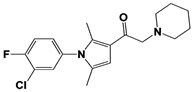
IU1-206 |
|
N/A |
Wang et al., 2018 [98] |
|
|
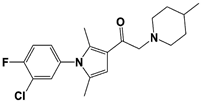
IU1-248 |
|
IC50: 0.83 μM (Ub-AMC) |
||
|
1B10 |
|
N/A |
Palmer et al., 2018 [99] |
|
|
1D18 |
|
N/A |
||
|
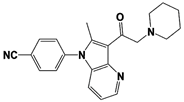
Compound 162 |
|
IC50: <0.5 μM (Ub-AMC) |
WO/2015/073528 [100] |
|
|
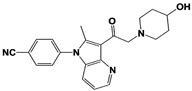
Compound 335 (SB1-B-57) |
|
IC50: <0.5 μM (Ub-AMC) |
||
|
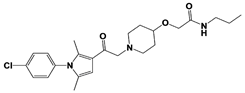
Compound 83 |
|
IC50: <0.5 μM (Ub-AMC) |
WO/2020/006269 [101] |
|
|
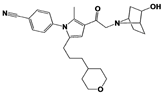
Compound 2B |
|
IC50: <0.05 μM (Ub-AMC) |
WO/2020/006296 [102] |
|
|
IU2-6 |
|
74% inhibition at 8 μM (Ub-AMC) |
WO/2012/012712 [103]; Kemp, 2016 [104] |
|
|
Compound 3 |
|
IC50: 0.5 μM (Ub-AMC) |
WO/2013/112651 [105]; Kemp, 2016 [104] |
|
|
RPN11 |
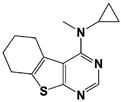
8-TQ * |
|
IC50: 2.4 μM (Ub4-pepOG) |
Li et al., 2017 [106]; Perez et al., 2017 [107] |
|
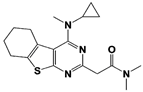
Capzimin |
|
IC50: 0.34 μM (Ub4-pepOG) |
||
|
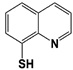
Thiolutin * |
|
IC50: 0.53 μM (Ub4-pepOG) |
Lauinger et al., 2017 [108] |
|
|
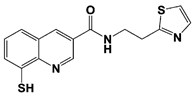
SOP6 * |
|
IC50: 3.8 μM (Fluorescent UbnGST-Wbp2) |
Li et al., 2018 [109] |
|
|
SOP11 * |
|
IC50: 1.3 μM (Fluorescent UbnGST-Wbp2) |
||
|
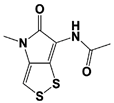
Promiscuous proteasomal DUB inhibitors |
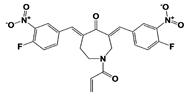
b-AP15 (USP14/ UCH37) |
|
19S RP IC50: 6.5 μM (Ub-Rho) |
D‘Arcy et al., 2011 [110]; Wang et al., 2015 [111] |
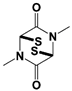
|
VLX1570 (USP14/ UCH37) |
|
19S RP IC50: 6.4 μM (Ub-Rho) |
Wang et al., 2015 [111] |
|
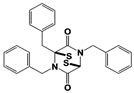
|
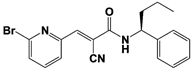
WP1130 (USP9x/USP5/ USP14/UCH37) |
|
IC50s: <5~10 μM (Ub-AMC & Ub-VS) |
Kapuria et al., 2010 [112] |
|
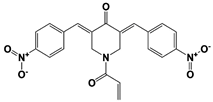
|
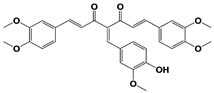
AC17 (19S RP) |
|
19S RP IC50: 4.23 μM (Ub-AMC) |
Zhou et al., 2013 [113] |
|
|
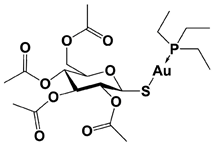
Auranofin ** (TrxR/19S RP; 19S RP at higher dosage than TrxR) |
|
TrxR system inhibition at ~1 μM (HCT-116 cell) Reduced 19S RP labeling at 5 μM (Ub-VS) |
Liu et al., 2014 [114]; Stafford et al., 2018 [115]; Zhang et al., 2019 [116] |
* These inhibitors also inhibit other JAMM metalloproteases. ** This inhibitor may have its non-DUB target in pharmacological dosage.
3.2. RPN11 Inhibitors
3.3. Other Proteasomal Deubiquitinase Inhibitors
References
- Hershko, A.; Ciechanover, A. The ubiquitin system for protein degradation. Annu. Rev. Biochem. 1992, 61, 761–807.
- Goldberg, A.L. Protein degradation and protection against misfolded or damaged proteins. Nature 2003, 426, 895–899.
- Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 2009, 78, 477–513.
- Mao, Y. Structure, Dynamics and Function of the 26S Proteasome. Subcell. Biochem. 2021, 96, 1–151.
- Finley, D.; Prado, M.A. The Proteasome and Its Network: Engineering for Adaptability. Cold Spring Harb. Perspect. Biol. 2020, 12, a033985.
- Greene, E.R.; Dong, K.C.; Martin, A. Understanding the 26S proteasome molecular machine from a structural and conformational dynamics perspective. Curr. Opin. Struct. Biol. 2020, 61, 33–41.
- Chen, X.; Htet, Z.M.; Lopez-Alfonzo, E.; Martin, A.; Walters, K.J. Proteasome interaction with ubiquitinated substrates: From mechanisms to therapies. FEBS J. 2020.
- Davis, C.; Spaller, B.L.; Matouschek, A. Mechanisms of substrate recognition by the 26S proteasome. Curr. Opin. Struct. Biol. 2020, 67, 161–169.
- Sahu, I.; Glickman, M.H. Proteasome in action: Substrate degradation by the 26S proteasome. Biochem. Soc. Trans. 2021, 49, 629–644.
- Mevissen, T.E.T.; Komander, D. Mechanisms of Deubiquitinase Specificity and Regulation. Annu. Rev. Biochem. 2017, 86, 159–192.
- De Poot, S.A.H.; Tian, G.; Finley, D. Meddling with Fate: The Proteasomal Deubiquitinating Enzymes. J. Mol. Biol. 2017, 429, 3525–3545.
- Shin, J.Y.; Muniyappan, S.; Tran, N.N.; Park, H.; Lee, S.B.; Lee, B.H. Deubiquitination Reactions on the Proteasome for Proteasome Versatility. Int. J. Mol. Sci. 2020, 21, 5312.
- Bard, J.A.M.; Goodall, E.A.; Greene, E.R.; Jonsson, E.; Dong, K.C.; Martin, A. Structure and Function of the 26S Proteasome. Annu. Rev. Biochem. 2018, 87, 697–724.
- Leggett, D.S.; Hanna, J.; Borodovsky, A.; Crosas, B.; Schmidt, M.; Baker, R.T.; Walz, T.; Ploegh, H.; Finley, D. Multiple associated proteins regulate proteasome structure and function. Mol. Cell 2002, 10, 495–507.
- Hanna, J.; Hathaway, N.A.; Tone, Y.; Crosas, B.; Elsasser, S.; Kirkpatrick, D.S.; Leggett, D.S.; Gygi, S.P.; King, R.W.; Finley, D. Deubiquitinating enzyme Ubp6 functions noncatalytically to delay proteasomal degradation. Cell 2006, 127, 99–111.
- Lee, B.H.; Lee, M.J.; Park, S.; Oh, D.C.; Elsasser, S.; Chen, P.C.; Gartner, C.; Dimova, N.; Hanna, J.; Gygi, S.P.; et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 2010, 467, 179–184.
- Lee, B.H.; Lu, Y.; Prado, M.A.; Shi, Y.; Tian, G.; Sun, S.; Elsasser, S.; Gygi, S.P.; King, R.W.; Finley, D. USP14 deubiquitinates proteasome-bound substrates that are ubiquitinated at multiple sites. Nature 2016, 532, 398–401.
- Muniyappan, S.; Lee, B.H. In vitro analysis of proteasome-associated USP14 activity for substrate degradation and deubiquitylation. Methods Enzymol. 2019, 619, 249–268.
- Verma, R.; Aravind, L.; Oania, R.; McDonald, W.H.; Yates, J.R., III; Koonin, E.V.; Deshaies, R.J. Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome. Science 2002, 298, 611–615.
- Yao, T.; Cohen, R.E. A cryptic protease couples deubiquitination and degradation by the proteasome. Nature 2002, 419, 403–407.
- Lam, Y.A.; Xu, W.; DeMartino, G.N.; Cohen, R.E. Editing of ubiquitin conjugates by an isopeptidase in the 26S proteasome. Nature 1997, 385, 737–740.
- Deol, K.K.; Crowe, S.O.; Du, J.; Bisbee, H.A.; Guenette, R.G.; Strieter, E.R. Proteasome-Bound UCH37/UCHL5 Debranches Ubiquitin Chains to Promote Degradation. Mol. Cell 2020, 80, 796–809.e9.
- Harrigan, J.A.; Jacq, X.; Martin, N.M.; Jackson, S.P. Deubiquitylating enzymes and drug discovery: Emerging opportunities. Nat. Rev. Drug Discov. 2018, 17, 57–78.
- Moon, S.; Lee, B.H. Chemically Induced Cellular Proteolysis: An Emerging Therapeutic Strategy for Undruggable Targets. Mol. Cells 2018, 41, 933–942.
- Ndubaku, C.; Tsui, V. Inhibiting the deubiquitinating enzymes (DUBs). J. Med. Chem. 2015, 58, 1581–1595.
- Gopinath, P.; Ohayon, S.; Nawatha, M.; Brik, A. Chemical and semisynthetic approaches to study and target deubiquitinases. Chem. Soc. Rev. 2016, 45, 4171–4198.
- Schauer, N.J.; Magin, R.S.; Liu, X.; Doherty, L.M.; Buhrlage, S.J. Advances in Discovering Deubiquitinating Enzyme (DUB) Inhibitors. J. Med. Chem. 2020, 63, 2731–2750.
- Wertz, I.E.; Wang, X. From Discovery to Bedside: Targeting the Ubiquitin System. Cell Chem. Biol. 2019, 26, 156–177.
- Cohen, P.; Tcherpakov, M. Will the Ubiquitin System Furnish as Many Drug Targets as Protein Kinases? Cell 2010, 143, 686–693.
- Popovic, D.; Vucic, D.; Dikic, I. Ubiquitination in disease pathogenesis and treatment. Nat. Med. 2014, 20, 1242–1253.
- Schmidt, M.; Finley, D. Regulation of proteasome activity in health and disease. BBA Mol. Cell Res. 2014, 1843, 13–25.
- Weathington, N.M.; Mallampalli, R.K. Emerging therapies targeting the ubiquitin proteasome system in cancer. J. Clin. Investig. 2014, 124, 6–12.
- Rape, M. Ubiquitylation at the crossroads of development and disease. Nat. Rev. Mol. Cell Biol. 2018, 19, 59–70.
- Richardson, P.G.; Hideshima, T.; Anderson, K.C. Bortezomib (PS-341): A novel, first-in-class proteasome inhibitor for the treatment of multiple myeloma and other cancers. Cancer Control 2003, 10, 361–369.
- Chen, D.; Frezza, M.; Schmitt, S.; Kanwar, J.; Dou, Q.P. Bortezomib as the First Proteasome Inhibitor Anticancer Drug: Current Status and Future Perspectives. Curr. Cancer Drug Targets 2011, 11, 239–253.
- Burslem, G.M.; Crews, C.M. Proteolysis-Targeting Chimeras as Therapeutics and Tools for Biological Discovery. Cell 2020, 181, 102–114.
- Verma, R.; Mohl, D.; Deshaies, R.J. Harnessing the Power of Proteolysis for Targeted Protein Inactivation. Mol. Cell 2020, 77, 446–460.
- Huang, X.D.; Dixit, V.M. Drugging the undruggables: Exploring the ubiquitin system for drug development. Cell Res. 2016, 26, 484–498.
- Kategaya, L.; Di Lello, P.; Rouge, L.; Pastor, R.; Clark, K.R.; Drummond, J.; Kleinheinz, T.; Lin, E.; Upton, J.P.; Prakash, S.; et al. USP7 small-molecule inhibitors interfere with ubiquitin binding. Nature 2017, 550, 534–538.
- Lamberto, I.; Liu, X.X.; Seo, H.S.; Schauer, N.J.; Iacob, R.E.; Hu, W.Y.; Das, D.; Mikhailova, T.; Weisberg, E.L.; Engen, J.R.; et al. Structure-Guided Development of a Potent and Selective Non-covalent Active-Site Inhibitor of USP7. Cell Chem. Biol. 2017, 24, 1490–1500.e11.
- Turnbull, A.P.; Ioannidis, S.; Krajewski, W.W.; Pinto-Fernandez, A.; Heride, C.; Martin, A.C.L.; Tonkin, L.M.; Townsend, E.C.; Buker, S.M.; Lancia, D.R.; et al. Molecular basis of USP7 inhibition by selective small-molecule inhibitors. Nature 2017, 550, 481–486.
- Gavory, G.; O’Dowd, C.R.; Helm, M.D.; Flasz, J.; Arkoudis, E.; Dossang, A.; Hughes, C.; Cassidy, E.; McClelland, K.; Odrzywol, E.; et al. Discovery and characterization of highly potent and selective allosteric USP7 inhibitors. Nat. Chem. Biol. 2018, 14, 118–125.
- Kluge, A.F.; Lagu, B.R.; Maiti, P.; Jaleel, M.; Webb, M.; Malhotra, J.; Mallat, A.; Srinivas, P.A.; Thompson, J.E. Novel highly selective inhibitors of ubiquitin specific protease 30 (USP30) accelerate mitophagy. Bioorganic Med. Chem. Lett. 2018, 28, 2655–2659.
- Li, P.; Liu, H.-M. Recent advances in the development of ubiquitin-specific-processing protease 7 (USP7) inhibitors. Eur. J. Med. Chem. 2020, 191, 112107.
- Borodovsky, A.; Kessler, B.; Casagrande, R.; Overkleeft, H.S.; Wilkinson, K.D.; Ploegh, H.L. A novel active site-directed probe specific for deubiquitylating enzymes reveals proteasome association of USP14. EMBO J. 2001, 20, 5187–5196.
- Aufderheide, A.; Beck, F.; Stengel, F.; Hartwig, M.; Schweitzer, A.; Pfeifer, G.; Goldberg, A.L.; Sakata, E.; Baumeister, W.; Förster, F. Structural characterization of the interaction of Ubp6 with the 26S proteasome. Proc. Natl. Acad. Sci. USA 2015, 112, 8626–8631.
- Huang, X.; Luan, B.; Wu, J.; Shi, Y. An atomic structure of the human 26S proteasome. Nat. Struct. Mol. Biol. 2016, 23, 778–785.
- Hanna, J.; Leggett, D.S.; Finley, D. Ubiquitin Depletion as a Key Mediator of Toxicity by Translational Inhibitors. Mol. Cell. Biol. 2003, 23, 9251–9261.
- Guterman, A.; Glickman, M. Complementary Roles for Rpn11 and Ubp6 in Deubiquitination and Proteolysis by the Proteasome. J. Biol. Chem. 2004, 279, 1729–1738.
- Hanna, J.; Meides, A.; Zhang, D.P.; Finley, D. A Ubiquitin Stress Response Induces Altered Proteasome Composition. Cell 2007, 129, 747–759.
- Koulich, E.; Li, X.; DeMartino, G.N. Relative Structural and Functional Roles of Multiple Deubiquitylating Proteins Associated with Mammalian 26S Proteasome. Mol. Biol. Cell 2008, 19, 1072–1082.
- Torres, E.M.; Dephoure, N.; Panneerselvam, A.; Tucker, C.M.; Whittaker, C.A.; Gygi, S.P.; Dunham, M.J.; Amon, A. Identification of Aneuploidy-Tolerating Mutations. Cell 2010, 143, 71–83.
- Oromendia, A.B.; Dodgson, S.E.; Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 2012, 26, 2696–2708.
- Dephoure, N.; Hwang, S.; O’Sullivan, C.; Dodgson, S.E.; Gygi, S.P.; Amon, A.; Torres, E.M. Quantitative proteomic analysis reveals posttranslational responses to aneuploidy in yeast. eLife 2014, 3, e03023.
- Wilson, S.; Bhattacharyya, B.; Rachel, R.A.; Coppola, V.; Tessarollo, L.; Householder, D.B.; Fletcher, C.F.; Miller, R.J.; Copeland, N.G.; Jenkins, N.A. Synaptic defects in ataxia mice result from a mutation in Usp14, encoding a ubiquitin-specific protease. Nat. Genet. 2002, 32, 420–425.
- Chen, P.-C.; Qin, L.-N.; Li, X.-M.; Walters, B.J.; Wilson, J.A.; Mei, L.; Wilson, S. The Proteasome-Associated Deubiquitinating Enzyme Usp14 Is Essential for the Maintenance of Synaptic Ubiquitin Levels and the Development of Neuromuscular Junctions. J. Neurosci. 2009, 29, 10909–10919.
- Walters, B.J.; Hallengren, J.J.; Theile, C.S.; Ploegh, H.L.; Wilson, S.M.; Dobrunz, L.E. A catalytic independent function of the deubiquitinating enzyme USP14 regulates hippocampal synaptic short-term plasticity and vesicle number. J. Physiol. 2014, 592, 571–586.
- Vaden, J.H.; Watson, J.A.; Howard, A.D.; Echen, P.-C.; Wilson, J.A.; Wilson, S.M. Distinct effects of ubiquitin overexpression on NMJ structure and motor performance in mice expressing catalytically inactive USP14. Front. Mol. Neurosci. 2015, 8, 11.
- Wertz, I.E.; Murray, J.M. Structurally-defined deubiquitinase inhibitors provide opportunities to investigate disease mechanisms. Drug Discov. Today Technol. 2019, 31, 109–123.
- Homma, T.; Ishibashi, D.; Nakagaki, T.; Fuse, T.; Mori, T.; Satoh, K.; Atarashi, R.; Nishida, N. Ubiquitin-specific protease 14 modulates degradation of cellular prion protein. Sci. Rep. 2015, 5, 11028.
- McKinnon, C.; Goold, R.G.; Andre, R.; Devoy, A.; Ortega, Z.; Moonga, J.; Linehan, J.M.; Brandner, S.; Lucas, J.J.; Collinge, J.; et al. Prion-mediated neurodegeneration is associated with early impairment of the ubiquitin–proteasome system. Acta Neuropathol. 2016, 131, 411–425.
- Boselli, M.; Lee, B.-H.; Robert, J.; Prado, M.A.; Min, S.-W.; Cheng, C.; Silva, M.C.; Seong, C.; Elsasser, S.; Hatle, K.M.; et al. An inhibitor of the proteasomal deubiquitinating enzyme USP14 induces tau elimination in cultured neurons. J. Biol. Chem. 2017, 292, 19209–19225.
- Xu, D.; Shan, B.; Lee, B.-H.; Zhu, K.; Zhang, T.; Sun, H.; Liu, M.; Shi, L.; Liang, W.; Qian, L.; et al. Phosphorylation and activation of ubiquitin-specific protease-14 by Akt regulates the ubiquitin-proteasome system. eLife 2015, 4, e10510.
- Chakraborty, J.; Von Stockum, S.; Marchesan, E.; Caicci, F.; Ferrari, V.; Rakovic, A.; Klein, C.; Antonini, A.; Bubacco, L.; Ziviani, E. USP14 inhibition corrects an in vivo model of impaired mitophagy. EMBO Mol. Med. 2018, 10, 11.
- Wu, N.; Liu, C.; Bai, C.; Han, Y.P.; Cho, W.C.S.; Li, Q. Over-Expression of Deubiquitinating Enzyme USP14 in Lung Adenocarcinoma Promotes Proliferation through the Accumulation of beta-Catenin. Int. J. Mol. Sci. 2013, 14, 10749–10760.
- Wang, Y.; Wang, J.; Zhong, J.; Deng, Y.; Xi, Q.; He, S.; Yang, S.; Jiang, L.; Huang, M.; Tang, C.; et al. Ubiquitin-specific protease 14 (USP14) regulates cellular proliferation and apoptosis in epithelial ovarian cancer. Med. Oncol. 2015, 32, 1–10.
- Zhu, Y.; Zhang, C.; Gu, C.; Li, Q.; Wu, N. Function of Deubiquitinating Enzyme USP14 as Oncogene in Different Types of Cancer. Cell. Physiol. Biochem. 2016, 38, 993–1002.
- Zhang, B.; Li, M.; Huang, P.; Guan, X.-Y.; Zhu, Y.-H. Overexpression of ubiquitin specific peptidase 14 predicts unfavorable prognosis in esophageal squamous cell carcinoma. Thorac. Cancer 2017, 8, 344–349.
- Hang, C.; Gong, C.; Fang, Y.; Chen, L.; Zhu, J. Ubiquitin-specific protease 14 (USP14) promotes proliferation and metastasis in pancreatic ductal adenocarcinoma. J. Mol. Histol. 2021, 52, 1–10.
- Lander, G.C.; Estrin, E.; Matyskiela, M.E.; Bashore, C.; Nogales, E.; Martin, A. Complete subunit architecture of the proteasome regulatory particle. Nat. Cell Biol. 2012, 482, 186–191.
- Chen, S.; Wu, J.; Lu, Y.; Ma, Y.-B.; Lee, B.-H.; Yu, Z.; Ouyang, Q.; Finley, D.J.; Kirschner, M.W.; Mao, Y. Structural basis for dynamic regulation of the human 26S proteasome. Proc. Natl. Acad. Sci. USA 2016, 113, 12991–12996.
- de la Peña, A.H.; Goodall, E.A.; Gates, S.N.; Lander, G.C.; Martin, A. Substrate-engaged 26S proteasome structures reveal mechanisms for ATP-hydrolysis–driven translocation. Science 2018, 362, eaav0725.
- Dong, Y.; Zhang, S.; Wu, Z.; Li, X.; Wang, W.L.; Zhu, Y.; Stoilova-McPhie, S.; Lu, Y.; Finley, D.; Mao, Y. Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nat. Cell Biol. 2018, 565, 49–55.
- Maytal-Kivity, V.; Reis, N.; Hofmann, K.; Glickman, M.H. MPN+, a putative catalytic motif found in a subset of MPN domain proteins from eukaryotes and prokaryotes, is critical for Rpn11 function. BMC Biochem. 2002, 3, 28.
- Lundgren, J.; Masson, P.; Realini, C.A.; Young, P. Use of RNA Interference and Complementation to Study the Function of the Drosophila and Human 26S Proteasome Subunit S13. Mol. Cell. Biol. 2003, 23, 5320–5330.
- Gallery, M.; Blank, J.L.; Lin, Y.; Gutierrez, J.A.; Pulido, J.C.; Rappoli, D.; Badola, S.; Rolfe, M.; Macbeth, K.J. The JAMM motif of human deubiquitinase Poh1 is essential for cell viability. Mol. Cancer Ther. 2007, 6, 262–268.
- Deshaies, R.J. Proteotoxic crisis, the ubiquitin-proteasome system, and cancer therapy. BMC Biol. 2014, 12, 94.
- Wang, B.; Ma, A.; Zhang, L.; Jin, W.-L.; Qiang, X.; Xu, G.; Qiu, B.; Yang, Z.; Liu, Y.; Xia, Q.; et al. POH1 deubiquitylates and stabilizes E2F1 to promote tumour formation. Nat. Commun. 2015, 6, 8704.
- Song, Y.; Ray, A.; Das, D.S.; Samur, M.K.; Carrasco, R.D.; Munshi, N.C.; Chauhan, D.; Anderson, K.C. Deubiquitylating Enzyme Rpn11/POH1/PSMD14 As Therapeutic Target in Multiple Myeloma. Blood 2016, 128, 4469.
- Luo, G.; Hu, N.; Xia, X.; Zhou, J.; Ye, C. RPN11 deubiquitinase promotes proliferation and migration of breast cancer cells. Mol. Med. Rep. 2017, 16, 331–338.
- Wang, C.-H.; Lu, S.-X.; Liu, L.-L.; Li, Y.; Yang, X.; He, Y.-F.; Chen, S.-L.; Cai, S.-H.; Wang, H.; Yun, J.-P. POH1 Knockdown Induces Cancer Cell Apoptosis via p53 and Bim. Neoplasia 2018, 20, 411–424.
- Wang, B.S.; Xu, X.L.; Yang, Z.J.; Zhang, L.; Liu, Y.; Ma, A.H.; Xu, G.Q.; Tang, M.; Jing, T.T.; Wu, L.; et al. POH1 contributes to hyperactivation of TGF-beta signaling and facilitates hepatocellular carcinoma metastasis through deubiquitinating TGF-beta receptors and caveolin-1. Ebiomedicine 2019, 41, 320–332.
- Yu, W.; Li, J.; Wang, Q.; Wang, B.; Zhang, L.; Liu, Y.; Tang, M.; Xu, G.; Yang, Z.; Wang, X.; et al. Targeting POH1 inhibits prostate cancer cell growth and enhances the suppressive efficacy of androgen deprivation and docetaxel. Prostate 2019, 79, 1304–1315.
- Hamazaki, J.; Iemura, S.-I.; Natsume, T.; Yashiroda, H.; Tanaka, K.; Murata, S. A novel proteasome interacting protein recruits the deubiquitinating enzyme UCH37 to 26S proteasomes. EMBO J. 2006, 25, 4524–4536.
- Qiu, X.B.; Ouyang, S.Y.; Li, C.J.; Miao, S.Y.; Wang, L.F.; Goldberg, A.L. hRpn13/ADRM1/GP110 is a novel proteasome subunit that binds the deubiquitinating enzyme, UCH37. EMBO J 2006, 25, 5742–5753.
- Yao, T.T.; Song, L.; Xu, W.; DeMartino, G.N.; Florens, L.; Swanson, S.K.; Washburn, M.P.; Conaway, R.C.; Conaway, J.W.; Cohen, R.E. Proteasome recruitment and activation of the Uch37 deubiquitinating enzyme by Adrm1. Nat. Cell Biol. 2006, 8, 994–1002.
- Yao, T.; Song, L.; Jin, J.; Cai, Y.; Takahashi, H.; Swanson, S.K.; Washburn, M.P.; Florens, L.; Conaway, R.C.; Cohen, R.E.; et al. Distinct Modes of Regulation of the Uch37 Deubiquitinating Enzyme in the Proteasome and in the Ino80 Chromatin-Remodeling Complex. Mol. Cell 2008, 31, 909–917.
- Sahtoe, D.D.; Van Dijk, W.J.; El Oualid, F.; Ekkebus, R.; Ovaa, H.; Sixma, T.K. Mechanism of UCH-L5 Activation and Inhibition by DEUBAD Domains in RPN13 and INO80G. Mol. Cell 2015, 57, 887–900.
- VanderLinden, R.T.; Hemmis, C.W.; Schmitt, B.; Ndoja, A.; Whitby, F.G.; Robinson, H.; Cohen, R.E.; Yao, T.; Hill, C.P. Structural Basis for the Activation and Inhibition of the UCH37 Deubiquitylase. Mol. Cell 2015, 57, 901–911.
- Zhang, N.Y.; Jacobson, A.D.; Macfadden, A.; Liu, C.W. Ubiquitin chain trimming recycles the substrate binding sites of the 26 S proteasome and promotes degradation of lysine 48-linked polyubiquitin conjugates. J. Biol. Chem. 2011, 286, 25540–25546.
- Chen, Y.; Fu, D.; Xi, J.; Ji, Z.; Liu, T.; Ma, Y.; Zhao, Y.; Dong, L.; Wang, Q.; Shen, X. Expression and Clinical Significance of UCH37 in Human Esophageal Squamous Cell Carcinoma. Dig. Dis. Sci. 2012, 57, 2310–2317.
- Fang, Y.; Fu, D.; Tang, W.; Cai, Y.; Ma, D.; Wang, H.; Xue, R.; Liu, T.; Huang, X.; Dong, L.; et al. Ubiquitin C-terminal Hydrolase 37, a novel predictor for hepatocellular carcinoma recurrence, promotes cell migration and invasion via interacting and deubiquitinating PRP19. Biochim. Biophys. Acta (BBA) Bioenerg. 2013, 1833, 559–572.
- Wang, L.; Chen, Y.-J.; Xu, K.; Wang, Y.-Y.; Shen, X.-Z.; Tu, R.-Q. High expression of UCH37 is significantly associated with poor prognosis in human epithelial ovarian cancer. Tumor Biol. 2014, 35, 11427–11433.
- Fang, Y.; Shen, X. Ubiquitin carboxyl-terminal hydrolases: Involvement in cancer progression and clinical implications. Cancer Metastasis Rev. 2017, 36, 669–682.
- Liu, D.; Song, Z.X.; Wang, X.Y.; Ouyang, L. Ubiquitin C-Terminal Hydrolase L5 (UCHL5) Accelerates the Growth of Endometrial Cancer via Activating the Wnt/beta-Catenin Signaling Pathway. Front. Oncol. 2020, 10, 865.
- Zhang, J.; Xu, H.; Yang, X.; Zhao, Y.; Xu, X.; Zhang, L.; Xuan, X.; Ma, C.; Qian, W.; Li, D. Deubiquitinase UCHL5 is elevated and associated with a poor clinical outcome in lung adenocarcinoma (LUAD). J. Cancer 2020, 11, 6675–6685.
- Lee, B.; Finley, D.; King, R. A High-Throughput Screening Method for Identification of Inhibitors of the Deubiquitinating Enzyme USP14. Curr. Protoc. Chem. Biol. 2012, 4, 311–330.
- Wang, Y.; Jiang, Y.; Ding, S.; Li, J.; Song, N.; Ren, Y.; Hong, D.; Wu, C.; Li, B.; Wang, F.; et al. Small molecule inhibitors reveal allosteric regulation of USP14 via steric blockade. Cell Res. 2018, 28, 1186–1194.
- Palmer, A.L.; De Jong, A.; Leestemaker, Y.; Geurink, P.P.; Wijdeven, R.H.; Ovaa, H.; Dolan, B.P. Inhibition of the Deubiquitinase Usp14 Diminishes Direct MHC Class I Antigen Presentation. J. Immunol. 2018, 200, 928–936.
- Cullen, M.; Hauck, S.; Geng, B.; Bastos, C.M.; Munoz, B.; Haeberlein, M. Proteasome Activity Enhancing Compounds. WO/2015/073528, 21 May 2015.
- Cullen, M.; Hauck, S.; Foley, M.; Tait, B.; Haeberlein, M. Proteasome Activity Enhancing Compounds. WO/2020/006269, 2 January 2020.
- Cullen, M.; Bastos, C.M.; Parks, D.; Munoz, B. Proteasome Activity Enhancing Compounds. WO/2020/006296, 2 January 2020.
- Finley, D.; King, R.W.; Lee, B.-H.; Lee, M.J.; Gahman, T.C. Tricyclic Proteasome Activity Enhancing Compounds. WO/2012/012712, 26 January 2012.
- Kemp, M. Recent Advances in the Discovery of Deubiquitinating Enzyme Inhibitors. Prog. Med. Chem. 2016, 55, 149–192.
- Chambers, R.J.; Foley, M.; Tait, B. Proteasome Activity Modulating Compounds. WO/2013/112651, 1 August 2013.
- Li, J.; Yakushi, T.; Parlati, F.; MacKinnon, A.L.; Perez, C.; Ma, Y.; Carter, K.P.; Colayco, S.; Magnuson, G.; Brown, B.; et al. Capzimin is a potent and specific inhibitor of proteasome isopeptidase Rpn11. Nat. Chem. Biol. 2017, 13, 486–493.
- Perez, C.; Li, J.; Parlati, F.; Rouffet, M.; Ma, Y.; MacKinnon, A.L.; Chou, T.-F.; Deshaies, R.J.; Cohen, S.M.; Parlati, F.; et al. Discovery of an Inhibitor of the Proteasome Subunit Rpn11. J. Med. Chem. 2017, 60, 1343–1361.
- Lauinger, L.; Li, J.; Shostak, A.; Cemel, I.A.; Ha, N.; Zhang, Y.; E Merkl, P.; Obermeyer, S.; Stankovic-Valentin, N.; Schafmeier, T.; et al. Thiolutin is a zinc chelator that inhibits the Rpn11 and other JAMM metalloproteases. Nat. Chem. Biol. 2017, 13, 709–714.
- Li, J.; Zhang, Y.; Santos, B.D.S.S.D.; Wang, F.; Ma, Y.; Perez, C.; Yang, Y.; Peng, J.; Cohen, S.M.; Chou, T.-F.; et al. Epidithiodiketopiperazines Inhibit Protein Degradation by Targeting Proteasome Deubiquitinase Rpn11. Cell Chem. Biol. 2018, 25, 1350–1358.e9.
- D’Arcy, P.; Brnjic, S.; Olofsson, M.H.; Fryknäs, M.; Lindsten, K.; De Cesare, M.; Perego, P.; Sadeghi, B.; Hassan, M.; Larsson, R.; et al. Inhibition of proteasome deubiquitinating activity as a new cancer therapy. Nat. Med. 2011, 17, 1636–1640.
- Wang, X.; D’Arcy, P.; Caulfield, T.R.; Paulus, A.; Chitta, K.; Mohanty, C.; Gullbo, J.; Chanan-Khan, A.; Linder, S. Synthesis and Evaluation of Derivatives of the Proteasome Deubiquitinase Inhibitor b-AP15. Chem. Biol. Drug Des. 2015, 86, 1036–1048.
- Kapuria, V.; Peterson, L.F.; Fang, D.; Bornmann, W.G.; Talpaz, M.; Donato, N.J. Deubiquitinase Inhibition by Small-Molecule WP1130 Triggers Aggresome Formation and Tumor Cell Apoptosis. Cancer Res. 2010, 70, 9265–9276.
- Zhou, B.; Zuo, Y.; Li, B.; Wang, H.; Liu, H.; Wang, X.; Qiu, X.; Hu, Y.; Wen, S.; Du, J.; et al. Deubiquitinase inhibition of 19S regulatory particles by 4-arylidene curcumin analog AC17 causes NF-kappaB inhibition and p53 reactivation in human lung cancer cells. Mol. Cancer Ther. 2013, 12, 1381–1392.
- Liu, N.; Li, X.; Huang, H.; Zhao, C.; Liao, S.; Yang, C.; Liu, S.; Song, W.; Lu, X.; Lan, X.; et al. Clinically used antirheumatic agent auranofin is a proteasomal deubiquitinase inhibitor and inhibits tumor growth. Oncotarget 2014, 5, 5453–5471.
- Stafford, W.C.; Peng, X.; Olofsson, M.H.; Zhang, X.; Luci, D.K.; Lu, L.; Cheng, Q.; Trésaugues, L.; Dexheimer, T.S.; Coussens, N.P.; et al. Irreversible inhibition of cytosolic thioredoxin reductase 1 as a mechanistic basis for anticancer therapy. Sci. Transl. Med. 2018, 10, eaaf7444.
- Zhang, X.; Selvaraju, K.; Saei, A.A.; D’Arcy, P.; Zubarev, R.; Arnér, E.S.; Linder, S. Repurposing of auranofin: Thioredoxin reductase remains a primary target of the drug. Biochimie 2019, 162, 46–54.
- Erdal, H.; Berndtsson, M.; Castro, J.; Brunk, U.; Shoshan, M.C.; Linder, S. Induction of lysosomal membrane permeabilization by compounds that activate p53-independent apoptosis. Proc. Natl. Acad. Sci. USA 2005, 102, 192–197.
- Berndtsson, M.; Beaujouin, M.; Rickardson, L.; Havelka, A.M.; Larsson, R.; Westman, J.; Liaudet-Coopman, E.; Linder, S. Induction of the lysosomal apoptosis pathway by inhibitors of the ubiquitin-proteasome system. Int. J. Cancer 2009, 124, 1463–1469.
- Wang, X.; Stafford, W.; Mazurkiewicz, M.; Fryknäs, M.; Brjnic, S.; Zhang, X.; Gullbo, J.; Larsson, R.; Arnér, E.S.J.; D’Arcy, P.; et al. The 19S Deubiquitinase Inhibitor b-AP15 Is Enriched in Cells and Elicits Rapid Commitment to Cell Death. Mol. Pharmacol. 2014, 85, 932–945.
- Wang, X.; Mazurkiewicz, M.; Hillert, E.K.; Olofsson, M.H.; Pierrou, S.; Hillertz, P.; Gullbo, J.; Selvaraju, K.; Paulus, A.; Akhtar, S.; et al. The proteasome deubiquitinase inhibitor VLX1570 shows selectivity for ubiquitin-specific protease-14 and induces apoptosis of multiple myeloma cells. Sci. Rep. 2016, 6, 1–15.