
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Dagmara Baraniak | + 4854 word(s) | 4854 | 2021-06-09 11:30:33 | | | |
| 2 | Camila Xu | + 98 word(s) | 4952 | 2021-06-17 10:35:21 | | | | |
| 3 | Camila Xu | + 98 word(s) | 4952 | 2021-06-17 10:36:25 | | |
Video Upload Options
Locked nucleic acids (LNAs) are widely used in RNA therapeutics as LNA-modified antisense oligonucleotides. From the structural point of view, LNA is an RNA derivative in which the ribose ring is constrained by a methylene linkage between the 2′-oxygen and the 4′-carbon. This conformational restriction causes an increased binding affinity to complementary DNA and RNA sequences, but the susceptibility to RNase H should be optimized for individual applications.
1. Introduction
The natural nucleic acids, DNA and RNA, are exquisitely suited to store and propagate genetic information [1][2]. It can mediate the heritable storage, recognize specific targets (aptamers), and even catalyze reactions [3]. Nowadays, RNA has become a focus of attention because it turned out that various RNAs are valuable tools in molecular biology and medicine. These molecules, including all classes of ncRNAs, are used in the design and development of drugs for fighting cancers, viruses, and other diseases at the nucleic acids level.
In the world of science, the key role for regulating life processes is ascribed to proteins. From the central dogma of molecular biology, it can be seen that the flow of genetic information, which takes place from DNA through RNA to protein, involves a molecule of RNA, and exactly the messenger RNA (mRNA). It follows that the mRNA molecule is coding RNA in transcription, because it rewrites information from DNA about proteins. The first one acts as a scaffold, and the second one is a courier in the delivery of amino acids that are linked into long chains to form a protein.
In turn, regulatory microRNAs (miRNAs) control gene expression, and are also involved in brain development and nerve cell specialization [4]. ; (c) catalytic (“ribozymes”)—they catalyze biochemical reactions, e.g., hammerhead, RNaseP; (d) regulatory functions—regulate gene expression, e.g., riboswitches; (e) control epigenetic phenomena [5][6]. In the treatment of human diseases, miRNA are regulators of endogenous gene expression, and thus biomarkers in cancer [7]. We see that the huge functional diversity of these molecules allows us to better understand gene expression, create new therapeutic strategies and better develop diagnostic methods for many diseases [8][9].
The concept of organic chemistry, introduced by Sharpless and Meldal, named click chemistry, lies in using simple and efficient reactions designed to bind various compounds in an easy way [10][11]. In recent years, hundreds of scientific works describing applications of click chemistry have been published, including extensive research papers. For us, the works in the field of biological chemistry are of particular importance, pertaining to synthesis of small molecules—clickable nucleosides and nucleotides [12][13][14][15][16], dimers derivatives—canonical [17][18] and non-canonical [19] linked dinucleosides, and modified oligodeoxynucleotides [20][21][22]. Among the discussed topics, other important applications can be listed, such as: receiving fluorescent markers [23][24][25][26][27][28][29][30], ligand design [31], crosslinking duplexes [32][33][34], labeling [35][36][37][38][39], conjugates [40][41][42], surface modification [43], artificial molecular machines [44], nanomaterials, and nanostructures [45][46][47].
Exploiting the CuAAC and RuAAC tools also opens access to complex architecture in biomedicinal science, for example: drug discovery [48][49][50][51], triazole-linked analogues of oligonucleotides [52], and therapeutic nucleic acids (“clicking” genes) [53][54][55] with different arrangement of the triazole linker [56], and enables gene editing by CRISP-Cas9 method [57][58][59][60].
There are many reviews discussing modified nucleic acids as therapeutic agents [61][62][63]. This results in the development of new potent antisense and antigen agents [64][65][66]; however, only extremely accurate chemistry and biochemistry can make genes and even genomes. Additionally, the standard chemical approach to synthesize oligonucleotides, namely the phosphoramidite chemistry cycle, is limited to around 100-base building blocks; beyond that length they are very difficult to make [67]. With the need for better gene synthesis, all innovative ideas attract a great deal of attention [68][69][70][71].
2. Triazole-Modified Oligonucleotides
Chemical modifications of oligonucleotides (ONs) came to prominence in the early 1990s with the advent of the antisense approach to control gene expression [72][73]. Antisense strategy as a part of controlling gene expression is a fast-growing research area. ONs can be used in antisense or triple helix therapy to inhibit RNA translation or DNA transcription. Thus, oligonucleotide analogues with triazole internucleotide linkages continue to attract significant attention on medical applications for gene silencing such as antisense, antigen [74][75], and siRNA
Therefore, its structure and possibilities of “improvement” have always intrigued scientists all over the world DNA usually forms a duplex, in other words double-stranded structure. Some elements of nucleic acids can take unusual structures, e.g., in the case of an excess of guanine base. Then, a four-strand nucleic acid structure called G-quadruplex is formed, characterized by high plasticity compared to the standard double-stranded form, which translates into their important biological functions.
Studies of the properties of modified oligonucleotides led to the development of unnatural oligomers with huge therapeutic potential, especially for the treatment of diseases characterized by the expression of unwanted genes. The most promising of such analogues are those exhibiting high binding affinity toward native DNA/RNA, since they may prove to be efficient antisense or siRNA agents [76]. Therefore, it is logical that researchers turned their attention toward the possible benefits of innovative and new synthetic approaches, such as the CuAAC methods [77].
This research presents synthesis of thymidine dinucleoside analogues with imidazole and triazole modified backbones before the era of click chemistry (Figure 1). The two communications described the detailed chemical synthesis of these compounds with an overview of the introduction into oligodeoxyribonucleotides strand in a DNA/RNA heteroduplex and evaluation of the influence of polarity and basicity of the modified backbone structure on RNA binding affinity, as well as thermodynamic properties. The backbone replacements evaluated lead to a destabilization of the duplexes formed by all modified sequences with their RNA complements. In all instances, the incorporation of heterocyclic backbone modifications leads to reduction of the thermal stability of duplexes.
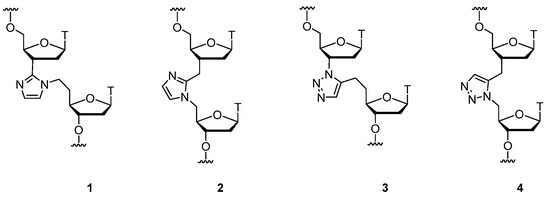
Figure 1. Thymidine dinucleoside analogues with imidazole and triazole modified backbones.
Another type of coupling between two pyrimidine analogues has been accomplished by Lazrek et al. The novel branched nucleoside dimers containing a 1,2,3-triazolyl linkage have been synthesized using 1,3-dipolar cycloaddition of N-3 or C-5 acetylene nucleosides with 3′-azido-3′-deoxythymidine (AZT). The coupling reactions under reflux always provide a mixture of the two possible 4- and 5-substituted isomers. As the authors suggest, such modified nucleosides may be of interest as potential candidates for nucleoside-based therapeutics.
Because of the important role of AZT in chemotherapy scientists are constantly looking for its valuable analogues. Also in this case, an inestimable support is given by click chemistry [77][31][55]. Dinucleosides 7 and 8, in which two thymidines are linked by triazole, have been synthesized by reacting AZT or 5′-azidothymidine with 3′-O-propargylthymidine in excellent yields. The reaction proceeds to completion at ambient temperature within 12 h.
In 2007, Nuzzi et al. published some model studies towards the synthesis of thymidine oligonucleotides with triazole internucleosidic linkages via Cu(I)-promoted ligation strategy [78]. These initial results have demonstrated a promising usefulness of such structures in search towards antisense exploitation. Also, an explorative study has been taken to prepare a trinucleotide—a triazole linked codon in a new class of oligothymidine analogues as shown by structures 9, 10, and 11 (Scheme 1). The authors the of work assure, that this study is a part of a longstanding research program to develop antisense nucleotides in their laboratory.
Subsequent applications of such interesting oligomers may be found in the literature already dedicated to modified ONs. The click reaction is used to ligation of two single stranded oligdeoxynucleotides (ODNs), one with a 5′-alkyne and the other with a 3′-azide (Scheme 2). A covalent catenation leads to intramolecular circularization and produces a DNA strand with an unnatural extended backbone at the ligation point. The product is obtained at room temperature within 2 h.
Another exciting example of CuAAC templated nucleic acid reactions uses 3′-O-propargyl 5-methyldeoxycytidine at the 3′-terminus of an ODN strand, and a 5′-azide at the 5′-strand terminus [79]. Thus, DNA strands with these terminal groups can be used to construct genes that are subsequently translated and transcribed in E. coli [80][79]. Moreover, using the CuAAC approach, epigenetic modifications can be incorporated into the gene to determine the influence of such trial [80]. This work indicates that CuAAC of 5′-azido and 3′-O-propargyl modifications might emerge as a non-enzymatic tool for ligation that could complement or replace ligase systems [79][81][82].
Cyclic DNA and RNA present unusual chemical and biological properties in comparison with linear counterparts, and have been evaluated for several biological applications, including antisense, triplex, and diagnostic ones. In turn, triazole-linked dumbbell ODNs can exist as potential decoy molecules [83]. The newly synthesized dumbbell ODNs showed excellent stability against 3′-exonuclease and high thermal stability, which are necessary for decoy molecules to achieve biological responses leading to alteration of gene expression. This click methodology can be valuable in nanotechnological applications involving DNA scaffolds [84][85].
Another scientific idea was presented in order to investigate the efficiency of sugar labeling of DNA under various conditions [86]. For this purpose, synthetic alkyne-modified nucleoside triphosphates (dU*TP) were incorporated into long DNA strands by the PCR reaction, as shown in Scheme 3. This result showed the decreased click reaction yield for DNA modified with short-chain alkyne nucleoside (ethynyluridine), rather than for DNA containing alkyne-modified uridine with an attached longer spacer. This work is of great practical value because labelled DNA are used in biomedical applications, for example in genetic analysis, genome screening, and high-throughput DNA sequencing.
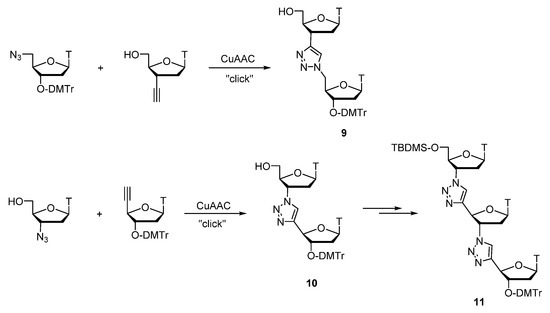
Scheme 1. Synthesis of non-natural dinucleotides and trinucleotide.
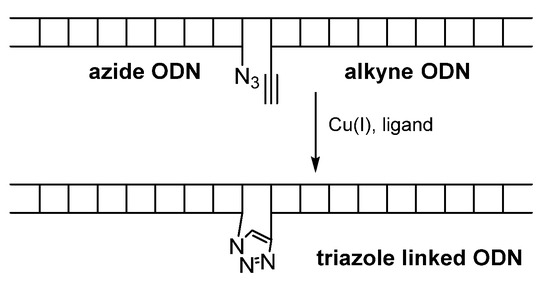
Scheme 2. Schematic template-mediated click-ligation of two oligonucleotides.
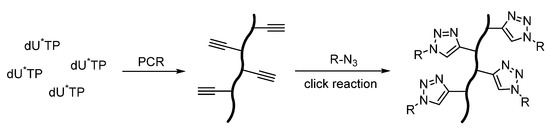
Scheme 3. Illustrative depiction of DNA functionalization using click chemistry. R–N3 any azide-molecule, dU∗TP is a uridine derivative modified with a terminal alkyne functionality.
Interesting methods of DNA modification can be achieved in post synthetic labeling [87]. That is why it is possible to attach alkynyl-labels, which results in DNA bearing various modifications; however, the copper ions damage DNA by breaking strands. The next limitation is that the direct incorporation of azides into synthetic DNA strands is intrinsically difficult, because the azide group reacts with the phosphorus(III) atom of the phosphoramidite group. Here, we can notice that properly prepared building blocks–nucleoside dimers may be very helpful, as new nucleic acid motifs earlier described in the literature [17][18][19][88][83][87][89][90][91][92].
As a part of the study of modified DNA, Seela and co-workers got interested in the use of CuAAC as an efficient way to label DNA [93]. One of such examples is presented below and applies in receiving oligonucleotides containing the octadiynyl derivative of dU with AZT In their research, the fluorescence properties of oligonucleotides for the visualization of DNA was under special emphasis [94][95][96]. In particular, the generation of fluorescence by the click reaction [88][94], and studying the mechanisms of its quenching, were presented [96][97].
This highly efficient method allowed for 1,3-dipolar cycloaddition reactions to be carried out on fully deprotected ONs, affording 1,4-regioisomeric 1,2,3-triazoles in a very short time (10–15 min). First examples of such reactions have been investigated for 3′,5′-dithymidine in order to reduce the reaction time [98][99]. Both systems led to products, but during reaction using CuI the formation of a side product was observed at room temperature. Click reaction using CuSO4 gave a single product, same as in the case of classical heating (80 °C, 5 h) and MW (80 °C, 200 W, 3 min), giving identical results (80% yields).
Further scientific reports show the research directions of triazole-linked nucleoside for longer chains design and synthesis, and even double-strand formation of such modified nucleic acids. These scientists managed to get the artificial 10-mer TLDNA on a solid-phase synthesis, that after examined could form a stable double strand with the complementary strand of natural DNA (Scheme 4). Because similar triazole linker with a longer methylene bridge destabilizes the double strand, the six-bond periodicity of TLDNA is crucial to form stable double strands. [100], which is aimed at preparation in a larger scale of triazole-oligomers, and even for other deoxyribonucleoside analogues bearing adenine, cytosine, and guanine nucleobases [101].
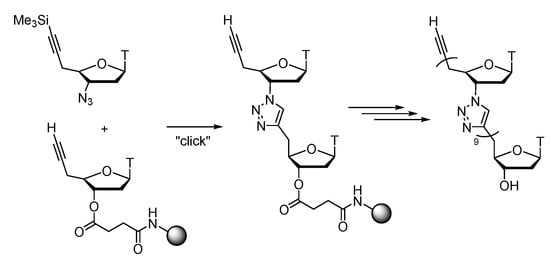
Scheme 4. Solid-phase synthesis of 10-mer TLDNA.
The same researchers obtained dithymidine phosphoramidite analogues with a triazole linker and an additional methylene group of type 13, which was then incorporated in oligonucleotides using solid-phase synthesis with standard phosphoramididte protocols [100]. They obtained a new dinucleotide with a triazole internucleotide modification type 14, that demonstrated DNA binding affinities similar to those of unmodified oligonucleotides, with simultaneous protection from nuclease hydrolysis due to the modification (Figure 2). Then, the UV-melting experiments proved a slight destabilization effect of duplexes (almost 4 times lower than for other triazole modifications). This finding shows a great potential of these new oligonucleotide analogues as the most promising triazole DNA mimics in terms of hybridization properties, compatibility with biosystems, and stability in biological liquids.
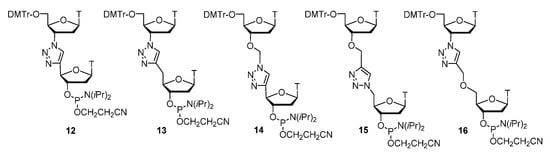
Figure 7. Phosphoramidite triazole-linked dimers T–T.
At the same time, El-Sagheer and Brown worked on DNA strands containing an unnatural triazole linkage, that had been synthesized by click ligation between ONs with AZT and 4′-propargyloamido-5′-derivative of 5′-deoxymethylthymidine (Scheme 5) [102]. Hence, the search for a polymerase-compatible triazole linkage with the initial experiments focused on non-enzymatic DNA strand ligation combined with reproducible amplification. An additional advantage of the triazole units is fact, that this chemical linkage is compatible with enzymatic processing, in particular PCR, and does not suffer from degradation due to endo- and exonucleases. This is the first time that highly efficient non-enzymatic DNA strand ligation has been combined with reproducible amplification.
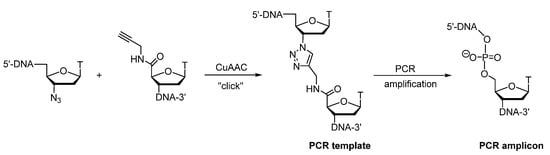
Scheme 5. Synthesis and polymerase chain reaction amplification of DNA strands containing an unnatural triazole linkage.
In recent years, this research group has dealt with the subject of click chemistry in the nucleic acids field in a diverse range of applications, namely: strand ligation [56][103] and cross-linking using CuAAC [104] or even receiving stable cyclic DNA miniduplexes [105]. So, within 10 years, they explained the correlation between structure and dynamics [106][107][108]; proved the biocompatibility of the artificial linker and the fact it could mimic the DNA phosphodiester group [109]; showed efficient RNA synthesis by a triazole-modified DNA template [110]; synthesized hairpin and hammerhead ribozymes with the triazole linkage [111]; obtained a click-linked gene that was functional in human cells [112], and thus confirmed that the importance of epigenetically modified genes and genomes increases [80]. It is astonishing in the case of triazole-modified oligodeoxynucleotides, that their influence on thermodynamic destabilization of the duplexes, despite the artificial linkage (that is produced by click ligation) is read through correctly and efficiently by DNA polymerases, and is functional in vivo. This is also the first demonstration of transcription through a heavily modified DNA backbone linkage and suggestion that click-ligated DNA could be useful for the direct synthesis of biologically active RNA and proteins.
Such an alkynylphosphonate clickable backbone has versatile possibilities of biological applications. In this way, it was possible to obtain oligonucleotide chimeras built from 16 to 23 mers with various modifications. Further studies allowed to state that such modified systems were resistant to 5′-and 3′-exonucleoases, and the stability of duplexes with RNA did not change significantly. The results of these experiments may encourage to continue the investigation of TP ODNs as potential antagomirs for targeting microRNA expression.
They can also act as valuable therapeutic tools in gene regulation, e.g., siRNA Additionally, oligonucleotides with propargylated nucleoside as clickable moieties and a bifunctional azide have been synthesized by the CuAAC reaction [113]. This strategy has been used to obtain cross-linking in DNA–DNA and DNA–RNA duplexes for study of their stability and hybridization properties. This procedure shows how to receive 2′-O-propargyl-2-(amino)adenosine phosphoramidite to the formation of homodimers and heterodimers while pointing to significant stability for duplexes with terminal ligation.
Using click chemistry, small molecules are applied for creating new RNA conjugates [114][115][116], for example, alkyne-modified ATP [117] or SAM-adenosine conjugates [118]. Other, less frequent uses of triazolo-modified RNAs are applied to searching for chemical probes in nucleic acid chemistry [119][120] and in physicochemical methods like EPR spectroscopy [121] and cat-ELCCA assays [122] in biomedical sciences.
Catalytic RNAs, named ribozymes, can catalyze cleavage or ligation of phosphodiester bonds, and many other reactions in all life forms [123]. They described a strategy for the synthesis of chemically modified RNA constructs exemplified by hairpin and hammerhead ribozymes and DNA– In the same study, the constructed DNA–RNA hybrid duplexes had the same conformation and stability as their natural counterparts. It proved the ability to synthesize long RNA strands by a combination of solid-phase support and click ligation, resulting in the synthesis of many biologically important RNA molecules.
As it turned out, click chemistry has enabled the large DNA strands to produce biologically active RNA and proteins. The modified nucleic acid containing triazole linkage is correctly read-through by DNA polymerases and is functional in bacteria [109]. This work shows that T7-RNAP enzyme (T7 RNA polymerase) can transcribe through modified DNA strands to synthesize RNA, which is evidence for the surprising biocompatibility of TLDNA. Therefore, click ligation is a strategy for overcoming the size limit of DNA and RNA obtained by solid-phase phosphoramidite chemistry (without the use of enzymes), and DNA constructs can be used directly to synthesize biologically active RNA constructs and proteins.
Chemical synthesis of RNA is less efficient than DNA owing to problems caused by the presence of the 2′-hydroxyl group of ribose, which requires selective protection during oligonucleotide assembly. Here, it is worth noting that most biologically important RNA molecules like ribozymes, aptamers, and riboswitches are significantly longer. Therefore, in most cases of oligonucleotide modification with CuAAC, nucleosides containing an alkyne substituent are introduced into the chain. The rare and exceptional 2′-azido modifications are compatible with 2′-fluoro and/or 2′-O-methyl modifications to achieve siRNAs of rich modification patterns and tunable properties.
To evaluate the effect of triazole linkage on thermal stability of RNA, they synthesized several self-complementary sequences. The UV thermal melting analysis, CD, and NMR spectra showed that the triazole linkage strongly destabilized RNA double helix by about 7–14 °C depending on the sequence and location of the modifications. In contrast, Isobe showed how long triazole linkage stabilized DNA [124][125]. On the other hand, RNA tolerates better the triazole linkage than DNA, despite being two atoms longer than the phosphate [126].
TLDNA has been demonstrated to be a competent enzymatic substrate, but the biologically non degradable linkage of TLRNA may provide an intriguing opportunity to explore the sophisticated function of RNA, for instance, RNA interference, that requires the enzymatic recognition of the oligonucleotides [127]. The design and synthesis of such constructs will certainly be investigated in the near future.
In bioconjugation chemistry the CuAAC allows for rapid labeling and ligation of RNA [128][129]. Once clickable groups are installed on RNA with free 2′-hydroxyl groups, they can be rapidly click labeled or conjugated together for biological studies. For example, triazole-linked lipid-oligonucleotide conjugates have been shown to be nontoxic, and this displays a broad applicability of click chemistry to any synthetic or natural RNA [40]. This strategy enables the capture dynamic aspects of RNA metabolism in living systems [129] and to study of the epitranscriptome marks such as new detection methods for m6A or mechanism-based trapping of an inosine-generating ADAR enzyme [130].
There are two major tools of biological significance in the current RNA research, namely RNA interference (RNAi) and RNA bioconjugation. Here, the structural and functional repertoire of RNAs can be significantly manipulated by chemical modifications. There are high expectations for the development of siRNA and miRNA as therapeutics agents to treat diseases [131][132]. as therapeutic agents, chemical modification is obligatory to enhance nuclease resistance, to prevent immune activation, to decrease off-target effects, and to improve pharmacokinetic and pharmacodynamic properties.
Utilizing the well-known CuAAC reaction, the triazole backbone dimers (UtU and CtU) based on a PNA-like structure were synthesized and incorporated into various siRNAs targeting representative gene transcripts of an exogenous and endogenous gene (Scheme 6). The triazole-modified siRNAs were found to be capable of gene silencing of reporter genes and thus proved the compatibility as substrates within the RNAi pathway. The modifications were designed to impart potentially favorable properties to siRNAs in order to target endogenous genes, for instance the oncogene BCL2. duplexes and mimics the silencing of gene expression when positioned within the internal Watson–Crick double-stranded region of siRNAs.
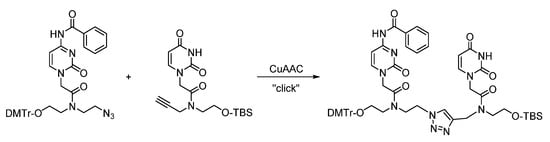
Scheme 6. Synthesis of triazole-linked CTRU dimer based on a PNA-type scaffold.
Short-interfering RNAs (siRNAs) are naturally occurring biomolecules used for post-transcriptional gene regulation, so they can be promising therapeutic agents by silencing gene expression of overexpressed deleterious genes. The chemically modified siRNAs enhance gene-silencing activity and biological stability with high nuclease resistance, and keep a similar immune response as unmodified siRNAs. RNA interference might be a promising alternative to conventional chemotherapy of genetic disorders, cancer, viral infections, and other diseases for which the target mRNA can be identified. However, the short interfering RNAs (siRNAs) need to be chemically modified to be efficient therapeutic agents, in order to improve nuclease resistance, crossing of cell membranes, biodistribution, and pharmacokinetics.
Chemically modified mRNA cap structures show very promising anti-cancer properties and therefore, are being extensively studied as a target for drug discovery. mRNA Large RNAs, such as long non-coding RNAs or mRNAs, have exciting potential as gene therapy drugs or vaccine antigens, and for such molecules post-transcriptional chemical modification will be crucial. The novel class of dinucleotide cap analogues containing a triazole ring within the oligophosphate chain are more durable and act with efficiency similar to their natural counterparts.
This technology has revolutionized the molecular biology—it creates new possibilities for plant breeding or is contributing to new innovative therapies for cancer and inherited diseases [133]. Additionally, in this case, scientists are looking for new solutions using click chemistry approach [60] It gives the opportunities to study pervasive antisense transcription, namely the use of CuAAC reaction to construct DNA templates for sgRNA expression, thus generating novel sense and antisense transcripts. Furthermore, click chemistry is an efficient method for sgRNA template construction which combined with dCas9 allows for advanced gene expression analysis.
Campbell and Wengel have found that oligonucleotides containing only LNA units form hybrids with RNA that activate RNase H to a lower degree than corresponding DNA oligonucleotides [134]. That is why one of the most promising modifications of antisense and/or antigen oligonucleotides is the introduction of triazole or triazole-bridged nucleic acids into LNAs [135][136]. It is worth emphasizing that bridged nucleic acids (BNAs) are attractive construction placed beyond LNAs and peptide nucleic acids (PNAs), with high binding affinity to ssRNA Novel attributes of LNAs and other classes of bridged nucleic acid analogues are extensively explored in different synthetic oligonucleotide-based therapeutics, so straightforward synthesis of their monomers could be very useful
First attempts towards design and synthesis of LNA-based nucleoside dimers using CuAAC reaction were done in the laboratory of Prasad [137][138] (Scheme 7). The optimized method can easily be adopted for automated synthesis of triazole-linked LNA-based oligonucleotides. for their biological and biophysical properties through incorporation into gene-silencing oligonucleotides, namely antisense oligonucleotides (ASOs) and double-stranded siRNAs [139]. They presented accurate analyzes of good hybridization affinity (when incorporated at the 3′ or 5′ termini of siRNAs, but with extremely low binding affinity at internal positions), outstanding nuclease stability, and high gene silencing activity by both RNase H and Ago2 based mechanisms.
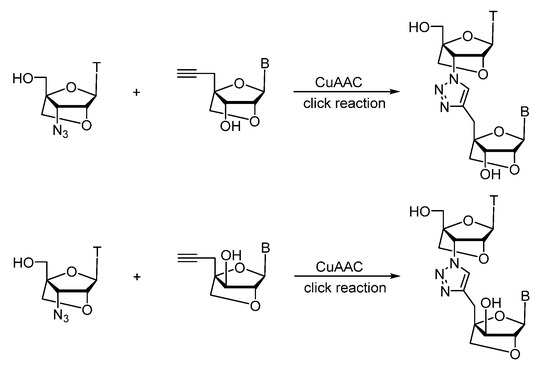
Scheme 7. Synthesis of triazole-linked LNA-dimers using Cu(I)-catalyzed click reaction.
Independently, Kumar et al. have studied the combination of triazole-linked LNAs and six-atom triazole linkage [140][141] It is known that oligonucleotides containing triazole linkage form less stable duplexes with complementary RNA/DNA targets compared to unmodified DNA strands; however (TL)LNAs with internal triazole-3′-LNA linkages bind to complementary RNA with similar affinity and specificity to unmodified oligonucleotides, and the introduction of triazole moiety significantly improves the thermal stability of the modified duplex. The experiment has also shown that the (TL)LNAs with LNA on either side of the triazole linkage have the highest stability against degradation by DNase, in addition to the strongest affinity for RNA targets. Thus, the combination of LNA and the triazole linkage provides a new class of potentially significant antisense oligonucleotide candidates.
From a chemical point of view, PNA molecules resemble more peptides than nucleic acids. Despite the differences in relation to nucleic acids, PNA molecules with a strictly defined sequence show the ability to bind to complementary DNA or RNA molecules using Watson–Crick hydrogen bonds, blocking or reducing in this way the expression of relevant genes [142][143]. These compounds, due to the formation of stronger complementary bonds with DNA or RNA, were considered as excellent candidates for antisense drugs in gene therapy since their discovery [144][145]. Therefore, prior to the introduction of such drugs, chemical modifications of PNAs or appropriate vectors should be developed to increase the bioavailability of these new antisense drugs [146][147].
PNA oligomers possessing an azide terminal group were obtained by standard Fmoc procedure [148], and then coupled with various alkyne monomers in the CuAAC reaction with various alkyne monomers. In the main product, dimer 20, the modification in the form of a triazole linker shows a little effect on the hybridization ability of the modified PNA strand and the sequence accuracy of the synthesized strand. These results show that the triazole group is a good substitute for an amide linker in PNA, and such clickable peptide nucleic acids (cPNA) are an important addition in nucleic acid applications.
Among others, two research groups have published synthetic protocols for new PNA oligonucleotides that contain a triazole linkage [149][150]. Efthymiou and Desaulniers showed a modified uracil–uracil dimer and its compatibility with hybridization to a complementary sequence when appended at the 5′ or 3′ end of DNA oligonucleotide. The second mentioned work, which has been performed in Zerrouki’s laboratory, displays the synthesis of a very interesting azido-heterotrimer, which may undergo further click reaction providing polytriazole PNA analogues. All these efforts have been made in order to synthesize chemically modified oligonucleotides with improved properties, and the results confirm that 1,4-disubstituted 1,2,3-triazole is a good mimic of a trans-amide bond.
G-quadruplexes (G4s) are a family of nucleic acid structures based on the formation of G-quarters of four guanines that are arranged in a plane to form several G-tetrad, stabilized by hydrogen bonds and a metal cation. G-quadruplexes represent unusual four-stranded structures found in the human DNA and RNA molecules which show unique properties including resistance to nucleases and fast folding. Interestingly, RNA and PNA also formed RNA–PNA hybrids of G-quadruplex structures [151]. Their main place of occurrence are genome regions with high transcriptional activity, the promoter regions and telomeres, so they undoubtedly take part in the regulation of gene expression.
G-quadruplexes are components of biologically important structures, they are commonly found in telomeres and oncogenes, which translates into their functions in ageing and disease development, including malignant tumors [152]. Such DNA–RNA hybrid G-quadruplex structure may also be a valuable target for anti-cancer agents directed against telomeres [153]. Telomeres are present at the ends of all eukaryotic chromosomes and play an important role in critical processes underlying genome stability, ageing, and cancer [154]. Understanding the chemistry of human telomere DNA and RNA biology will allow for the developing of chemical approaches to discover anticancer agents [155].
The related enzyme, telomerase, was first recognized as a unique and exciting anticancer target about five years after its discovery. No other tumor-associated gene is as widely expressed as telomerase in cancers. He has stated that in the future it will be possible to create synthetic molecules that block the proliferation of cells during the growth of cancerous tissue. The clinical experiments’ evidence for use of telomerase or telomeres as targets in cancer therapy are encouraging and are exciting prospects in future medical strategies.
DNA– RNA hybrid G-quadruplex structure is technically difficult to study by traditional methods, such as NMR and crystallography, since various G-quadruplexes may coexist as a mixture. This reaction can trap a particular species or produce a snapshot of various structures that are present in a complex solution [156] (Scheme 17). The detected “azido–alkyne cycloaddition” product of the alkyne- and azido-labeled RNA and DNA coupling identifies that DNA–RNA hybrid G-quadruplex structure can be formed from human telomeric DNA and RNA sequences.
Click chemistry can be a very useful method for a search, identification, and designing therapeutics, and above all, studying the functions of these biomolecules [156][157]. DNA aptamers bearing triazole internucleotide linkages can be also found in such applications as novel thrombin-binding aptamers for anticoagulant effects [158], human telomere biology [155][159][160], and furthermore, to design and synthesize a click-light-up probe to investigate whether DNA-RNA G-quadruplexes exist in living cells [161]. Human telomere structures are difficult to probe in living cells because the concentration of G-quadruplexes is too low to be detected in the presence of only a few dozen chromosome ends, so employing a light-switching probe is possible to find the process of DNA–RNA G-quadruplex formation [159]. Owing to the fact that DNA aptamers are increasingly recognized as drug candidates, herein click chemistry also enables the synthesis and development of small molecules that target G-quadruplex nucleic acids to elucidate drug specificity [157].
3. Conclusions
In conclusion, oligonucleotides modified at the phosphodiester linkage are important class of compounds, especially in the field of antisense and antigen strategies. We are interested in the replacement of the phosphodiester linkage of the natural oligonucleotides with novel, neutral, and heterocyclic linkages based on their stability and rigidity, and therefore, report here an efficient synthesis of the 1,2,3-triazole units to prepare triazolo-modified oligonucleotides.
In this review we have demonstrated the usefulness of the CuAAC or RuAAC ligation strategy in nucleic acid chemistry. This shows the importance of non-natural oligonucleotides acting as inhibitors of target deleterious genes responsible for various diseases. The main goal of this work was to present efficient routes to design nucleic acids and their components dedicated for gene expression control. The third generation of nucleic acids therapeutics—newest and most promising with enhanced binding affinity and biostability—are PNAs, LNAs, tcDNA, and CeNA.
Fundamental knowledge from many fields, including chemistry, structural biology, cell biology, and medicine, has opened up the possibility of development of novel strategies against cancer and other diseases. This review highlights the contribution of click chemistry to drug development, but also to labelling and imaging for better visualization. The click chemistry covers chemical synthesis of small molecules as well as oligonucleotides by the bio-orthogonal ligation process and their application as drugs and tools in molecular medicine. Thus, click reaction is one of the most popular interdisciplinary reaction that bridges the gap between chemistry and biology.
References
- Becker, S.; Schneider, C.; Crisp, A.; Carell, T. Non-canonical nucleosides and chemistry of the emergence of life. Nat. Commun. 2018, 9, 5174.
- Lönnberg, H. Chemistry of Nucleic Acids; De Gruyter: Berlin, Germany, 2020.
- Mattick, J.S. The Genetic Signatures of Noncoding RNAs. PLoS Genet. 2009, 5, 1–12.
- Turner, J.D.; Williamson, R.; Almefty, K.K.; Nakaji, P.; Porter, R.; Tse, V.; Kalani, M.Y. The many roles of microRNAs in brain tumor biology. Neurosurg. Focus 2010, 28, E3.
- Zhang, J.; Lau, M.W.; Ferré-D’Amaré, A.R. Ribozymes and Riboswitches: Modulation of RNA Function by Small Molecules. Biochemistry 2010, 49, 9123–9131.
- Garst, A.D.; Edwards, A.L.; Batey, R.T. Riboswitches: Structures and mechanisms. Cold Spring Harb. Perspect. Biol. 2011, 3, a003533.
- Conde, J.; Edelman, E.R.; Artzi, N. Target-responsive DNA/RNA nanomaterials for microRNA sensing and inhibition: The jack-of-all-trades in cancer nanotheranostics? Adv. Drug Deliv. Rev. 2015, 81, 169–183.
- Storz, G. An Expanding Universe of Noncoding RNAs. Science 2002, 296, 1260–1263.
- Kuhlmann, J.D.; Baraniskin, A.; Hahn, S.A.; Mosel, F.; Bredemeier, M.; Wimberger, P.; Kimmig, R.; Kasimir-Bauer. S. Circulating U2 small nuclear RNA fragments as a novel diagnostic tool for patients with epithelial ovarian cancer. Clin. Chem. 2014, 60, 206–213.
- Rostovtsev, V.V.; Green, L.G.; Fokin, V.V.; Sharpless, K.B. A Stepwise Huisgen Cycloaddition Process: Copper(I)-Catalyzed Regioselective “Ligation” of Azides and Terminal Alkynes. Angew. Chem. Int. Ed. 2002, 41, 2596–2599.
- Tornøe, C.W.; Christensen, C.; Meldal, M. Peptidotriazoles on solid phase: [1–3]-triazoles by regiospecific copper(i)-catalyzed 1,3-dipolar cycloadditions of terminal alkynes to azides. J. Org. Chem. 2002, 67, 3057–3064.
- Zavgorodny, S.G.; Pechenov, A.E.; Shvets, V.I.; Miroshnikov, A.I. S,X-Acetals in Nucleoside Chemistry. III1. Synthesis of 2′-and 3′-O-Azidomethyl Derivatives of Ribonucleosides. Nucleos. Nucleot. Nucl. 2000, 19, 1977–1991.
- Baraniak, D.; Baranowski, D.; Ruszkowski, P.; Boryski, J. 3’-O- and 5’-O-Propargyl derivatives of 5-fluoro-2’-deoxyuridine: Synthesis, cytotoxic evaluation and conformational analysis. Nucleos. Nucleot. Nucl. 2016, 35, 178–194.
- Castro, V.; Rodríguez, H.; Albericio, F. CuAAC: An Efficient Click Chemistry Reaction on Solid Phase. ACS Comb. Sci. 2016, 18, 1–14.
- Kropp, H.M.; Dürr, S.L.; Peter, C.; Diederichs, K.; Marx, A. Snapshots of a modified nucleotide moving through the confines of a DNA polymerase. Proc. Natl. Acad. Sci. USA 2018, 115, 9992–9997.
- Lingala, S.; Nordstrøm, L.U.; Mallikaratchy, P.R. Synthesis of stable azide and alkyne functionalized phosphoramidite nucleosides. Tetrahedron Lett. 2019, 60, 211–213.
- Michalska, L.; Wawrzyniak, D.; Szymańska-Michalak, A.; Barciszewski, J.; Boryski, J.; Baraniak, D. Synthesis and biological assay of new 2′-deoxyuridine dimers containing a 1,2,3-triazole linker. Part, I. Nucleos. Nucleot. Nucl. 2019, 38, 218–235.
- Baraniak, D.; Baranowski, D.; Ruszkowski, P.; Boryski, J. Nucleoside dimers analogues with 1,2,3-triazole linkage: Conjugation of floxuridine and thymidine provides novel tools for cancer treatment. Part II. Nucleos. Nucleot. Nucl. 2019, 38, 807–835.
- Baraniak, D.; Ruszkowski, P.; Baranowski, D.; Framski, G.; Boryski, J. Nucleoside dimers analogs containing floxuridine and thymidine with unnatural linker groups: Synthesis and cancer line studies. Part III. Nucleos. Nucleot. Nucl. 2019, 38, 980–1005.
- Jawalekar, A.M.; Malik, S.; Verkade, J.M.M.; Gibson, B.; Barta, N.S.; Hodges, J.C.; Rowan, A.; van Delft, F.L. Oligonucleotide Tagging for Copper-Free Click Conjugation. Molecules 2013, 18, 7346–7363.
- Sharma, V.K.; Sharma, R.K.; Singh, S.K. Antisense oligonucleotides: Modifications and clinical trials. Med. Chem. Commun. 2014, 5, 1454–1471.
- Guo, F.; Li, Q.; Zhou, C. Synthesis and biological applications of fluoro-modified nucleic acids. Org. Biomol. Chem. 2017, 15, 9552–9565.
- Chittepu, P.; Sirivolu, V.R.; Seela, F. Nucleosides and oligonucleotides containing 1,2,3-triazole residues with nucleobase tethers: Synthesis via the azide-alkyne ‘click’ reaction. Bioorg. Med. Chem. 2008, 16, 8427–8439.
- Guo, J.; Xu, N.; Li, Z.; Zhang, S.; Wu, J.; Kim, D.H.; Marma, M.S.; Meng, Q.; Cao, H.; Li, X.; et al. Four-color DNA sequencing with 3′-O-modified nucleotide reversible terminators and chemically cleavable fluorescent dideoxynucleotides. Proc. Natl. Acad. Sci. USA 2008, 105, 9145–9150.
- Okholm, A.; Kjems, J.; Astakhova, K. Fluorescence detection of natural RNA using rationally designed “clickable” oligonucleotide probes. RSC Adv. 2014, 4, 45653–45656.
- Horisawa, K. Specific and quantitative labeling of biomolecules using click chemistry. Front. Physiol. 2014, 5, 457.
- Saito, Y.; Hudson, R.H.E. Base-modified fluorescent purine nucleosides and nucleotides for use in oligonucleotide probes. J. Photochem. Photobiol. C 2018, 36, 48–73.
- Venkatesham, A.; Pillalamarri, S.R.; Wit, F.D.; Lescrinier, E.; Debyser, Z.; Van Aerschot, A. Propargylated Purine Deoxynucleosides: New Tools for Fluorescence Imaging Strategies. Molecules 2019, 24, 468.
- Bag, S.S.; Das, S.K. Triazolyl C-nucleosides via the intermediacy of β-1′-ethynyl-2′-deoxyribose derived from a Nicholas reaction: Synthesis, photophysical properties and interaction with BSA. Tetrahedron 2019, 75, 3024–3037.
- Klimkowski, P.; De Ornellas, S.; Singleton, D.; El-Sagheer, A.H.; Brown, T. Design of thiazole orange oligonucleotide probes for detection of DNA and RNA by fluorescence and duplex melting. Org. Biomol. Chem. 2019, 17, 5943–5950.
- Prasher, P.; Sharma, M. Tailored therapeutics based on 1,2,3-1H-triazoles: A mini review. Med. Chem. Commun. 2019, 10, 1302–1328.
- Pujari, S.S.; Seela, F. Parallel Stranded DNA Stabilized with Internal Sugar Cross-Links: Synthesis and Click Ligation of Oligonucleotides Containing 2′-Propargylated Isoguanosine. J. Org. Chem. 2013, 78, 8545–8561.
- Xiong, H.; Seela, F. Cross-Linked DNA: Site-Selective “Click” Ligation in Duplexes with Bis-Azides and Stability Changes Caused by Internal Cross-Links. Bioconjug. Chem. 2012, 23, 1230–1243.
- Tera, M.; Taji, Z.H.; Luedtke, N.W. Intercalation-enhanced “Click” Crosslinking of DNA. Angew. Chem. Int. Ed. 2018, 130, 15631–15635.
- Best, M.D. Click Chemistry and Bioorthogonal Reactions: Unprecedented Selectivity in the Labeling of Biological Molecules. Biochemistry 2009, 48, 6571–6584.
- Grammel, M.; Hang, H. Chemical reporters for biological discovery. Nat. Chem. Biol. 2013, 9, 475–484.
- Ren, X.; Gerowska, M.; El-Sagheerac, A.H.; Brown, T. Enzymatic incorporation and fluorescent labelling of cyclooctyne-modified deoxyuridine triphosphates in DNA. Bioorg. Med. Chem. 2014, 22, 4384–4390.
- Seo, S.; Onizuka, K.; Nishioka, C.; Takahashi, E.; Tsuneda, S.; Abe, H.; Ito, Y. Phosphorylated 5-ethynyl-2′-deoxyuridine for advanced DNA labeling. Org. Biomol. Chem. 2015, 13, 4589–4595.
- Sallustrau, A.; Bregant, S.; Chollet, C.; Audisio, D.; Taran, F. Scalable and practical synthesis of clickable Cu-chelating azides. Chem. Commun. 2017, 53, 7890–7893.
- Godeau, G.; Staedel, C.; Barthélémy, P. Lipid-Conjugated Oligonucleotides via “Click Chemistry” Efficiently Inhibit Hepatitis C Virus Translation. J. Med. Chem. 2008, 51, 4374–4376.
- Singh, Y.; Murat, P.; Defrancq, E. Recent developments in oligonucleotide conjugation. Chem. Soc. Rev. 2010, 39, 2054–2070.
- Zheng, M.; Zheng, L.; Zhang, P.; Li, J.; Zhang, Y. Development of Bioorthogonal Reactions and Their Applications in Bioconjugation. Molecules 2015, 20, 3190–3205.
- Neumann, S.; Biewend, M.; Rana, S.; Binder, W.H. The CuAAC: Principles, Homogeneous and Heterogeneous Catalysts, and Novel Developments and Applications. Macromol. Rapid Commun. 2020, 41, 1900359.
- Erbas-Cakmak, S.; Leigh, D.A.; McTernan, C.T.; Nussbaumer, A.L. Artificial Molecular Machines. Chem. Rev. 2015, 115, 10081–10206.
- Chandrasekaran, A.R.; Rusling, D.A. Triplex-forming oligonucleotides: A third strand for DNA nanotechnology. Nucleic Acids Res. 2018, 46, 1021–1037.
- Valsangkar, V.; Chandrasekaran, A.R.; Wang, R.; Haruehanroengra, P.; Levchenko, O.; Halvorsen, K.; Sheng, J. Click-based functionalization of a 2′-O-propargyl-modified branched DNA nanostructure. J. Mater. Chem. B 2017, 5, 2074–2077.
- Takezawa, Y.; Shionoya, M. Supramolecular DNA Three-Way Junction Motifs with a Bridging Metal Center. Front. Chem. 2020, 7, 925.
- Thirumurugan, P.; Matosiuk, D.; Jozwiak, K. Click chemistry for drug development and diverse chemical-biology applications. Chem. Rev. 2013, 113, 4905–4979.
- Garner, A.L. at-ELCCA: Catalyzing Drug Discovery Through Click Chemistry. Chem. Commun. 2018, 54, 6531–6539.
- Corso, A.D.; Pignataro, L.; Belvisi, L.; Gennari, C. Innovative Linker Strategies for Tumor-Targeted Drug Conjugates. Chem. Eur. J. 2019, 25, 14740–14757.
- Rodrigues, T.; Bernardes, G.J.L. Machine learning for target discovery in drug development. Curr. Opin. Chem. Biol. 2020, 56, 16–22.
- Isobe, H.; Fujino, T. Triazole-linked analogues of DNA and RNA (TLDNA and TLRNA): Synthesis and functions. Chem. Rec. 2014, 14, 41–51.
- Astakhova, K.; Ray, R.; Taskova, M.; Uhd, J.; Carstens, A.; Morris, K. “Clicking” Gene Therapeutics: A Successful Union of Chemistry and Biomedicine for New Solutions. Mol. Pharm. 2018, 15, 2892–2899.
- Fairbanks, B.D.; Culver, H.R.; Mavila, S.; Bowman, C.N. Towards High-Efficiency Synthesis of Xenonucleic Acids. Trends Chem. 2019, 2, 43–56.
- Fantoni, N.Z.; El-Sagheer, A.H.; Brown, T. A Hitchhiker’s Guide to Click-Chemistry with Nucleic Acids. Chem. Rev. 2021.
- Brown, T.; El-Sagheer, A.H. Oligonucleotide ligation. U.S. Patent 8,846,883 B2, 30 September 2014.
- Howe, F.S.; Russell, A.; Lamstaes, A.R.; El-Sagheer, A.; Nair, A.; Brown, T.; Mellor, J. CRISPRi is not strand-specific at all loci and redefines the transcriptional landscape. eLife 2017, 6, e29878.
- Taemaitree, L.; Shivalingam, A.; El-Sagheer, A.H.; Brown, T. An artificial triazole backbone linkage provides a split-and-click strategy to bioactive chemically modified CRISPR sgRNA. Nat. Commun. 2019, 10, 1610.
- Filippova, J.; Matveeva, A.; Zhuravlev, E.; Stepanov, G. Guide RNA modification as a way to improve CRISPR/Cas9-based genome-editing systems. Biochimie 2019, 167, 49–60.
- Chen, Q.; Zhang, Y.; Yin, H. Recent advances in chemical modifications of guide RNA, mRNA and donor template for CRISPR-mediated genome editing. Adv. Drug Deliv. Rev. 2021, 168, 246–258.
- Micklefield, J. Backbone modification of nucleic acids: Synthesis, structure and therapeutic applications. Curr. Med. Chem. 2001, 8, 1157–1179.
- Alvarez-Salas, L.M. Nucleic Acids as Therapeutic Agents. Curr. Top. Med. Chem. 2008, 8, 1379–1404.
- Ochoa, S.; Milam, V.T. Modified Nucleic Acids: Expanding the Capabilities of Functional Oligonucleotides. Molecules 2020, 25, 4659.
- Opalinska, J.B.; Gewirtz, A.M. Nucleic-acid therapeutics: Basic principles and recent applications. Nat. Rev. Drug Discov. 2002, 1, 503–514.
- Sharma, V.K.; Rungta, P.; Prasad, A.K. Nucleic acid therapeutics: Basic concepts and recent developments. RSC Adv. 2014, 4, 16618–16631.
- Sharma, V.K.; Watts, J.K. Oligonucleotide therapeutics: Chemistry, delivery and clinical progres. Future Med. Chem. 2015, 7, 2221–2242.
- Hughes, R.A.; Ellington, A.D. Synthetic DNA Synthesis and Assembly: Putting the Synthetic in Synthetic Biology. Cold Spring Harb. Perspect. Biol. 2017, 9, a023812.
- Bonandi, L.; Christodoulou, M.S.; Fumagalli, G.; Perdicchia, D.; Rastelli, G.; Passarella, D. The 1,2,3-triazole ring as a bioisostere in medicinal chemistry. Drug Discov. Today 2017, 22, 1572–1581.
- Eremeeva, E.; Herdewijn, P. Non canonical genetic material. Curr. Opin. Biotechnol. 2019, 57, 25–33.
- Oliveira, B.L.; Guo, Z.; Bernardes, G.J.L. Inverse electron demand Diels–Alder reactions in chemical biology. Chem. Soc. Rev. 2017, 46, 4895–4950.
- Quadrelli, P. Modern Applications of Cycloaddition Chemistry; Elsevier: Amsterdam, The Netherlands, 2019.
- Uhlmann, E.; Peyman, A. Antisense Oligonucleotides: A New Therapeutic Principle. Chem. Rev. 1990, 90, 543–584.
- Kurreck, J. Antisense technologies. Improvement through novel chemical modifications. Eur. J. Biochem. 2003, 270, 1628–1644.
- Dean, N.M.; Bennett, C.F. Antisense oligonucleotide-based therapeutics for cancer. Oncogene 2003, 22, 9087–9096.
- Tamm, I.; Wagner, M. Antisense therapy in clinical oncology. Mol. Biotechnol. 2006, 33, 221–238.
- Wan, W.B.; Seth, P.P. The Medicinal Chemistry of Therapeutic Oligonucleotides. J. Med. Chem. 2016, 59, 9645–9667.
- Amblard, F.; Cho, J.H.; Schinazi, R.F. Cu(I)-Catalyzed Huisgen Azide−Alkyne 1,3-Dipolar Cycloaddition Reaction in Nucleoside, Nucleotide, and Oligonucleotide Chemistry. Chem. Rev. 2009, 109, 4207–4220.
- Nuzzi, A.; Massi, A.; Dondoni, A. Model Studies Toward the Synthesis of Thymidine Oligonucleotides with Triazole Internucleosidic Linkages Via Iterative Cu(I)-Promoted Azide–Alkyne Ligation Chemistry. QSAR Comb. Sci. 2007, 26, 1191–1199.
- Sanzone, A.P.; El-Sagheer, A.H.; Brown, T.; Tavassoli, A. Assessing the biocompatibility of click-linked DNA in Escherichia coli. Nucleic Acids Res. 2012, 40, 10567–10575.
- Kukwikila, M.; Gale, N.; El-Sagheer, A.H.; Brown, T.; Tavassoli, A. Assembly of a biocompatible triazole-linked gene by one-pot click-DNA ligation. Nat. Chem. 2017, 9, 1089–1098.
- Routh, A.; Head, S.R.; Ordoukhanian, P.; Johnson, J.E. ClickSeq: Fragmentation-Free Next-Generation Sequencing via Click Ligation of Adaptors to Stochastically Terminated 3’-Azido cDNAs. J. Mol. Biol. 2015, 427, 2610–2616.
- Miura, F.; Fujino, T.; Kogashi, K.; Shibata, Y.; Miura, M.; Isobe, H.; Ito, T. Triazole linking for preparation of a next-generation sequencing library from single-stranded DNA. Nucleic Acids Res. 2018, 46, e95.
- Nakane, M.; Ichikawa, S.; Matsuda, A. Triazole-Linked Dumbbell Oligodeoxynucleotides with NF-κB Binding Ability as Potential Decoy Molecules. J. Org. Chem. 2008, 73, 1842–1851.
- Vasilyeva, S.V.; Levina, A.S.; Li-Zhulanov, N.S.; Shatskaya, N.V.; Baiborodin, S.I.; Repkova, M.N.; Zarytova, V.F.; Mazurkova, N.A.; Silnikov, V.N. SiO2 nanoparticles as platform for delivery of 3′-triazole analogues of AZT-triphosphate into cells. Bioorg. Med. Chem. 2015, 23, 2168–2175.
- Cassinelli, V.; Oberleitner, B.; Sobotta, J.; Nickels, P.; Grossi, G.; Kempter, S.; Frischmuth, T.; Liedl, T.; Manetto, A. One-Step Formation of “Chain-Armor”-Stabilized DNA Nanostructures. Angew. Chem. Int. Ed. 2015, 54, 7795–7798.
- Wirges, C.T.; Gramlich, P.M.E.; Gutsmiedl, K.; Gierlich, J.; Burley, G.A.; Carell, T. Pronounced Effect of DNA Hybridization on Click Reaction Efficiency. QSAR Comb. Sci. 2007, 26, 1159–1164.
- Gramlich, P.M.E.; Wirges, C.T.; Manetto, A.; Carell, T. Postsynthetic DNA modification through the copper-catalyzed azide-alkyne cycloaddition reaction. Angew. Chem. Int. Ed. 2008, 47, 8350–8358.
- Xiong, H.; Seela, F. Stepwise “Click” Chemistry for the Template Independent Construction of a Broad Variety of Cross-Linked Oligonucleotides: Influence of Linker Length, Position, and Linking Number on DNA Duplex Stability. J. Org. Chem. 2011, 76, 5584–5597.
- O’Mahony, G.; Ehrman, E.; Grøtli, M. Synthesis of adenosine-based fluorosides containing a novel heterocyclic ring system. Tetrahedron Lett. 2005, 46, 6745–6748.
- Mathew, S.C.; By, Y.; Berthault, A.; Virolleaud, M.-A.; Carrega, L.; Chouraqui, G.; Commeiras, L.; Condo, J.; Attolini, M.; Gaudel-Siri, A.; et al. Expeditious synthesis and biological evaluation of new C-6 1,2,3-triazole adenosine derivatives A1 receptorantagonists or agonists. Org. Biomol. Chem. 2010, 8, 3874–3881.
- Baker, Y.R.; Traoré, D.; Wanat, P.; Tyburn, A.; El-Sagheer, A.H.; Brown, T. Searching for the ideal triazole: Investigating the 1,5-triazole as a charge neutral DNA backbone mimic. Tetrahedron 2020, 76, 130914.
- Kumar, A.S. Design and synthesis of double-headed nucleosides by using click chemistry approach. Chem. Data Collect. 2020, 28, 100468.
- Seela, F.; Sirivolu, V.R. DNA Containing Side Chains with Terminal Triple Bonds: Base-Pair Stability and Functionalization of Alkynylated Pyrimidines and 7-Deazapurines. Chem. Biodivers. 2006, 3, 509–514.
- Sirivou, V.R.; Chittepu, P.; Seela, F. DNA with Branched Internal Side Chains: Synthesis of 5-Tripropargylamine-dU and Conjugation by an Azide-Alkyne Double Click Reaction. ChemBioChem 2008, 9, 2305–2316.
- Seela, F.; Sirivolu, V.R. Pyrrolo-dColigonucleotides bearing alkynyl side chains with terminal triple bonds: Synthesis, base pairing and fluorescent dye conjugates prepared by the azide–alkyne “click” reaction. Org. Biomol. Chem. 2008, 6, 1674–1687.
- Seela, F.; Sirivolu, V.R.; Chittepu, P. Modification of DNA with Octadiynyl Side Chains: Synthesis, Base Pairing, and Formation of Fluorescent Coumarin Dye Conjugates of Four Nucleobases by the Alkyne−Azide “Click” Reaction. Bioconjugate Chem. 2008, 19, 211–224.
- El-Sagheer, A.H.; Brown, T. Click chemistry with DNA. Chem. Soc. Rev. 2010, 39, 1388–1405.
- Lucas, R.; Neto, V.; Bouazza, A.H.; Zerrouki, R.; Granet, R.; Krausz, P.; Champavier, Y. Microwave-assisted synthesis of a triazole-linked 3′–5′ dithymidine using click chemistry. Tetrahedron Lett. 2008, 49, 1004–1007.
- Lucas, R.; Zerrouki, R.; Granet, R.; Krausz, P.; Champavier, Y. A rapid efficient microwave-assisted synthesis of a 3′,5′-pentathymidine by copper(I)-catalyzed [3 + 2] cycloaddition. Tetrahedron 2008, 64, 5467–5471.
- Varizhuk, A.; Chizhov, A.; Smirnov, I.; Kaluzhny, D.; Florentiev, V. Triazole-Linked Oligonucleotides with Mixed-Base Sequences: Synthesis and Hybridization Properties. Eur. J. Org. Chem. 2012, 11, 2173–2179.
- Fujino, T.; Tsunaka, N.; Guillot-Nieckowski, M.; Nakanishi, W.; Iwamoto, T.; Nakamura, E.; Isobe, H. Synthesis and structures of deoxyribonucleoside analogues for triazole-linked DNA (TLDNA). Tetrahedron Lett. 2010, 51, 2036–2038.
- El-Sagheer, A.; Brown, T. Synthesis and Polymerase Chain Reaction Amplification of DNA Strands Containing an Unnatural Triazole Linkage. J. Am. Chem. Soc. 2009, 131, 3958–3964.
- Kumar, R.; El-Sagheer, A.; Tumpane, J.; Lincoln, P.; Wilhelmsson, L.M.; Brown, T. Template-Directed Oligonucleotide Strand Ligation, Covalent Intramolecular DNA Circularization and Catenation Using Click Chemistry. J. Am. Chem. Soc. 2007, 129, 6859–6864.
- Kočalka, P.; El-Sagheer, A.H.; Brown, T. Rapid and Efficient DNA Strand Cross-Linking by Click Chemistry. ChemBioChem 2008, 9, 1280–1285.
- El-Sagheer, A.H.; Kumar, R.; Findlow, S.; Werner, J.M.; Lane, A.N.; Brown, T. A Very Stable Cyclic DNA Miniduplex with Just Two Base Pairs. ChemBioChem 2008, 9, 50–52.
- Dallmann, A.; El-Sagheer, A.H.; Dehmel, L.; Mügge, C.; Griesinger, C.; Ernsting, N.P.; Brown, T. Structure and Dynamics of Triazole-Linked DNA: Biocompatibility Explained. Chem. Eur. J. 2011, 17, 14714–14717.
- El-Sagheer, A.H.; Brown, T. Combined nucleobase and backbone modifications enhance DNA duplex stability and preserve biocompatibility. Chem. Sci. 2014, 5, 253–259.
- Shivalingam, A.; Tyburn, A.E.S.; El-Sagheer, A.H.; Brown, T. Molecular Requirements of High-Fidelity Replication-Competent DNA Backbones for Orthogonal Chemical Ligation. J. Am. Chem. Soc. 2017, 139, 1575–1583.
- El-Sagheer, A.H.; Sanzone, A.P.; Gao, R.; Tavassoli, A.; Brown, T. Biocompatible artificial DNA linker that is read through by DNA polymerases and is functional in Escherichia coli. Proc. Natl. Acad. Sci. USA 2011, 108, 11338–11343.
- El-Sagheer, A.H.; Brown, T. Efficient RNA synthesis by in vitro transcription of a triazole-modified DNA template. Chem. Commun. 2011, 47, 12057–12058.
- El-Sagheer, A.H.; Brown, T. Click Nucleic Acid Ligation: Applications in Biology and Nanotechnology. Acc. Chem. Res. 2012, 45, 1258–1267.
- Birts, C.N.; Sanzone, A.P.; El-Sagheer, A.H.; Blaydes, J.P.; Brown, T.; Tavassoli, A. Transcription of click-linked DNA in human cells. Angew. Chem. Int. Ed. 2014, 53, 2362–2365.
- Pujari, S.S.; Leonard, P.; Seela, F. Oligonucleotides with “Clickable” Sugar Residues: Synthesis, Duplex Stability, and Terminal versus Central Interstrand Cross-Linking of 2′-O-Propargylated 2-Aminoadenosine with a Bifunctional Azide. J. Org. Chem. 2014, 79, 4423–4437.
- Wang, W.; Chen, K.; Qu, D.; Chi, W.; Xiong, W.; Huang, Y.; Wen, J.; Feng, S.; Zhang, B. One pot conjugation of small molecules to RNA using click chemistry. Tetrahedron Lett. 2012, 53, 6747–6750.
- Peel, B.J.; Hagen, G.; Krishnamurthy, K.; Desaulniers, J.-P. Conjugation and Evaluation of Small Hydrophobic Molecules to Triazole-Linked siRNAs. ACS Med. Chem. Lett. 2015, 6, 117–122.
- Costales, M.G.; Rzuczek, S.G.; Disney, M.D. Comparison of small molecules and oligonucleotides that target a toxic, non-coding RNA. Bioorganic Med. Chem. Lett. 2016, 26, 2605–2609.
- Zheng, Y.; Beal, P.A. Synthesis and evaluation of an alkyne-modified ATP analog for enzymatic incorporation into RNA. Bioorganic Med. Chem. Lett. 2016, 26, 1799–1802.
- Atdjian, C.; Coelho, D.; Iannazzo, L.; Ethève-Quelquejeu, M.; Braud, E. Synthesis of Triazole-Linked SAM-Adenosine Conjugates: Functionalization of Adenosine at N-1 or N-6 Position without Protecting Groups. Molecules 2020, 25, 3241.
- Sau, S.P.; Hrdlicka, P.J. C2′-Pyrene-Functionalized Triazole-Linked DNA: Universal DNA/RNA Hybridization Probes. J. Org. Chem. 2012, 77, 5–16.
- Velema, W.A.; Kool, E.T. The chemistry and applications of RNA 2′-OH acylation. Nat. Rev. Chem. 2020, 4, 22–37.
- Haugland, M.M.; El-Sagheer, A.; Porter, R.J.; Peña, J.; Brown, T.; Anderson, E.A.; Lovett, J.E. 2′-Alkynylnucleotides: A Sequence- and Spin Label-Flexible Strategy for EPR Spectroscopy in DNA. J. Am. Chem. Soc. 2016, 138, 9069–9072.
- Lorenz, D.A.; Garner, A.L. A click chemistry-based microRNA maturation assay optimized for high-throughput screening. Chem. Commun. 2016, 52, 8267–8270.
- Walter, N.G.; Engelke, D.R. Ribozymes: Catalytic RNAs that cut things, make things, and do odd and useful jobs. Biologist 2002, 49, 199–203.
- Isobe, H.; Fujino, T.; Yamazaki, N.; Guillot-Nieckowski, M.; Nakamura, E. Triazole-Linked Analogue of Deoxyribonucleic Acid (TLDNA): Design, Synthesis, and Double-Strand Formation with Natural DNA. Org. Lett. 2008, 10, 3729–3732.
- Fujino, T.; Yamazaki, N.; Isobe, H. Convergent synthesis of oligomers of triazole-linked DNA analogue (TLDNA) in solution phase. Tetrahedron Lett. 2009, 50, 4101–4103.
- El-Sagheer, A.H.; Brown, T. New strategy for the synthesis of chemically modified RNA constructs exemplified by hairpin and hammerhead ribozymes. Proc. Natl. Acad. Sci. USA 2010, 107, 15329–15334.
- Fujino, T.; Endo, K.; Yamazaki, N.; Isobe, H. Synthesis of Triazole-linked Analogues of RNA (TLRNA). Chem. Lett. 2012, 41, 403–405.
- Paredes, E.; Das, S.R. Click chemistry for rapid labeling and ligation of RNA. ChemBioChem 2011, 12, 125–131.
- Muthmann, N.; Hartstock, K.; Rentmeister, A. Chemo-enzymatic treatment of RNA to facilitate analyses. WIREs RNA 2020, 11, e1561.
- Palumbo, C.M.; Beal, P.A. Nucleoside analogs in the study of the epitranscriptome. Methods 2019, 156, 46–52.
- Aigner, A.; Hartl, M.; Fauster, K.; Steger, J.; Bister, K.; Micura, R. Chemical Synthesis of Site-Specifically 2′-Azido-Modified RNA and Potential Applications for Bioconjugation and RNA Interference. ChemBioChem 2011, 12, 47–51.
- Meter, E.N.V.; Onyango, J.A.; Teske, K.A. A review of currently identified small molecule modulators of microRNA function. Eur. J. Med. Chem. 2020, 188, 112008.
- Fernholm, A. Genetic Scissors: A Tool for Rewriting the Code of Life; The Royal Swedish Academy of Sciences: Stockholm, Sweden, 2020.
- Campbell, M.A.; Wengel, J. Locked vs. unlocked nucleic acids (LNA vs.UNA): Contrasting structures work towards common therapeutic goals. Chem. Soc. Rev. 2011, 40, 5680–5689.
- Imanishi, T.; Obika, S. BNAs: Novel nucleic acid analogs with a bridged sugar moiety. Chem. Commun. 2002, 1653–1659.
- Mitsuoka, Y.; Yamamoto, T.; Kugimiya, A.; Waki, R.; Wada, F.; Tahara, S.; Sawamura, M.; Noda, M.; Fujimura, Y.; Kato, Y.; et al. Triazole- and Tetrazole-Bridged Nucleic Acids: Synthesis, Duplex Stability, Nuclease Resistance, and in Vitro and in Vivo Antisense Potency. J. Org. Chem. 2017, 82, 12–24.
- Singh, S.K.; Sharma, V.K.; Bohra, K.; Olsen, C.E.; Prasad, A.K. Synthesis of Triazole-linked LNA-based Non-ionic Nucleoside Dimers Using Cu(I)- Catalyzed ‘Click’ Reaction. Curr. Org. Synth. 2014, 11, 757–766.
- Srivastava, S.; Singh, S.K.; Sharma, V.K.; Mangla, P.; Olsen, C.E.; Prasad, A.K. Design and Synthesis of Triazole-Linked xylo-Nucleoside Dimers. Nucleos. Nucleot. Nucl. 2015, 34, 388–399.
- Sharma, V.K.; Singh, S.K.; Krishnamurthy, P.M.; Alterman, J.F.; Haraszti, R.A.; Khvorova, A.; Prasad, A.K.; Watts, J.K. Synthesis and biological properties of triazole-linked locked nucleic acid. Chem. Commun. 2017, 53, 8906–8909.
- Kumar, P.; El-Sagheer, A.H.; Truong, L.; Brown, T. Locked nucleic acid (LNA) enhances binding affinity of triazole-linked DNA towards RNA. Chem. Commun. 2017, 53, 8910–8913.
- Kumar, P.; Truong, L.; Baker, Y.R.; El-Sagheer, A.H.; Brown, T. ; Synthesis, Affinity for Complementary RNA and DNA, and Enzymatic Stability of Triazole-Linked Locked Nucleic Acids (t-LNAs). ACS Omega 2018, 3, 6976–6987.
- Larsen, H.J.; Bentin, T.; Nielsen, P.E. Antisense properties of peptide nucleic acid. Biochim. Biophys. Acta 1999, 1489, 159–166.
- Soomets, U.; Hällbrink, M.; Langel, Ü. Antisense properties of peptide nucleic acid. Front. Biosci. 1999, 4, 782–786.
- Četojević-Simin, D.; Jakimov, D.; Mrđanović, J.; Bogdanović, V.; Kojić, V.; Andrijević, L.; Bogdanović, G. Peptide nucleic acid: Sequence specific recognition in cancer diagnostics and gene therapy. Arch. Oncol. 2001, 9, 33–37.
- Piacenti, V.; Langella, E.; Autiero, I.; Nolan, J.C.; Piskareva, O.; Adamo, M.F.A.; Saviano, M.; Moccia, M. A combined experimental and computational study on peptide nucleic acid (PNA) analogues of tumor suppressive miRNA-34a. Bioorg. Chem. 2019, 91, 103165.
- Holub, J.M.; Kirshenbaum, K. Tricks with clicks: Modification of peptidomimetic oligomers via copper-catalyzed azide-alkyne [3 + 2] cycloaddition. Chem. Soc. Rev. 2010, 39, 1325–1337.
- Amant, A.H.S.; Engbers, C.; Hudson, R.H.E. A solid-phase CuAAC strategy for the synthesis of PNA containing nucleobase surrogates. Artif. Dna Pna Xna 2013, 4, 4–10.
- Thomson, S.A.; Josey, J.A.; Cadilla, R.; Gaul, M.D.; Hassman, C.F.; Luzzio, M.J.; Pipe, A.J.; Reed, K.L.; Ricca, D.J.; Wiethe, R.W.; et al. Fmoc mediated synthesis of Peptide Nucleic Acids. Tetrahedron 1995, 51, 6179–6194.
- Efthymiou, T.C.; Desaulniers, J.-P. Synthesis and properties of oligonucleotides that contain a triazole-linked nucleic acid dimer. J. Heterocyclic Chem. 2011, 48, 533–539.
- Vergnaud, J.; Faugerasa, P.-A.; Chaleix, V.; Champavier, Y.; Zerrouki, R. Design of a new oligotriazole peptide nucleic acid analogue (oT-PNA). Tetrahedron Lett. 2011, 52, 6185–6189.
- Marin, V.L.; Armitage, B.A. RNA Guanine Quadruplex Invasion by Complementary and Homologous PNA Probes. J. Am. Chem. Soc. 2005, 127, 8032–8033.
- Gudanis, D.; Kaniowski, D.; Kulik, K.; Baranowski, D.; Gdaniec, Z.; Nawrot, B. Formation of an RNA Quadruplex-Duplex Hybrid in Living Cells between mRNA of the Epidermal Growth Factor Receptor (EGFR) and a G-Rich Antisense Oligoribonucleotide. Cells 2020, 9, 2375.
- Solier, S.; Müller, S.; Rodriguez, R. Whole-genome mapping of small-molecule targets for cancer medicine. Curr. Opin. Chem. Biol. 2020, 56, 42–50.
- Schärer, O.D. Chemistry and biology of DNA repair. Angew. Chem. Int. Ed. 2003, 42, 2946–2974.
- Xu, Y. Chemistry in human telomere biology: Structure, function and targeting of telomere DNA/RNA. Chem. Soc. Rev. 2011, 40, 2719–2740.
- Xu, Y.; Suzuki, Y.; Komiyama, M. Click chemistry for the identification of G-quadruplex structures: Discovery of a DNA-RNA G-quadruplex. Angew. Chem. Int. Ed. 2009, 48, 3281–3284.
- Saha, P.; Panda, D.; Dash, J. The application of click chemistry for targeting quadruplex nucleic acids. Chem. Commun. 2019, 55, 731–750.
- Varizhuk, A.M.; Tsvetkov, V.B.; Tatarinova, O.N.; Kaluzhny, D.N.; Florentiev, V.L.; Timofeev, E.N.; Shchyolkina, A.K.; Borisova, O.F.; Smirnov, I.P.; Grokhovsky, S.L.; et al. Synthesis, characterization and in vitro activity of thrombin-binding DNA aptamers with triazole internucleotide linkages. Eur. J. Med. Chem. 2013, 67, 90–97.
- Xu, Y.; Ishizuka, T.; Yang, J.; Ito, K.; Katada, H.; Komiyama, M.; Hayashi, T. Oligonucleotide Models of Telomeric DNA and RNA Form a Hybrid G-quadruplex Structure as a Potential Component of Telomeres. J. Biol. Chem. 2012, 287, 41787–41796.
- Xu, Y.; Suzuki, Y.; Ishizuka, T.; Xiao, C.-D.; Liu, X.; Hayashi, T. Finding a human telomere DNA–RNA hybrid G-quadruplex formed by human telomeric 6-mer RNA and 16-mer DNA using click chemistry: A protective structure for telomere end. Bioorg. Med. Chem. 2014, 22, 4419–4421.
- Ishizuka, T.; Xu, Y. Click Chemistry Takes a Snapshot of DNA-RNA Hybrid G-Quadruplex in Living Cells. J. Nat. Sci. 2016, 2, e237.




