Pathogen-associated molecular patterns (PAMPs) and danger-associated molecular patterns (DAMPs) induce NLRP3 inflammasome activation, and subsequent formation of active caspase-1 as well as the maturation of interleukin-1β (IL-1β) and gasdermin D (GSDMD), mediating the occurrence of pyroptosis and inflammation. Aberrant NLRP3 inflammasome activation causes a variety of diseases. Therefore, the NLRP3 inflammasome pathway is a target for prevention and treatment of relative diseases.
- :inflammasome
- NLRP3
- post-translational modification
- inflammasome
1. Introduction
2. NLRP3 Inflammasome Activation
Two signals are required for NLRP3 inflammasome activation. Signal I involves the priming signal, induces IL-1β expression and upregulates NLRP3 expression through activating TLR and NF-κB pathways [17,18][17][18] as well as NLRP3 phosphorylation. In addition, signal II, transduced by PAMPs and host-derived DAMPs, triggers the assembly and activation of the NLRP3 inflammasome [19]. Mechanisms by which the NLRP3 inflammasome is activated are extensively explored. At least four models were proposed: K+ efflux [20], the generation of mitochondrial reactive oxygen species (mROS) [21], cathepsin B release from damaged lysosomes [22] and Ca2+ mobilization [23]. NLRP3 oligomerization via its NACHT domain leads to PYD clustering, which elicits recruitment and clustering of ASC through PYD–PYD interaction. ASC clustering subsequently provokes caspase-1 recruitment and assembly of the inflammasome complex. Then, caspase-1 undergoes autocleavage and formation of active p10/p20 tetramer, which cleaves proinflammatory cytokines such as IL-1β and IL-18 into their active molecules [24]. Active caspase-1 also cleaves GSDMD. The GSDMD N-terminal fragment (GSDMD-N) oligomerizes in the plasma membrane to generate approximately 21 nm-diameter GSDMD pores, leading to osmotic imbalance and cell death called pyroptosis (Figure 1) [24,25][24][25].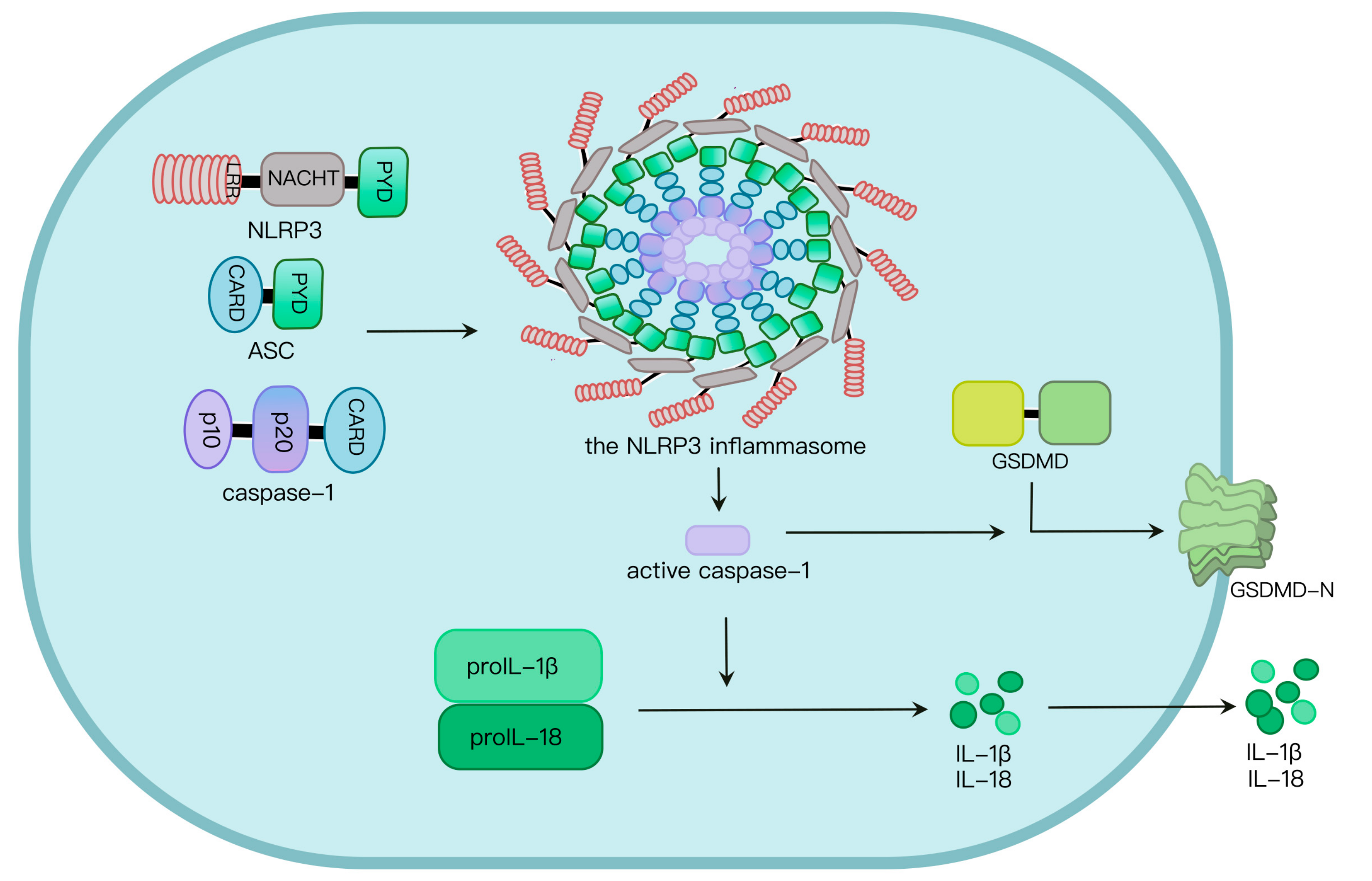
3. Regulation of Ubiquitination and Deubiquitination in NLRP3 Inflammasome Activation
Ubiquitin, a highly conserved small regulatory eukaryotic protein, contains 76 amino acids and 7 lysine residues, including K6, K11, K27, K29, K33, K48 and K63. It can be covalently attached to target proteins through a cascade of enzymatic reactions catalyzed by ubiquitin-activating enzymes (E1), ubiquitin-conjugating enzymes (E2) and ubiquitin ligases (E3) [26]. Ubiquitin is bound to the target substrates via an isopeptide bond formed between the C-terminal glycine of ubiquitin and the ε-amino group of lysine in the substrate [27]. Similar isopeptide bonds can be formed from linkage of the C-terminus of one ubiquitin to one of the seven lysine residues or the N terminal methionine on another ubiquitin to form ubiquitin chains [28]. Deubiquitinases remove conjugated ubiquitin from the substrates [29].
3.1. Ubiquitination of NLRP3
NLRP3 ubiquitination plays a critical role in the regulation of its activity. The effects of different types of ubiquitin chains on the NLRP3 activity vary. Among the three enzymes participating in ubiquitination, E3 ubiquitin ligases act as the major proteins involved in the orchestration of NLRP3 inflammasome activity. Its regulation in NLRP3 inflammasome activation is mainly mediated by K48-, K63- and K27-linked ubiquitination. K48-linked ubiquitin chains employ a closed conformation with the hydrophobic residues at the interdomain interface exposed to ubiquitin chain recognition factors, serving as a signal for degradation by the 26S proteasome [51][30]. K63-linked ubiquitin chains act as a signal for altering the functions of the modified protein, including signaling transduction, DNA repair and intracellular trafficking [52][31].3.2. Deubiquitination of NLRP3
Deubiquitinases remove K48- and K11-linked ubiquitin chains, contributing to NLRP3 inflammasome activation. This is consistent with the effect of K48-linked ubiquitination on NLRP3 inflammasome activity. Upon infection with the DNA virus herpes simplex virus type 1 (HSV-1), the stimulator of interferon genes (STING) binds to NLRP3, attenuates K48- and K63-linked ubiquitination of NLRP3 and increases its protein expression on the endoplasmic reticulum, facilitating NLRP3 inflammasome activation and the subsequent release of proinflammatory cytokines [44][32]. Ubiquitin-specific peptidase 1 (USP1)-associated factor 1 (UAF1, also called WDR48 or p80) is a stoichiometric binding partner of USP1 [68][33]. The UAF1-USP1 deubiquitinase complex selectively eliminates K48-linked ubiquitin chains of NLRP3 to stabilize its expression via interaction with the LRR and NACHT domains, which in turn promotes NLRP3 inflammasome activation [45][34].3.3. Ubiquitination and Deubiquitination of ASC and Caspase-1
The influence of K63-linked chains on ASC still remains to be investigated. Both the addition and elimination of K63 ubiquitin chains promote NLRP3 inflammasome activation. This may be associated with different modification sites or regulation by other signaling pathways triggered by ubiquitin enzymes or deubiquitinases. On the one hand, mitochondrial antiviral signaling protein (MAVS) facilitates interaction between the tumor necrosis factor receptor-associated factor (TRAF3) and ASC, provoking K63-linked ubiquitination of ASC at Lys174 and contributing to NLRP3 inflammasome activation [48,72][35][36]. Pellino E3 ubiquitin protein ligase 1 (Peli1) mediates both K48- and K63-ubiquitination [73,74][37][38].4. Regulation of Phosphorylation and Dephosphorylation in NLRP3 Inflammasome Activation
Phosphorylation modulates protein function and controls the turnover of its targets and subcellular localization by altering protein conformation or influencing protein–protein interaction. Protein kinases mediate the phosphate group transfer from ATP to serine, threonine and tyrosine residues of the substrates, while phosphatases removes the phosphate group of a phosphorylated protein substrate [80][39]. Phosphorylation and dephosphorylation of the components of the NLRP3 inflammasome control its activity.4.1. NLRP3 Phosphorylation
NLRP3 phosphorylation at Ser725 [81][40], Ser194 [82][41] or tyrosines in the PYD-NACHT polybasic linker [89][42] contribute to NLRP3 inflammasome activation. Misshapen-like kinase 1 (MINK1), a member of the mammalian germinal center kinase (GCK) family [90][43], binds to the NLRP3 LRR domain, and phosphorylates NLRP3 at Ser725, which are critical for the priming step of NLRP3 inflammasome activation [81][40]. ROS serves only as a priming signal, but fails to contribute to the activation step of the NLRP3 inflammasome [91][44]. It is able to increase the kinase activity of MINK1 and facilitate NLRP3 phosphorylation at Ser725. This subsequently promotes inflammasome priming [81][40].4.2. NLRP3 Dephosphorylation
NLPR3 dephosphorylation at Ser5 [84][45], Ser803 [95][46] or Tyr861 [85][47] enhances NLRP3 inflammasome activation. The phosphatase protein, phosphatase 2A (PP2A), dephosphorylates NLRP3 at Ser5 to allow its activation, and the regulation of PP2A in NLRP3 inflammasome activation is controlled by BTK [84,96][45][48]. Protein tyrosine phosphatase non-receptor type 22 (PTPN22) interacts with and dephosphorylates NLRP3 at Tyr861, decreasing NLRP3 inflammasome-mediated IL-1β secretion [85][47].4.3. Phosphorylation of ASC and Caspase-1
The phosphorylation and dephosphorylation of NLRP3 at different sites play distinct roles in the regulation of NLRP3 inflammasome activation, while the phosphorylation of ASC and caspase-1 promotes NLRP3 inflammasome activation. Pyruvate kinase (Pyk2) [87][49] and JNK [99][50] are involved in human ASC Tyr146 (mouse ASC Tyr144) phosphorylation, facilitating ASC oligomerization and NLRP3 inflammasome assembly. Spleen tyrosine kinase (Syk) contributes to NLRP3 agonist-mediated ASC self-association and inflammasome assembly through phosphorylating ASC at Tyr146 and Tyr187 residues [99,100][50][51]. LPS treatment induces interaction between p21 (Rac family small GTPase 1) activated kinase 1 (PAK1) and caspase-1, mediating caspase-1 phosphorylation at Ser376 and its activation [88][52].5. Regulation of SUMOylation in NLRP3 Inflammasome Activation
The small ubiquitin-like modifier (SUMO) protein is evolutionarily conserved and ubiquitously expressed in eukaryotes. It belongs to the ubiquitin-like family and alters the properties and functions of modified proteins via PTMs [101,102,103][53][54][55]. Four SUMO proteins have been identified in humans, SUMO1–4. SUMO2 and -3 are highly homologous [104][56]. SUMOylation is a reversible PTM process. SUMO binds to a lysine residue of a substrate and is removed from the modified protein through SUMO-specific peptidase-mediated deSUMOylation. SUMO is expressed as a C-terminally extended precursor, and then processed to generate the active form. The SUMO-activating enzyme SAE1/SAE2 covalently links to the C-terminus of SUMO via the sulfhydryl group of a cysteine residue; then, SUMO is transferred to the SUMO-conjugating enzyme Ubc9, and finally conjugated to a lysine side chain of the target protein mediated by a SUMO ligase. SUMO chains are able to assemble on substrates [105,106][57][58]. NLRP3 Lys204 SUMOylation facilitates its activation [107][59], while Lys689 SUMOylation suppresses its activation [108][60]. Ubc9 binds to NLRP3 to mediate its SUMOylation at Lys204 induced by SUMO1, which facilitates ASC oligomerization and inflammasome assembly. Interaction between SUMO-specific protease 3 (SENP3) and NLRP3 results in NLRP3 deSUMOylation and attenuates ASC speck formation, as well as inflammasome activation [107][59]. Mitochondrial-anchored protein ligase (MAPL), also called mitochondrial E3 ubiquitin protein ligase 1 (MUL1), mediates RING-finger-dependent NLRP3 Lys689 SUMOylation, and enhances caspase-1 maturation and IL-1β release in murine macrophages and human cells. NLRP3 Lys689 deSUMOylation induced by SENP6 and 7 promotes NLRP3-dependent ASC oligomerization and caspase-1 activation. MAPL has no effect on absent in melanoma 2 (AIM2) inflammasome activation [108][60].6. Regulation of Alkylation in NLRP3 Inflammasome Activation
Alkylation targeting ATPase activity of NLRP3 negatively controls the inflammasome activation through a decreasing affinity for ATP. The NACHT domain, carrying ATPase activity, adopts an ADP/ATP switch mechanism to orchestrate NLRP3 activation [110][61]. The NACHT domain is composed of four subdomains, including a nucleotide-binding domain (NBD, aa131–373 in human NLRP3), a helical domain 1 (HD1, aa374–434), a winged-helix domain (WHD, aa435–538) and helical domain 2 (HD2, aa539–676). Interaction between WHD His522 and ADP keeps NLRP3 in a closed and inactive conformation [111,112][62][63]. Treatment with NLRP3 agonists leads to the exit of ADP, the binding of ATP with the Walker A motif in NBD and ATP hydrolysis [110][61]. MCC950 suppresses ATP hydrolysis and NLRP3 inflammasome activation via interaction with the Walker B motif in NBD [113][64]. 3, 4-methylenedioxy-β-nitrostyrene (MNS) [114][65], 2-cyclohexylimino-6-methyl-6,7-dihydro-5H-benzo [1[1][3],3], oxathiol-4-one (BOT-4-one) [115][66] and vanillylacetone [116][67] mediate NLRP3 alkylation and reduce its ATPase activity.7. Regulation of S-Nitrosylation in NLRP3 Inflammasome Activation
Similar to NLRP3 alkylation, NLRP3 S-nitrosylation plays an inhibitory role in the inflammasome activity. S-nitrosylation refers to the covalent binding of a NO group to a protein cysteine thiol to form S-nitrosothiols [125][68]. S-nitroso-N-acetylpenicillamine (SNAP), an NO donor, dampens nigericin-induced capase-1 maturation, as well as release of IL-1β and IL-18, in the TLR1/2 agonist PAM3CSK4-primed murine peritoneal macrophages. Priming with PAM3CSK4 causes S-nitrosylation of NLRP3 and caspase-1, and the C-terminus of NLRP3 is more susceptible to S-nitrosylation than its N-terminus. Inflammasomes of AIM2 and NLRC4 are partially inhibited [126][69]. HEK293T cells transfected with NLRP3 or caspase-1 expression plasmids were treated with SNAP or not, and lysed to obtain lysate 1. Independent cultures transfected with plasmids expressing the other components of the NLRP3 inflammasome and IL-1β were lysed to obtain lysate 2. Mature IL-1β was assessed in the mixed lysates with lysate 1 and lysate 2.8. Regulation of Acetylation in NLRP3 Inflammasome Activation
In contrast to alkylation and S-nitrosylation of NLRP3, NLRP3 acetylation boosts the inflammasome activation. Lysine acetyltransferase 5 (KAT5), also called Tat-interactive protein 60 kDa (TIP60), belongs to MOZ-Ybf2/Sas3-Sas2-TIP60 (MYST) family [129][70]. KAT5 mediates NLRP3 acetylation at Lys24, facilitating its interaction with NEK7 and oligomerization. The KAT5 inhibitor NU9056 restricts NLRP3-dependent caspase-1 maturation and IL-1β release both in vivo and in vitro [130][71]. Sirtuin 2 (SIRT2), an NAD+-dependent deacetylase and a metabolic sensor, targets NLRP3 for deacetylation in BMDMs. Sirt2 deletion or treatment with the SIRT2 inhibitor AGK2 results in increased IL-1β production and cleaved caspase-1, but has no effect on pro-IL-1β expression. Sirt2 knockout makes no change to caspase-1 maturation and IL-1β secretion following stimulation with flagellin, a NLRC4 inducer, or poly(dA:dT), an AIM2 inducer.9. Regulation of S-Glutathionylation in NLRP3 Inflammasome Activation
Protein S-glutathionylation, an oxidative PTM, is a reversible formation of mixed disulfides between tripeptide glutathione and low-pKa cysteine [132][72]. Glutathione transferase Omega 1 (GSTO1-1) belongs to the cytosolic glutathione transferase (GST) super family [133][73]. Gsto1-1 deletion reduces proinflammatory cytokine expression and ameliorates the inflammatory response in response to LPS in mice. GSTO1-1 deglutathionylates NEK7 Cys253 to promote its interaction with NLRP3 and the inflammasome activation [134][74]. Superoxide dismutase 1 (SOD1) contributes to caspase-1 activation via inhibiting its glutathionylation. Upon stimulation with LPS + ATP, SOD1 deficiency leads to increased ROS generation, decreased cellular redox potential and reversible oxidation and glutathionylation of caspase-1 Cys397 and Cys362, eliciting the inhibition of caspase-1 activity. Caspase-1 is activated and not glutathionylated in WT murine peritoneal macrophages [135][75].10. The NLRP3 Inflammasome and Cancers
The NLRP3 inflammasome plays dual roles in the pathogenesis of cancers. It has a protective anti-tumorigenic effect in colitis-associated cancer, colorectal cancer, hepatocellular carcinoma and melanoma, and plays a pro-tumorigenic role in breast cancer, colon cancer, colorectal cancer, epithelial skin cancer, fibrosarcoma and gastric cancer [138,139][76][77]. Regulation of NLRP3 inflammasome activity via PTMs is essential for cancer development and progression. TRIM31 is upregulated at the protein level in human hepatocellular carcinoma and colorectal cancer, and promotes invasion and metastasis [140,141][78][79]. It mediates the K48-linked ubiquitination and degradation of NLRP3, restricting NLRP3 inflammasome activity [35][80]. Alpinumisoflavone and estrogen suppress the proliferation and metastasis of hepatocellular carcinoma cells via enhancing NLRP3 inflammasome activation [142,143][81][82]. Caffeic acid phenethyl ester (CAPE) enhances NLRP3 ubiquitination via facilitating NLRP3–Cullin1 interaction and suppresses NLRP3 inflammasome activation, which protects mice from azoxymethane/dextran sulfate sodium-induced colon cancer [39,144][83][84].References
- Brubaker, S.W.; Bonham, K.S.; Zanoni, I.; Kagan, J.C. Innate immune pattern recognition: A cell biological perspective. Annu. Rev. Immunol. 2015, 33, 257–290.
- Broz, P.; Dixit, V.M. Inflammasomes: Mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 2016, 16, 407–420.
- Yang, Q.; Yu, C.; Yang, Z.; Wei, Q.; Mu, K.; Zhang, Y.; Zhao, W.; Wang, X.; Huai, W.; Han, L. Deregulated NLRP3 and NLRP1 inflammasomes and their correlations with disease activity in systemic lupus erythematosus. J. Rheumatol. 2014, 41, 444–452.
- Guo, C.; Fu, R.; Wang, S.; Huang, Y.; Li, X.; Zhou, M.; Zhao, J.; Yang, N. NLRP3 inflammasome activation contributes to the pathogenesis of rheumatoid arthritis. Clin. Exp. Immunol. 2018, 194, 231–243.
- Clavijo-Cornejo, D.; Lopez-Reyes, A.; Cruz-Arenas, E.; Jacobo-Albavera, L.; Francisco-Balderas, A.; Dominguez-Perez, M.; Mellado, J.V.; Torre, L.H.S.; Pineda, C.; Martinez-Nava, G.; et al. The Role of Nlrp3 Inflammasome Polymorphisms in the Gout Susceptibility. Ann. Rheum. Dis. 2022, 81, 1142.
- Hoseini, Z.; Sepahvand, F.; Rashidi, B.; Sahebkar, A.; Masoudifar, A.; Mirzaei, H. NLRP3 inflammasome: Its regulation and involvement in atherosclerosis. J. Cell. Physiol. 2018, 233, 2116–2132.
- Tan, M.S.; Yu, J.T.; Jiang, T.; Zhu, X.C.; Tan, L. The NLRP3 inflammasome in Alzheimer’s disease. Mol. Neurobiol. 2013, 48, 875–882.
- Irrera, N.; Russo, M.; Pallio, G.; Bitto, A.; Mannino, F.; Minutoli, L.; Altavilla, D.; Squadrito, F. The Role of NLRP3 Inflammasome in the Pathogenesis of Traumatic Brain Injury. Int. J. Mol. Sci. 2020, 21, 6204.
- Li, S.J.; Zhang, Y.F.; Ma, S.H.; Yi, Y.; Yu, H.Y.; Pei, L.; Feng, D. The role of NLRP3 inflammasome in stroke and central poststroke pain. Medicine (Baltimore) 2018, 97, e11861.
- Schroder, K.; Tschopp, J. The Inflammasomes. Cell 2010, 140, 821–832.
- Srinivasula, S.M.; Poyet, J.L.; Razmara, M.; Datta, P.; Zhang, Z.; Alnemri, E.S. The PYRIN-CARD protein ASC is an activating adaptor for caspase-1. J. Biol. Chem. 2002, 277, 21119–21122.
- Lavrik, I.N.; Golks, A.; Krammer, P.H. Caspases: Pharmacological manipulation of cell death. J. Clin. Investig. 2005, 115, 2665–2672.
- Ramazi, S.; Zahiri, J. Posttranslational modifications in proteins: Resources, tools and prediction methods. Database 2021, 2021, baab012.
- Pan, S.; Chen, R. Pathological implication of protein post-translational modifications in cancer. Mol. Aspects Med. 2022, 86, 101097.
- Jennings, E.Q.; Fritz, K.S.; Galligan, J.J. Biochemical genesis of enzymatic and non-enzymatic post-translational modifications. Mol. Aspects Med. 2022, 86, 101053.
- Qin, Y.; Li, Q.; Liang, W.; Yan, R.; Tong, L.; Jia, M.; Zhao, C.; Zhao, W. TRIM28 SUMOylates and stabilizes NLRP3 to facilitate inflammasome activation. Nat. Commun. 2021, 12, 4794.
- Netea, M.G.; Nold-Petry, C.A.; Nold, M.F.; Joosten, L.A.; Opitz, B.; van der Meer, J.H.; van de Veerdonk, F.L.; Ferwerda, G.; Heinhuis, B.; Devesa, I.; et al. Differential requirement for the activation of the inflammasome for processing and release of IL-1beta in monocytes and macrophages. Blood 2009, 113, 2324–2335.
- Bauernfeind, F.G.; Horvath, G.; Stutz, A.; Alnemri, E.S.; MacDonald, K.; Speert, D.; Fernandes-Alnemri, T.; Wu, J.; Monks, B.G.; Fitzgerald, K.A.; et al. Cutting edge: NF-kappaB activating pattern recognition and cytokine receptors license NLRP3 inflammasome activation by regulating NLRP3 expression. J. Immunol. 2009, 183, 787–791.
- Jo, E.K.; Kim, J.K.; Shin, D.M.; Sasakawa, C. Molecular mechanisms regulating NLRP3 inflammasome activation. Cell. Mol. Immunol. 2016, 13, 148–159.
- Petrilli, V.; Papin, S.; Dostert, C.; Mayor, A.; Martinon, F.; Tschopp, J. Activation of the NALP3 inflammasome is triggered by low intracellular potassium concentration. Cell Death Differ. 2007, 14, 1583–1589.
- Heid, M.E.; Keyel, P.A.; Kamga, C.; Shiva, S.; Watkins, S.C.; Salter, R.D. Mitochondrial reactive oxygen species induces NLRP3-dependent lysosomal damage and inflammasome activation. J. Immunol. 2013, 191, 5230–5238.
- Okada, M.; Matsuzawa, A.; Yoshimura, A.; Ichijo, H. The lysosome rupture-activated TAK1-JNK pathway regulates NLRP3 inflammasome activation. J. Biol. Chem. 2014, 289, 32926–32936.
- Horng, T. Calcium signaling and mitochondrial destabilization in the triggering of the NLRP3 inflammasome. Trends Immunol. 2014, 35, 253–261.
- He, Y.; Hara, H.; Nunez, G. Mechanism and Regulation of NLRP3 Inflammasome Activation. Trends Biochem. Sci. 2016, 41, 1012–1021.
- Santa Cruz Garcia, A.B.; Schnur, K.P.; Malik, A.B.; Mo, G.C.H. Gasdermin D pores are dynamically regulated by local phosphoinositide circuitry. Nat. Commun. 2022, 13, 52.
- Deng, L.; Meng, T.; Chen, L.; Wei, W.; Wang, P. The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct. Target. Ther. 2020, 5, 11.
- Nandi, D.; Tahiliani, P.; Kumar, A.; Chandu, D. The ubiquitin-proteasome system. J. Biosci. 2006, 31, 137–155.
- Liwocha, J.; Krist, D.T.; van der Heden van Noort, G.J.; Hansen, F.M.; Truong, V.H.; Karayel, O.; Purser, N.; Houston, D.; Burton, N.; Bostock, M.J.; et al. Linkage-specific ubiquitin chain formation depends on a lysine hydrocarbon ruler. Nat. Chem. Biol. 2021, 17, 272–279.
- Schauer, N.J.; Magin, R.S.; Liu, X.; Doherty, L.M.; Buhrlage, S.J. Advances in Discovering Deubiquitinating Enzyme (DUB) Inhibitors. J. Med. Chem. 2020, 63, 2731–2750.
- Varadan, R.; Walker, O.; Pickart, C.; Fushman, D. Structural properties of polyubiquitin chains in solution. J. Mol. Biol. 2002, 324, 637–647.
- Deshaies, R.J.; Joazeiro, C.A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 2009, 78, 399–434.
- Wang, W.; Hu, D.; Wu, C.; Feng, Y.; Li, A.; Liu, W.; Wang, Y.; Chen, K.; Tian, M.; Xiao, F.; et al. STING promotes NLRP3 localization in ER and facilitates NLRP3 deubiquitination to activate the inflammasome upon HSV-1 infection. PLoS Pathog. 2020, 16, e1008335.
- Murai, J.; Yang, K.; Dejsuphong, D.; Hirota, K.; Takeda, S.; D’Andrea, A.D. The USP1/UAF1 complex promotes double-strand break repair through homologous recombination. Mol. Cell. Biol. 2011, 31, 2462–2469.
- Song, H.; Zhao, C.; Yu, Z.; Li, Q.; Yan, R.; Qin, Y.; Jia, M.; Zhao, W. UAF1 deubiquitinase complexes facilitate NLRP3 inflammasome activation by promoting NLRP3 expression. Nat. Commun. 2020, 11, 6042.
- Guan, K.; Wei, C.; Zheng, Z.; Song, T.; Wu, F.; Zhang, Y.; Cao, Y.; Ma, S.; Chen, W.; Xu, Q.; et al. MAVS Promotes Inflammasome Activation by Targeting ASC for K63-Linked Ubiquitination via the E3 Ligase TRAF3. J. Immunol. 2015, 194, 4880–4890.
- Siu, K.L.; Yuen, K.S.; Castano-Rodriguez, C.; Ye, Z.W.; Yeung, M.L.; Fung, S.Y.; Yuan, S.; Chan, C.P.; Yuen, K.Y.; Enjuanes, L.; et al. Severe acute respiratory syndrome coronavirus ORF3a protein activates the NLRP3 inflammasome by promoting TRAF3-dependent ubiquitination of ASC. FASEB J. 2019, 33, 8865–8877.
- Jin, W.; Chang, M.; Sun, S.C. Peli: A family of signal-responsive E3 ubiquitin ligases mediating TLR signaling and T-cell tolerance. Cell. Mol. Immunol. 2012, 9, 113–122.
- Moynagh, P.N. The Pellino family: IRAK E3 ligases with emerging roles in innate immune signalling. Trends Immunol. 2009, 30, 33–42.
- Humphrey, S.J.; James, D.E.; Mann, M. Protein Phosphorylation: A Major Switch Mechanism for Metabolic Regulation. Trends Endocrinol. Metab. 2015, 26, 676–687.
- Zhu, K.; Jin, X.; Chi, Z.; Chen, S.; Wu, S.; Sloan, R.D.; Lin, X.; Neculai, D.; Wang, D.; Hu, H.; et al. Priming of NLRP3 inflammasome activation by Msn kinase MINK1 in macrophages. Cell. Mol. Immunol. 2021, 18, 2372–2382.
- Song, N.; Liu, Z.S.; Xue, W.; Bai, Z.F.; Wang, Q.Y.; Dai, J.; Liu, X.; Huang, Y.J.; Cai, H.; Zhan, X.Y.; et al. NLRP3 Phosphorylation Is an Essential Priming Event for Inflammasome Activation. Mol. Cell 2017, 68, 185–197.e6.
- Bittner, Z.A.; Liu, X.; Mateo Tortola, M.; Tapia-Abellan, A.; Shankar, S.; Andreeva, L.; Mangan, M.; Spalinger, M.; Kalbacher, H.; Duwell, P.; et al. BTK operates a phospho-tyrosine switch to regulate NLRP3 inflammasome activity. J. Exp. Med. 2021, 218, e20201656.
- Dan, I.; Watanabe, N.M.; Kobayashi, T.; Yamashita-Suzuki, K.; Fukagaya, Y.; Kajikawa, E.; Kimura, W.K.; Nakashima, T.M.; Matsumoto, K.; Ninomiya-Tsuji, J.; et al. Molecular cloning of MINK, a novel member of mammalian GCK family kinases, which is up-regulated during postnatal mouse cerebral development. FEBS Lett. 2000, 469, 19–23.
- Bauernfeind, F.; Bartok, E.; Rieger, A.; Franchi, L.; Nunez, G.; Hornung, V. Cutting edge: Reactive oxygen species inhibitors block priming, but not activation, of the NLRP3 inflammasome. J. Immunol. 2011, 187, 613–617.
- Stutz, A.; Kolbe, C.C.; Stahl, R.; Horvath, G.L.; Franklin, B.S.; van Ray, O.; Brinkschulte, R.; Geyer, M.; Meissner, F.; Latz, E. NLRP3 inflammasome assembly is regulated by phosphorylation of the pyrin domain. J. Exp. Med. 2017, 214, 1725–1736.
- Niu, T.; De Rosny, C.; Chautard, S.; Rey, A.; Patoli, D.; Groslambert, M.; Cosson, C.; Lagrange, B.; Zhang, Z.; Visvikis, O.; et al. NLRP3 phosphorylation in its LRR domain critically regulates inflammasome assembly. Nat. Commun. 2021, 12, 5862.
- Spalinger, M.R.; Kasper, S.; Gottier, C.; Lang, S.; Atrott, K.; Vavricka, S.R.; Scharl, S.; Raselli, T.; Frey-Wagner, I.; Gutte, P.M.; et al. NLRP3 tyrosine phosphorylation is controlled by protein tyrosine phosphatase PTPN22. J. Clin. Investig. 2016, 126, 1783–1800.
- Fischer, F.A.; Mies, L.F.M.; Nizami, S.; Pantazi, E.; Danielli, S.; Demarco, B.; Ohlmeyer, M.; Lee, M.S.J.; Coban, C.; Kagan, J.C.; et al. TBK1 and IKKepsilon act like an OFF switch to limit NLRP3 inflammasome pathway activation. Proc. Natl. Acad. Sci. USA 2021, 118, e2009309118.
- Chung, I.C.; OuYang, C.N.; Yuan, S.N.; Li, H.P.; Chen, J.T.; Shieh, H.R.; Chen, Y.J.; Ojcius, D.M.; Chu, C.L.; Yu, J.S.; et al. Pyk2 activates the NLRP3 inflammasome by directly phosphorylating ASC and contributes to inflammasome-dependent peritonitis. Sci. Rep. 2016, 6, 36214.
- Hara, H.; Tsuchiya, K.; Kawamura, I.; Fang, R.; Hernandez-Cuellar, E.; Shen, Y.; Mizuguchi, J.; Schweighoffer, E.; Tybulewicz, V.; Mitsuyama, M. Phosphorylation of the adaptor ASC acts as a molecular switch that controls the formation of speck-like aggregates and inflammasome activity. Nat. Immunol. 2013, 14, 1247–1255.
- Lin, Y.C.; Huang, D.Y.; Wang, J.S.; Lin, Y.L.; Hsieh, S.L.; Huang, K.C.; Lin, W.W. Syk is involved in NLRP3 inflammasome-mediated caspase-1 activation through adaptor ASC phosphorylation and enhanced oligomerization. J. Leukoc. Biol. 2015, 97, 825–835.
- Basak, C.; Pathak, S.K.; Bhattacharyya, A.; Mandal, D.; Pathak, S.; Kundu, M. NF-kappaB- and C/EBPbeta-driven interleukin-1beta gene expression and PAK1-mediated caspase-1 activation play essential roles in interleukin-1beta release from Helicobacter pylori lipopolysaccharide-stimulated macrophages. J. Biol. Chem. 2005, 280, 4279–4288.
- Hay, R.T. SUMO: A history of modification. Mol. Cell 2005, 18, 1–12.
- Lv, Z.; Yuan, L.; Atkison, J.H.; Williams, K.M.; Vega, R.; Sessions, E.H.; Divlianska, D.B.; Davies, C.; Chen, Y.; Olsen, S.K. Molecular mechanism of a covalent allosteric inhibitor of SUMO E1 activating enzyme. Nat. Commun. 2018, 9, 5145.
- Hay, R.T. Decoding the SUMO signal. Biochem. Soc. Trans. 2013, 41, 463–473.
- Wang, L.; Wansleeben, C.; Zhao, S.; Miao, P.; Paschen, W.; Yang, W. SUMO2 is essential while SUMO3 is dispensable for mouse embryonic development. EMBO Rep. 2014, 15, 878–885.
- Ryu, H.Y.; Ahn, S.H.; Hochstrasser, M. SUMO and cellular adaptive mechanisms. Exp. Mol. Med. 2020, 52, 931–939.
- Adorisio, S.; Fierabracci, A.; Muscari, I.; Liberati, A.M.; Ayroldi, E.; Migliorati, G.; Thuy, T.T.; Riccardi, C.; Delfino, D.V. SUMO proteins: Guardians of immune system. J. Autoimmun. 2017, 84, 21–28.
- Shao, L.; Liu, Y.; Wang, W.; Li, A.; Wan, P.; Liu, W.; Shereen, M.A.; Liu, F.; Zhang, W.; Tan, Q.; et al. SUMO1 SUMOylates and SENP3 deSUMOylates NLRP3 to orchestrate the inflammasome activation. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2020, 34, 1497–1515.
- Barry, R.; John, S.W.; Liccardi, G.; Tenev, T.; Jaco, I.; Chen, C.H.; Choi, J.; Kasperkiewicz, P.; Fernandes-Alnemri, T.; Alnemri, E.; et al. SUMO-mediated regulation of NLRP3 modulates inflammasome activity. Nat. Commun. 2018, 9, 3001.
- Duncan, J.A.; Bergstralh, D.T.; Wang, Y.; Willingham, S.B.; Ye, Z.; Zimmermann, A.G.; Ting, J.P. Cryopyrin/NALP3 binds ATP/dATP, is an ATPase, and requires ATP binding to mediate inflammatory signaling. Proc. Natl. Acad. Sci. USA 2007, 104, 8041–8046.
- Dekker, C.; Mattes, H.; Wright, M.; Boettcher, A.; Hinniger, A.; Hughes, N.; Kapps-Fouthier, S.; Eder, J.; Erbel, P.; Stiefl, N.; et al. Crystal Structure of NLRP3 NACHT Domain With an Inhibitor Defines Mechanism of Inflammasome Inhibition. J. Mol. Biol. 2021, 433, 167309.
- Danot, O.; Marquenet, E.; Vidal-Ingigliardi, D.; Richet, E. Wheel of Life, Wheel of Death: A Mechanistic Insight into Signaling by STAND Proteins. Structure 2009, 17, 172–182.
- Coll, R.C.; Hill, J.R.; Day, C.J.; Zamoshnikova, A.; Boucher, D.; Massey, N.L.; Chitty, J.L.; Fraser, J.A.; Jennings, M.P.; Robertson, A.A.B.; et al. MCC950 directly targets the NLRP3 ATP-hydrolysis motif for inflammasome inhibition. Nat. Chem. Biol. 2019, 15, 556–559.
- He, Y.; Varadarajan, S.; Munoz-Planillo, R.; Burberry, A.; Nakamura, Y.; Nunez, G. 3,4-methylenedioxy-beta-nitrostyrene inhibits NLRP3 inflammasome activation by blocking assembly of the inflammasome. J. Biol. Chem. 2014, 289, 1142–1150.
- Shim, D.W.; Shin, W.Y.; Yu, S.H.; Kim, B.H.; Ye, S.K.; Koppula, S.; Won, H.S.; Kang, T.B.; Lee, K.H. BOT-4-one attenuates NLRP3 inflammasome activation: NLRP3 alkylation leading to the regulation of its ATPase activity and ubiquitination. Sci. Rep. 2017, 7, 15020.
- Kim, H.-B.; Kwon, S.-C.; Sun, X.; Akther, M.; Han, J.-H.; Kim, T.-Y.; Kang, T.-B.; Lee, K.-H. Vanillylacetone attenuates NLRP3 inflammasome mediated immune responses in murine bone marrow derived macrophages via NLRP3 alkylation. J. Funct. Foods 2020, 64, 103655.
- Sengupta, R.; Holmgren, A. The role of thioredoxin in the regulation of cellular processes by S-nitrosylation. Biochim. Biophys. Acta 2012, 1820, 689–700.
- Hernandez-Cuellar, E.; Tsuchiya, K.; Hara, H.; Fang, R.; Sakai, S.; Kawamura, I.; Akira, S.; Mitsuyama, M. Cutting edge: Nitric oxide inhibits the NLRP3 inflammasome. J. Immunol. 2012, 189, 5113–5117.
- Sapountzi, V.; Logan, I.R.; Robson, C.N. Cellular functions of TIP60. Int. J. Biochem. Cell Biol. 2006, 38, 1496–1509.
- Zhao, K.; Zhang, Y.; Xu, X.; Liu, L.; Huang, L.; Luo, R.; Li, J.; Zhang, N.; Lu, B. Acetylation is required for NLRP3 self-aggregation and full activation of the inflammasome. bioRxiv 2019.
- Ullevig, S.L.; Kim, H.S.; Short, J.D.; Tavakoli, S.; Weintraub, S.T.; Downs, K.; Asmis, R. Protein S-Glutathionylation Mediates Macrophage Responses to Metabolic Cues from the Extracellular Environment. Antioxid. Redox Signal. 2016, 25, 836–851.
- Menon, D.; Innes, A.; Oakley, A.J.; Dahlstrom, J.E.; Jensen, L.M.; Brustle, A.; Tummala, P.; Rooke, M.; Casarotto, M.G.; Baell, J.B.; et al. GSTO1-1 plays a pro-inflammatory role in models of inflammation, colitis and obesity. Sci. Rep. 2017, 7, 17832.
- Hughes, M.M.; Hooftman, A.; Angiari, S.; Tummala, P.; Zaslona, Z.; Runtsch, M.C.; McGettrick, A.F.; Sutton, C.E.; Diskin, C.; Rooke, M.; et al. Glutathione Transferase Omega-1 Regulates NLRP3 Inflammasome Activation through NEK7 Deglutathionylation. Cell Rep. 2019, 29, 151–161.e5.
- Meissner, F.; Molawi, K.; Zychlinsky, A. Superoxide dismutase 1 regulates caspase-1 and endotoxic shock. Nat. Immunol. 2008, 9, 866–872.
- Hamarsheh, S.; Zeiser, R. NLRP3 Inflammasome Activation in Cancer: A Double-Edged Sword. Front. Immunol. 2020, 11, 1444.
- Sharma, B.R.; Kanneganti, T.D. NLRP3 inflammasome in cancer and metabolic diseases. Nat. Immunol. 2021, 22, 550–559.
- Guo, P.; Ma, X.; Zhao, W.; Huai, W.; Li, T.; Qiu, Y.; Zhang, Y.; Han, L. TRIM31 is upregulated in hepatocellular carcinoma and promotes disease progression by inducing ubiquitination of TSC1-TSC2 complex. Oncogene 2018, 37, 478–488.
- Wang, H.; Yao, L.; Gong, Y.; Zhang, B. TRIM31 regulates chronic inflammation via NF-kappaB signal pathway to promote invasion and metastasis in colorectal cancer. Am. J. Transl. Res. 2018, 10, 1247–1259.
- Song, H.; Liu, B.; Huai, W.; Yu, Z.; Wang, W.; Zhao, J.; Han, L.; Jiang, G.; Zhang, L.; Gao, C.; et al. The E3 ubiquitin ligase TRIM31 attenuates NLRP3 inflammasome activation by promoting proteasomal degradation of NLRP3. Nat. Commun. 2016, 7, 13727.
- Wei, Q.; Guo, P.; Mu, K.; Zhang, Y.; Zhao, W.; Huai, W.; Qiu, Y.; Li, T.; Ma, X.; Liu, Y.; et al. Estrogen suppresses hepatocellular carcinoma cells through ERbeta-mediated upregulation of the NLRP3 inflammasome. Lab. Investig. 2015, 95, 804–816.
- Zhang, Y.; Yang, H.; Sun, M.; He, T.; Liu, Y.; Yang, X.; Shi, X.; Liu, X. Alpinumisoflavone suppresses hepatocellular carcinoma cell growth and metastasis via NLRP3 inflammasome-mediated pyroptosis. Pharmacol. Rep. 2020, 72, 1370–1382.
- Wan, P.; Zhang, Q.; Liu, W.; Jia, Y.; Ai, S.; Wang, T.; Wang, W.; Pan, P.; Yang, G.; Xiang, Q.; et al. Cullin1 binds and promotes NLRP3 ubiquitination to repress systematic inflammasome activation. FASEB J. 2019, 33, 5793–5807.
- Dai, G.; Jiang, Z.; Sun, B.; Liu, C.; Meng, Q.; Ding, K.; Jing, W.; Ju, W. Caffeic Acid Phenethyl Ester Prevents Colitis-Associated Cancer by Inhibiting NLRP3 Inflammasome. Front. Oncol. 2020, 10, 721.
