In November of 2021, a recently evolved variant of SARS-CoV-2, omicron, was discovered. In just one month, omicron has spread to more than 89 countries resulting in a rapid rise in cases and a new wave of infections. With over 46 mutations, omicron brings concern to the public health and may be able to infect at a greater capacity than previous strains. Although able to infect double vaccinated and previously infected individuals, the booster vaccine may prove promising. However, more research is needed to fully elucidate the key function of each mutation and to better develop effective drugs. Marine resources may be a promising drug discovery avenue to investigate. Through viral entry blockade and preventing viral replication and protein synthesis, metabolites produced from marine organisms may be promising against the evolving SARS-CoV-2.
- omicron
- marine resources
- COVID-19
1. Introduction
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) first emerged almost two years ago and has been ongoing since [1]. It is without a doubt that the SARS-CoV-2 pandemic has led to frustration and prevalent negative effects on the mental wellbeing of individuals as well as the economy [2][3][4]. Vaccinations were effective at first, however, the solution was temporary, and the single-stranded SARS-CoV-2 virus is continuously evolving through mutations in key domains [5][6][7][8]. Several SARS-CoV-2 variants including the alpha, beta and delta strains were associated with new waves of COVID-19 infection [9][10]. The delta variant was shown to have a higher transmissible and infectious risk than other strains with reports of having a higher viral load and mutations granting a better immune response escape [11][12][13]. On 24 November 2021, a new SARS-CoV-2 variant, known as omicron (B.1.1.529) was reported to World Health Organization (WHO) [14]. The omicron variant was first reported on 11 November 2021, in Botswana and just a few days later, was identified in South Africa [15][16]. To date (January 2022), omicron has been reported in more than 57 countries, with South Korea reporting its highest number of COVID-19 cases in December 2021, since the start of the pandemic [17]. It has not been elucidated whether the transmissibility and severity status of the omicron variant is greater than the previous strains [18][19]. Over 45 mutations and several gene deletions have been identified, some in the same genes as the previous variants and others being novel (Figure 1) [20]. Together, these mutations in key regions give the omicron strain the ability to escape the host immune system, rendering vaccinations less effective [20][21]. Furthermore, it is not clear yet what the effects of several mutations are, and this serves as a barrier for developing new vaccinations or drugs for prevention and treatment of omicron viral infection [22][23].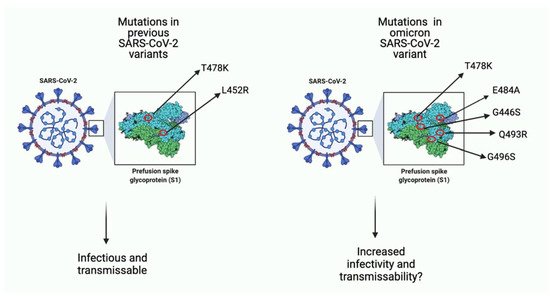
2. The Potential for Prevention/Treatment of SARS-CoV-2 Variants Using Marine Resources
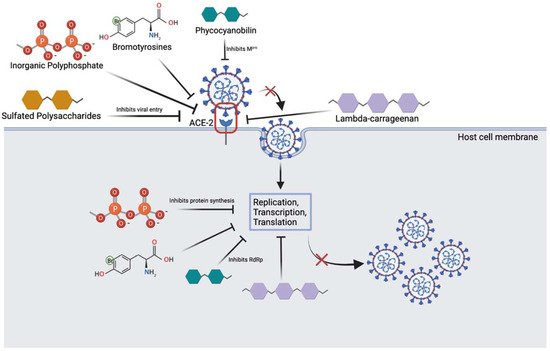
| Marine Compound | Sources | Mechanism of Action |
|---|---|---|
| Inorganic polyphosphate (polyP) [26][27] | Marine bacteria and sponges |
|
| Lambda-carrageenan [24][28] | Marine algae |
|
| Phycocyanobilins (PCB) [40][41] | Cyanobacteria and algae rhodophytes |
|
| Sulfated polysaccharides [42][43][44] | Cyanobacteria, sea cucumber, microalgae |
|
| Bromotyrosines [29][46][47][50] | Marine sponges |
|
| Caffeic acid hexoside, phloretin, cholestan-3-ol, 2-methylene [48][49] | Marine seaweed |
|
References
- Yang, L.; Liu, S.; Liu, J.; Zhang, Z.; Wan, X.; Huang, B.; Chen, Y.; Zhang, Y. COVID-19: Immunopathogenesis and Immunotherapeutics. Sig. Transduct. Target. Ther. 2020, 5, 128.
- Talevi, D.; Socci, V.; Carai, M.; Carnaghi, G.; Faleri, S.; Trebbi, E.; di Bernardo, A.; Capelli, F.; Pacitti, F. Mental Health Outcomes of the COVID-19 Pandemic. Riv. Psichiatr. 2020, 55, 137–144.
- Jin, H.; Wang, H.; Li, X.; Zheng, W.; Ye, S.; Zhang, S.; Zhou, J.; Pennington, M. Economic Burden of COVID-19, China, January–March, 2020: A Cost-of-Illness Study. Bull. World Health Organ. 2021, 99, 112–124.
- Oberndorfer, M.; Dorner, T.E.; Brunnmayr, M.; Berger, K.; Dugandzic, B.; Bach, M. Health-related and Socio-economic Burden of the COVID-19 Pandemic in Vienna. Health Soc. Care Commun. 2021.
- Rahman, M.S.; Islam, M.R.; Alam, A.S.M.R.U.; Islam, I.; Hoque, M.N.; Akter, S.; Rahaman, M.M.; Sultana, M.; Hossain, M.A. Evolutionary Dynamics of SARS-CoV-2 Nucleocapsid Protein and Its Consequences. J. Med. Virol. 2021, 93, 2177–2195.
- Giovanetti, M.; Benedetti, F.; Campisi, G.; Ciccozzi, A.; Fabris, S.; Ceccarelli, G.; Tambone, V.; Caruso, A.; Angeletti, S.; Zella, D.; et al. Evolution Patterns of SARS-CoV-2: Snapshot on Its Genome Variants. Biochem. Biophys. Res. Commun. 2021, 538, 88–91.
- Shah, M.; Woo, H.G. Omicron: A Heavily Mutated SARS-CoV-2 Variant Exhibits Stronger Binding to ACE2 and Potently Escape Approved COVID-19 Therapeutic Antibodies. Genomics 2021.
- Spratt, A.N.; Kannan, S.R.; Woods, L.T.; Weisman, G.A.; Quinn, T.P.; Lorson, C.L.; Sönnerborg, A.; Byrareddy, S.N.; Singh, K. Evolution, Correlation, Structural Impact and Dynamics of Emerging SARS-CoV-2 Variants. Comput. Struct. Biotechnol. J. 2021, 19, 3799–3809.
- Duong, D. Alpha, Beta, Delta, Gamma: What’s Important to Know about SARS-CoV-2 Variants of Concern? CMAJ 2021, 193, E1059–E1060.
- Cele, S.; Karim, F.; Lustig, G.; San, J.E.; Hermanus, T.; Tegally, H.; Snyman, J.; Moyo-Gwete, T.; Wilkinson, E.; Bernstein, M.; et al. SARS-CoV-2 Evolved during Advanced HIV Disease Immunosuppression Has Beta-like Escape of Vaccine and Delta Infection Elicited Immunity. Infect. Dis. 2021.
- Wang, Y.; Chen, R.; Hu, F.; Lan, Y.; Yang, Z.; Zhan, C.; Shi, J.; Deng, X.; Jiang, M.; Zhong, S.; et al. Transmission, Viral Kinetics and Clinical Characteristics of the Emergent SARS-CoV-2 Delta VOC in Guangzhou, China. EClinicalMedicine 2021, 40, 101129.
- van Kampen, J.J.A.; van de Vijver, D.A.M.C.; Fraaij, P.L.A.; Haagmans, B.L.; Lamers, M.M.; Okba, N.; van den Akker, J.P.C.; Endeman, H.; Gommers, D.A.M.P.J.; Cornelissen, J.J.; et al. Duration and Key Determinants of Infectious Virus Shedding in Hospitalized Patients with Coronavirus Disease-2019 (COVID-19). Nat. Commun. 2021, 12, 267.
- Farinholt, T.; Doddapaneni, H.; Qin, X.; Menon, V.; Meng, Q.; Metcalf, G.; Chao, H.; Gingras, M.-C.; Avadhanula, V.; Farinholt, P.; et al. Transmission Event of SARS-CoV-2 Delta Variant Reveals Multiple Vaccine Breakthrough Infections. BMC Med. 2021, 19, 255.
- World Health Organization. Classification of Omicron (B.1.1.529): SARS-CoV-2 Variant of Concern. Available online: https://www.who.int/news/item/26-11-2021-classification-of-omicron-(b.1.1.529)-sars-cov-2-variant-of-concern (accessed on 19 December 2021).
- Dyer, O. COVID-19: South Africa’s Surge in Cases Deepens Alarm over Omicron Variant. BMJ 2021, 375, n3013.
- Wang, L.; Cheng, G. Sequence Analysis of the Emerging SARS-CoV-2 Variant Omicron in South Africa. J. Med. Virol. 2021, 94, 1728–1733.
- Mahase, E. COVID-19: Do Vaccines Work against Omicron—and Other Questions Answered. BMJ 2021, 375, n3062.
- Ingraham, N.E.; Ingbar, D.H. The Omicron Variant of SARS-CoV-2: Understanding the Known and Living with Unknowns. Clin. Transl. Med. 2021, 11, e685.
- Karim, S.S.A.; Karim, Q.A. Omicron SARS-CoV-2 Variant: A New Chapter in the COVID-19 Pandemic. Lancet 2021, 398, 2126–2128.
- Kannan, S.R.; Spratt, A.N.; Sharma, K.; Chand, H.S.; Byrareddy, S.N.; Singh, K. Omicron SARS-CoV-2 Variant: Unique Features and Their Impact on Pre-Existing Antibodies. J. Autoimmun. 2022, 126, 102779.
- Cao, Y.; Wang, J.; Jian, F.; Xiao, T.; Song, W.; Yisimayi, A.; Huang, W.; Li, Q.; Wang, P.; An, R.; et al. B.1.1.529 Escapes the Majority of SARS-CoV-2 Neutralizing Antibodies of Diverse Epitopes. Immunology 2021.
- Kumar, S.; Thambiraja, T.S.; Karuppanan, K.; Subramaniam, G. Omicron and Delta Variant of SARS-CoV-2: A Comparative Computational Study of Spike Protein. J. Med. Virol. 2022, 94, 1641–1649.
- Cevik, M.; Grubaugh, N.D.; Iwasaki, A.; Openshaw, P. COVID-19 Vaccines: Keeping Pace with SARS-CoV-2 Variants. Cell 2021, 184, 5077–5081.
- Geahchan, S.; Ehrlich, H.; Rahman, M.A. The Anti-Viral Applications of Marine Resources for COVID-19 Treatment: An Overview. Mar. Drugs 2021, 19, 409.
- Khan, M.T.; Ali, A.; Wang, Q.; Irfan, M.; Khan, A.; Zeb, M.T.; Zhang, Y.-J.; Chinnasamy, S.; Wei, D.-Q. Marine Natural Compounds as Potents Inhibitors against the Main Protease of SARS-CoV-2—A Molecular Dynamic Study. J. Biomol. Struct. 2021, 39, 3627–3637.
- Ferrucci, V.; Kong, D.-Y.; Asadzadeh, F.; Marrone, L.; Boccia, A.; Siciliano, R.; Criscuolo, G.; Anastasio, C.; Quarantelli, F.; Comegna, M.; et al. Long-Chain Polyphosphates Impair SARS-CoV-2 Infection and Replication. Sci. Signal. 2021, 14, eabe5040.
- Neufurth, M.; Wang, X.; Tolba, E.; Lieberwirth, I.; Wang, S.; Schröder, H.C.; Müller, W.E.G. The Inorganic Polymer, Polyphosphate, Blocks Binding of SARS-CoV-2 Spike Protein to ACE2 Receptor at Physiological Concentrations. Biochem. Pharmacol. 2020, 182, 114215.
- Jang, Y.; Shin, H.; Lee, M.K.; Kwon, O.S.; Shin, J.S.; Kim, Y.; Kim, C.W.; Lee, H.-R.; Kim, M. Antiviral Activity of Lambda-Carrageenan against Influenza Viruses and Severe Acute Respiratory Syndrome Coronavirus 2. Sci. Rep. 2021, 11, 821.
- Muzychka, L.; Voronkina, A.; Kovalchuk, V.; Smolii, O.B.; Wysokowski, M.; Petrenko, I.; Youssef, D.T.A.; Ehrlich, I.; Ehrlich, H. Marine Biomimetics: Bromotyrosines Loaded Chitinous Skeleton as Source of Antibacterial Agents. Appl. Phys. A 2021, 127, 15.
- Asif, M.; Saleem, M.; Yaseen, H.S.; Yehya, A.H.; Saadullah, M.; Zubair, H.M.; Oon, C.E.; Khaniabadi, P.M.; Khalid, S.H.; Khan, I.U.; et al. Potential Role of Marine Species-Derived Bioactive Agents in the Management of SARS-CoV-2 Infection. Future Microbiol. 2021, 16, 1289–1301.
- El-Demerdash, A.; Al-Karmalawy, A.A.; Abdel-Aziz, T.M.; Elhady, S.S.; Darwish, K.M.; Hassan, A.H.E. Investigating the Structure–Activity Relationship of Marine Natural Polyketides as Promising SARS-CoV-2 Main Protease Inhibitors. RSC Adv. 2021, 11, 31339–31363.
- Singh, R.; Chauhan, N.; Kuddus, M. Exploring the Therapeutic Potential of Marine-Derived Bioactive Compounds against COVID-19. Environ. Sci. Pollut. Res. 2021, 28, 52798–52809.
- Lira, S.P.D.; Seleghim, M.H.R.; Williams, D.E.; Marion, F.; Hamill, P.; Jean, F.; Andersen, R.J.; Hajdu, E.; Berlinck, R.G.S. A SARS-Coronovirus 3CL Protease Inhibitor Isolated from the Marine Sponge Axinella Cf. Corrugata: Structure Elucidation and Synthesis. J. Braz. Chem. Soc. 2007, 18, 440–443.
- Drechsel, A.; Helm, J.; Ehrlich, H.; Pantovic, S.; Bornstein, S.R.; Bechmann, N. Anti-Tumor Activity vs. Normal Cell Toxicity: Therapeutic Potential of the Bromotyrosines Aerothionin and Homoaerothionin In Vitro. Mar. Drugs 2020, 18, 236.
- Lianingsih, F. In Silico Analysis of Sponges Compound Against Mpro COVID-19: A Review. Proc. Int. Conf. Eng. 2021, 4, 296–300.
- Bechmann, N.; Ehrlich, H.; Eisenhofer, G.; Ehrlich, A.; Meschke, S.; Ziegler, C.; Bornstein, S. Anti-Tumorigenic and Anti-Metastatic Activity of the Sponge-Derived Marine Drugs Aeroplysinin-1 and Isofistularin-3 against Pheochromocytoma In Vitro. Mar. Drugs 2018, 16, 172.
- Binnewerg, B.; Schubert, M.; Voronkina, A.; Muzychka, L.; Wysokowski, M.; Petrenko, I.; Djurović, M.; Kovalchuk, V.; Tsurkan, M.; Martinovic, R.; et al. Marine Biomaterials: Biomimetic and Pharmacological Potential of Cultivated Aplysina Aerophoba Marine Demosponge. Mater. Sci. Eng. C 2020, 109, 110566.
- Ehrlich, H.; Bazhenov, V.; Meschke, S.; Bürger, M.; Ehrlich, A.; Petovic, S.; Durovic, M. Marine Invertebrates of Boka Kotorska Bay Unique Sources for Bioinspired Materials Science. In The Boka Kotorska Bay Environment; Joksimović, A., Djurović, M., Semenov, A.V., Zonn, I.S., Kostianoy, A.G., Eds.; The Handbook of Environmental Chemistry; Springer International Publishing: Cham, Switzerland, 2016; Volume 54, pp. 313–334. ISBN 978-3-319-51613-4.
- Hamoda, A.M.; Fayed, B.; Ashmawy, N.S.; El-Shorbagi, A.-N.A.; Hamdy, R.; Soliman, S.S.M. Marine Sponge Is a Promising Natural Source of Anti-SARS-CoV-2 Scaffold. Front. Pharmacol. 2021, 12, 666664.
- Pendyala, B.; Patras, A. In Silico Screening of Food Bioactive Compounds to Predict Potential Inhibitors of COVID-19 Main Protease (Mpro) and RNA-Dependent RNA Polymerase (RdRp). Chemistry 2020.
- Petit, L.; Vernès, L.; Cadoret, J.-P. Docking and in Silico Toxicity Assessment of Arthrospira Compounds as Potential Antiviral Agents against SARS-CoV-2. J. Appl. Phycol. 2021, 33, 1579–1602.
- Song, S.; Peng, H.; Wang, Q.; Liu, Z.; Dong, X.; Wen, C.; Ai, C.; Zhang, Y.; Wang, Z.; Zhu, B. Inhibitory Activities of Marine Sulfated Polysaccharides against SARS-CoV-2. Food Funct. 2020, 11, 7415–7420.
- Grassauer, A.; Weinmuellner, R.; Meier, C.; Pretsch, A.; Prieschl-Grassauer, E.; Unger, H. Iota-Carrageenan Is a Potent Inhibitor of Rhinovirus Infection. Virol. J. 2008, 5, 107.
- Morokutti-Kurz, M.; Fröba, M.; Graf, P.; Große, M.; Grassauer, A.; Auth, J.; Schubert, U.; Prieschl-Grassauer, E. Iota-Carrageenan Neutralizes SARS-CoV-2 and Inhibits Viral Replication in Vitro. PLoS ONE 2021, 16, e0237480.
- Schubert, M.; Binnewerg, B.; Voronkina, A.; Muzychka, L.; Wysokowski, M.; Petrenko, I.; Kovalchuk, V.; Tsurkan, M.; Martinovic, R.; Bechmann, N.; et al. Naturally Prefabricated Marine Biomaterials: Isolation and Applications of Flat Chitinous 3D Scaffolds from Ianthella Labyrinthus (Demospongiae: Verongiida). Int. J. Mol. Sci. 2019, 20, 5105.
- Fromont, J.; Żółtowska-Aksamitowska, S.; Galli, R.; Meissner, H.; Erpenbeck, D.; Vacelet, J.; Diaz, C.; Tsurkan, M.V.; Petrenko, I.; Youssef, D.T.A.; et al. New Family and Genus of a Dendrilla-like Sponge with Characters of Verongiida. Part II. Discovery of Chitin in the Skeleton of Ernstilla Lacunosa. Zool. Anz. 2019, 280, 21–29.
- Kovalchuk, V.; Voronkina, A.; Binnewerg, B.; Schubert, M.; Muzychka, L.; Wysokowski, M.; Tsurkan, M.V.; Bechmann, N.; Petrenko, I.; Fursov, A.; et al. Naturally Drug-Loaded Chitin: Isolation and Applications. Mar. Drugs 2019, 17, 574.
- Bharathi, M.; Sivamaruthi, B.S.; Kesika, P.; Thangaleela, S.; Chaiyasut, C. In Silico Screening of Bioactive Compounds of Representative Seaweeds to Inhibit SARS-CoV-2 ACE2-Bound Omicron B.1.1.529 Spike Protein Trimer. Mar. Drugs 2022, 20, 148.
- Behzad, S.; Sureda, A.; Barreca, D.; Nabavi, S.F.; Rastrelli, L.; Nabavi, S.M. Health Effects of Phloretin: From Chemistry to Medicine. Phytochem. Rev. 2017, 16, 527–533.
- Schubert, M.; Bertoglio, F.; Steinke, S.; Heine, P.A.; Ynga-Durand, M.A.; Zuo, F.; Du, L.; Korn, J.; Milošević, M.; Wenzel, E.V.; et al. Human Serum from SARS-CoV-2 Vaccinated and COVID-19 Patients Shows Reduced Binding to the RBD of SARS-CoV-2 Omicron Variant. BMC Med. 2022, 20, 102.
