| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Manuel Piñeiro Fiel | + 1465 word(s) | 1465 | 2021-03-15 08:55:11 | | | |
| 2 | Vicky Zhou | Meta information modification | 1465 | 2021-03-25 07:01:39 | | |
Video Upload Options
PET radiomics is a new medical imaging field exploiting image features to develop novel diagnostic, predictive and prognostic multiparametric models to support personalized clinical decisions and improve individualized treatment selection.
1. Introduction
Decades of research on cancer biology have revealed that tumors are heterogeneous entities at all scales (macroscopic, physiological, microscopic and genetic) [1][2][3][4], with different regions showing distinct morphological and phenotypic profiles [5][6][7]. Nowadays, it is widely accepted that tumor heterogeneity has profound implications in tumor development, therapeutic outcomes and survival [8][9][10][11], making it essential to develop methods for studying tumor heterogeneity in vivo [12].
In this context, non-invasive imaging techniques, such as magnetic resonance (MR), computed tomography (CT) and positron emission tomography (PET) become relevant due to their ability to provide information on the whole tumor in one acquisition [13]. Nowadays, imaging is central to cancer management, having applications in screening, diagnosis, staging, prognosis and treatment response, among others [14][15][16][17][18]. Mainly, PET has emerged as the predominant imaging modality, overperforming conventional imaging techniques for the evaluation of blood [19], head and neck [20] or lung cancer [21]; and it has been increasingly proposed as an ideal tool for characterizing the biology of tumors at the macroscopic scale [13][22][23][24][25]. Over the last years, there has been an increasing interest in extracting quantitative information from PET images using image analysis [26][27]. Thus, semi-quantitative parameters such as the standard uptake value (SUV), the metabolic tumor volume (MTV) or the total lesion glycolysis (TLG) [28] obtained from 18F-fluorodeoxyglucose PET (FDG-PET) images, have been demonstrated to provide relatively objective information useful for the diagnosis, earlier evaluation and monitoring of treatment response [24][25][28]. These parameters are now fully incorporated into clinical guidelines and commonly measured at most hospitals in developed countries [29].
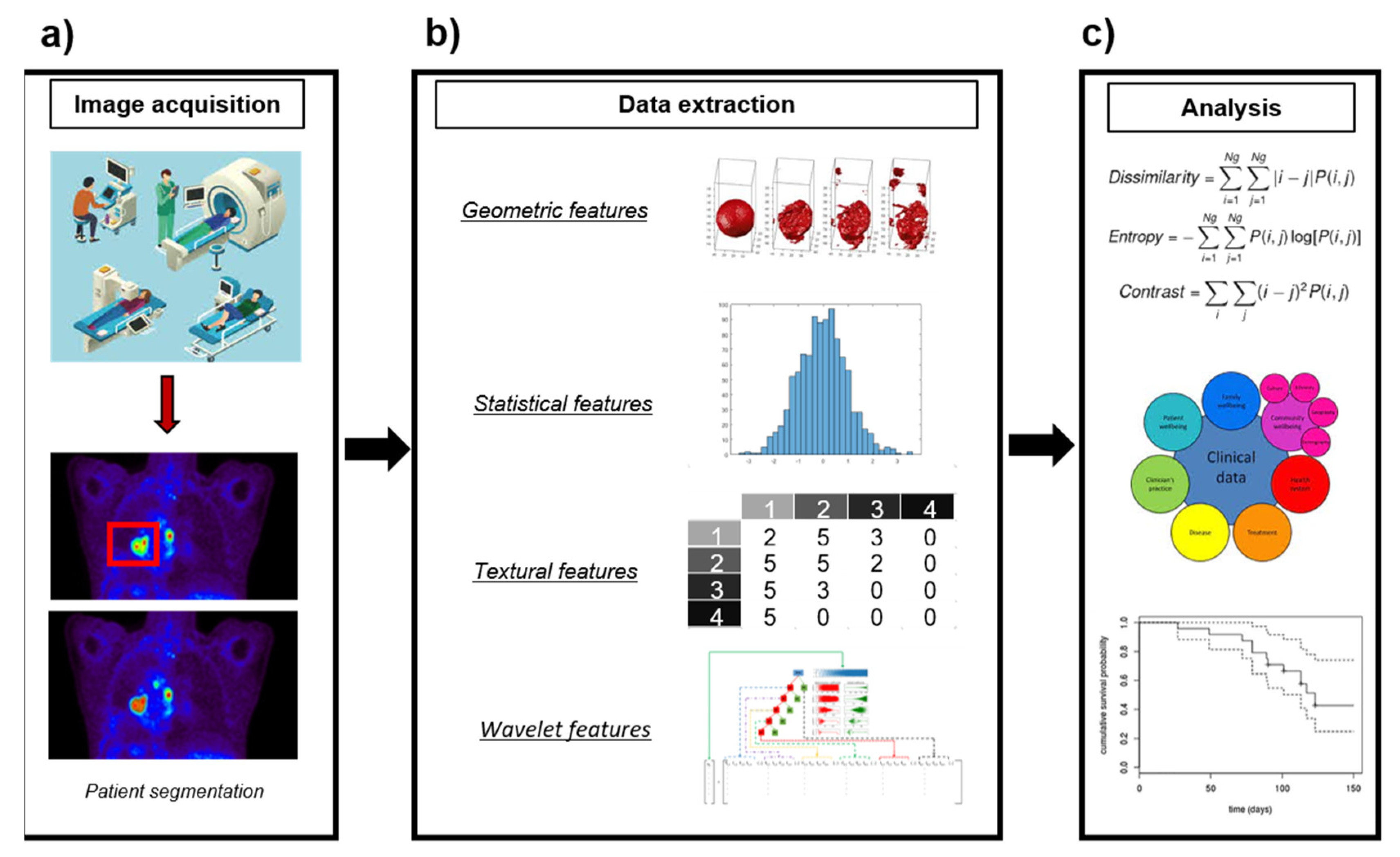
Several research studies have pointed out that high-order textural features derived from PET images can provide information about tumor heterogeneity, expanding the information available from clinical reports, laboratory tests and genomic or proteomic assays [26][27][30]. This has led to the incorporation of PET imaging to radiomics, a new medical imaging field exploiting image features to develop novel diagnostic, predictive and prognostic multiparametric models to support personalized clinical decisions and improve individualized treatment selection [27][31][32] (Figure 1). Textural analysis has long been applied in CT [33] and MRI [34], but it had not been introduced in PET until the last decade. Since then, increasing numbers of studies have suggested that PET textural features would be correlated with tumor biology and heterogeneity [35][36][37][38][39][40][41][42], providing valuable information for tailoring individual treatments [13][18][23][43][44][45][46].
Despite the promising early reports, numerous studies have highlighted the challenges to be addressed before the use of PET radiomics becomes reliable and interpretable [46]. In addition to issues common to PET imaging itself, such as noise or partial volume effects [47][48][49], radiomics must deal with standardization issues related to the differences in acquisition and reconstruction parameters, post-processing techniques, tumor segmentation methods or even texture calculations [46][50][51][52][53]. The complex formulation of radiomics makes it challenging to explain pervasive findings such as the correlations displayed between different texture indices [54][55][56][57][58][59][60], as well as strong correlations between textural indices and tumor volume [61][62][63], which compromise the added value of these parameters in comparison with SUV or MTV. Furthermore, the variability in definitions and nomenclature of heterogeneity metrics themselves complicate any evaluation and comparison of published results [64]. Finally, there is no consensus about how many patients are needed, which textures must be extracted, or what methodologies should be applied for proper validation [24][46][52].
Due to the prominent role of quantitative image analysis in the diagnosis, tumor characterization and prognosis of cancer patients, the development of reliable and well-validated image analysis methods is of paramount importance. Particularly, parallel with the popularization of PET radiomics, several authors have highlighted the methodological and statistical issues in their works.
2. PET Textural Analysis and Radiomics in Cancer
Lung, head and neck, breast and gynecological cancers emerged as the most studied cancer types. It is well-known that radiomics is a demanding technique in terms of data, and thus we assume that this is a consequence of the prevalence of these types of cancer [65]. The average number of patients per study was 114, and we observed that 65% of the evaluated publications included data from less than 100 patients (29%, less than 50 patients). Previous works evaluating radiomic methodologies suggested a proportion of five patients per evaluated feature to avoid model overfitting [52][66] or a minimum of 100 patients [31]. Based on this and considering that the average number of high-order features was 31 (median = 26, range, 1–286), we would recommend including larger numbers of patients in further works. In addition to this, a limited number of publications (8%) provided a validation of the proposed models using independent databases or a validation in an independent subsample of the initial dataset (28%). We assume that this fact is related to the small amount of data available. Publications with higher numbers of patients used resources such as The Cancer Imaging Archive (TCIA) [67]. We expect that the continuous growth of public imaging databases will improve these numbers in the future and provide a common playground where algorithms could be compared and validated. The rise of collaborative models such as federated learning [68][69] where different centers share models validated across centers while keeping their own data anonymous will also generate more robust algorithms in the near future.
Furthermore, we observed that very few papers aimed at externally validating previously developed models [70][71]. We believe that this should be a common practice in the field. Nevertheless, we observed that most of the publications did not provide easy access to the developed models. As for data sharing, accessibility of software is a substantial concern, reducing the applicability and potential impact of published studies and models [72]. The collaboration between centers and researchers should be improved for further model validation, which is needed to generalize PET radiomics. In this regards, van Griethuysen et al. recently released PyRadiomics, an open modular platform to provide the community with standard tools for promoting further independent development and evaluation [73].
Regarding the objectives of the evaluated publications, the evaluation of prognosis or treatment response was the most common (46%), followed by diagnosis and staging (21%), tumor characterization (articles aimed at predicting the biological characteristics of tumors traditionally obtained by other means (i.e., histology, genetics)) (18%) and the evaluation of technical factors (15%). This result is not surprising, as textures are intended to measure heterogeneity, well-known to be related to aggressiveness and poor outcomes [74][75][76]. We believe that more papers focused on tumor characterization and technical factors would be useful in this regard. On the one hand, although heterogeneity itself was proposed as a biomarker in the past [77][78] it has been reported that heterogeneity is related to biological characteristics such as tumor microenvironment [79][80], genetic expression [81] and, macroscopically, tumor grades or cancer subtypes [82]; thus, linking radiomic features with particular biological characteristics could provide additional evidence for the field. On the other hand, further work is still needed to evaluate the impact of image acquisition, reconstruction, post-processing and feature calculation [83]. In an exemplary work, Bodowicz et al. investigated an association of PET radiomics with local tumor control after radiochemotherapy in head and neck cancer, developing radiomic implementations with two different software packages. Of the 649 features calculated, only 12% were reproducible between the two software implementations, and, although both models were similarly predictive, they included different sets of features, pointing to the need of further harmonization on feature calculation. Harmonization initiatives, such as the Image Biomarker Standardization Initiative [64], would be extremely helpful for this process, as reliable and reproducible measurements are of paramount importance for biomarkers to progress on their validation.
Most of the evaluated studies (78%) concluded that the inclusion of textural features improved the results of models developed with clinical or first-order imaging metrics, which is a powerful conclusion. Nevertheless, it must be interpreted with caution since, as mentioned above, most of the evaluated studies included an insufficient number of patients or were not extensively validated. Many studies reported models including both first and high-order features, which were compared usually with counterparts including a reduced number of features, so an improvement of the result can be caused by overfitting [84][85]. Several papers from our group [63] and others [60][86][87] have suggested a strong correlation between the most used textural features and conventional parameters such as MTV and SUV, which must be untangled.
In summary, PET radiomics is an up-and-coming field, and PET radiomics might have a role in clinical practice in the future. In this regard, the results of the most relevant papers are very appealing. Nevertheless, we have identified several methodological concerns related to the validation of the purposed algorithms, the number of patients included, the lack of data and software accessibility and a need for further methodological standardization. We have also observed that the community is already developing solutions to overcome these limitations.
References
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; Tarpey, P.; et al. Intratumor Heterogeneity and Branched Evolution Revealed by Multiregion Sequencing. N. Engl. J. Med. 2012, 366, 883–892.
- McGranahan, N.; Swanton, C. Biological and Therapeutic Impact of Intratumor Heterogeneity in Cancer Evolution. Cancer Cell 2015, 27, 15–26.
- Leskela, S.; Pérez-Mies, B.; Rosa-Rosa, J.M.; Cristobal, E.; Biscuola, M.; Palacios-Berraquero, M.L.; Ong, S.; Guia, X.M.-G.; Palacios, J. Molecular Basis of Tumor Heterogeneity in Endometrial Carcinosarcoma. Cancers 2019, 11, 964.
- Hass, R.; von der Ohe, J.; Ungefroren, H. Impact of the Tumor Microenvironment on Tumor Heterogeneity and Consequences for Cancer Cell Plasticity and Stemness. Cancers 2020, 12, 3716.
- Tellez-Gabriel, M.; Ory, B.; Lamoureux, F.; Heymann, M.-F.; Heymann, D. Tumor Heterogeneity: The Key Advantages of Single-Cell Analysis. Int. J. Mol. Sci 2016, 17, 2142.
- Michor, F.; Polyak, K. The Origins and Implications of Intratumor Heterogeneity. Cancer Prev Res. 2010, 3, 1361–1364.
- Visvader, J.E. Cells of Origin in Cancer. Nature 2011, 469, 314–322.
- Marusyk, A.; Polyak, K. Tumor Heterogeneity: Causes and Consequences. Biochim. Biophys. Acta 2010, 1805, 105–117.
- Roma-Rodrigues, C.; Mendes, R.; Baptista, P.V.; Fernandes, A.R. Targeting Tumor Microenvironment for Cancer Therapy. Int. J. Mol. Sci. 2019, 20, 840.
- Lim, Z.-F.; Ma, P.C. Emerging Insights of Tumor Heterogeneity and Drug Resistance Mechanisms in Lung Cancer Targeted Therapy. J. Hematol. Oncol. 2019, 12, 134.
- Baliu-Piqué, M.; Pandiella, A.; Ocana, A. Breast Cancer Heterogeneity and Response to Novel Therapeutics. Cancers 2020, 12, 3271.
- Bonin, S.; Stanta, G. Pre-Analytics and Tumor Heterogeneity. New Biotechnol. 2020, 55, 30–35.
- Davnall, F.; Yip, C.S.P.; Ljungqvist, G.; Selmi, M.; Ng, F.; Sanghera, B.; Ganeshan, B.; Miles, K.A.; Cook, G.J.; Goh, V. Assessment of Tumor Heterogeneity: An Emerging Imaging Tool for Clinical Practice? Insights Imaging 2012, 3, 573–589.
- Fass, L. Imaging and Cancer: A Review. Mol. Oncol. 2008, 2, 115–152.
- Emaminejad, N.; Qian, W.; Guan, Y.; Tan, M.; Qiu, Y.; Liu, H.; Zheng, B. Fusion of Quantitative Image and Genomic Biomarkers to Improve Prognosis Assessment of Early Stage Lung Cancer Patients. IEEE Trans. Biomed. Eng. 2016, 63, 1034–1043.
- Popovici, V.; Budinská, E.; Dušek, L.; Kozubek, M.; Bosman, F. Image-Based Surrogate Biomarkers for Molecular Subtypes of Colorectal Cancer. Bioinformatics 2017, 33, 2002–2009.
- Scalco, E.; Rizzo, G. Texture Analysis of Medical Images for Radiotherapy Applications. Br. J. Radiol. 2016, 90.
- Aerts, H.J.W.L.; Velazquez, E.R.; Leijenaar, R.T.H.; Parmar, C.; Grossmann, P.; Carvalho, S.; Cavalho, S.; Bussink, J.; Monshouwer, R.; Haibe-Kains, B.; et al. Decoding Tumor Phenotype by Noninvasive Imaging Using a Quantitative Radiomics Approach. Nat. Commun. 2014, 5, 4006.
- Trotman, J.; Luminari, S.; Boussetta, S.; Versari, A.; Dupuis, J.; Tychyj, C.; Marcheselli, L.; Berriolo-Riedinger, A.; Franceschetto, A.; Julian, A.; et al. Prognostic Value of PET-CT after First-Line Therapy in Patients with Follicular Lymphoma: A Pooled Analysis of Central Scan Review in Three Multicentre Studies. Lancet Haematol. 2014, 1, e17–e27.
- Szyszko, T.A.; Cook, G.J.R. PET/CT and PET/MRI in Head and Neck Malignancy. Clin. Radiol. 2018, 73, 60–69.
- Al-Jahdali, H.; Khan, A.N.; Loutfi, S.; Al-Harbi, A.S. Guidelines for the Role of FDG-PET/CT in Lung Cancer Management. J. Infect. Public Health 2012, 5 (Suppl. S1), S35–S40.
- Krause, B.J.; Schwarzenböck, S.; Souvatzoglou, M. FDG PET and PET/CT. Recent Results Cancer Res. 2013, 187, 351–369.
- Chicklore, S.; Goh, V.; Siddique, M.; Roy, A.; Marsden, P.K.; Cook, G.J.R. Quantifying Tumor Heterogeneity in 18F-FDG PET/CT Imaging by Texture Analysis. Eur. J. Nucl. Med. Mol. Imaging 2013, 40, 133–140.
- Fonti, R.; Conson, M.; Del Vecchio, S. PET/CT in Radiation Oncology. Semin. Oncol. 2019, 46, 202–209.
- Miller, T.R.; Pinkus, E.; Dehdashti, F.; Grigsby, P.W. Improved Prognostic Value of 18F-FDG PET Using a Simple Visual Analysis of Tumor Characteristics in Patients with Cervical Cancer. J. Nucl. Med. 2003, 44, 192–197.
- Avanzo, M.; Stancanello, J.; El Naqa, I. Beyond Imaging: The Promise of Radiomics. Phys. Med. 2017, 38, 122–139.
- Kumar, V.; Gu, Y.; Basu, S.; Berglund, A.; Eschrich, S.A.; Schabath, M.B.; Forster, K.; Aerts, H.J.W.L.; Dekker, A.; Fenstermacher, D.; et al. Radiomics: The Process and the Challenges. Magn. Reson. Imaging 2012, 30, 1234–1248.
- Fujima, N.; Hirata, K.; Shiga, T.; Yasuda, K.; Onimaru, R.; Tsuchiya, K.; Kano, S.; Mizumachi, T.; Homma, A.; Kudo, K.; et al. Semi-Quantitative Analysis of Pre-Treatment Morphological and Intratumoral Characteristics Using 18F-Fluorodeoxyglucose Positron-Emission Tomography as Predictors of Treatment Outcome in Nasal and Paranasal Squamous Cell Carcinoma. Quant. Imaging Med. Surg. 2018, 8, 788–795.
- Joo Hyun, O.; Lodge, M.A.; Wahl, R.L. Practical PERCIST: A Simplified Guide to PET Response Criteria in Solid Tumors 1.0. Radiology 2016, 280, 576–584.
- Parekh, V.; Jacobs, M.A. Radiomics: A New Application from Established Techniques. Expert Rev. Precis. Med. Drug Dev. 2016, 1, 207–226.
- Gillies, R.J.; Kinahan, P.E.; Hricak, H. Radiomics: Images Are More than Pictures, They Are Data. Radiology 2016, 278, 563–577.
- Lambin, P.; Rios-Velazquez, E.; Leijenaar, R.; Carvalho, S.; van Stiphout, R.G.P.M.; Granton, P.; Zegers, C.M.L.; Gillies, R.; Boellard, R.; Dekker, A.; et al. Radiomics: Extracting More Information from Medical Images Using Advanced Feature Analysis. Eur. J. Cancer 2012, 48, 441–446.
- Lubner, M.G.; Smith, A.D.; Sandrasegaran, K.; Sahani, D.V.; Pickhardt, P.J. CT Texture Analysis: Definitions, Applications, Biologic Correlates, and Challenges. RadioGraphics 2017, 37, 1483–1503.
- Larroza, A.; Bodí, V.; Moratal, D. Texture Analysis in Magnetic Resonance Imaging: Review and Considerations for Future Applications. In Assessment of Cellular and Organ Function and Dysfunction using Direct and Derived MRI Methodologies; Constantinides, C., Ed.; InTech: London, UK, 2016; ISBN 978-953-51-2722-2.
- Bailly, C.; Bodet-Milin, C.; Bourgeois, M.; Gouard, S.; Ansquer, C.; Barbaud, M.; Sébille, J.-C.; Chérel, M.; Kraeber-Bodéré, F.; Carlier, T. Exploring Tumor Heterogeneity Using PET Imaging: The Big Picture. Cancers 2019, 11, 1282.
- Hatt, M.; Tixier, F.; Visvikis, D.; Cheze Le Rest, C. Radiomics in PET/CT: More Than Meets the Eye? J. Nucl. Med. 2017, 58, 365–366.
- Tixier, F.; Hatt, M.; Valla, C.; Fleury, V.; Lamour, C.; Ezzouhri, S.; Ingrand, P.; Perdrisot, R.; Visvikis, D.; Le Rest, C.C. Visual versus Quantitative Assessment of Intratumor 18F-FDG PET Uptake Heterogeneity: Prognostic Value in Non-Small Cell Lung Cancer. J. Nucl. Med. 2014, 55, 1235–1241.
- Haralick, R.M.; Shanmugam, K.; Dinstein, I. Textural Features for Image Classification. IEEE Trans. Syst. Man Cybern. 1973, 6, 610–621.
- Amadasun, M.; King, R. Textural Features Corresponding to Textural Properties. IEEE Trans. Syst. Man Cybern. 1989, 19, 1264–1274.
- Alic, L.; Niessen, W.J.; Veenland, J.F. Quantification of Heterogeneity as a Biomarker in Tumor Imaging: A Systematic Review. PLoS ONE 2014, 9, e110300.
- Materka, A.; Strzelecki, M. Texture Analysis Methods—A Review; COST B11 Report; Technical University of Lodz, Institute of Electronics: Lodz, Poland, 1998; 33p.
- Castellano, G.; Bonilha, L.; Li, L.M.; Cendes, F. Texture Analysis of Medical Images. Clin. Radiol. 2004, 59, 1061–1069.
- Alobaidli, S.; McQuaid, S.; South, C.; Prakash, V.; Evans, P.; Nisbet, A. The Role of Texture Analysis in Imaging as an Outcome Predictor and Potential Tool in Radiotherapy Treatment Planning. Br. J. Radiol. 2014, 87, 20140369.
- Mattonen, S.A.; Ward, A.D.; Palma, D.A. Pulmonary Imaging after Stereotactic Radiotherapy—Does RECIST Still Apply? Br. J. Radiol. 2016, 89.
- Lu, W.; Wang, J.; Zhang, H.H. Computerized PET/CT Image Analysis in the Evaluation of Tumor Response to Therapy. Br. J. Radiol. 2015, 88, 20140625.
- Hatt, M.; Tixier, F.; Pierce, L.; Kinahan, P.E.; Le Rest, C.C.; Visvikis, D. Characterization of PET/CT Images Using Texture Analysis: The Past, the Present… Any Future? Eur. J. Nucl. Med. Mol. Imaging 2017, 44, 151–165.
- Silva-Rodríguez, J.; Tsoumpas, C.; Domínguez-Prado, I.; Pardo-Montero, J.; Ruibal, Á.; Aguiar, P. Impact and Correction of the Bladder Uptake on 18F-FCH PET Quantification: A Simulation Study Using the XCAT2 Phantom. Phys. Med. Biol. 2016, 61, 758.
- Boellaard, R.; Krak, N.C.; Hoekstra, O.S.; Lammertsma, A.A. Effects of Noise, Image Resolution, and ROI Definition on the Accuracy of Standard Uptake Values: A Simulation Study. J. Nucl. Med. 2004, 45, 1519–1527.
- Silva-Rodríguez, J.; Aguiar, P.; Domínguez-Prado, I.; Fierro, P.; Ruibal, Á. Simulated FDG-PET Studies for the Assessment of SUV Quantification Methods. Rev. Esp. Med. Nucl. Imagen Mol. 2015, 34, 13–18.
- Depeursinge, A.; Foncubierta-Rodriguez, A.; Van De Ville, D.; Müller, H. Three-Dimensional Solid Texture Analysis in Biomedical Imaging: Review and Opportunities. Med. Image Anal. 2014, 18, 176–196.
- Altman, D.G.; Lausen, B.; Sauerbrei, W.; Schumacher, M. Dangers of Using “Optimal” Cutpoints in the Evaluation of Prognostic Factors. J. Natl. Cancer Inst. 1994, 86, 829–835.
- Chalkidou, A.; O’Doherty, M.J.; Marsden, P.K. False Discovery Rates in PET and CT Studies with Texture Features: A Systematic Review. PLoS ONE 2015, 10, e0124165.
- Yan, J.; Chu-Shern, J.L.; Loi, H.Y.; Khor, L.K.; Sinha, A.K.; Quek, S.T.; Tham, I.W.K.; Townsend, D. Impact of Image Reconstruction Settings on Texture Features in 18F-FDG PET. J. Nucl. Med. 2015, 56, 1667–1673.
- Moon, S.H.; Kim, J.; Joung, J.-G.; Cha, H.; Park, W.-Y.; Ahn, J.S.; Ahn, M.-J.; Park, K.; Choi, J.Y.; Lee, K.-H.; et al. Correlations between Metabolic Texture Features, Genetic Heterogeneity, and Mutation Burden in Patients with Lung Cancer. Eur. J. Nucl. Med. Mol. Imaging 2019, 46, 446–454.
- Scrivener, M.; de Jong, E.E.C.; van Timmeren, J.E.; Pieters, T.; Ghaye, B.; Geets, X. Radiomics Applied to Lung Cancer: A Review. Transl. Cancer Res. 2016, 5, 398–409.
- Han, S.; Woo, S.; Suh, C.H.; Kim, Y.J.; Oh, J.S.; Lee, J.J. A Systematic Review of the Prognostic Value of Texture Analysis in 18F-FDG PET in Lung Cancer. Ann. Nucl. Med. 2018, 32, 602–610.
- Jensen, G.L.; Yost, C.M.; Mackin, D.S.; Fried, D.V.; Zhou, S.; Court, L.E.; Gomez, D.R. Prognostic Value of Combining a Quantitative Image Feature from Positron Emission Tomography with Clinical Factors in Oligometastatic Non-Small Cell Lung Cancer. Radiother. Oncol. 2018, 126, 362–367.
- Kirienko, M.; Cozzi, L.; Antunovic, L.; Lozza, L.; Fogliata, A.; Voulaz, E.; Rossi, A.; Chiti, A.; Sollini, M. Prediction of Disease-Free Survival by the PET/CT Radiomic Signature in Non-Small Cell Lung Cancer Patients Undergoing Surgery. Eur. J. Nucl. Med. Mol. Imaging 2018, 45, 207–217.
- Lemarignier, C.; Martineau, A.; Teixeira, L.; Vercellino, L.; Espie, M.; Merlet, P.; Groheux, D. Correlation between Tumor Characteristics, SUV Measurements, Metabolic Tumor Volume, TLG and Textural Features Assessed with 18F-FDG PET in a Large Cohort of Oestrogen Receptor-Positive Breast Cancer Patients. Eur. J. Nucl. Med. Mol. Imaging 2017, 44.
- Orlhac, F.; Soussan, M.; Maisonobe, J.-A.; Garcia, C.A.; Vanderlinden, B.; Buvat, I. Tumor Texture Analysis in 18F-FDG PET: Relationships between Texture Parameters, Histogram Indices, Standardized Uptake Values, Metabolic Volumes, and Total Lesion Glycolysis. J. Nucl. Med. 2014, 55, 414–422.
- Brooks, F.J.; Grigsby, P.W. The Effect of Small Tumor Volumes on Studies of Intratumoral Heterogeneity of Tracer Uptake. J. Nucl. Med. 2014, 55, 37–42.
- Hatt, M.; Majdoub, M.; Vallières, M.; Tixier, F.; Le Rest, C.C.; Groheux, D.; Hindié, E.; Martineau, A.; Pradier, O.; Hustinx, R.; et al. 18F-FDG PET Uptake Characterization through Texture Analysis: Investigating the Complementary Nature of Heterogeneity and Functional Tumor Volume in a Multi-Cancer Site Patient Cohort. J. Nucl. Med. 2015, 56, 38–44.
- Piñeiro-Fiel, M.; Moscoso, A.; Lado-Cacheiro, L.; Pombo-Pasín, M.; Rey-Bretal, D.; Gómez-Lado, N.; Mondelo-García, C.; Silva-Rodríguez, J.; Pubul, V.; Sánchez, M.; et al. Is FDG-PET Texture Analysis Related to Intratumor Biological Heterogeneity in Lung Cancer? Eur. Radiol. 2020.
- Zwanenburg, A.; Vallières, M.; Abdalah, M.A.; Aerts, H.J.W.L.; Andrearczyk, V.; Apte, A.; Ashrafinia, S.; Bakas, S.; Beukinga, R.J.; Boellaard, R.; et al. The Image Biomarker Standardization Initiative: Standardized Quantitative Radiomics for High-Throughput Image-Based Phenotyping. Radiology 2020, 295, 328–338.
- WHO | World Health Organization. Available online: (accessed on 25 November 2020).
- Papanikolaou, N.; Matos, C.; Koh, D.M. How to Develop a Meaningful Radiomic Signature for Clinical Use in Oncologic Patients. Cancer Imaging 2020, 20, 33.
- Clark, K.; Vendt, B.; Smith, K.; Freymann, J.; Kirby, J.; Koppel, P.; Moore, S.; Phillips, S.; Maffitt, D.; Pringle, M.; et al. The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository. J. Digit. Imaging 2013, 26, 1045–1057.
- Kaissis, G.A.; Makowski, M.R.; Rückert, D.; Braren, R.F. Secure, Privacy-Preserving and Federated Machine Learning in Medical Imaging. Nat. Mach. Intell. 2020, 2, 305–311.
- Sheller, M.J.; Edwards, B.; Reina, G.A.; Martin, J.; Pati, S.; Kotrotsou, A.; Milchenko, M.; Xu, W.; Marcus, D.; Colen, R.R.; et al. Federated Learning in Medicine: Facilitating Multi-Institutional Collaborations without Sharing Patient Data. Sci. Rep. 2020, 10, 12598.
- Carvalho, S.; Leijenaar, R.T.H.; Troost, E.G.C.; van Timmeren, J.E.; Oberije, C.; van Elmpt, W.; de Geus-Oei, L.-F.; Bussink, J.; Lambin, P. 18F-Fluorodeoxyglucose Positron-Emission Tomography (FDG-PET)-Radiomics of Metastatic Lymph Nodes and Primary Tumor in Non-Small Cell Lung Cancer (NSCLC)—A Prospective Externally Validated Study. PLoS ONE 2018, 13, e0192859.
- Lucia, F.; Visvikis, D.; Vallières, M.; Desseroit, M.-C.; Miranda, O.; Robin, P.; Bonaffini, P.A.; Alfieri, J.; Masson, I.; Mervoyer, A.; et al. External Validation of a Combined PET and MRI Radiomics Model for Prediction of Recurrence in Cervical Cancer Patients Treated with Chemoradiotherapy. Eur. J. Nucl. Med. Mol. Imaging 2019, 46, 864–877.
- Welch, M.L.; McIntosh, C.; Haibe-Kains, B.; Milosevic, M.F.; Wee, L.; Dekker, A.; Huang, S.H.; Purdie, T.G.; O’Sullivan, B.; Aerts, H.J.W.L.; et al. Vulnerabilities of Radiomic Signature Development: The Need for Safeguards. Radiother. Oncol. 2019, 130, 2–9.
- Van Griethuysen, J.J.M.; Fedorov, A.; Parmar, C.; Hosny, A.; Aucoin, N.; Narayan, V.; Beets-Tan, R.G.H.; Fillion-Robin, J.-C.; Pieper, S.; Aerts, H.J.W.L. Computational Radiomics System to Decode the Radiographic Phenotype. Cancer Res. 2017, 77, e104–e107.
- Martens, R.M.; Koopman, T.; Noij, D.P.; Pfaehler, E.; Übelhör, C.; Sharma, S.; Vergeer, M.R.; Leemans, C.R.; Hoekstra, O.S.; Yaqub, M.; et al. Predictive Value of Quantitative 18F-FDG-PET Radiomics Analysis in Patients with Head and Neck Squamous Cell Carcinoma. EJNMMI Res. 2020, 10, 102.
- Folkert, M.R.; Setton, J.; Apte, A.P.; Grkovski, M.; Young, R.J.; Schöder, H.; Thorstad, W.L.; Lee, N.Y.; Deasy, J.O.; Oh, J.H. Predictive Modeling of Outcomes Following Definitive Chemoradiotherapy for Oropharyngeal Cancer Based on FDG-PET Image Characteristics. Phys. Med. Biol. 2017, 62, 5327–5343.
- Bogowicz, M.; Leijenaar, R.T.H.; Tanadini-Lang, S.; Riesterer, O.; Pruschy, M.; Studer, G.; Unkelbach, J.; Guckenberger, M.; Konukoglu, E.; Lambin, P. Post-Radiochemotherapy PET Radiomics in Head and Neck Cancer—The Influence of Radiomics Implementation on the Reproducibility of Local Control Tumor Models. Radiother. Oncol. 2017, 125, 385–391.
- Van Duijn, P.W.; Marques, R.B.; Ziel-van der Made, A.C.J.; van Zoggel, H.J.A.A.; Aghai, A.; Berrevoets, C.; Debets, R.; Jenster, G.; Trapman, J.; van Weerden, W.M. Tumor Heterogeneity, Aggressiveness, and Immune Cell Composition in a Novel Syngeneic PSA-Targeted Pten Knockout Mouse Prostate Cancer (MuCaP) Model. Prostate 2018, 78, 1013–1023.
- Januškevičienė, I.; Petrikaitė, V. Heterogeneity of Breast Cancer: The Importance of Interaction between Different Tumor Cell Populations. Life Sci. 2019, 239, 117009.
- Cherezov, D.; Goldgof, D.; Hall, L.; Gillies, R.; Schabath, M.; Müller, H.; Depeursinge, A. Revealing Tumor Habitats from Texture Heterogeneity Analysis for Classification of Lung Cancer Malignancy and Aggressiveness. Sci. Rep. 2019, 9, 4500.
- Marusyk, A.; Almendro, V.; Polyak, K. Intra-Tumor Heterogeneity: A Looking Glass for Cancer? Nat. Rev. Cancer 2012, 12, 323–334.
- Junttila, M.R.; de Sauvage, F.J. Influence of Tumor Micro-Environment Heterogeneity on Therapeutic Response. Nature 2013, 501, 346–354.
- Robinson, M.H.; Vasquez, J.; Kaushal, A.; MacDonald, T.J.; Velázquez Vega, J.E.; Schniederjan, M.; Dhodapkar, K. Subtype and Grade-Dependent Spatial Heterogeneity of T-Cell Infiltration in Pediatric Glioma. J. Immunother. Cancer 2020, 8, e001066.
- Traverso, A.; Wee, L.; Dekker, A.; Gillies, R. Repeatability and Reproducibility of Radiomic Features: A Systematic Review. Int. J. Radiat. Oncol. Biol. Phys. 2018, 102, 1143–1158.
- Avanzo, M.; Wei, L.; Stancanello, J.; Vallières, M.; Rao, A.; Morin, O.; Mattonen, S.A.; El Naqa, I. Machine and Deep Learning Methods for Radiomics. Med. Phys. 2020, 47.
- Mayerhoefer, M.E.; Materka, A.; Langs, G.; Häggström, I.; Szczypiński, P.; Gibbs, P.; Cook, G. Introduction to Radiomics. J. Nucl. Med. 2020, 61, 488–495.
- Konert, T.; Everitt, S.; La Fontaine, M.D.; van de Kamer, J.B.; MacManus, M.P.; Vogel, W.V.; Callahan, J.; Sonke, J.-J. Robust, Independent and Relevant Prognostic 18F-Fluorodeoxyglucose Positron Emission Tomography Radiomics Features in Non-Small Cell Lung Cancer: Are There Any? PLoS ONE 2020, 15, e0228793.
- Hatt, M.; Tixier, F.; Cheze Le Rest, C.; Pradier, O.; Visvikis, D. Robustness of Intratumor 18F-FDG PET Uptake Heterogeneity Quantification for Therapy Response Prediction in Oesophageal Carcinoma. Eur. J. Nucl. Med. Mol. Imaging 2013, 40, 1662–1671.