| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Anna Allue Guardia | + 3484 word(s) | 3484 | 2021-02-08 04:19:44 | | | |
| 2 | Peter Tang | -1 word(s) | 3483 | 2021-02-24 03:51:54 | | | | |
| 3 | Peter Tang | Meta information modification | 3483 | 2021-03-01 10:54:03 | | | | |
| 4 | Peter Tang | Meta information modification | 3483 | 2021-03-02 02:21:57 | | |
Video Upload Options
The current emergence of multi-, extensively-, extremely-, and total-drug resistant strains of Mycobacterium tuberculosis poses a major health, social, and economic threat, and stresses the need to develop new therapeutic strategies. Mycobacteriophages are genetically diverse viruses that specifically infect mycobacterial hosts, including members of the M. tuberculosis complex. Here, we will review general features of mycobacteriophages and their mechanisms of M.tb killing, as well as their advantages and limitations as therapeutic and prophylactic agents against drug-resistant M.tb.
1. Introduction
Tuberculosis (TB) is the leading cause of mortality worldwide due to a single infectious disease, with an estimate of ~1.4 million attributed deaths in 2019 [1]. It is caused by the airborne pathogen Mycobacterium tuberculosis (M. tuberculosis), which upon inhalation is deposited in the lung alveolar space. When active TB disease is developed, the standard treatment consists of a 6-month regimen using a combination of four first-line drugs that, although very efficient, can promote the emergence of resistance in the absence of sufficient healthcare infrastructure and patient poor adherence to therapy [2]. As a direct consequence, in the past two decades, multi (MDR), extensively (XDR), extremely (XXDR), and total (TDR) drug-resistant M. tuberculosis strains have emerged worldwide as a threat to public health and TB control [3][4][5], stressing the need to develop new drugs and/or alternative anti-TB therapies [6][7].
The idea of using bacteriophages (virus that infect bacteria) to treat infectious diseases was first introduced at the beginning of the 20th century, shortly after their discovery by Felix d’Herelle [8][9]. During the 1920’s, 30’s and 40’s bacteriophages were commercially produced in several countries for the treatment or prophylaxis of several diseases such as dysentery, cholera, skin lesions or respiratory tract infections, among others [10][11][12]. However, with the introduction of penicillin and other antibiotics in the 1940’s, the use of phage therapy was abandoned in Western countries. To date, phage therapy is still considered an experimental treatment in Western countries and has not been approved for human use yet [13][14][15]. However, with the worldwide emergence of multidrug-resistant strains for most bacterial diseases, including TB, antibiotics are becoming ineffective, with only a few new drugs in the antibiotic pipeline expected to be available in the next few years [16]. As a consequence, these past few years have seen a renewed interest in the use of bacteriophages to treat multidrug-resistant bacteria, as well as chronic and persistent infections [17][18][19][20][21][22], with some experimental studies being conducted in the TB field [18].
2. Mycobacteriophages: mode of action against mycobacterial hosts
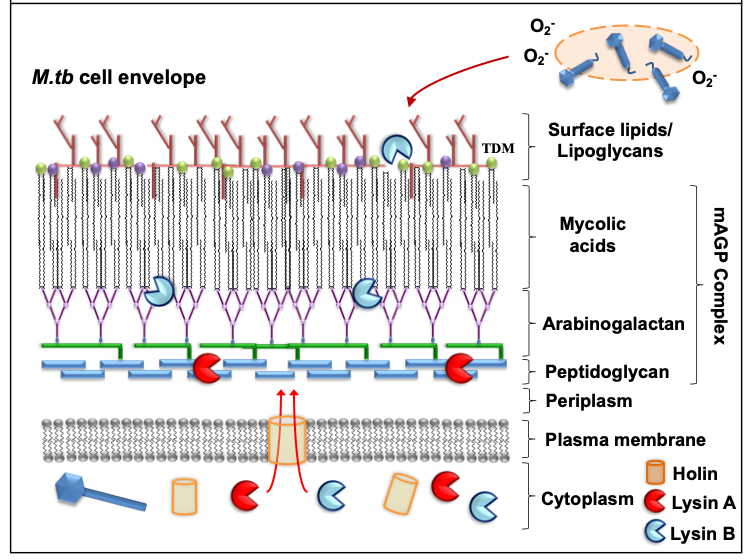
Mycobacteriophages are virus that specifically infect mycobacterial hosts, first isolated in the 1940s [23]. Mycobacteriophages are equipped with the machinery to efficiently lyse mycobacteria. That includes encoding for lysins or endolysins (LysA), enzymes with the ability to cleave the peptidoglycan (PG) layer present in the mycobacterial cell envelope [24][25][26][27] (Figure 1). Further, most mycobacteriophages rely on endolysin-holin systems to degrade the PG layer. While endolysins target the integrity of the M. tuberculosis cell envelope, holins are membrane proteins that help translocate endolysins to reach their target or trigger the activation of the enzymes at a defined time [24] (Figure 1). LysA holin-independent lysis mechanisms are also described in some mycobacteriophages [28]. However, the M. tuberculosis cell envelope is a much more complex and dynamic structure compared to gram-negative and -positive pathogens, featuring a thick mycolyl-arabinogalactan-peptidoglycan complex core (mAGP) involved in virulence and persistence [29]. To go past this natural barrier, most mycobacteriophages encode another enzyme located downstream of lysA, named Lysin B (LysB), an esterase that degrades this complex cell envelope structure by cleaving the ester bonds between the mycolic acids and the arabinogalactan [30][31]. LysB also degrades trehalose dimycolate (TDM), a glycolipid that plays an important role in mycobacterial pathogenesis [32][33] (Figure 1). Recently, the antimicrobial activity of phage D29 LysB against M. ulcerans has been demonstrated in a mouse model [34]. Besides the lysis as a primary mechanism of mycobacterial death, other authors have suggested the existence of secondary mechanisms, such as the production of superoxide radicals by lysed cells, or the induction of programmed cell death by mycobacteriophages [35] (Figure 1). Phages can also stimulate increased phagocytosis of bacteria through the opsonization of bacterial cells [36], where the phage coats the bacteria and makes it more recognizable by macrophages and other innate immune cells.
Figure 1. Mode of action against mycobacterial hosts. Most mycobacteriophages rely on endolysin-holin systems to kill their hosts. Secondary mechanisms (e.g. release of superoxide (O2-) radicals from lysed bacilli) might also contribute in the elimination of M.tb.
Phage-derived lytic enzymes such as endolysins are being studied and bioengineered as potential antimicrobial agents to combat infections caused by different gram-negative and -positive pathogens [34][37][38][39][40][41]. Thus, “enzybiotic” is defined as a new and promising class of drugs derived from phage endolysins [42]. Even though the use of phage-derived products has the potential to treat drug-resistant TB, there are some limitations when compared to the use of live mycobacteriophages, such as these phage enzymes lack the capacity of self-replication at the site of the infection, they cannot adapt to the environment, and similar to drugs, the pathogen can adapt to select resistant clones, although it is not expected to be at a high rate [43].
3. Phage Therapy to Treat Multidrug-Resistant TB
So far, only a few mycobacteriophages have been investigated as potential therapeutic options to treat TB in different in vitro studies (reviewed in detail by [44]), with only two in vivo studies using the guinea pig model [45][46]. However, to date, there is still limited data regarding the use of mycobacteriophages in the treatment of MDR-mycobacterial infections [18].
There are several general phage features that make mycobacteriophages suitable as therapeutic agents against drug-resistant TB [47][48]: a) Phages only infect and replicate inside their bacterial hosts without harming the human eukaryotic cells [49]; b) Their host range is mostly limited to Mycobacterium spp., which allows the development of targeted therapies with no apparent collateral damage to the human microbiota [50]; c) Phages are the most abundant organisms in the biosphere [51][52], with new mycobacteriophages sequenced and characterized these past few years [53][54]; d) Easy to engineer to contain suitable features for phage therapy; e) Because phages have the ability to replicate inside the pathogen using the bacterial cell machinery to generate more viral particles, less number of doses are required when compared to drug treatments [50][55]; and lastly f) Phages are easy to propagate at large-scale in an in vitro setting, significantly reducing the production costs when compared to drugs [47].
Nonetheless, there are still challenges to overcome before mycobacteriophages could be widely used to treat MDR-, XDR-, and XXDR-TB [56]. These are discussed below and summarized in Table 1.
Table 1. Phage therapy challenges in drug-resistant TB treatment
|
Challenges and limitations |
Potential solutions |
|
Host specificity |
· Global phage database screening. · Phage’s host range expansion using directed phage evolution and/or bioengineering. · Development of screening bioinformatics tools to identify targeted M.tb host virulent factor epitopes (e.g. efflux pump). |
|
Unknown impact of human ALF on the M.tb cell envelope |
· Identify how the M.tb cell envelope adapts (changes) to the different environments that encounters at different stages of infection [e.g. contact with ALF, within the phagosome, extracellular, within granulomas or cavities, or when being transmitted (in sputum)]. |
|
Phages access to intracellular M.tb |
· Novel phage delivery systems [e.g. M. smegmatis (Trojan horse concept), phage microencapsulation]. · Phage bioengineering to recognize well-defined macrophage receptors (the mannose receptor or MR). |
|
M.tb resistance to phages |
· Use of different phage cocktails. · Phage-drug combined treatment (phage-drug synergy) in combination with the mammalian host immune response. · Phage sequential treatment. · Phage personalized treatment. |
|
Overactivation of the mammalian host immune system and risk of anaphylaxis |
· Optimize phage delivery routes. · Establish phage dosage and frequency. · Maximize synergy between phages and the mammalian host immune system. |
|
Lack of phage therapy regulations |
· Standardize global regulations for phage production (under GMP conditions). |
|
Phage cytotoxicity to the human host |
· Use of highly lytic phages that do not integrate into the M.tb genome. · Targeted phage genetic bioengineering to remove potential phage virulent factors to the mammalian host. · Define function of unknown phage genes. |
3.1. Mycobacteriophage host specificity
Mycobacteriophages host range is restricted to Mycobacterium and some other members of the Actinobacteria phylum [57][58]. Most mycobacteriophages have been isolated using the non-pathogenic M. smegmatis as a host, and only a few of them are shown to also efficiently infect M. tuberculosis [57][58][59][60][61]. Mycobacteriophage specific host range could be considered as a double-edge sword in regards to potential applications in phage therapy: on one hand, phage specificity is required for successful clearance of a particular pathogen without affecting the overall human microbiome composition. On the other hand, it also represents a significant limitation, as this high specificity might create a bottleneck in terms of identifying potential phage candidates to efficiently kill M. tuberculosis. In this regard, mycobacteriophages have the ability to switch or expand their host range frequently among different mycobacterial strains [62][63].
The recognition of M. tuberculosis cell envelope surface receptors by phage tail proteins and subsequent binding will likely determine phage-bacterial host specificity and infection efficiency, although not much is known about phage binding receptors on the M. tuberculosis cell envelope surface [64], where only a few mycobacteriophage targets have been described for non-tuberculous mycobacteria [65][66]. Recently, BSL-2 safe strains of MDR-M. tuberculosis that are triple auxotrophic (pantothenate, leucine and either arginine or methionine) have been developed for performing experiments in laboratories that do not have BSL-3 facilities [67]. These strains seem to be the most promising candidates to test mycobacteriophages for activity against MDR strains. Further, several studies have shown that phage specificity can be altered through genetic engineering of their RBPs [68][69], which could open the avenue for the custom design of M. tuberculosis-specific mycobacteriophages for the treatment of drug-resistant TB. It would also allow to modify well-characterized lytic phages and expand its host range to efficiently infect M. tuberculosis.
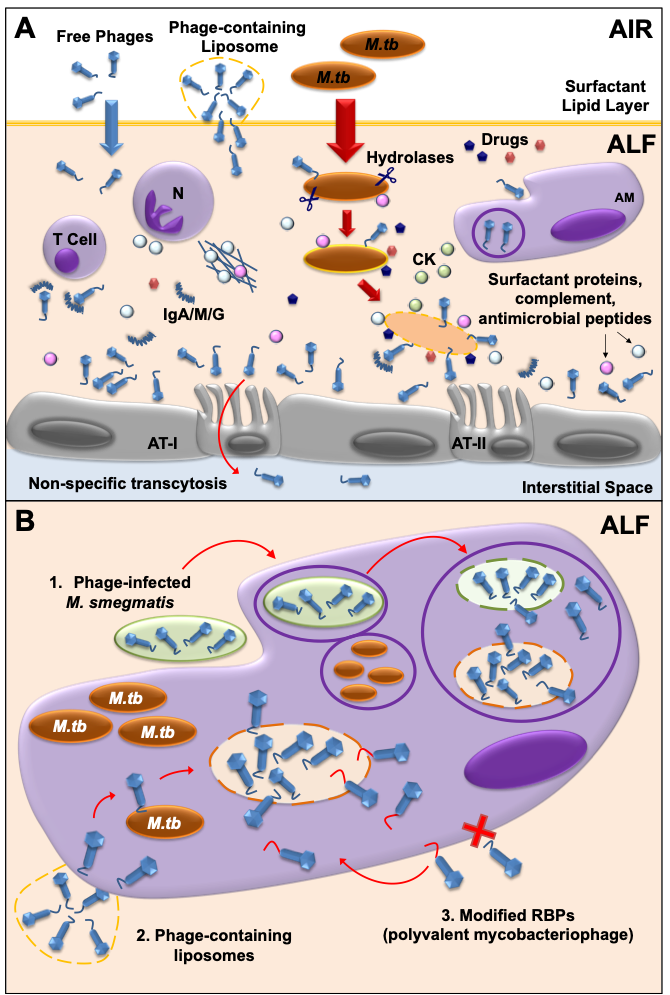
Another key factor to consider in mycobacteriophage-M. tuberculosis interactions is the impact of the human lung environment in shaping the M. tuberculosis cell envelope. Upon infection, M. tuberculosis is deposited in the lung alveolar space where it comes in close contact with the lung mucosa for an undetermined period of time, including an aqueous hypophase called alveolar lining fluid (or ALF). ALF contains a series of innate soluble molecules, including hydrolases, shown to significantly modify the M.tb cell envelope upon contact (Figure 2A) [70][71][72][73][74][75]. Thus, the in vitrorepertoire of potential exposed receptors on the M. tuberculosis cell envelope surface might be modified during the initial stages of infection depending on the host ALF status, potentially directing phage-M. tuberculosis interactions in vivo. As most in vitro studies use M. tuberculosis grown in solid or liquid medium, where bacteria have not been exposed to ALF, there could be well-characterized mycobacteriophages efficient in recognizing specific M. tuberculosis strains that have been discarded because their testing was performed using in vitro models that do not reflect the impact of the lung environment. Definitely, the lung environment is a critical factor to take into account that requires further study when designing phage therapies to treat drug-resistant TB.
Figure 2. Mycobacteriophage-M. tuberculosis interactions in the human lung alveolar environment. (A) Free and/or encapsulated mycobacteriophages (e.g.in liposomes) reach the lung alveolar space and are bathed in ALF, where they generate a local immune response. Some phages will be cleared by phagocytic cells [e.g. alveolar macrophages (AMs)], or by neutralizing antibodies (IgA/M/G). If delivered in sufficient numbers, mycobacteriophages are capable of binding glycan residues in the mucus layer secreted by alveolar epithelial cells type II (AT-II) through Ig-like domains in their capsids, providing an extra non-host antimicrobial protection layer against M.tb infection. Mycobacteriophages can also be translocated to the interstitial space through the epithelium via a non-specific transcytosis mechanism. In the alveolar space, innate soluble molecules such as hydrolases (represented as scissors), modify theM. tuberculosis cell envelope, and will define de novo exposed M. tuberculosis surface receptors to be recognized by mycobacteriophages. In a combined mycobacteriophage-drug therapy, drugs and mycobacteriophages work together with an activated mammalian immune response [including neutrophil (N) degranulation and neutrophil extracellular traps secretion, AMs activation, and cytokine and chemokine secretion] to kill M. tuberculosis and control the infection. (B) Different strategies can be used to deliver mycobacteriophages inside mammalian cells (e.g. AMs) to access intracellular M. tb : 1) Mycobacteriophage-infected M. smegmatis bacilli acting as vehicles are phagocytized by AMs and delivered into M.tb -containing phagosomes, where they access the lumen and infect M.tb; 2) Liposome-associated mycobacteriophages; 3) Generation of polyvalent mycobacteriophages through receptor binding protein (RBP) engineering to recognize mammalian immune cells.
3.2. Mycobacteriophage-M. tuberculosis interactions within mammalian host cells
Since mycobacteriophages do not seem to have a natural ability to cross mammalian cell membranes, the fact that M. tuberculosis can be found inside macrophages and/or granulomas once the infection is established, represents an added challenge for the development of TB phage therapy strategies [44]. These past few years, several strategies have been developed to overcome this issue. As an example, mycobacteriophage delivery inside macrophages has been achieved using M. smegmatis, which can serve as host bacterial reservoir for mycobacteriophage proliferation to increase phage titers before reaching the targeted M. tuberculosis pathogen [76][77][78] (Figure 2B). The use of liposomes as a non-bacterial vector proved that liposome-associated TM4 phages infect mammalian cells more efficiently than free phages [79] (Figure 2B). An alternative strategy could be to engineer mycobacteriophages to recognize and bind host cells, so these can be internalized and have access to intracellular M. tuberculosis (Figure 2B). Mycobacteriophages which are engineered in either capsid or tail proteins to carry ligands such as mannose-6-phosphate could be targeted towards intracellular compartments of macrophages.
3.3. Bacterial resistance to mycobacteriophages
One of the major factors to consider in mycobacteriophage therapy is the potential development of bacterial resistance against phages over time [43][80][81][82], similar to what happens with drugs. But because phages are dynamic and constant evolving entities, they are also capable of developing counter-defense mechanisms to avoid bacterial blockade [83][84][85][86][87]. In the case of M. tuberculosis, it should be considered that this pathogen is characterized by a low mutation rate of ~ 2 x 10-10 mutations/bp/generation [88] that, in combination with a long replication time of approximately 20 h, highly decreases the probability to develop resistance to mycobacteriophages, which favors the use of phage therapy for the treatment of drug-resistant TB when compared to other pathogens. Still, several strategies can be applied to shift the balance towards a sustainable phage infection reducing the risk of resistance. For example, the use of phage cocktails with a mixture of genetically diverse phages against several strains of M. tuberculosis [80] can minimize the occurrence of cross-resistance, where the development of resistance to all phages in the mix would come with a high fitness cost for the bacterium [89][90][91][92]. Indeed, a three-phage cocktail was used in the treatment of disseminated drug-resistant M. abscessus subsp. massiliense, a non-tuberculous mycobacterium, in a 15-year-old patient with cystic fibrosis [18], reducing infected skin lesions and improved liver function. Other strategies to reduce the appearance of phage resistance are the use of sequential treatments, in which individual phages with different characteristics are administered one after the other [93], or a combination therapy of phage and drugs [94][95][96][97] (Figure 2A), which has been shown to be effective thanks to a phenomenon called Phage-Antibiotic Synergy (PAS). Finally, for specific settings and cases, the use of personalized treatments could be used to avoid the overuse of generalized phage or drug therapies [98]. Overall, more in vitro and in vivo studies exploring the mechanisms of bacterial resistance to phages and mycobacteriophage counter-defenses are needed in order to develop successful phage therapy strategies to treat drug-resistant TB, as very little is known about those mechanisms in M. tuberculosis [64].
3.4. Mycobacteriophage interactions with the mammalian immune system and the lung virome
Recent metagenomic studies have described the ubiquitous presence of viruses as part of the human microbiome [99][100][101]. Phages (“phageome”) are typically the most abundant type, and have special importance in modulating bacterial communities [102]. Further, the impact of the phageome in the host immune system needs careful consideration. Indeed, phages can interact directly or indirectly with mammalian cells, altering host immune responses [36][103]. As an example of a direct interaction, phages are capable of binding glycan residues present in the mucus layer secreted by the epithelium through Ig-like domains in their capsids [104] (Figure 2A). This could be of particular interest for the use of mycobacteriophages as prophylactic agents, where phages could be administered in sufficient amounts to bind to the lung mucosal surfaces, providing an extra layer of local innate immune protection against M. tuberculosis infections [60], with major implications in TB transmission, especially in TB endemic areas.
More recently, it was demonstrated that phages can be translocated through epithelial cells by a non-specific transcytosis mechanism, including intestinal and lung epithelium [105] (Figure 2A), which might explain a systemic presence of phages after oral administration. Systemic mycobacteriophage delivery may be feasible to treat TB, although phage self-replication might not be enough if bacterial loads are low at the infection site [106]. Instead, phages delivered through intranasal or endotracheal routes could probably reach the lungs at higher numbers with limiting systemic effects, being more adequate to treat lung infections such as TB at their active stage [60][61][107][108]. The reduction of phage numbers due to the activation of mammalian host innate and adaptive immune responses is also important when determining the optimal dose/s for a particular phage treatment. Phages are cleared from the human body by phagocytes [108][109] (Figure 2A), although the production of neutralizing antibodies against phages has also been described (Figure 2A). Repeated doses of high phage concentrations might be required for successful treatment; however, multiple phage doses and/or high phage titers may over activate the immune system, leading to phage clearance from the tissue [43][107].
In this process named “Immunophage Synergy”, the action of the immune system is necessary and complements the phage antimicrobial activity [110] (Figure 2A). Only a few studies detailing the use of mycobacteriophages to eliminate M. tuberculosis complex are available, with limited data regarding the immunological effects of phage therapeutics [44][111]. Although no documented cases of anaphylaxis have been reported in humans, as opposed to drugs [112][113], it is critical to define the exact immunological responses in order to predict the success of new potential phage therapies for drug resistant-TB.
4. Concluding remarks
In a “Dry Pipeline” era of drugs, where only a few new compounds are being discovered, the need for alternative strategies to treat emergent MDR, XDR, XXDR and/or TDR M.tb strains is a priority. The notion of phage therapy against bacterial infections has been around for almost a century due to its numerous advantages. The vast diversity observed in almost 2,000 characterized mycobacteriophages opens the possibility of finding highly specific and efficient phages for the elimination of drug-resistant M. tuberculosis strains, otherwise associated with untreatable infections. An important concept that has been overlooked in most in vitro mycobacteriophage-bacterial host specificity studies is the impact of the lung environment in shaping the M. tuberculosis cell envelope. This could be further exploited to uncover novel mycobacteriophage receptors and to revisit previously discovered mycobacteriophages that could be highly specific when taking into account the impact of the human lung environment.
Despite the obvious advantages of using phages as therapeutic agents and promising results from recent studies [114], there are still several challenges that need to be addressed in order for phage therapy to be widely implemented in modern medicine [115]. For example, the exact immunomodulatory properties of phages and interactions within the human host are still largely unknown. A mycobacteriophage used in phage therapy should reach the lungs in sufficient numbers and be able to work synergistically with the host innate immune system (and drugs, in a combined phage-drug therapy) [95] to clear the infection, but without causing a dysregulated systemic immune reaction or adverse effects. A concept to explore would be the use of phages to generate a low-level protective immune response to prevent M. tuberculosis infections. TB comorbidities such as aging, HIV co-infection, and chronic disease such as diabetes alter the host’s immune response [73] and consequently, a particular phage treatment that would work for a healthy individual might not work in these cases, stressing the need for more personalized treatments.
Currently, due to mycobacteriophage therapy clinical practicality, this may be only used on the most difficult or virtually incurable cases of drug-resistant TB, where drug therapy does not work or is insufficient. Phage therapy alone may never completely replace the use of traditional drugs, but combined mycobacteriophage-drug therapies working together and enhancing the stimulation of the host immune response might prove to be the most efficient in order to shorten TB treatment and/or reduction in drug dosage. The use of combined therapies in MDR patients worldwide could reduce the conversion and cases of XDR and XXDR-TB. Using mycobacteriophages as prophylactic instead of therapeutic agents might be a better strategy in TB endemic areas. In this regard, regular mycobacteriophage administration in high-risk individuals could be exploited as a prophylactic measure to prevent onset of TB disease instead of preventive TB therapy, although the impact of the constant presence of mycobacteriophages in the lung microbiome and the host immunity generated would need to be well-established prior to their therapeutic use.
References
- WHO, Global Tuberculosis Report 2020. 2020.
- Zaman, K., Tuberculosis: a global health problem. J Health Popul Nutr 2010, 28, (2), 111-3.
- Ellner, J. J., The emergence of extensively drug-resistant tuberculosis: a global health crisis requiring new interventions: part I: the origins and nature of the problem. Clin Transl Sci 2008, 1, (3), 249-54.
- Loewenberg, S., India reports cases of totally drug-resistant tuberculosis. Lancet 2012, 379, (9812), 205.
- Velayati, A. A.; Masjedi, M. R.; Farnia, P.; Tabarsi, P.; Ghanavi, J.; ZiaZarifi, A. H.; Hoffner, S. E., Emergence of New Forms of Totally Drug-Resistant Tuberculosis Bacilli Super Extensively Drug-Resistant Tuberculosis or Totally Drug-Resistant Strains in Iran. Chest 2009, 136, (2), 420-425.
- Burki, T. K., The uphill battle to find new TB treatments. Lancet Respir Med 2017.
- Lange, C.; Chesov, D.; Heyckendorf, J.; Leung, C. C.; Udwadia, Z.; Dheda, K., Drug-resistant tuberculosis: An update on disease burden, diagnosis and treatment. Respirology 2018, 23, (7), 656-673.
- Wilkinson, L., Felix d'Herelle and the origins of molecular biology. Med Hist 2001, 45, (2), 294-295.
- Chanishvili, N., Phage Therapy-History from Twort and d'Herelle Through Soviet Experience to Current Approaches. Adv Virus Res 2012, 83, 3-40.
- Sulakvelidze, A.; Alavidze, Z.; Morris, J. G., Jr., Bacteriophage therapy. Antimicrob Agents Chemother 2001, 45, (3), 649-59.
- Dublanchet, A.; Bourne, S., The epic of phage therapy. Can J Infect Dis Med Microbiol 2007, 18, (1), 15-8.
- Kokin, G. A., Phage therapy and phage prophylaxis of gas gangrene. In Experience of the Soviet Military Medicine during the Great Patriotic War 1941–1945, Medgiz, Moscow, 1946; Vol. 3.
- Gorski, A.; Miedzybrodzki, R.; Borysowski, J.; Weber-Dabrowska, B.; Lobocka, M.; Fortuna, W.; Letkiewicz, S.; Zimecki, M.; Filby, G., Bacteriophage therapy for the treatment of infections. Curr Opin Invest Dr 2009, 10, (8), 766-774.
- Furfaro, L. L.; Payne, M. S.; Chang, B. J., Bacteriophage Therapy: Clinical Trials and Regulatory Hurdles. Front Cell Infect Microbiol 2018, 8, 376.
- Wienhold, S. M.; Lienau, J.; Witzenrath, M., Towards Inhaled Phage Therapy in Western Europe. Viruses 2019, 11, (3).
- Lohrasbi, V.; Talebi, M.; Bialvaei, A. Z.; Fattorini, L.; Drancourt, M.; Heidary, M.; Darban-Sarokhalil, D., Trends in the discovery of new drugs for Mycobacterium tuberculosis therapy with a glance at resistance. Tuberculosis (Edinb) 2018, 109, 17-27.
- LaVergne, S.; Hamilton, T.; Biswas, B.; Kumaraswamy, M.; Schooley, R. T.; Wooten, D., Phage Therapy for a Multidrug-Resistant Acinetobacter baumannii Craniectomy Site Infection. Open Forum Infect Dis 2018, 5, (4), ofy064.
- Dedrick, R. M.; Guerrero-Bustamante, C. A.; Garlena, R. A.; Russell, D. A.; Ford, K.; Harris, K.; Gilmour, K. C.; Soothill, J.; Jacobs-Sera, D.; Schooley, R. T.; Hatfull, G. F.; Spencer, H., Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat Med 2019, 25, (5), 730-733.
- Jeon, J.; Park, J. H.; Yong, D., Efficacy of bacteriophage treatment against carbapenem-resistant Acinetobacter baumannii in Galleria mellonella larvae and a mouse model of acute pneumonia. BMC Microbiol 2019, 19, (1), 70.
- Tkhilaishvili, T.; Winkler, T.; Muller, M.; Perka, C.; Trampuz, A., Bacteriophages as adjuvant to antibiotics for the treatment of periprosthetic joint infection caused by multidrug-resistant Pseudomonas aeruginosa. Antimicrob Agents Chemother 2019.
- Lin, D. M.; Koskella, B.; Lin, H. C., Phage therapy: An alternative to antibiotics in the age of multi-drug resistance. World J Gastrointest Pharmacol Ther 2017, 8, (3), 162-173.
- Kortright, K. E.; Chan, B. K.; Koff, J. L.; Turner, P. E., Phage Therapy: A Renewed Approach to Combat Antibiotic-Resistant Bacteria. Cell Host Microbe 2019, 25, (2), 219-232.
- Gardner, G. M.; Weiser, R. S., A bacteriophage for Mycobacterium smegmatis. Proc Soc Exp Biol Med 1947, 66, (1), 205.
- Pimentel, M., Genetics of Phage Lysis. Microbiol Spectr 2014, 2, (1), MGM2-0017-2013.
- Joshi, H.; Nair, G.; Gangakhedkar, R.; Jain, V., Understanding the role of the lysozyme-like domain of D29 mycobacteriophage-encoded endolysin in host cell lysis and phage propagation. Microbiology 2019, 165, (9), 1013-1023.
- Catalao, M. J.; Pimentel, M., Mycobacteriophage Lysis Enzymes: Targeting the Mycobacterial Cell Envelope. Viruses 2018, 10, (8).
- Joshi, H.; Seniya, S. P.; Suryanarayanan, V.; Patidar, N. D.; Singh, S. K.; Jain, V., Dissecting the structure-function relationship in lysozyme domain of mycobacteriophage D29-encoded peptidoglycan hydrolase. FEBS Lett 2017, 591, (20), 3276-3287.
- Catalao, M. J.; Gil, F.; Moniz-Pereira, J.; Pimentel, M., The mycobacteriophage Ms6 encodes a chaperone-like protein involved in the endolysin delivery to the peptidoglycan. Mol Microbiol 2010, 77, (3), 672-86.
- Dulberger, C. L.; Rubin, E. J.; Boutte, C. C., The mycobacterial cell envelope - a moving target. Nat Rev Microbiol 2020, 18, (1), 47-59.
- Gigante, A. M.; Hampton, C. M.; Dillard, R. S.; Gil, F.; Catalao, M. J.; Moniz-Pereira, J.; Wright, E. R.; Pimentel, M., The Ms6 Mycolyl-Arabinogalactan Esterase LysB is Essential for an Efficient Mycobacteriophage-Induced Lysis. Viruses 2017, 9, (11).
- Payne, K.; Sun, Q.; Sacchettini, J.; Hatfull, G. F., Mycobacteriophage Lysin B is a novel mycolylarabinogalactan esterase. Mol Microbiol 2009, 73, (3), 367-81.
- Gil, F.; Grzegorzewicz, A. E.; Catalao, M. J.; Vital, J.; McNeil, M. R.; Pimentel, M., Mycobacteriophage Ms6 LysB specifically targets the outer membrane of Mycobacterium smegmatis. Microbiology 2010, 156, (Pt 5), 1497-504.
- Garcia-Vilanova, A.; Chan, J.; Torrelles, J. B., Underestimated Manipulative Roles of Mycobacterium tuberculosis Cell Envelope Glycolipids During Infection. Front Immunol 2019, 10, 2909.
- Fraga, A. G.; Trigo, G.; Murthy, R. K.; Akhtar, S.; Hebbur, M.; Pacheco, A. R.; Dominguez, J.; Silva-Gomes, R.; Goncalves, C. M.; Oliveira, H.; Castro, A. G.; Sharma, U.; Azeredo, J.; Pedrosa, J., Antimicrobial activity of Mycobacteriophage D29 Lysin B during Mycobacterium ulcerans infection. PLoS Negl Trop Dis 2019, 13, (8), e0007113.
- Samaddar, S.; Grewal, R. K.; Sinha, S.; Ghosh, S.; Roy, S.; Das Gupta, S. K., Dynamics of Mycobacteriophage-Mycobacterial Host Interaction: Evidence for Secondary Mechanisms for Host Lethality. Appl Environ Microbiol 2016, 82, (1), 124-33.
- Van Belleghem, J. D.; Dabrowska, K.; Vaneechoutte, M.; Barr, J. J.; Bollyky, P. L., Interactions between Bacteriophage, Bacteria, and the Mammalian Immune System. Viruses 2018, 11, (1).
- Lood, R.; Winer, B. Y.; Pelzek, A. J.; Diez-Martinez, R.; Thandar, M.; Euler, C. W.; Schuch, R.; Fischetti, V. A., Novel phage lysin capable of killing the multidrug-resistant gram-negative bacterium Acinetobacter baumannii in a mouse bacteremia model. Antimicrob Agents Chemother 2015, 59, (4), 1983-91.
- Stone, E.; Campbell, K.; Grant, I.; McAuliffe, O., Understanding and Exploiting Phage-Host Interactions. Viruses 2019, 11, (6).
- Diez-Martinez, R.; De Paz, H. D.; Garcia-Fernandez, E.; Bustamante, N.; Euler, C. W.; Fischetti, V. A.; Menendez, M.; Garcia, P., A novel chimeric phage lysin with high in vitro and in vivo bactericidal activity against Streptococcus pneumoniae. J Antimicrob Chemother 2015, 70, (6), 1763-73.
- Yan, G.; Yang, R.; Fan, K.; Dong, H.; Gao, C.; Wang, S.; Yu, L.; Cheng, Z.; Lei, L., External lysis of Escherichia coli by a bacteriophage endolysin modified with hydrophobic amino acids. AMB Express 2019, 9, (1), 106.
- Zampara, A.; Sorensen, M. C. H.; Grimon, D.; Antenucci, F.; Vitt, A. R.; Bortolaia, V.; Briers, Y.; Brondsted, L., Exploiting phage receptor binding proteins to enable endolysins to kill Gram-negative bacteria. Sci Rep 2020, 10, (1), 12087.
- Dams, D.; Briers, Y., Enzybiotics: Enzyme-Based Antibacterials as Therapeutics. Adv Exp Med Biol 2019, 1148, 233-253.
- Casey, E.; van Sinderen, D.; Mahony, J., In Vitro Characteristics of Phages to Guide 'Real Life' Phage Therapy Suitability. Viruses-Basel 2018, 10, (4).
- Azimi, T.; Mosadegh, M.; Nasiri, M. J.; Sabour, S.; Karimaei, S.; Nasser, A., Phage therapy as a renewed therapeutic approach to mycobacterial infections: a comprehensive review. Infect Drug Resist 2019, 12, 2943-2959.
- Sula, L.; Sulova, J.; Stolcpartova, M., Therapy of experimental tuberculosis in guinea pigs with mycobacterial phages DS-6A, GR-21 T, My-327. Czech Med 1981, 4, (4), 209-14.
- Peng Li, L. Y., Chen BaoWen, Li YouLun, Shen XiaoBing, Su Cheng, and Wang GuoZhi, Therapeutic effect of bacteriophage D29 in the treatment for guinea pigs infected with sensitive strain of Mycobacterium tuberculosis. Chinese Journal of Zoonoses 2009, 25(8), 733-736.
- Principi, N.; Silvestri, E.; Esposito, S., Advantages and Limitations of Bacteriophages for the Treatment of Bacterial Infections. Front Pharmacol 2019, 10, 513.
- Gordillo Altamirano, F. L.; Barr, J. J., Phage Therapy in the Postantibiotic Era. Clin Microbiol Rev 2019, 32, (2).
- Ooi, M. L.; Drilling, A. J.; Morales, S.; Fong, S.; Moraitis, S.; Macias-Valle, L.; Vreugde, S.; Psaltis, A. J.; Wormald, P. J., Safety and Tolerability of Bacteriophage Therapy for Chronic Rhinosinusitis Due to Staphylococcus aureus. Jama Otolaryngol 2019, 145, (8), 723-729.
- Bogovazova, G. G.; Voroshilova, N. N.; Bondarenko, V. M., [The efficacy of Klebsiella pneumoniae bacteriophage in the therapy of experimental Klebsiella infection]. Zh Mikrobiol Epidemiol Immunobiol 1991, (4), 5-8.
- Aziz, R. K.; Dwivedi, B.; Akhter, S.; Breitbart, M.; Edwards, R. A., Multidimensional metrics for estimating phage abundance, distribution, gene density, and sequence coverage in metagenomes. Front Microbiol 2015, 6, 381.
- Hatfull, G. F., Bacteriophage genomics. Curr Opin Microbiol 2008, 11, (5), 447-53.
- Bajpai, U.; Mehta, A. K.; Eniyan, K.; Sinha, A.; Ray, A.; Virdi, S.; Ahmad, S.; Shah, A.; Arora, D.; Marwaha, D.; Chauhan, G.; Saraswat, P.; Bathla, P.; Singh, R., Isolation and characterization of bacteriophages from India, with lytic activity against Mycobacterium tuberculosis. Can J Microbiol 2018, 64, (7), 483-491.
- Caratenuto, R. A., 3rd; Ciabattoni, G. O.; DesGranges, N. J.; Drost, C. L.; Gao, L.; Gipson, B.; Kahler, N. C.; Kirven, N. A.; Melehani, J. C.; Patel, K.; Rokes, A. B.; Seth, R. A.; West, M. C.; Alhout, A. A.; Akoto, F. F.; Capogna, N.; Cudkevich, N.; Graham, L. H.; Grapel, M. S.; Haleem, M. M.; Korenberg, J. B.; Lichak, B. P.; McKinley, L. N.; Mendello, K. R.; Murphy, C. E.; Pyfer, L. M.; Ramirez, W. A.; Reisner, J. R.; Swope, R. H.; Thoonkuzhy, M. J.; Vargas, L. A.; Veliz, C. A.; Volpe, K. R.; Zhang, K. D.; Faltine-Gonzalez, D. Z.; Zuilkoski, C. M.; Mageeney, C. M.; Mohammed, H. T.; Kenna, M. A.; Ware, V. C., Genome Sequences of Six Cluster N Mycobacteriophages, Kevin1, Nenae, Parmesanjohn, ShrimpFriedEgg, Smurph, and SpongeBob, Isolated on Mycobacterium smegmatis mc(2)155. Microbiol Resour Announc 2019, 8, (22).
- Levin, B. R.; Bull, J. J., Phage therapy revisited: The population biology of a bacterial infection and its treatment with bacteriophage and antibiotics. Am Nat 1996, 147, (6), 881-898.
- Young, R.; Gill, J. J., MICROBIOLOGY. Phage therapy redux--What is to be done? Science 2015, 350, (6265), 1163-4.
- Dedrick, R. M.; Guerrero Bustamante, C. A.; Garlena, R. A.; Pinches, R. S.; Cornely, K.; Hatfull, G. F., Mycobacteriophage ZoeJ: A broad host-range close relative of mycobacteriophage TM4. Tuberculosis (Edinb) 2019, 115, 14-23.
- Mayer, O.; Jain, P.; Weisbrod, T. R.; Biro, D.; Ho, L.; Jacobs-Sera, D.; Hatfull, G. F.; Jacobs, W. R., Jr., Fluorescent Reporter DS6A Mycobacteriophages Reveal Unique Variations in Infectibility and Phage Production in Mycobacteria. J Bacteriol 2016, 198, (23), 3220-3232.
- Lapenkova, M. B.; Smirnova, N. S.; Rutkevich, P. N.; Vladimirsky, M. A., Evaluation of the Efficiency of Lytic Mycobacteriophage D29 on the Model of M. tuberculosis-Infected Macrophage RAW 264 Cell Line. Bull Exp Biol Med 2018, 164, (3), 344-346.
- Carrigy, N. B.; Larsen, S. E.; Reese, V.; Pecor, T.; Harrison, M.; Kuehl, P. J.; Hatfull, G. F.; Sauvageau, D.; Baldwin, S. L.; Finlay, W. H.; Coler, R. N.; Vehring, R., Prophylaxis of Mycobacterium tuberculosis H37Rv Infection in a Preclinical Mouse Model via Inhalation of Nebulized Bacteriophage D29. Antimicrob Agents Chemother 2019.
- Liu, K. Y.; Yang, W. H.; Dong, X. K.; Cong, L. M.; Li, N.; Li, Y.; Wen, Z. B.; Yin, Z.; Lan, Z. J.; Li, W. P.; Li, J. S., Inhalation Study of Mycobacteriophage D29 Aerosol for Mice by Endotracheal Route and Nose-Only Exposure. J Aerosol Med Pulm Drug Deliv 2016, 29, (5), 393-405.
- Hatfull, G. F., Mycobacteriophages. Microbiol Spectr 2018, 6, (5).
- Jacobs-Sera, D.; Marinelli, L. J.; Bowman, C.; Broussard, G. W.; Guerrero Bustamante, C.; Boyle, M. M.; Petrova, Z. O.; Dedrick, R. M.; Pope, W. H.; Science Education Alliance Phage Hunters Advancing, G.; Evolutionary Science Sea-Phages, P.; Modlin, R. L.; Hendrix, R. W.; Hatfull, G. F., On the nature of mycobacteriophage diversity and host preference. Virology 2012, 434, (2), 187-201.
- Chen, J.; Kriakov, J.; Singh, A.; Jacobs, W. R., Jr.; Besra, G. S.; Bhatt, A., Defects in glycopeptidolipid biosynthesis confer phage I3 resistance in Mycobacterium smegmatis. Microbiology 2009, 155, (Pt 12), 4050-7.
- Furuchi, A.; Tokunaga, T., Nature of the receptor substance of Mycobacterium smegmatis for D4 bacteriophage adsorption. J Bacteriol 1972, 111, (2), 404-11.
- Khoo, K. H.; Suzuki, R.; Dell, A.; Morris, H. R.; McNeil, M. R.; Brennan, P. J.; Besra, G. S., Chemistry of the lyxose-containing mycobacteriophage receptors of Mycobacterium phlei/Mycobacterium smegmatis. Biochemistry 1996, 35, (36), 11812-9.
- Vilcheze, C.; Copeland, J.; Keiser, T. L.; Weisbrod, T.; Washington, J.; Jain, P.; Malek, A.; Weinrick, B.; Jacobs, W. R., Jr., Rational Design of Biosafety Level 2-Approved, Multidrug-Resistant Strains of Mycobacterium tuberculosis through Nutrient Auxotrophy. mBio 2018, 9, (3).
- de Jonge, P. A.; Nobrega, F. L.; Brouns, S. J. J.; Dutilh, B. E., Molecular and Evolutionary Determinants of Bacteriophage Host Range. Trends Microbiol 2019, 27, (1), 51-63.
- Dunne, M.; Rupf, B.; Tala, M.; Qabrati, X.; Ernst, P.; Shen, Y.; Sumrall, E.; Heeb, L.; Pluckthun, A.; Loessner, M. J.; Kilcher, S., Reprogramming Bacteriophage Host Range through Structure-Guided Design of Chimeric Receptor Binding Proteins. Cell Rep 2019, 29, (5), 1336-1350 e4.
- Arcos, J.; Sasindran, S. J.; Fujiwara, N.; Turner, J.; Schlesinger, L. S.; Torrelles, J. B., Human lung hydrolases delineate Mycobacterium tuberculosis-macrophage interactions and the capacity to control infection. J Immunol 2011, 187, (1), 372-81.
- Scordo, J. M.; Olmo-Fontanez, A. M.; Kelley, H. V.; Sidiki, S.; Arcos, J.; Akhter, A.; Wewers, M. D.; Torrelles, J. B., The human lung mucosa drives differential Mycobacterium tuberculosis infection outcome in the alveolar epithelium. Mucosal Immunol 2019, 12, (3), 795-804.
- Arcos, J.; Sasindran, S. J.; Moliva, J. I.; Scordo, J. M.; Sidiki, S.; Guo, H.; Venigalla, P.; Kelley, H. V.; Lin, G.; Diangelo, L.; Silwani, S. N.; Zhang, J.; Turner, J.; Torrelles, J. B., Mycobacterium tuberculosis cell wall released fragments by the action of the human lung mucosa modulate macrophages to control infection in an IL-10-dependent manner. Mucosal Immunol 2017, 10, (5), 1248-1258.
- Moliva, J. I.; Duncan, M. A.; Olmo-Fontanez, A.; Akhter, A.; Arnett, E.; Scordo, J. M.; Ault, R.; Sasindran, S. J.; Azad, A. K.; Montoya, M. J.; Reinhold-Larsson, N.; Rajaram, M. V. S.; Merrit, R. E.; Lafuse, W. P.; Zhang, L.; Wang, S. H.; Beamer, G.; Wang, Y.; Proud, K.; Maselli, D. J.; Peters, J.; Weintraub, S. T.; Turner, J.; Schlesinger, L. S.; Torrelles, J. B., The Lung Mucosa Environment in the Elderly Increases Host Susceptibility to Mycobacterium tuberculosis Infection. J Infect Dis 2019, 220, (3), 514-523.
- Moliva, J. I.; Hossfeld, A. P.; Canan, C. H.; Dwivedi, V.; Wewers, M. D.; Beamer, G.; Turner, J.; Torrelles, J. B., Exposure to human alveolar lining fluid enhances Mycobacterium bovis BCG vaccine efficacy against Mycobacterium tuberculosis infection in a CD8(+) T-cell-dependent manner. Mucosal Immunol 2018, 11, (3), 968-978.
- Torrelles, J. B.; Schlesinger, L. S., Integrating Lung Physiology, Immunology, and Tuberculosis. Trends Microbiol 2017, 25, (8), 688-697.
- Broxmeyer, L.; Sosnowska, D.; Miltner, E.; Chacon, O.; Wagner, D.; McGarvey, J.; Barletta, R. G.; Bermudez, L. E., Killing of Mycobacterium avium and Mycobacterium tuberculosis by a mycobacteriophage delivered by a nonvirulent mycobacterium: a model for phage therapy of intracellular bacterial pathogens. J Infect Dis 2002, 186, (8), 1155-60.
- Danelishvili, L.; Young, L. S.; Bermudez, L. E., In vivo efficacy of phage therapy for Mycobacterium avium infection as delivered by a nonvirulent mycobacterium. Microb Drug Resist 2006, 12, (1), 1-6.
- Sweeney, K. A.; Dao, D. N.; Goldberg, M. F.; Hsu, T.; Venkataswamy, M. M.; Henao-Tamayo, M.; Ordway, D.; Sellers, R. S.; Jain, P.; Chen, B.; Chen, M.; Kim, J.; Lukose, R.; Chan, J.; Orme, I. M.; Porcelli, S. A.; Jacobs, W. R., Jr., A recombinant Mycobacterium smegmatis induces potent bactericidal immunity against Mycobacterium tuberculosis. Nat Med 2011, 17, (10), 1261-8.
- Nieth, A.; Verseux, C.; Barnert, S.; Suss, R.; Romer, W., A first step toward liposome-mediated intracellular bacteriophage therapy. Expert Opin Drug Deliv 2015, 12, (9), 1411-24.
- Rohde, C.; Resch, G.; Pirnay, J. P.; Blasdel, B. G.; Debarbieux, L.; Gelman, D.; Gorski, A.; Hazan, R.; Huys, I.; Kakabadze, E.; Lobocka, M.; Maestri, A.; Almeida, G. M. F.; Makalatia, K.; Malik, D. J.; Maslanova, I.; Merabishvili, M.; Pantucek, R.; Rose, T.; Stverakova, D.; Van Raemdonck, H.; Verbeken, G.; Chanishvili, N., Expert Opinion on Three Phage Therapy Related Topics: Bacterial Phage Resistance, Phage Training and Prophages in Bacterial Production Strains. Viruses 2018, 10, (4).
- Kim, J. S., Microbial warfare against viruses. Science 2018, 359, (6379), 993.
- Seed, K. D., Battling Phages: How Bacteria Defend against Viral Attack. Plos Pathog 2015, 11, (6).
- Labrie, S. J.; Samson, J. E.; Moineau, S., Bacteriophage resistance mechanisms. Nat Rev Microbiol 2010, 8, (5), 317-27.
- Hampton, H. G.; Watson, B. N. J.; Fineran, P. C., The arms race between bacteria and their phage foes. Nature 2020, 577, (7790), 327-336.
- Maxwell, K. L., Phages Fight Back: Inactivation of the CRISPR-Cas Bacterial Immune System by Anti-CRISPR Proteins. Plos Pathog 2016, 12, (1), e1005282.
- Goldfarb, T.; Sberro, H.; Weinstock, E.; Cohen, O.; Doron, S.; Charpak-Amikam, Y.; Afik, S.; Ofir, G.; Sorek, R., BREX is a novel phage resistance system widespread in microbial genomes. Embo J 2015, 34, (2), 169-183.
- Ofir, G.; Melamed, S.; Sberro, H.; Mukamel, Z.; Silverman, S.; Yaakov, G.; Doron, S.; Sorek, R., DISARM is a widespread bacterial defence system with broad anti-phage activities. Nature Microbiology 2018, 3, (1).
- Nguyen, Q. H.; Contamin, L.; Nguyen, T. V. A.; Banuls, A. L., Insights into the processes that drive the evolution of drug resistance in Mycobacterium tuberculosis. Evolutionary Applications 2018, 11, (9), 1498-1511.
- Capparelli, R.; Nocerino, N.; Lanzetta, R.; Silipo, A.; Amoresano, A.; Giangrande, C.; Becker, K.; Blaiotta, G.; Evidente, A.; Cimmino, A.; Iannaccone, M.; Parlato, M.; Medaglia, C.; Roperto, S.; Roperto, F.; Ramunno, L.; Iannelli, D., Bacteriophage-resistant Staphylococcus aureus mutant confers broad immunity against staphylococcal infection in mice. Plos One 2010, 5, (7), e11720.
- Evans, T. J.; Trauner, A.; Komitopoulou, E.; Salmond, G. P., Exploitation of a new flagellatropic phage of Erwinia for positive selection of bacterial mutants attenuated in plant virulence: towards phage therapy. J Appl Microbiol 2010, 108, (2), 676-85.
- Chan, B. K.; Sistrom, M.; Wertz, J. E.; Kortright, K. E.; Narayan, D.; Turner, P. E., Phage selection restores antibiotic sensitivity in MDR Pseudomonas aeruginosa. Sci Rep 2016, 6, 26717.
- Wright, R. C. T.; Friman, V. P.; Smith, M. C. M.; Brockhurst, M. A., Cross-resistance is modular in bacteria-phage interactions. Plos Biol 2018, 16, (10).
- Hall, A. R.; De Vos, D.; Friman, V. P.; Pirnay, J. P.; Buckling, A., Effects of sequential and simultaneous applications of bacteriophages on populations of Pseudomonas aeruginosa in vitro and in wax moth larvae. Appl Environ Microbiol 2012, 78, (16), 5646-52.
- Chaudhry, W. N.; Concepcion-Acevedo, J.; Park, T.; Andleeb, S.; Bull, J. J.; Levin, B. R., Synergy and Order Effects of Antibiotics and Phages in Killing Pseudomonas aeruginosa Biofilms. Plos One 2017, 12, (1), e0168615.
- Torres-Barcelo, C.; Hochberg, M. E., Evolutionary Rationale for Phases as Complements of Antibiotics. Trends in Microbiology 2016, 24, (4), 249-256.
- Oechslin, F.; Piccardi, P.; Mancini, S.; Gabard, J.; Moreillon, P.; Entenza, J. M.; Resch, G.; Que, Y. A., Synergistic Interaction Between Phage Therapy and Antibiotics Clears Pseudomonas Aeruginosa Infection in Endocarditis and Reduces Virulence. J Infect Dis 2017, 215, (5), 703-712.
- Zhang, Q. G.; Buckling, A., Phages limit the evolution of bacterial antibiotic resistance in experimental microcosms. Evol Appl 2012, 5, (6), 575-82.
- Trasta, A., Personalized medicine and proper dosage: Over- and undertreatment of chronic diseases endanger patients' health and strain public health systems. EMBO Rep 2018, 19, (4).
- Pride, D. T.; Salzman, J.; Haynes, M.; Rohwer, F.; Davis-Long, C.; White, R. A.; Loomer, P.; Armitage, G. C.; Relman, D. A., Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. Isme J 2012, 6, (5), 915-926.
- Santiago-Rodriguez, T. M.; Ly, M.; Bonilla, N.; Pride, D. T., The human urine virome in association with urinary tract infections. Front Microbiol 2015, 6, 14.
- Navarro, F.; Muniesa, M., Phages in the Human Body. Front Microbiol 2017, 8, 566.
- Lim, E. S.; Zhou, Y.; Zhao, G.; Bauer, I. K.; Droit, L.; Ndao, I. M.; Warner, B. B.; Tarr, P. I.; Wang, D.; Holtz, L. R., Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat Med 2015, 21, (10), 1228-34.
- Krut, O.; Bekeredjian-Ding, I., Contribution of the Immune Response to Phage Therapy. J Immunol 2018, 200, (9), 3037-3044.
- Barr, J. J.; Auro, R.; Furlan, M.; Whiteson, K. L.; Erb, M. L.; Pogliano, J.; Stotland, A.; Wolkowicz, R.; Cutting, A. S.; Doran, K. S.; Salamon, P.; Youle, M.; Rohwer, F., Bacteriophage adhering to mucus provide a non-host-derived immunity. Proc Natl Acad Sci U S A 2013, 110, (26), 10771-6.
- Nguyen, S.; Baker, K.; Padman, B. S.; Patwa, R.; Dunstan, R. A.; Weston, T. A.; Schlosser, K.; Bailey, B.; Lithgow, T.; Lazarou, M.; Luque, A.; Rohwer, F.; Blumberg, R. S.; Barr, J. J., Bacteriophage Transcytosis Provides a Mechanism To Cross Epithelial Cell Layers. mBio 2017, 8, (6).
- Levin, B. R.; Bull, J. J., Population and evolutionary dynamics of phage therapy. Nat Rev Microbiol 2004, 2, (2), 166-73.
- Huh, H.; Wong, S.; St Jean, J.; Slavcev, R., Bacteriophage interactions with mammalian tissue: Therapeutic applications. Adv Drug Deliv Rev 2019, 145, 4-17.
- Dabrowska, K., Phage therapy: What factors shape phage pharmacokinetics and bioavailability? Systematic and critical review. Med Res Rev 2019, 39, (5), 2000-2025.
- Jonczyk-Matysiak, E.; Weber-Dabrowska, B.; Owczarek, B.; Miedzybrodzki, R.; Lusiak-Szelachowska, M.; Lodej, N.; Gorski, A., Phage-Phagocyte Interactions and Their Implications for Phage Application as Therapeutics. Viruses-Basel 2017, 9, (6).
- Roach, D. R.; Leung, C. Y.; Henry, M.; Morello, E.; Singh, D.; Di Santo, J. P.; Weitz, J. S.; Debarbieux, L., Synergy between the Host Immune System and Bacteriophage Is Essential for Successful Phage Therapy against an Acute Respiratory Pathogen. Cell Host Microbe 2017, 22, (1), 38-+.
- Xiong, X.; Zhang, H. M.; Wu, T. T.; Xu, L.; Gan, Y. L.; Jiang, L. S.; Zhang, L.; Guo, S. L., Titer dynamic analysis of D29 within MTB-infected macrophages and effect on immune function of macrophages. Exp Lung Res 2014, 40, (2), 86-98.
- McCallin, S.; Sarker, S. A.; Sultana, S.; Oechslin, F.; Brussow, H., Metagenome analysis of Russian and Georgian Pyophage cocktails and a placebo-controlled safety trial of single phage versus phage cocktail in healthy Staphylococcus aureus carriers. Environ Microbiol 2018, 20, (9), 3278-3293.
- Sarker, S. A.; McCallin, S.; Barretto, C.; Berger, B.; Pittet, A. C.; Sultana, S.; Krause, L.; Huq, S.; Bibiloni, R.; Bruttin, A.; Reuteler, G.; Brussow, H., Oral T4-like phage cocktail application to healthy adult volunteers from Bangladesh. Virology 2012, 434, (2), 222-32.
- Petrovic Fabijan, A.; Lin, R. C. Y.; Ho, J.; Maddocks, S.; Ben Zakour, N. L.; Iredell, J. R.; Westmead Bacteriophage Therapy, T., Safety of bacteriophage therapy in severe Staphylococcus aureus infection. Nat Microbiol 2020.
- Pelfrene, E.; Willebrand, E.; Cavaleiro Sanches, A.; Sebris, Z.; Cavaleri, M., Bacteriophage therapy: a regulatory perspective. J Antimicrob Chemother 2016, 71, (8), 2071-4.