
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Joshua Waxman | + 4769 word(s) | 4769 | 2021-02-17 04:39:46 | | | |
| 2 | Vivi Li | Meta information modification | 4769 | 2021-02-23 04:12:13 | | |
Video Upload Options
Proper development and function of the vertebrate heart is vital for embryonic and postnatal life. Many congenital heart defects in humans are associated with disruption of genes that direct the formation or maintenance of atrial and pacemaker cardiomyocytes at the venous pole of the heart. Zebrafish are an outstanding model for studying vertebrate cardiogenesis, due to the conservation of molecular mechanisms underlying early heart development, external development, and ease of genetic manipulation.
1. Introduction
All vertebrate hearts comprise the same fundamental structural units: the atrial and ventricular chambers, which are specialized to reflect their functions receiving and expelling blood, respectively. The coordinated assembly of these chambers into a functional organ is an intricate process that is critical for normal embryonic development and throughout life of all vertebrates. In humans, mutations in genes that disrupt chamber formation and maintenance are associated with congenital heart defects (CHDs), which are the most common type of birth defect [1][2][3][4][5][6][7][8][9][10]. Many of these signals affect proper development of the atrial chamber and its ability to function, with atrial septal defects (ASDs) making up as much as 20% of CHDs found in children and adults [5][6][7][8][9]. Moreover, mutations affecting development of the sinoatrial node (SAN) can result in cardiac pacing defects, such as sick sinus syndrome, arrhythmias, and atrial fibrillation [11][12][13][14][15].
While proper blood circulation is necessary in all vertebrate organisms, the number of atrial and ventricular chambers within vertebrate hearts can vary. Mammals have a four-chambered heart with two atria and two ventricles, which are necessary for circulation to the lungs and the rest of the body. However, teleosts, such as the common model zebrafish, have a relatively simple two-chambered heart consisting of only a single atrium and ventricle. Despite these differences, the underlying molecular signals and morphogenetic processes, particularly those that govern the specification and maintenance of cardiac chambers during heart development, are conserved across species. Moreover, due to their external development, the ease of genetic manipulation, and their ability to survive the first week post fertilization without a functional cardiovascular system, zebrafish have become an excellent tool for understanding the early processes underlying vertebrate cardiogenesis [16][17][18][19][20].
Proper heart function relies on the coordinated contraction of the atrial and ventricular chambers. Importantly, zebrafish cardiomyocytes share key electrophysiological properties with mammalian cardiomyocytes, allowing them to also serve as a model for human conduction defects [16][21]. Cardiac contraction is initiated by a population of specialized cardiomyocytes in the SAN, which is located at the base of the right atrial chamber in mammals and the venous pole of the single atrial chamber in zebrafish. The electrical signal then propagates across the atrium, where it pauses at the atrioventricular canal (AVC) before moving rapidly across the ventricle [22]. The cardiomyocytes within the ventricular and atrial chambers and the SAN are functionally distinct with characteristics conferred by the expression of specific gene programs, contractile proteins, and ion channels [23][24][25][26][27]. The thin-walled atrium possesses cardiomyocytes that have a squamous morphology, disorganized sarcomeres, and a triangular action potential [23][24][25][28][29]. Cardiomyocytes of the thick-walled, highly trabeculated ventricle are more cuboidal, with organized sarcomeres, and a flat action potential plateau [22][24][25]. SAN cardiomyocytes have a slower rate of depolarization and are able to generate spontaneous action potentials [22][26][30][31]. In this review, we highlight mechanisms governing the development of the atrium and the SAN within the venous pole of the zebrafish heart and explore how information from the zebrafish can be used as a model to study human CHDs and arrhythmias.
2. Mechanisms of Atrial Chamber Development and Chamber Identity Maintenance in the Zebrafish Heart
2.1. Cardiac Progenitor Location and Morphology of the Developing Zebrafish Heart
Like all vertebrates, the zebrafish heart is the first organ to form and function during embryogenesis. The initial stages of zebrafish heart development involve the specification, migration, and differentiation of cardiomyocyte progenitor cells as they are integrated into the forming heart tube [18][19][32][33][34]. Cell lineage tracing studies with caged-fluorescein have produced fate maps of the early zebrafish cardiac progenitors prior to gastrulation, which show that precursors for both atrial and ventricular cardiomyocytes first reside in bilateral populations in the lateral marginal zone by 5 h post-fertilization (hpf) [32][35]. Within the late blastula embryos, atrial cardiomyocyte progenitors are located more ventrally and slightly farther from the margin, the region of the embryo where cells will involute during gastrulation. Ventricular cardiomyocyte precursors are located more dorsally and closer to the margin. While there is a small region of overlap, these progenitor populations appear to be largely distinct; labeled cells predominantly give rise to only atrial or ventricular cardiomyocytes [32][33][35] (Figure 1A). During gastrulation, cardiomyocyte progenitor cells within the lateral mesoderm involute and migrate anteriorly to form bilateral progenitor fields in the anterior lateral plate mesoderm (ALPM) (Figure 1B). As indicated from the earlier fate maps, lineage tracing of the ALPM at later stages suggest that atrial and ventricular cardiomyocyte precursor populations retain predominantly adjacent positions: ventricular progenitors reside more medially and atrial progenitors lay more laterally [35]. Once in the ALPM, the cardiac progenitors begin to differentiate, as indicated by the expression of chamber-specific myosin genes. Expression of ventricular myosin heavy chain (vmhc), also referred to as myosin heavy chain 7 (myh7), can be detected in the medial ventricular progenitors as early as the 13 somite stage [36]. Expression of atrial myosin heavy chain (amhc), also named myosin heavy chain 6 (myh6), begins slightly later at 19 somites [33] (Figure 1C). While different myosins are some of the gene expression differences that define atrial and ventricular cardiomyocytes, one study has indicated that left–right asymmetry, reminiscent of the left and right atrial chambers in mammals, can also be seen as early as the 19 somite stage [37]. It was shown that pitx2c and meis2b are upregulated on the left and downregulated on the right in the zebrafish atrium, consistent with the left atrial expression pattern of Pitx2 in mammals [37][38]. As the cardiac progenitor cells are differentiating, they migrate to the midline, begin to fuse more posteriorly (Figure 1C), and then form the cardiac disc in which differentiating atrial cardiomyocytes lie peripherally to ventricular cardiomyocytes (Figure 1D).
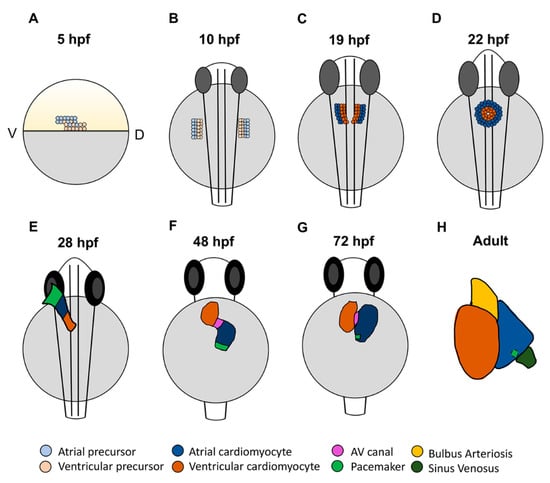
Figure 1. Stages of zebrafish heart development. (A) At 5 h post-fertilization (hpf), cardiac progenitors are located in the lateral marginal zone, with atrial progenitors located more ventrally than ventricular progenitors. (B) Following gastrulation at the tailbud stage (10 hpf), cardiac progenitors migrate to the anterior lateral plate mesoderm (ALPM). (C) In the ALPM, progenitors begin to differentiate and express chamber-specific genes. (D) Cells then migrate to the midline and fuse, forming the cardiac disc where atrial cardiomyocytes surround ventricular cardiomyocytes. (E) The disc elongates to form the linear heart tube, which begins beating by 24 hpf. At 28 hpf, the dominant pacemaker covers a large area at the venous pole. (F) By 48 hpf, the heart has finished looping and the two chambers have formed. Here, the pacemaker is a ring at the venous pole. (G) By 72 hpf, the dominant pacemaker is restricted to a small population of cells in the inner curvature at the venous pole of the atrium. (H) In the adult heart, the bulbus arteriosus and sinus venosus, which serve as the outflow and inflow tracts, respectively, have matured. The dominant pacemaker is located at the sinus venosus–atrial junction.
Importantly, vertebrate hearts are formed from the progressive differentiation of progenitor populations that grow the heart at the poles. Thus, the zebrafish cardiac disc then elongates over the yolk to form the linear heart tube (Figure 1E). The earlier-differentiating cells that comprise the initial heart tube are termed the first heart field (FHF) [39][40]. Populations of later differentiating cells, referred to as the second heart field (SHF), then contribute to both the venous and arterial poles as the heart tube elongates and grows [41][42][43]. For comparison, in mammals, the SHF contributes to the right ventricle, while the left ventricle is primarily derived from the FHF. Although both heart fields contribute to the left and right atrial chambers, it has been suggested that as many as two thirds of atrial cardiomyocytes are derived from the SHF [44]. In zebrafish, the SHF primarily contributes to the outflow tract and half of the ventricle, with a small number of posterior SHF cells populating the venous pole of the atrium [18][34][42][45]. As the heart tube elongates, it migrates left and begins looping, where by 48 hpf the physically distinct chambers have formed: the venous left-sided atrium and arterial right-sided ventricle are easily distinguishable and separated by the AVC [18][34] (Figure 1F). From this point, the heart will continue to mature and develop the additional structures that are characteristic of each chamber [19][46] (Figure 1G,H). Trabeculation in the ventricle and valve leaflet development are evident by three days post-fertilization (dpf). The mature ventricle comprises a cortical cardiomyocyte and deeper layer that make up the trabeculae, which form through the delamination of cardiomyocytes from the compact layer beginning at 3 dpf [47][48]. Conversely, the atrial myocardium becomes pectinate in structure by about 14 dpf [46][49]. Genetic lineage tracing has shown that pectinate atrial cardiomyocytes in the mature atrium form through the proliferation and branching of existing cardiomyocytes [49], suggesting that there are different mechanisms driving the maturation of the atrial and ventricular chambers. Thus, the heart undergoes extensive morphological changes throughout development that establish each of the cardiac chambers.
2.2. Signals Regulating the Specification of Atrial Progenitors
Numerous early signaling pathways are involved in the regulation of cardiomyocyte specification as part of their requirements in the establishment of the body plan, which factors into determination of the number of atrial cardiomyocytes within the zebrafish heart. Signals including Hedgehog (HH), canonical Wnt/β-catenin (Wnt), and Nodal exert similar effects promoting the specification of both atrial and ventricular cardiomyocytes. Increasing HH signaling through the injection of sonic hedgehog mRNA promotes atrial and ventricular specification and yields hearts with increased cardiomyocytes, while loss of HH signaling through the genetic and pharmacological inhibition of the Smoothened receptor reduces cardiac progenitor specification, consequently resulting in hearts with decreased atrial and ventricular cardiomyocytes [50]. Similarly, using heat-shock inducible transgenes to increase and decrease canonical Wnt signaling by the initiation of gastrulation leads to corresponding increases and decreases in cardiomyocyte progenitor specification, respectively [51][52], due to effects on the induction of mesoderm. Nodal signaling, which is required for mesoderm specification, is also required for cardiomyocyte specification. Specifically, mutations in the Apelin receptor, which modulates Nodal signaling, also lead to a loss of both atrial and ventricular cardiomyocytes [53][54] (Figure 2A).
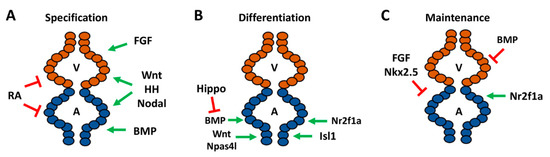
Figure 2. Signaling pathways required for different stages of atrial development and cardiac maintenance in zebrafish. (A) Pathways required for specification of chamber progenitors in the early embryo. (B) Factors responsible for differentiation of atrial cardiomyocytes within the ALPM and at the venous pole. (C) Genes shown to promote or repress atrial identity within differentiated embryonic ventricular cardiomyocytes and repress ventricular gene expression in atrial cardiomyocytes. A—atrium, V—ventricle.
Although initial studies in mice and zebrafish concluded that retinoic acid (RA) signaling promotes atrial cardiomyocyte specification [55][56], pivotal work in zebrafish showed that its earliest requirement is to restrict cardiac specification of both atrial and ventricular progenitors within the ALPM [57][58]. Specifically, inhibition of RA signaling through use of the neckless (nls) mutant, which harbors a mutation in the major embryonic producer of RA aldh1a2 (formerly called raldh2), and treatment with RA signaling antagonists results in enlarged hearts with an increase in the number of atrial and ventricular cardiomyocytes [57][58]. Excess RA signaling can also inhibit the specification of both atrial and ventricular cardiomyocytes, while intermediate increases in RA signaling can differentially affect the specification of cardiomyocytes, which likely led to the initial interpretations that RA signaling promotes atrial specification [59][60]. Thus, the specification of both atrial and ventricular progenitors is effectively balanced through opposing roles of early HH, Wnt, and Nodal, which promote cardiomyocyte specification, and the restrictive role of RA signaling (Figure 2A).
In contrast to signals that have similar consequences on the specification of atrial and ventricular cardiomyocytes, early signals that affect dorsal–ventral patterning have differential effects on chamber specification. The dorsalizing pathway fibroblast growth factor (FGF) signaling preferentially affects the specification of ventricular progenitors with minimal effect on the atrium [61]. Conversely, the ventralizing bone morphogenic protein (BMP) signaling pathway is necessary and sufficient to promote atrial specification. Loss of BMP signaling in the alk8/lost-a-fin (laf) mutants, treatment with pharmacological BMP signaling inhibitors, and an inducible dominant-negative Alk8 transgene all demonstrated that BMP signaling is required early for atrial progenitor specification during gastrulation and results in hearts with decreased atrial cardiomyocytes. Consistently, increasing BMP signaling through the injection of constitutively-active alk8 mRNA results in increased atrial cardiomyocytes, reinforcing the role of BMP in atrial specification [62][63]. Thus, early signaling that ventralizes the embryo is required for the specification of atrial precursors (Figure 2A).
2.3. Signals Directing the Differentiation of Atrial Cardiomyocytes
Subsequent to the signals that modulate atrial cardiomyocyte number and chamber size through establishment of cardiomyocyte progenitor populations, signals then control atrial chamber size through influencing cardiomyocyte differentiation from within the ALPM. While RA signaling may restrict the number of both atrial and ventricular cardiomyocytes, the lineage tracing analysis suggested that signals downstream of RA may differentially affect atrial and ventricular productions [58]. Depletion of hoxb5b, an RA-responsive gene within the ALPM which is posterior to the cardiac progenitors, has been shown to result in excess atrial cardiomyocytes, suggesting it may function downstream of RA to restrict the number of atrial cardiomyocytes [58]. However, these results require confirmation with genetic mutants. Additionally, cloche mutants, which we now understand are caused by mutation of the transcription factor Npas4l [64], and downstream factors Scl (Tal1) and Etv2 (Etsrp) function to restrict heart size through promoting hematopoietic and vascular lineages within the ALPM. Cloche mutants or concurrent depletion of Scl and Etv2 with morpholinos both produce enlarged hearts due to an increase in atrial cardiomyocytes [65] (Figure 2B). In addition to its early role, canonical Wnt signaling functions reiteratively in later roles. At the tailbud stage, excess Wnt signaling can potently inhibit the differentiation, but not the specification, of both atrial and ventricular progenitors [51]. However, in contrast to earlier requirements that have similar effects on cardiomyocyte progenitors, during later stages of somitogenesis Wnt signaling specifically affects the differentiation of atrial cardiomyocytes. Using heat-shock inducible transgenes to modulate Wnt signaling at the 16 somite stage, when the cardiac progenitors reside within bilateral populations in the ALPM, showed that increased Wnt signaling produces an increase in the number of atrial cardiomyocytes and decreased Wnt signaling produces fewer atrial cardiomyocytes [51]. Similarly, while BMP signaling is predominantly required for atrial cardiomyocyte specification throughout gastrulation, it has also been shown to play reiterative roles in atrial differentiation. Blocking BMP signaling using a heat-shock inducible Noggin3 transgene at 16 hpf causes a specific decrease in atrial cardiomyocytes that is independent of its earlier role ventralizing embryos [63]. In contrast to the positive influences of Wnt and BMP signaling on atrial differentiation during late somitogenesis, research has shown that, at these stages, Hippo signaling restricts the number of atrial cardiomyocytes added to the venous pole. Large tumor suppressor kinase (Lats) proteins are critical regulators of the Hippo pathway [66], because activation of Lats1/2 is required for proper Hippo signaling. In lats1; lats2 mutants, it was shown that decreased Hippo signaling results in an increased number of atrial cardiomyocytes, supporting that Hippo signaling restricts atrial size. Subsequent analysis revealed that Hippo functions upstream of BMP signaling to restrict the number of SHF-derived cardiomyocytes that arise from the ALPM and contribute to the venous pole of the heart [67] (Figure 2B), providing insight into the interplay between these signaling pathways in heart development.
In addition to the aforementioned signaling pathways, specific members of the Islet (Isl) and Nuclear receptor 2 (Nr2f/Coup-tf) transcription factor families have been shown to be necessary to promote the differentiation of atrial cardiomyocytes at the venous pole. Isl1 marks the venous SHF in zebrafish. Consistent with this expression, isl1 mutants have decreased atrial cardiomyocytes, which temporal differentiation assays have shown to be due to reduced differentiation [41] (Figure 2B). Interestingly, Isl1 is specifically required for the differentiation of cardiomyocytes at the venous pole in zebrafish embryos, contrasting with the requirement of Isl1 in mice, which is required for the differentiation of all SHF progenitors at both the posterior venous and anterior arterial poles [41][44]. However, it has been shown that isl2b is required for development of the arterial pole in zebrafish, suggesting that in teleosts Islet family members have distinct roles in progenitor differentiation at the different poles of the heart [68].
Nr2f transcription factors are the proteins that are most prominently associated with the determination of atrial identity in vertebrates. Global knockout of Nr2f2 in mice results in embryos with small, dysmorphic atria [69]. In zebrafish, our lab has shown that Nr2f1a is the functional equivalent of mammalian Nr2f2 with respect to heart development. Within the embryonic zebrafish heart, Nr2f1a is expressed specifically within atrial cardiomyocytes and progenitors at the venous pole. Nr2f1a mutant zebrafish develop smaller atria due to decreased differentiation of atrial cardiomyocytes at the venous pole [70]. Thus, Wnt, BMP, and Hippo signaling and the Hoxb5b, Isl1, and Nr2f1a transcription factors all influence the proper differentiation of atrial cardiomyocytes within the zebrafish heart (Figure 2B).
2.4. Maintenance and Plasticity of Cardiomyocyte Identity
Although the lineage tracing of blastula cell embryos has suggested that normally the fates of cardiomyocytes are largely distinct [35], studies have indicated that there is plasticity between atrial and ventricular progenitors and that chamber identity must be continually reinforced in zebrafish embryos after the cardiomyocytes have overtly differentiated [25][27][28][71][72][73][74]. A clue that there is greater plasticity between atrial and ventricular cardiomyocytes in normal zebrafish heart development was indicated from genetic labeling of embryonic cardiomyocytes with amhc:CreERT2 and loxP-mediated color-switch reporter transgenes [49]. Some amhc-labeled cardiomyocytes within the embryos differentiate as ventricular cardiomyocytes, suggesting that there are initially cardiomyocytes which have begun to differentiate with the potential to become either atrial or ventricular cardiomyocytes, but then differentiate as ventricular cardiomyocytes as the embryonic heart matures [49]. The necessity to maintain ventricular identity at the expense of atrial identity is illustrated by the Nkx2.5 and Nkx2.7 transcription factors [73][75]. Zebrafish nkx2.5 mutants initially have an equivalent number of atrial and ventricular cardiomyocytes as their wild-type siblings. However, they progressively develop an enlarged atrium and diminutive ventricle as the ventricular cardiomyocytes take on atrial identity (Figure 2C). While Nkx2.5 is the primary factor necessary to maintain ventricular identity, concomitant loss of Nkx2.7 has been shown to exacerbate the fate transformation, suggesting some limited redundancy between these transcription factors. Nkx2.5/7 maintain ventricular identity by promoting the expression of ventricular-specific genes, irx4 and hey2 [73][75]. Remarkably, induction of Nkx2.5 with a heat-shock inducible transgene at late somitogenesis is sufficient to rescue Nkx2.5 mutants to adulthood, which suggests that the requirement of Nkx2.5 in repressing atrial identity in ventricular cardiomyocytes occurs within a specific developmental window, but that it does not perform this role throughout life [75].
Fgf8a is expressed within the zebrafish ventricle [76]. It has been shown that the FGF pathway sits at the top of a signaling cascade to reinforce ventricular identity at the expense of atrial identity [74]. Inhibition of FGF signaling with a dominant negative transgene or pharmacological inhibitors beginning at the 18 somite stage results in ectopic atrial gene expression within the ventricle [74]. Interestingly, it was shown that overexpression of Nkx2.5 in embryos while concurrently inhibiting FGF signaling resulted in a partial rescue of this fate transformation, indicating that FGF signaling functions upstream of Nkx2.5 in maintaining ventricular identity (Figure 2C), but that other unidentified factors likely also play a role in the repression of atrial identity within ventricular cardiomyocytes [74]. While FGF signaling is necessary to maintain ventricular identity, BMP signaling may also need to be actively repressed in ventricular cardiomyocytes to prevent the acquisition of atrial cardiomyocyte identity (Figure 2C). Increasing BMP signaling at the 18 somite stage with a heat-shock inducible BMP2b in embryos depleted of smad6a, which represses BMP signaling, results in both fewer ventricular cardiomyocytes and some ventricular cardiomyocytes that ectopically express Amhc [63]. However, the mechanisms by which excess BMP signaling promotes atrial identity in the ventricle still needs to be elucidated. The plasticity between atrial and ventricular cardiomyocyte identities can also be seen in the regenerative response to cardiac injury in embryonic zebrafish hearts. Upon ablation of ventricular cardiomyocytes in 5 dpf larval zebrafish, atrial cardiomyocytes proliferate and transdifferentiate into ventricular cardiomyocytes to regenerate the ventricle [77]. This atrial-to-ventricular injury response is diminished in adult zebrafish, suggesting that, like the requirement of Nkx2.5 in maintaining ventricular identity, there is an age-dependent developmental window to this plasticity [77].
Despite the evidence that factors maintain ventricular identity in embryonic zebrafish hearts, it is not known if there are genes responsible for maintaining atrial identity. Thus far, factors have not been identified that clearly show differentiated atrial cardiomyocytes need to repress ventricular cardiomyocyte identity in zebrafish. However, Nr2f transcription factors have been shown to maintain atrial cardiomyocyte identity in mice, in addition to their requirement in atrial cardiomyocyte differentiation. In mice, cardiac-specific knockout of Nr2f2 with Myh6-Cre results in ectopic expression of ventricular genes within atrial cardiomyocytes, because Nr2f2 represses ventricular-identity genes Irx4 and Hey2 [78]. Consistently, overexpression of Nr2f2 was sufficient to confer atrial identity within ventricular cardiomyocytes [78]. In addition to reduced atrial cardiomyocyte number, nr2f1a mutant zebrafish display an expansion of AVC markers into the atrium, suggesting that Nr2f1a is required to limit the size of the AVC [70]. Nr2f1a mutants also have an expansion of the ventricular differentiation marker vmhc into the atrium (Figure 2C), although there was no apparent effect on ventricular cardiomyocyte number at least through 48 hpf [70]. It is not clear, however, that these effects represent a transdifferentiation of the atrial cardiomyocytes comparable to what is found in mice. Additional studies are needed to determine if atrial cardiomyocytes in Nr2f1a mutant zebrafish undergo the same fate transformation that is seen in the Nr2f2 conditional knockout mice at later stages of development. Overall, these studies have shown that differentiated embryonic cardiomyocytes in zebrafish maintain a degree of plasticity and the ventricle will adopt atrial identity in the absence of critical maintenance factors (Figure 2C).
2.5. Conserved Transcriptional Networks Promoting Sinoatrial Node Development
The zebrafish heart first begins beating at 24 hpf with the formation of the linear heart tube. Initial contractions are slow and peristaltic, originating at the venous pole. As the heart loops, the electrophysiology changes and a conduction delay in the AVC leads to the sequential atrial–ventricular contraction pattern [22]. With the use of optogenetics, it has been shown that the initial pacemaker region is a large area at the venous pole of the heart tube, which then condenses to form a ring by 48 hpf (Figure 1E,F). By 72 hpf, the dominant pacemaker has narrowed further to a small population of cells in the inner curvature of the atrium [79] (Figure 1G). Interestingly, this location is analogous to the location of the pacemaker in the right atrium of the mammalian heart.
The genetic mechanisms underlying development of the SAN have been extensively studied in mice, but thus far they have not been as well characterized in zebrafish. We will cover specifically what is known about the development of the SAN in zebrafish in comparison to mammals. Multiple reviews have already detailed the development of the cardiac conduction system as a whole [31][80][81]. Work in mice has shown that information from a complex genetic network of transcription factors is integrated to establish SAN identity within the atrial myocardium at the venous pole. T-box transcription factors Tbx3 and Tbx18 have been shown to be vital for proper SAN development, although they have different functions [82]. Tbx18 deficient mice develop a severely hypoplastic SAN, however the SAN gene program is maintained within these cells and the mice do not develop overt bradycardia. Conversely, Tbx3 deficiency leads to a morphologically normal SAN that ectopically expresses atrial myocardium genes. This suggests that Tbx18 is required for SAN morphogenesis and Tbx3 is required for pacemaker differentiation [82]. Interestingly, overexpression of Tbx3 in the mouse atrial myocardium is sufficient to confer pacemaker identity, highlighting its role as a key driver of SAN identity [83]. In contrast to Tbx3 and Tbx18, Nkx2.5 represses SAN identity within atrial cardiomyocytes. Nkx2.5 mutant mice have an enlarged SAN, while Nkx2.5 overexpression leads to SAN hypoplasia and decreased expression of Tbx3 [84][85][86]. Isl1 and Shox2 have also been shown to be expressed in and required for pacemaker development [87][88][89][90][91]. Loss of function of either of these transcription factors results in a hypoplastic SAN and decreased expression of Tbx3 [90][91]. Shox2 also functions upstream of Nkx2.5, with Shox2 mutants developing ectopic expression of Nkx2.5 within the pacemaker. Overexpression of Shox2 is also able to partially repress Nkx2.5 [84][89][91][92]. Thus, these studies in mice have established a multi-tier genetic network in which cross regulation of the transcription factors Tbx3, Isl1, and Shox2 promote SAN identity and oppose the repressive action of Nkx2.5 to confer SAN identity within a proper number of atrial cardiomyocytes at the venous pole of the heart.
2.6. Development of the Zebrafish Sinoatrial Node
While earlier studies in zebrafish indicated the presence of specialized pacemaker tissue in the heart [22][79], the molecular regulation of the zebrafish SAN has historically been largely understudied, in part due to the lack of reliable SAN markers. Isl1 was the first molecular marker identified for the zebrafish SAN. Isl1-expressing cells form a ring at the venous pole of the atrium in embryonic hearts [26]. Its expression is maintained in a ring at the base of the atrium and junction with the sinus venosus through adulthood. In addition to reduced atrial cardiomyocytes, isl1 mutants have an abnormal heartbeat, highlighting its conserved importance in pacemaker development and function [26][41]. It has been indicated that Isl1 promotes SAN development upstream of Wnt/β-catenin, because Wnt signaling is lost in the SAN of isl1 mutants [93]. Furthermore, at these later stages, Wnt signaling controls heart rate through modulating the response of the pacemaker cardiomyocytes to parasympathetic input [93]. In contrast to mice, which have lost Shox and only have Shox2, humans and zebrafish have retained both Shox and Shox2. Interestingly, by replacing Shox2 with Shox in mice, it has been shown that human Shox and mouse Shox2 can similarly promote SAN development [92]. In zebrafish, Shox2 has been shown to promote SAN identity, with shox2 morpholino-depleted zebrafish developing bradycardia [89]. Overexpression of Isl1 in Shox2 morphants can rescue the bradycardia phenotype, suggesting that Shox2 functions upstream of Isl1 in SAN development [94] (Figure 3). However, the requirement of Shox2 in zebrafish and the relationship to Isl1 within the SAN should be verified through the use of genetic mutants. Consistent with conserved requirements for Nkx2.5 repressing SAN identity within working atrial cardiomyocytes, studies have shown that Nkx2.5 limits Isl1 expression to the venous pole of the atrium [95] (Figure 3). Nkx2.5 zebrafish mutants have a progressive expansion of Isl1+ atrial cardiomyocytes. Conversely, Nkx2.5 overexpression leads to diminished Isl1 expression at the venous pole [84][85][95]. Nkx2.5 expression is largely absent from the SAN cells, although Nkx2.5 is expressed throughout the zebrafish heart at earlier stages; therefore, it was questioned if the Nkx2.5–SAN cardiomyocytes are derived from an Nkx2.5+ progenitor population [96][97]. Lineage tracing in zebrafish embryos confirmed that the pacemaker cardiomyocytes are derived from Nkx2.5+ progenitors that come from the most lateral regions of the cardiac mesoderm [98]. Furthermore, in this population, Wnt5b initiates canonical Wnt signaling, which silences Nkx2.5 and activates Isl1 and Tbx18 (Figure 3). Consistent with this role in cardiomyocyte specification, loss of Wnt5b function results in a decrease in pacemaker cardiomyocytes and corresponding increase in the number of atrial cardiomyocytes. Conversely, overexpression of Wnt5b produces the opposite effect, with increased pacemaker cardiomyocytes and decreased atrial cardiomyocytes [98]. Interestingly, Wnt5b mutant embryos develop slower heart rates compared to wild-type, however inhibition of Wnt signaling by overexpressing Axin1 between 36 and 52 hpf results in an increase in heart rate, emphasizing that canonical Wnt signaling first helps establish the SAN earlier in cardiogenesis and then modulates pacemaker function later [93][98]. Consistently, while previous work did not investigate the differentiation of pacemaker cells specifically, the early requirement of Wtn5b is consistent with the previously identified role for Wnt signaling in promoting the differentiation of venous atrial cardiomyocytes [51]. Canonical Wnt signaling as a driver of SAN identity has also been shown in mouse and human embryonic stem cells, indicating its conserved role in pacemaker cardiomyocyte differentiation [99]. Thus, while investigation of the genetic networks driving SAN formation in zebrafish to this point is less extensive than mice, the current data support that a fundamentally conserved network of transcription factors are necessary to establish the proper number of pacemaker cardiomyocytes within the venous pole of the zebrafish heart (Figure 3).
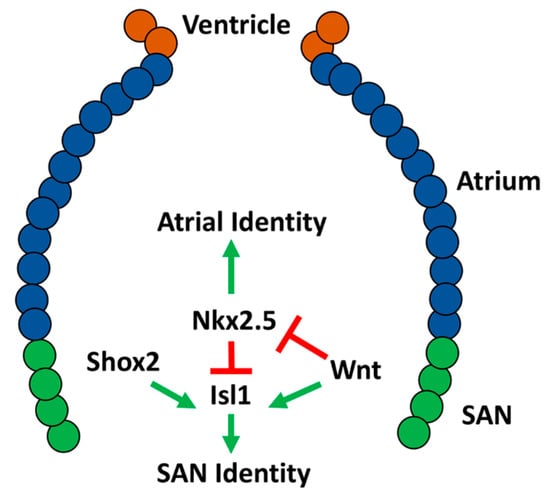
Figure 3. Molecular network shown to be responsible for differentiation of the sinoatrial node (SAN) in embryonic zebrafish.
References
- Al Turki, S.; Manickaraj, A.K.; Mercer, C.L.; Gerety, S.S.; Hitz, M.P.; Lindsay, S.; D’Alessandro, L.C.A.; Swaminathan, G.J.; Bentham, J.; Arndt, A.K.; et al. Rare variants in NR2F2 cause congenital heart defects in humans. Am. J. Hum. Genet. 2014, 94, 574–585.
- Nakamura, E.; Makita, Y.; Okamoto, T.; Nagaya, K.; Hayashi, T.; Sugimoto, M.; Manabe, H.; Taketazu, G.; Kajino, H.; Fujieda, K. 5.78 Mb terminal deletion of chromosome 15q in a girl, evaluation of NR2F2 as candidate gene for congenital heart defects. Eur. J. Med. Genet. 2011, 54, 354–356.
- Benson, D.W.; Silberbach, G.M.; Kavanaugh-McHugh, A.; Cottrill, C.; Zhang, Y.; Riggs, S.; Smalls, O.; Johnson, M.C.; Watson, M.S.; Seidman, J.G.; et al. Mutations in the cardiac transcription factor NKX2.5 affect diverse cardiac developmental pathways. J. Clin. Investig. 1999, 104, 1567–1573.
- Schott, J.J.; Benson, D.W.; Basson, C.T.; Pease, W.; Silberbach, G.M.; Moak, J.P.; Maron, B.J.; Seidman, C.E.; Seidman, J.G. Congenital heart disease caused by mutations in the transcription factor NKX2-5. Science 1998, 281, 108–111.
- Hoffman, J.I.E.; Kaplan, S. The incidence of congenital heart disease. J. Am. Coll. Cardiol. 2002, 39, 1890–1900.
- Loffredo, C.A. Epidemiology of cardiovascular malformations: Prevalence and risk factors. Am. J. Med. Genet. 2000, 97, 319–325.
- Van der Linde, D.; Konings, E.E.M.; Slager, M.A.; Witsenburg, M.; Helbing, W.A.; Takkenberg, J.J.M.; Roos-Hesselink, J.W. Birth prevalence of congenital heart disease worldwide. J. Am. Coll. Cardiol. 2011, 58, 2241–2247.
- Benjamin, E.J.; Virani, S.S.; Callaway, C.W.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Chiuve, S.E.; Cushman, M.; Delling, F.N.; Deo, R.; et al. Heart disease and stroke statistics-2018 update: A report from the American Heart Association. Circulation 2018, 137, E67–E492.
- Marelli, A.J.; Mackie, A.S.; Ionescu-Ittu, R.; Rahme, E.; Pilote, L. Congenital heart disease in the general population: Changing prevalence and age distribution. Circulation 2007, 115, 163–172.
- Bruneau, B.G.; Logan, M.; Davis, N.; Levi, T.; Tabin, C.J.; Seidman, J.G.; Seidman, C.E. Chamber-specific cardiac expression of Tbx5 and heart defects in Holt- Oram syndrome. Dev. Biol. 1999, 211, 100–108.
- John, R.M.; Kumar, S. Sinus node and atrial arrhythmias. Circulation 2016, 133, 1892–1900.
- Choudhury, M.; Boyett, M.R.; Morris, G.M. Biology of the sinus node and its disease. Arrhythmia Electrophysiol. Rev. 2015, 4, 28–34.
- Hoffmann, S.; Paone, C.; Sumer, S.A.; Diebold, S.; Weiss, B.; Roeth, R.; Clauss, S.; Klier, I.; Kääb, S.; Schulz, A.; et al. Functional characterization of rare variants in the SHOX2 gene identified in sinus node dysfunction and atrial fibrillation. Front. Genet. 2019, 10.
- Milanesi, R.; Baruscotti, M.; Gnecchi-Ruscone, T.; DiFrancesco, D. Familial sinus bradycardia associated with a mutation in the cardiac pacemaker channel. N. Engl. J. Med. 2006, 354, 151–157.
- Jhaveri, S.; Aziz, P.F.; Saarel, E. Expanding the electrical phenotype of NKX2-5 mutations: Ventricular tachycardia, atrial fibrillation, and complete heart block within one family. Hear. Case Rep. 2018, 4, 530–533.
- Vornanen, M.; Hassinen, M. Zebrafish heart as a model for human cardiac electrophysiology. Channels 2016, 10, 101–110.
- Giardoglou, P.; Beis, D. On zebrafish disease models and matters of the heart. Biomedicines 2019, 7, 15.
- Bakkers, J. Zebrafish as a model to study cardiac development and human cardiac disease. Cardiovasc. Res. 2011, 91, 279–288.
- Brown, D.; Samsa, L.; Qian, L.; Liu, J. Advances in the study of heart development and disease using zebrafish. J. Cardiovasc. Dev. Dis. 2016, 3, 13.
- Poon, K.L.; Brand, T. The zebrafish model system in cardiovascular research: A tiny fish with mighty prospects. Glob. Cardiol. Sci. Pract. 2013, 2013, 4.
- Ravens, U. Ionic basis of cardiac electrophysiology in zebrafish compared to human hearts. Prog. Biophys. Mol. Biol. 2018, 138, 38–44.
- Chi, N.C.; Shaw, R.M.; Jungblut, B.; Huisken, J.; Ferrer, T.; Arnaout, R.; Scott, I.; Beis, D.; Xiao, T.; Baier, H.; et al. Genetic and physiologic dissection of the vertebrate cardiac conduction system. PLoS Biol. 2008, 6, 1006–1019.
- Brandenburg, S.; Kohl, T.; Williams, G.S.B.; Gusev, K.; Wagner, E.; Rog-Zielinska, E.A.; Hebisch, E.; Dura, M.; Didié, M.; Gotthardt, M.; et al. Axial tubule junctions control rapid calcium signaling in atria. J. Clin. Investig. 2016, 126, 3999–4015.
- Smyrnias, I.; Mair, W.; Harzheim, D.; Walker, S.A.; Roderick, H.L.; Bootman, M.D. Comparison of the T-tubule system in adult rat ventricular and atrial myocytes, and its role in excitation-contraction coupling and inotropic stimulation. Cell Calcium 2010, 47, 210–223.
- Bloomekatz, J.; Galvez-Santisteban, M.; Chi, N.C. Myocardial plasticity: Cardiac development, regeneration and disease. Curr. Opin. Genet. Dev. 2016, 40, 120–130.
- Tessadori, F.; van Weerd, J.H.; Burkhard, S.B.; Verkerk, A.O.; de Pater, E.; Boukens, B.J.; Vink, A.; Christoffels, V.M.; Bakkers, J. Identification and functional characterization of cardiac pacemaker cells in zebrafish. PLoS ONE 2012, 7.
- Tabibiazar, R.; Wagner, R.A.; Liao, A.; Quertermous, T. Transcriptional profiling of the heart reveals chamber-specific gene expression patterns. Circ. Res. 2003, 93, 1193–1201.
- Ng, S.Y.; Wong, C.K.; Tsang, S.Y. Differential gene expressions in atrial and ventricular myocytes: Insights into the road of applying embryonic stem cell-derived cardiomyocytes for future therapies. Am. J. Physiol. Physiol. 2010, 299, C1234–C1249.
- Bootman, M.D.; Higazi, D.R.; Coombes, S.; Roderick, H.L. Calcium signalling during excitation-contraction coupling in mammalian atrial myocytes. J. Cell Sci. 2006, 119, 3915–3925.
- Nof, E.; Antzelevitch, C.; Glikson, M. The contribution of HCN4 to normal sinus node function in humans and animal models. Pacing Clin. Electrophysiol. 2010, 33, 100–106.
- Christoffels, V.M.; Smits, G.J.; Kispert, A.; Moorman, A.F.M. Development of the pacemaker tissues of the heart. Circ. Res. 2010, 106, 240–254.
- Stainier, D.Y.; Lee, R.K.; Fishman, M.C. Cardiovascular development in the zebrafish. I. Myocardial fate map and heart tube formation. Development 1993, 119, 31–40.
- Berdougo, E.; Coleman, H.; Lee, D.H.; Stainier, D.Y.R.; Yelon, D. Mutation of weak atrium/atrial myosin heavy chain disrupts atrial function and influences ventricular morphogenesis in zebrafish. Development 2003, 130, 6121–6129.
- Staudt, D.; Stainier, D. Uncovering the molecular and cellular mechanisms of heart development using the zebrafish. Annu. Rev. Genet. 2012, 46, 397–418.
- Keegan, B.R.; Meyer, D.; Yelon, D. Organization of cardiac chamber progenitors in the zebrafish blastula. Development 2004, 131, 3081–3091.
- Yelon, D.; Stainier, D.Y.R. Patterning during organogenesis: Genetic analysis of cardiac chamber formation. Semin. Cell Dev. Biol. 1999, 10, 93–98.
- Guerra, A.; Germano, R.F.V.; Stone, O.; Arnaout, R.; Guenther, S.; Ahuja, S.; Uribe, V.; Vanhollebeke, B.; Stainier, D.Y.R.; Reischauer, S. Distinct myocardial lineages break atrial symmetry during cardiogenesis in zebrafish. Elife 2018, 7.
- Tessari, A.; Pietrobon, M.; Notte, A.; Cifelli, G.; Gage, P.J.; Schneider, M.D.; Lembo, G.; Campione, M. Myocardial Pitx2 differentially regulates the left atrial identity and ventricular asymmetric remodeling programs. Circ. Res. 2008, 102, 813–822.
- Rohr, S.; Otten, C.; Abdelilah-Seyfried, S. Asymmetric involution of the myocardial field drives heart tube formation in zebrafish. Circ. Res. 2008, 102.
- Yelon, D.; Horne, S.A.; Stainier, D.Y.R. Restricted expression of cardiac myosin genes reveals regulated aspects of heart tube assembly in zebrafish. Dev. Biol. 1999, 214, 23–37.
- De Pater, E.; Clijsters, L.; Marques, S.R.; Lin, Y.F.; Garavito-Aguilar, Z.V.; Yelon, D.; Bakkers, J. Distinct phases of cardiomyocyte differentiation regulate growth of the zebrafish heart. Development 2009, 136, 1633–1641.
- Liu, J.; Stainier, D.Y.R. Zebrafish in the study of early cardiac development. Circ. Res. 2012, 110, 870–874.
- Hami, D.; Grimes, A.C.; Tsai, H.J.; Kirby, M.L. Zebrafish cardiac development requires a conserved secondary heart field. Development 2011, 138, 2389–2398.
- Cai, C.L.; Liang, X.; Shi, Y.; Chu, P.H.; Pfaff, S.L.; Chen, J.; Evans, S. Isl1 identifies a cardiac progenitor population that proliferates prior to differentiation and contributes a majority of cells to the heart. Dev. Cell 2003, 5, 877–889.
- Lazic, S.; Scott, I.C. Mef2cb regulates late myocardial cell addition from a second heart field-like population of progenitors in zebrafish. Dev. Biol. 2011, 354, 123–133.
- Hu, N.; Sedmera, D.; Yost, H.J.; Clark, E.B. Structure and function of the developing zebrafish heart. Anat. Rec. 2000, 260, 148–157.
- Gupta, V.; Poss, K.D. Clonally dominant cardiomyocytes direct heart morphogenesis. Nature 2012, 484, 479–484.
- Liu, J.; Bressan, M.; Hassel, D.; Huisken, J.; Staudt, D.; Kikuchi, K.; Poss, K.D.; Mikawa, T.; Stainier, D.Y.R. A dual role for ErbB2 signaling in cardiac trabeculation. Development 2010, 137, 3867–3875.
- Foglia, M.J.; Cao, J.; Tornini, V.A.; Poss, K.D. Multicolor mapping of the cardiomyocyte proliferation dynamics that construct the atrium. Development 2016, 143, 1688–1696.
- Thomas, N.A.; Koudijs, M.; van Eeden, F.J.M.; Joyner, A.L.; Yelon, D. Hedgehog signaling plays a cell-autonomous role in maximizing cardiac developmental potential. Development 2008, 135, 3789–3799.
- Dohn, T.E.; Waxman, J.S. Distinct phases of Wnt/β-catenin signaling direct cardiomyocyte formation in zebrafish. Dev. Biol. 2012, 361, 364–376.
- Ueno, S.; Weidinger, G.; Osugi, T.; Kohn, A.D.; Golob, J.L.; Pabon, L.; Reinecke, H.; Moon, R.T.; Murry, C.E. Biphasic role for Wnt/β-catenin signaling in cardiac specification in zebrafish and embryonic stem cells. Proc. Natl. Acad. Sci. USA 2007, 104, 9685–9690.
- Deshwar, A.R.; Chng, S.C.; Ho, L.; Reversade, B.; Scott, I.C. The Apelin receptor enhances Nodal/TGFβ signaling to ensure proper cardiac development. Elife 2016, 5.
- Scott, I.C.; Masri, B.; D’Amico, L.A.; Jin, S.W.; Jungblut, B.; Wehman, A.M.; Baier, H.; Audigier, Y.; Stainier, D.Y.R. The G Protein-Coupled receptor agtrl1b regulates early development of myocardial progenitors. Dev. Cell 2007, 12, 403–413.
- Stainier, D.Y.R.; Fishman, M.C. Patterning the zebrafish heart tube: Acquisition of anteroposterior polarity. Dev. Biol. 1992, 153, 91–101.
- Hochgreb, T.; Linhares, V.L.; Menezes, D.C.; Sampaio, A.C.; Yan, C.Y.I.; Cardoso, W.V.; Rosenthal, N.; Xavier-Neto, J. A caudorostral wave of RALDH2 conveys anteroposterior information to the cardiac field. Development 2003, 130, 5363–5374.
- Keegan, B.R.; Feldman, J.L.; Begemann, G.; Ingham, P.W.; Yelon, D. Retinoic acid signaling restricts the cardiac progenitor pool. Science 2005, 307, 247–249.
- Waxman, J.S.; Keegan, B.R.; Roberts, R.W.; Poss, K.D.; Yelon, D. Hoxb5b Acts Downstream of retinoic acid signaling in the forelimb field to restrict heart field potential in zebrafish. Dev. Cell 2008, 15, 923–934.
- Waxman, J.S.; Yelon, D. Increased hox activity mimics the teratogenic effects of excess retinoic acid signaling. Dev. Dyn. 2009, 238, 1207–1213.
- D’Aniello, E.; Rydeen, A.B.; Anderson, J.L.; Mandal, A.; Waxman, J.S. Depletion of retinoic acid receptors initiates a novel positive feedback mechanism that promotes teratogenic increases in retinoic acid. PLoS Genet. 2013, 9, 1003689.
- Marques, S.R.; Lee, Y.; Poss, K.D.; Yelon, D. Reiterative roles for FGF signaling in the establishment of size and proportion of the zebrafish heart. Dev. Biol. 2008, 321, 397–406.
- Marques, S.R.; Yelon, D. Differential requirement for BMP signaling in atrial and ventricular lineages establishes cardiac chamber proportionality. Dev. Biol. 2009, 328, 472–482.
- De Pater, E.; Ciampricotti, M.; Priller, F.; Veerkamp, J.; Strate, I.; Smith, K.; Lagendijk, A.K.; Schilling, T.F.; Herzog, W.; Abdelilah-Seyfried, S.; et al. Bmp signaling exerts opposite effects on cardiac differentiation. Circ. Res. 2012, 110, 578–587.
- Reischauer, S.; Stone, O.A.; Villasenor, A.; Chi, N.; Jin, S.W.; Martin, M.; Lee, M.T.; Fukuda, N.; Marass, M.; Witty, A.; et al. Cloche is a bHLH-PAS transcription factor that drives haemato-vascular specification. Nature 2016, 535, 294–298.
- Schoenebeck, J.J.; Keegan, B.R.; Yelon, D. Vessel and blood specification override cardiac potential in anterior mesoderm. Dev. Cell 2007, 13, 254–267.
- Zhou, Q.; Li, L.; Zhao, B.; Guan, K.-L. The hippo pathway in heart development, regeneration, and diseases. Circ. Res. 2015, 116, 1431–1447.
- Fukui, H.; Miyazaki, T.; Chow, R.W.Y.; Ishikawa, H.; Nakajima, H.; Vermot, J.; Mochizuki, N. Hippo signaling determines the number of venous pole cells that originate from the anterior lateral plate mesoderm in zebrafish. Elife 2018, 7.
- Witzel, H.R.; Cheedipudi, S.; Gao, R.; Stainier, D.Y.R.; Dobreva, G.D. Isl2b regulates anterior second heart field development in zebrafish. Sci. Rep. 2017, 7, 1–9.
- Pereira, F.A.; Yuhong, Q.; Zhou, G.; Tsai, M.J.; Tsai, S.Y. The orphan nuclear receptor COUP-TFII is required for angiogenesis and heart development. Genes Dev. 1999, 13, 1037–1049.
- Duong, T.B.; Ravisankar, P.; Song, Y.C.; Gafranek, J.T.; Rydeen, A.B.; Dohn, T.E.; Barske, L.A.; Crump, J.G.; Waxman, J.S. Nr2f1a balances atrial chamber and atrioventricular canal size via BMP signaling-independent and -dependent mechanisms. Dev. Biol. 2018, 434, 7–14.
- Barth, A.S.; Merk, S.; Arnoldi, E.; Zwermann, L.; Kloos, P.; Gebauer, M.; Steinmeyer, K.; Bleich, M.; Kääb, S.; Pfeufer, A.; et al. Functional profiling of human atrial and ventricular gene expression. Pflugers Arch. Eur. J. Physiol. 2005, 450, 201–208.
- Xin, M.; Small, E.M.; Van Rooij, E.; Qi, X.; Richardson, J.A.; Srivastava, D.; Nakagawa, O.; Olson, E.N. Essential roles of the bHLH transcription factor Hrt2 in repression of atrial gene expression and maintenance of postnatal cardiac function. Proc. Natl. Acad. Sci. USA 2007, 104, 7975–7980.
- Targoff, K.L.; Colombo, S.; George, V.; Schell, T.; Kim, S.H.; Solnica-Krezel, L.; Yelon, D. Nkx genes are essential for maintenance of ventricular identity. Development 2013, 140, 4203–4213.
- Pradhan, A.; Zeng, X.X.I.; Sidhwani, P.; Marques, S.R.; George, V.; Targoff, K.L.; Chi, N.C.; Yelon, D. FGF signaling enforces cardiac chamber identity in the developing ventricle. Development 2017, 144, 1328–1338.
- George, V.; Colombo, S.; Targoff, K.L. An early requirement for nkx2.5 ensures the first and second heart field ventricular identity and cardiac function into adulthood. Dev. Biol. 2015, 400, 10–22.
- Reifers, F.; Walsh, E.C.; Léger, S.; Stainier, D.Y.R.; Brand, M. Induction and differentiation of the zebrafish heart requires fibroblast growth factor 8 (fgf8/acerebellar). Development 2000, 127, 225–235.
- Zhang, R.; Han, P.; Yang, H.; Ouyang, K.; Lee, D.; Lin, Y.F.; Ocorr, K.; Kang, G.; Chen, J.; Stainier, D.Y.R.; et al. In vivo cardiac reprogramming contributes to zebrafish heart regeneration. Nature 2013, 498, 497–501.
- Wu, S.P.; Cheng, C.M.; Lanz, R.B.; Wang, T.; Respress, J.L.; Ather, S.; Chen, W.; Tsai, S.J.; Wehrens, X.H.T.; Tsai, M.J.; et al. Atrial identity is determined by a COUP-TFII regulatory network. Dev. Cell 2013, 25, 417–426.
- Arrenberg, A.B.; Stainier, Y.R.; Baier, H.; Huisken, J. Optogenetic control of cardiac function. Source Sci. 2010, 330, 971–974.
- Van Weerd, J.H.; Christoffels, V.M. The formation and function of the cardiac conduction system. Development 2016, 143, 197–210.
- Burkhard, S.; van Eif, V.; Garric, L.; Christoffels, V.; Bakkers, J. On the evolution of the cardiac pacemaker. J. Cardiovasc. Dev. Dis. 2017, 4, 4.
- Wiese, C.; Grieskamp, T.; Airik, R.; Mommersteeg, M.T.M.; Gardiwal, A.; De Gier-De Vries, C.; Schuster-Gossler, K.; Moorman, A.F.M.; Kispert, A.; Christoffels, V.M. Formation of the sinus node head and differentiation of sinus node myocardium are independently regulated by Tbx18 and Tbx3. Circ. Res. 2009, 104, 388–397.
- Hoogaars, W.M.H.; Engel, A.; Brons, J.F.; Verkerk, A.O.; De Lange, F.J.; Wong, L.Y.E.; Bakker, M.L.; Clout, D.E.; Wakker, V.; Barnett, P.; et al. Tbx3 controls the sinoatrial node gene program and imposes pacemaker function on the atria. Genes Dev. 2007, 21, 1098–1112.
- Espinoza-Lewis, R.A.; Liu, H.; Sun, C.; Chen, C.; Jiao, K.; Chen, Y.P. Ectopic expression of Nkx2.5 suppresses the formation of the sinoatrial node in mice. Dev. Biol. 2011, 356, 359–369.
- Nakashima, Y.; Yanez, D.A.; Touma, M.; Nakano, H.; Jaroszewicz, A.; Jordan, M.C.; Pellegrini, M.; Roos, K.P.; Nakano, A. Nkx2-5 suppresses the proliferation of atrial myocytes and conduction system. Circ. Res. 2014, 114, 1103–1113.
- Mommersteeg, M.T.M.; Hoogaars, W.M.H.; Prall, O.W.J.; De Gier-De Vries, C.; Wiese, C.; Clout, D.E.W.; Papaioannou, V.E.; Brown, N.A.; Harvey, R.P.; Moorman, A.F.M.; et al. Molecular pathway for the localized formation of the sinoatrial node. Circ. Res. 2007, 100, 354–362.
- Sun, Y.; Liang, X.; Najafi, N.; Cass, M.; Lin, L.; Cai, C.L.; Chen, J.; Evans, S.M. Islet 1 is expressed in distinct cardiovascular lineages, including pacemaker and coronary vascular cells. Dev. Biol. 2007, 304, 286–296.
- Weinberger, F.; Mehrkens, D.; Friedrich, F.W.; Stubbendorff, M.; Hua, X.; Müller, J.C.; Schrepfer, S.; Evans, S.M.; Carrier, L.; Eschenhagen, T. Localization of islet-1-positive cells in the healthy and infarcted adult murine heart. Circ. Res. 2012, 110, 1303–1310.
- Blaschke, R.J.; Hahurij, N.D.; Kuijper, S.; Just, S.; Wisse, L.J.; Deissler, K.; Maxelon, T.; Anastassiadis, K.; Spitzer, J.; Hardt, S.E.; et al. Targeted mutation reveals essential functions of the homeodomain transcription factor Shox2 in sinoatrial and pacemaking development. Circulation 2007, 115, 1830–1838.
- Liang, X.; Zhang, Q.; Cattaneo, P.; Zhuang, S.; Gong, X.; Spann, N.J.; Jiang, C.; Cao, X.; Zhao, X.; Zhang, X.; et al. Transcription factor ISL1 is essential for pacemaker development and function. J. Clin. Investig. 2015, 125, 3256–3268.
- Espinoza-Lewis, R.A.; Yu, L.; He, F.; Liu, H.; Tang, R.; Shi, J.; Sun, X.; Martin, J.F.; Wang, D.; Yang, J.; et al. Shox2 is essential for the differentiation of cardiac pacemaker cells by repressing Nkx2-5. Dev. Biol. 2009, 327, 376–385.
- Liu, H.; Chen, C.H.; Espinoza-Lewis, R.A.; Jiao, Z.; Sheu, I.; Hu, X.; Lin, M.; Zhang, Y.; Chen, Y.P. Functional redundancy between human SHOX and mouse Shox2 genes in the regulation of sinoatrial node formation and pacemaking function. J. Biol. Chem. 2011, 286, 17029–17038.
- Burkhard, S.B.; Bakkers, J. Spatially resolved RNA-sequencing of the embryonic heart identifies a role for Wnt/β-catenin signaling in autonomic control of heart rate. Elife 2018, 7.
- Hoffmann, S.; Berger, I.M.; Glaser, A.; Bacon, C.; Li, L.; Gretz, N.; Steinbeisser, H.; Rottbauer, W.; Just, S.; Rappold, G. Islet1 is a direct transcriptional target of the homeodomain transcription factor Shox2 and rescues the Shox2-mediated bradycardia. Basic Res. Cardiol. 2013, 108, 339.
- Colombo, S.; De Sena-Tomaś, C.; George, V.; Werdich, A.A.; Kapur, S.; Macrae, C.A.; Targoff, K.L. Nkx genes establish second heart field cardiomyocyte progenitors at the arterial pole and pattern the venous pole through isl1 repression. Development 2018, 145.
- Bressan, M.; Liu, G.; Mikawa, T. Early mesodermal cues assign avian cardiac pacemaker fate potential in a tertiary heart field. Science 2013, 340, 744–748.
- Mommersteeg, M.T.M.; Domínguez, J.N.; Wiese, C.; Norden, J.; De Gier-De Vries, C.; Burch, J.B.E.; Kispert, A.; Brown, N.A.; Moorman, A.F.M.; Christoffels, V.M. The sinus venosus progenitors separate and diversify from the first and second heart fields early in development. Cardiovasc. Res. 2010, 87, 92–101.
- Ren, J.; Han, P.; Ma, X.; Farah, E.N.; Bloomekatz, J.; Zeng, X.X.I.; Zhang, R.; Swim, M.M.; Witty, A.D.; Knight, H.G.; et al. Canonical Wnt5b Signaling Directs Outlying Nkx2.5+ Mesoderm into Pacemaker Cardiomyocytes. Dev. Cell 2019, 50, 729–743.
- Liang, W.; Han, P.; Kim, E.H.; Mak, J.; Zhang, R.; Torrente, A.G.; Goldhaber, J.I.; Marbán, E.; Cho, H.C. Canonical Wnt signaling promotes pacemaker cell specification of cardiac mesodermal cells derived from mouse and human embryonic stem cells. Stem Cells 2020, 38, 352–368.





