
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Valentina Daponte | + 2613 word(s) | 2613 | 2021-02-13 03:12:16 | | | |
| 2 | Vicky Zhou | + 1 word(s) | 2614 | 2021-02-22 10:55:42 | | |
Video Upload Options
Appendage regeneration in vertebrates means the ability to regenerate amputated or injured tissues and organs, which is a fascinating property shared by several invertebrates and, interestingly, some vertebrates.
1. Introduction
Regeneration is the capacity of an organism to regrow a part of the body after injury. Many species of anamniote vertebrates are capable of spectacular accomplishment of regeneration. Among them, the best known is salamander, which is able to restore limb, tail, eye, jaw, and heart [1]. Regeneration is also present in various members of the class of actinopterygians [2]. The adult zebrafish (Danio rerio), a small teleost, exhibits remarkable regeneration of fins, central nervous system structures, or entire organs including heart, pancreas, liver, and kidney [3]. The ability to activate the regeneration morphogenetic program is almost completely lost mainly in terrestrial animals, such as amniotes, in which nearly all structures are unable to regenerate, with some exceptions such as the regeneration of digit tips in children [4] and young mice [5], ear pinna in the spiny mouse [6], or the antler regeneration in deer [7]. Lizards are the only terrestrial amniotes that can regenerate a structure long and complex as the tail [8].
The distribution of the regenerative capacity along the phylogenetic tree is haphazard and a common pattern is difficult to identify (Figure 1). Regeneration appeared very early during evolution, likely coinciding with the origin of multicellularity [9]. The similarities between regeneration and development may suggest that the former originated as an epiphenomenon of an ontogenetic program that an organism accesses and employs when a structure is lost. Based on these observations, regeneration could be an ancestral property, lost due to higher energetic maintenance costs (for a detailed review, see [10]).
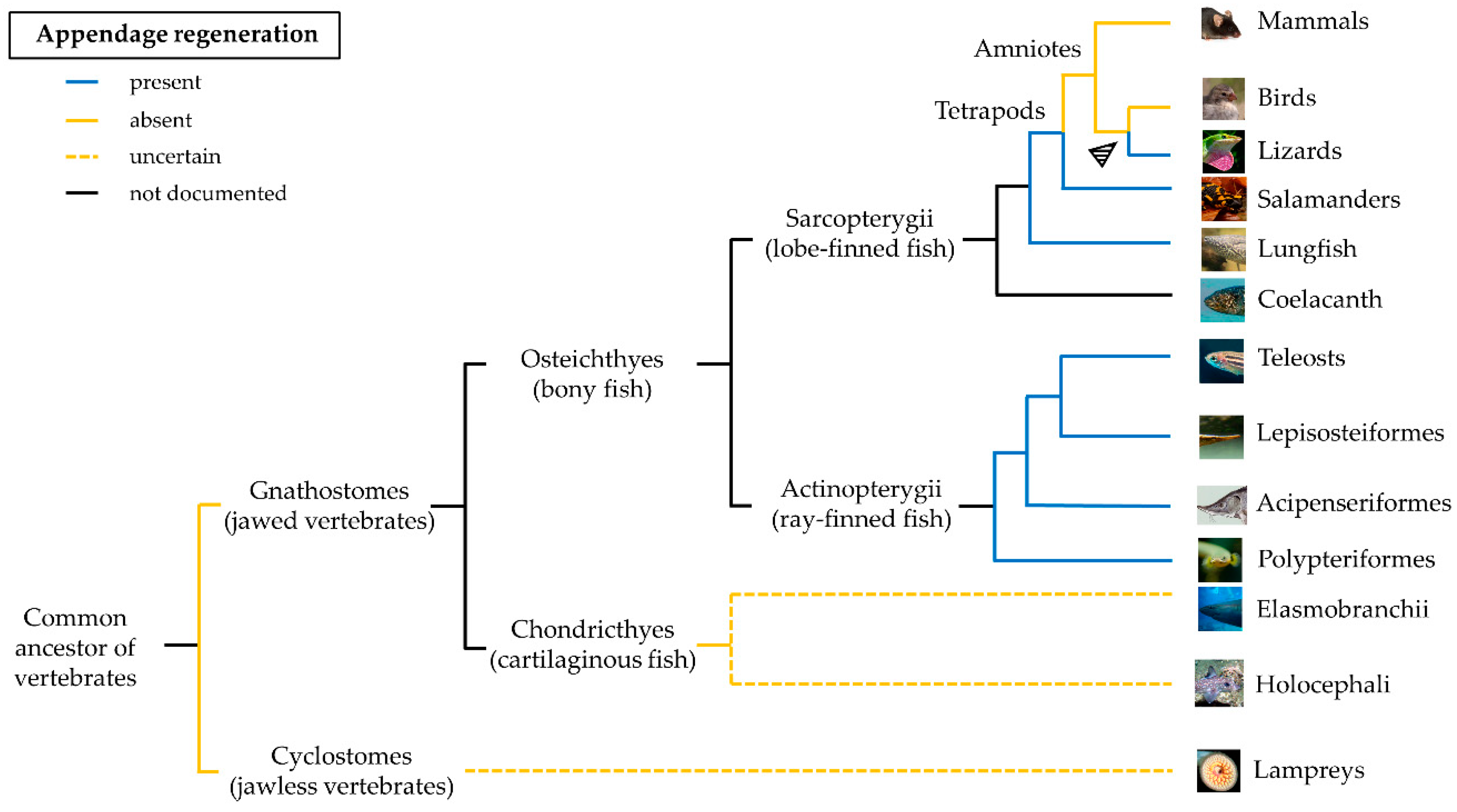
In salamanders, several genes are associated with the regeneration ability, one of them is the one coding for the glycerophosphatidyl (GPI)-anchored cell surface Prod1, which mediates positional memory through a proximal-distal gradient of cell adhesion [11]. Despite a protein with similar functions being recently identified in the regenerating tail of the lizard Gekko japonicus [12], the Prod1 gene has not been found in any other species [13].
The ability to regenerate paired and unpaired fins was documented in many ray-finned fish (Actinopterygii). Teleosts possess a wide range of regenerative abilities, from complete absence of regeneration in some members of the family Blennidae [14] to a high regenerative ability of zebrafish. However, this trait is limited to the bony fin rays and does not include the proximal structures such as musculature and endoskeleton. The only exception are Polypteriformes, one of the basal lineages of Actinopterygii [15][16]. The strong regenerative potential found in this taxon suggests that regeneration in this case has emerged early during evolution, long before the appearance of Teleostei. In terrestrial amniotes, the evolution of a strong adaptive immune system could have favored a more defensive role of macrophages during the early phases of regeneration to the expenses of their ability of tissue remodeling [17]. The extreme “non-lethal predation” of lizards’ tails in the wild could instead explain the presence of a regenerative program in these amniotes [18].[1][2][3][4][5][6][7][8][9][10]
Many factors have contributed to the loss of regeneration in the evolution, and further phylogenetic and comparative studies are necessary to unravel the mechanisms regulating the process. The investigation of the regeneration of different organs, from the simple tail to the more complex limb in different organisms, represents a powerful tool to shed light on the process.
2. Common Morphological Aspects of Early Regenerative Response
Lizard tail, salamander limb, and zebrafish fin regeneration are examples of epimorphosis. The term epimorphic regeneration, coined by Thomas H. Morgan in 1901 [19], describes the restoration of a part of the organism without remodeling, characterized by the formation of the blastema. This mass of undifferentiated cells at the wound site mediates tissue differentiation [20][21]. Epimorphosis is characterized by a high degree of cellular differentiation and stands in contrast with morphallaxis, usually observed in hydras, in which blastema is not formed and the majority of regeneration is taking place by reorganization of the remaining parts of the body [20][21].
Lizard, salamander, and zebrafish share several common morphological aspects of epimorphic regeneration that distinguish this process from the limited regeneration in mammals. Some differences were however identified. For example, lizards, unlike salamanders, are unable to regenerate amputated limbs. Also, after autotomy or amputation, lizard tail undergoes an “imperfect regeneration” in which the bony vertebral column is replaced with a hollow cone of cartilage and the fracture planes, that are structures evolved to allow autotomy at precise locations, are not regenerated [22]. On the contrary, salamanders and zebrafish can restore the original appendage. Lastly, while lizard and salamander appendage regeneration occur following cartilage formation, zebrafish caudal fin rays are directly generated as bone.
The regenerative process begins immediately after an appendage loss and consists of three phases: wound healing, blastema formation and regenerative outgrowth (Figure 2).
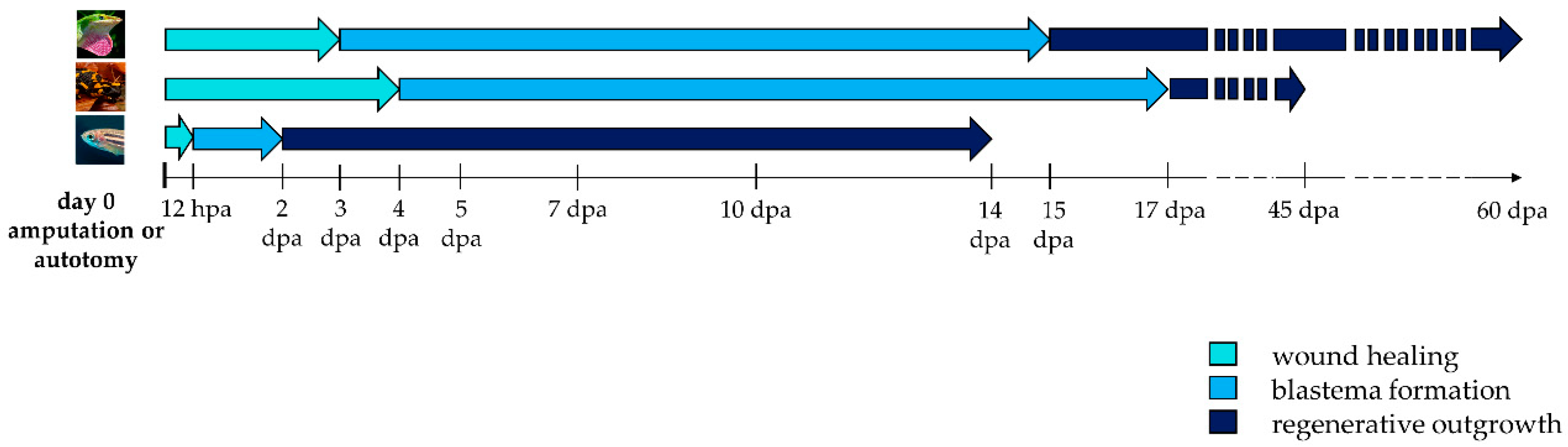
Figure 2. Timeline activation of wound healing, blastema formation and regenerative outgrowth during appendage regeneration in vertebrates. Arrows indicate the duration of each regenerative step in lizard, salamander, and zebrafish. Hpa: hours post-amputation; dpa: days post-amputation.
Initially, after autotomy or amputation, a blood clot forms. Lizard’s tail stump has evolved the presence of vascular sphincters located proximal to fracture planes that contract to avoid excessive bleeding and to facilitate hemostasis [23]. Within a few hours, epithelial cells begin to migrate to cover the wound, forming an epithelial layer called the wound epidermis. The integrity of the wound epidermis is essential to accomplish regeneration in lizard and salamander. Impeding its formation by excision or suture prevents the appendage to regenerate [24][25]. Repetitive amputations seem to trigger a persistent wound healing response, and result in a decline in regenerative fidelity in lizard and salamander [26][27]. In the newt Notophthalmus viridescens, repeated amputation of the limb can lead to severe morphological abnormalities [28]. To the contrary, in zebrafish, repeated amputations do not perturb regeneration and instead lead to an increase in the dermal bone thickness [29].
After several days, in salamander and zebrafish, the thick wound epidermis forms a specialized structure, the apical epidermal cap (AEC) that acts as a layer of signaling cells coordinating the regenerative process [23][30]. For this reason, AEC needs to be continuously in contact, and to communicate with, the underneath mesenchymal cells. Interposition of dermis or the formation of basement membrane between the two structures inhibits blastema formation and regeneration [31]. This property suggests that diffusible signaling factors are produced in the AEC and signal to the underlying mesenchyme. In lizards, this thick epithelium has a different morphology, owing a discontinuous basement membrane, and is often referred to as an apical epidermal peg (AEP) [32].
As quickly as the wound epidermis thickens, the tissues below undergo degeneration and histolysis. The dynamic remodeling of extracellular matrix (ECM) plays a crucial role in this process. The secretion of acid hydrolases and proteinases of the matrix metalloproteinases (MMPs) family at the level of the basal layer of the wound epidermis helps degrading ECM components, such as collagens and laminins. This process is likely facilitating cell migration and cell communication by disturbing basement membrane re-assembly [33][34][35]. MMPs also favor the elimination of cellular residues and debris generated by tissue destruction and by the bactericidal activity of neutrophils and macrophages [31]. In lizard and salamander, the degradation involves primarily bone and muscle [36][37]. In zebrafish, in which the caudal fin is mainly composed of bone, the establishment of a cryoinjury-mediated tissue damage triggers a similar osteolytic process [38]. The degeneration of stump tissues, or “histolysis”, favors the release of progenitor cells that will contribute to the formation of blastema. Comparative anatomy of regenerating appendages in lizard, salamander, and zebrafish is depicted in Figure 3.
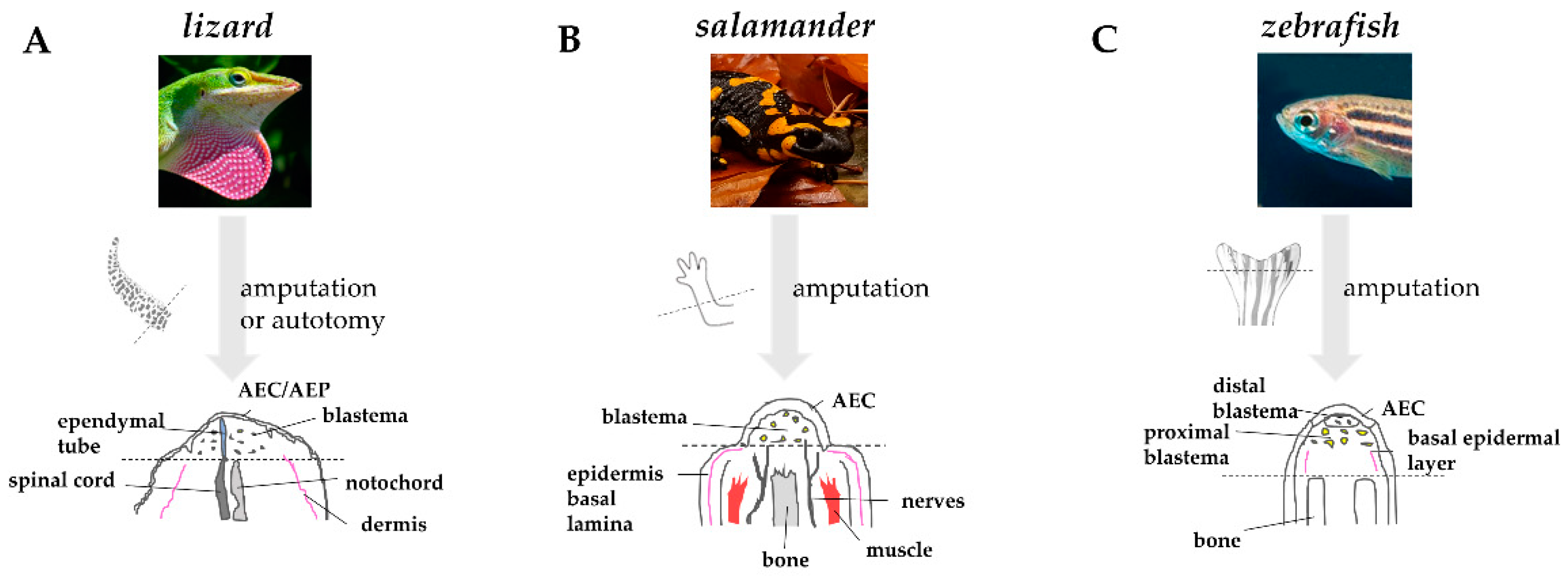
Figure 3. Comparative anatomy of regenerating appendages in lizard, salamander, and zebrafish. (A) Amputation or autotomy of lizard tail induces the formation of a cone-like shaped blastema, covered by a thick wound epidermis that will form the apical epidermal peg(AEP), whose correspondence to the apical epidermal cap (AEC) found in amphibians and fish is still uncertain. During blastema formation, likely in response to AEP signals, the central canal of the original spinal cord, the ependymal tube, elongates and infiltrates the proliferating tail blastema. (B) After amputation of salamander limb, the wound epidermis quickly covers the stump. Within days, the wound epithelium becomes innervated, thickens, and becomes a specialized AEC. The AEC induces dedifferentiation in the underlying stump tissue and attracts cells, which accumulate below the AEC to form the blastema. Modified from Payzin-Dogru and Whited, 2018. (C) After zebrafish fin amputation, epithelial cells migrate to cover the wound, forming the AEC. Under the AEC, stump tissues dedifferentiate to form the blastema. Within 24 h, blastemal cells segregate into two compartments: the distal blastema, populated by slowly proliferating cells, and the proximal blastema, in which cells rapidly proliferate and differentiate to replace the amputated tissue. Dotted lines indicate the site of amputation/autotomy.
3. Immune System and Regeneration
The strong, scar-free regenerative potential in selected vertebrate species has raised the question whether differences in immune systems can affect the outcome of tissue regeneration. One of the indications of a negative effect of the immune system on regeneration was described in anuran amphibians which lose their robust regeneration ability after metamorphosis when the immune response is enhanced [29].
Lizard, salamander, and zebrafish heavily rely on an efficient innate immunity, consisting of diverse components such as antimicrobial peptides, neutrophils, macrophages, and the complement system [39][40][41][42]. Their adaptive immunity, however, differs from that of mammals by missing lymphoid organs such as Peyer’s patches and lymph nodes [39][43][44].
Lizards, of the three species the closest related to mammals, share similarities with them in terms of generation of antibody diversity, but produce slower and less robust adaptive humoral responses [39]. Their B cells are characterized by phagocytic activity, a feature shared with salamanders and still under debate for zebrafish [43][45][46], and act as a component of innate immunity [39]. The strong innate immune system seen in this group of vertebrates could be sufficient to protect from microbial infections present in the first stages of life, making efficient adaptive immunity non-essential.
Salamanders are generally considered “immunodeficient” since they have a weak cell-mediated immune response, and their humoral immunity is based only on IgM production and is apparently amnestic [40]. Moreover, they have a restricted MHC class II diversity, likely resulting in a poor T-helper stimulation and a low cytokine synthesis [47] that could in part explain the low inflammatory response seen during limb regeneration.
Zebrafish develop their adaptive immunity later, at the end of their embryo phase [48]. Their T-cell responses are negatively affected by low body temperature [49]. RAG1 protein, essential for antibody and T-cell receptor V(D)J recombination, is conserved in zebrafish. However, homozygous rag1 mutant fish, lacking functional T and B lymphocytes, reach adulthood and are fertile [50][51], suggesting that these cells are likely not necessary for an efficient immune response.
An ultrastructural study of the regenerating tail of the lizard Podarcis muralis showed that repeated amputations or cauterization trigger an increase in the number of immune cells such as granulocytes, macrophages, and lymphocytes within the blastema and led to the deposition of fibrinoid material with scarring [26]. The study supports the hypothesis that the evolution of the immune system may be associated to the reduced regeneration ability in lizards, and in amniotes in general. Further studies are needed to clarify how immune cells and mesenchymal cells of the blastema communicate and interact during regeneration.
4. Conclusions
Mammals, humans included, are incapable of regenerating amputated or lost limbs. Damage or injury can be sometimes life threatening and is reasonable to wonder how we might be able to enhance our own regenerative potential. For this purpose, the understanding of cellular and molecular mechanisms orchestrating regeneration is crucial. In the past years, the field of regenerative biology has seen considerable progress, particularly thanks to the study of organisms with a high regenerative potential. Although morphological mechanisms and cell contributors of regeneration can vary widely even among close species, the comparison of the three vertebrate models here discussed allows to identify an overall common strategy to successfully restore a lost appendage. So, what makes this possible? As it seems, a combination of multiple factors. First, the formation of a thick wound epidermis/AEC that supports the regenerative process without scarring. The formation of a blastema, which provides the source of differentiated cells that will restore the lost tissues is the key process present in vertebrates with regenerative ability. Proper activation of specific molecular pathways has been proven necessary for successful regeneration. The data collected on the three described models unequivocally points to the role of Wnt/β-catenin and FGFs as master regulators of the process. The expression of these key molecules is not only necessary for regeneration to occur but is also able to promote it in organs with limited regenerative potential, such as lizard limb. Blastema growth is also strictly related to the level of immunosurveillance. Lizard tail, salamander limb and zebrafish caudal fin can be considered immuno-privileged organs, in which inflammation triggered by injury equilibrates with the presence of healing, anti-inflammatory macrophages.
Each model presents advantages and limitations. Salamanders are considered the master regenerators being the only vertebrate able to regenerate a full limb. Since its discovery in the 18th century , the regenerating limb system was used to perform fundamental experiments to delineate the basic properties of regeneration. However, many aspects of the urodelian life cycle are at odds with those of higher vertebrates. The axolotl, the most used salamander for regenerative studies, is neotenic and retains all its juvenile features even when it reaches adulthood. Lizards, on the other hand, follow a similar development as mammals and therefore are more appealing from a developmental point of view. However, as amniotes, their regenerative ability is restricted to the complex structure of the tail, and their lost appendages undergo an imperfect regeneration.
Given the simplicity of using powerful genetic tools, zebrafish has quickly emerged as a model of choice to study regeneration. Zebrafish present several advantages, such as the external fertilization and the transparency of embryos that makes them particularly appealing for forward genetic approaches allowing to investigate regeneration in health and disease.
Our knowledge of regenerative mechanisms is strictly related to the tools we can use. Thus, the possibility to apply more advanced genetic tools also on powerful regenerators such as salamander and lizard could represent a key step forward. The recent reports of Iberian ribbed newt (Pleurodeles waltl) and axolotl (Ambystoma mexicanum) complete genomes, together with the recent generation of a CRISPR/Cas9 Anolis lizard, will hopefully provide new tools to increase the use of these organisms to find the answer to regenerative questions still unsolved.
No unique model for regeneration exists, and the comparative study of vertebrates with high regenerative ability is fundamental to achieve significant insights on what are the factors driving regeneration, aiming to translate them into the field of regenerative medicine. Indeed, the field of regenerative biology has made an enormous step forward in the past decade, taking advance of imaging, genomics, and genome editing to identify key cell types and molecules involved in the regeneration of many model organisms. Yet, it can be difficult to foresee when and how findings from these studies will really advance regenerative medicine. The identification of genes modulating the origins and fates of blastemal progenitor cells will likely be the key to achieve appendage regeneration, and it will provide novel targets for gene manipulation in mammals.
References
- Joven, A.; Elewa, A.; Simon, A. Model systems for regeneration: Salamanders. Development 2019, 146.
- Darnet, S.; Dragalzew, A.C.; Amaral, D.B.; Sousa, J.F.; Thompson, A.W.; Cass, A.N.; Lorena, J.; Pires, E.S.; Costa, C.M.; Sousa, M.P.; et al. Deep evolutionary origin of limb and fin regeneration. Proc. Natl. Acad. Sci. USA 2019, 116, 15106–15115.
- Gemberling, M.; Bailey, T.J.; Hyde, D.R.; Poss, K.D. The zebrafish as a model for complex tissue regeneration. Trends Genet. 2013, 29, 611–620.
- Jafari, P.; Muller, C.; Grognuz, A.; Applegate, L.A.; Raffoul, W.; di Summa, P.G.; Durand, S. First Insights into Human Fingertip Regeneration by Echo-Doppler Imaging and Wound Microenvironment Assessment. Int. J. Mol. Sci. 2017, 18, 1054.
- Seifert, A.W.; Muneoka, K. The blastema and epimorphic regeneration in mammals. Dev. Biol. 2018, 433, 190–199.
- Matias Santos, D.; Rita, A.M.; Casanellas, I.; Brito Ova, A.; Araújo, I.M.; Power, D.; Tiscornia, G. Ear wound regeneration in the African spiny mouse Acomys cahirinus. Regeneration 2016, 3, 52–61.
- Kierdorf, U.; Kierdorf, H.; Szuwart, T. Deer antler regeneration: Cells, concepts, and controversies. J. Morphol. 2007, 268, 726–738.
- Jacyniak, K.; McDonald, R.P.; Vickaryous, M.K. Tail regeneration and other phenomena of wound healing and tissue restoration in lizards. J. Exp. Biol. 2017, 220, 2858–2869.
- Bely, A.E.; Nyberg, K.G. Evolution of animal regeneration: Re-emergence of a field. Trends Ecol. Evol. 2010, 25, 161–170.
- Maginnis, T.L. The costs of autotomy and regeneration in animals: A review and framework for future research. Behav. Ecol. 2006, 17, 857–872.
- Da Silva, S.M.; Gates, P.B.; Brockes, J.P. The Newt Ortholog of CD59 Is Implicated in Proximodistal Identity during Amphibian Limb Regeneration. Dev. Cell 2002, 3, 547–555.
- Wang, Y.; Wang, R.; Jiang, S.; Zhou, W.; Liu, Y.; Wang, Y.; Gu, Q.; Gu, Y.; Dong, Y.; Liu, M.; et al. Gecko CD59 is implicated in proximodistal identity during tail regeneration. PLoS ONE 2011, 6, e17878.
- Kumar, A.; Gates, P.B.; Brockes, J.P. Positional identity of adult stem cells in salamander limb regeneration. C R Biol. 2007, 330, 485–490.
- Wagner, G.P.; Misof, B.Y. Evolutionary modification of regenerative capability in vertebrates: A comparative study on teleost pectoral fin regeneration. J. Exp. Zool. 1992, 261, 62–78.
- Cuervo, R.; Hernández-Martínez, R.; Chimal-Monroy, J.; Merchant-Larios, H.; Covarrubias, L. Full regeneration of the tribasal Polypterus fin. Proc. Natl. Acad. Sci. USA 2012, 109, 3838–3843.
- Noack, K.; Zardoya, R.; Meyer, A. The complete mitochondrial DNA sequence of the bichir (Polypterus ornatipinnis), a basal ray-finned fish: Ancient establishment of the consensus vertebrate gene order. Genetics 1996, 144, 1165–1180.
- Mescher, A.L. Macrophages and fibroblasts during inflammation and tissue repair in models of organ regeneration. Regeneration (Oxf) 2017, 4, 39–53.
- Slack, J.M. Animal regeneration: Ancestral character or evolutionary novelty? EMBO Rep. 2017, 18, 1497–1508.
- Morgan, T.H. Regeneration; Macmillan: New York, NY, USA, 1901.
- Agata, K.; Saito, Y.; Nakajima, E. Unifying principles of regeneration I: Epimorphosis versus morphallaxis. Dev. Growth Differ. 2007, 49, 73–78.
- Carlson, B.M. Chapter 1—An Introduction to Regeneration. In Principles of Regenerative Biology; Carlson, B.M., Ed.; Academic Press: Burlington, VT, USA, 2007; pp. 1–29.
- Fisher, R.E.; Geiger, L.A.; Stroik, L.K.; Hutchins, E.D.; George, R.M.; Denardo, D.F.; Kusumi, K.; Rawls, J.A.; Wilson-Rawls, J. A histological comparison of the original and regenerated tail in the green anole, Anolis carolinensis. Anat. Rec. 2012, 295, 1609–1619.
- Londono, R.; Sun, A.X.; Tuan, R.S.; Lozito, T.P. Tissue Repair and Epimorphic Regeneration: An Overview. Curr. Pathobiol. Rep. 2018, 6, 61–69.
- Thornton, C.S. The effect of apical cap removal on limb regeneration in Amblystoma larvae. J. Exp. Zool. 1957, 134, 357–381.
- Mescher, A.L. Effects on adult newt limb regeneration of partial and complete skin flaps over the amputation surface. J. Exp. Zool. 1976, 195, 117–128.
- Alibardi, L. Tail regeneration reduction in lizards after repetitive amputation or cauterization reflects an increase of immune cells in blastemas. Zool. Res. 2018, 39, 413–423.
- Bryant, D.M.; Sousounis, K.; Payzin-Dogru, D.; Bryant, S.; Sandoval, A.G.W.; Martinez Fernandez, J.; Mariano, R.; Oshiro, R.; Wong, A.Y.; Leigh, N.D.; et al. Identification of regenerative roadblocks via repeat deployment of limb regeneration in axolotls. NPJ Regen. Med. 2017, 2, 30.
- Dearlove, G.E.; Dresden, M.H. Regenerative abnormalities in Notophthalmus viridescens induced by repeated amputations. J. Exp. Zool. 1976, 196, 251–262.
- Azevedo, A.S.; Grotek, B.; Jacinto, A.; Weidinger, G.; Saúde, L. The Regenerative Capacity of the Zebrafish Caudal Fin Is Not Affected by Repeated Amputations. PLoS ONE 2011, 6, e22820.
- Pfefferli, C.; Jazwinska, A. The art of fin regeneration in zebrafish. Regeneration 2015, 2, 72–83.
- Stocum, D.L. Mechanisms of urodele limb regeneration. Regeneration 2017, 4, 159–200.
- Alibardi, L. Review: Biological and Molecular Differences between Tail Regeneration and Limb Scarring in Lizard: An Inspiring Model Addressing Limb Regeneration in Amniotes. J. Exp. Zool. B Mol. Dev. Evol. 2017, 328, 493–514.
- Bai, S.; Thummel, R.; Godwin, A.R.; Nagase, H.; Itoh, Y.; Li, L.; Evans, R.; McDermott, J.; Seiki, M.; Sarras, M.P., Jr. Matrix metalloproteinase expression and function during fin regeneration in zebrafish: Analysis of MT1-MMP, MMP2 and TIMP2. Matrix Biol. 2005, 24, 247–260.
- Delorme, S.L.; Lungu, I.M.; Vickaryous, M.K. Scar-free wound healing and regeneration following tail loss in the leopard gecko, Eublepharis macularius. Anat. Rec. 2012, 295, 1575–1595.
- Stocum, D.L. Chapter 8—Regeneration of Appendages. In Regenerative Biology and Medicine, 2nd ed.; Stocum, D.L., Ed.; Academic Press: San Diego, CA, USA, 2012; pp. 183–226.
- Thornton, C.S. The histogenesis of the regenerating fore limb of larval Amblystoma after exarticulation of the humerus. J. Morphol. 1938, 62, 219–241.
- Campbell, L.J.; Crews, C.M. Molecular and Cellular Basis of Regeneration and Tissue Repair. Cell. Mol. Life Sci. 2007, 65, 73.
- Chassot, B.; Pury, D.; Jazwinska, A. Zebrafish fin regeneration after cryoinjury-induced tissue damage. Biol. Open 2016, 5, 819–828.
- Zimmerman, L.M.; Vogel, L.A.; Bowden, R.M. Understanding the vertebrate immune system: Insights from the reptilian perspective. J. Exp. Biol. 2010, 213, 661–671.
- Harty, M.; Neff, A.W.; King, M.W.; Mescher, A.L. Regeneration or scarring: An immunologic perspective. Dev. Dyn. 2003, 226, 268–279.
- Rauta, P.R.; Nayak, B.; Das, S. Immune system and immune responses in fish and their role in comparative immunity study: A model for higher organisms. Immunol. Lett. 2012, 148, 23–33.
- Alibardi, L. Hyaluronic acid in the tail and limb of amphibians and lizards recreates permissive embryonic conditions for regeneration due to its hygroscopic and immunosuppressive properties. J. Exp. Zool. B Mol. Dev. Evol. 2017, 328, 760–771.
- Parra, D.; Takizawa, F.; Sunyer, J.O. Evolution of B cell immunity. Annu. Rev. Anim. Biosci. 2013, 1, 65–97.
- Renshaw, S.A.; Trede, N.S. A model 450 million years in the making: Zebrafish and vertebrate immunity. Dis. Model. Mech. 2012, 5, 38–47.
- Li, J.; Barreda, D.R.; Zhang, Y.A.; Boshra, H.; Gelman, A.E.; Lapatra, S.; Tort, L.; Sunyer, J.O. B lymphocytes from early vertebrates have potent phagocytic and microbicidal abilities. Nat. Immunol. 2006, 7, 1116–1124.
- Page, D.M.; Wittamer, V.; Bertrand, J.Y.; Lewis, K.L.; Pratt, D.N.; Delgado, N.; Schale, S.E.; McGue, C.; Jacobsen, B.H.; Doty, A.; et al. An evolutionarily conserved program of B-cell development and activation in zebrafish. Blood 2013, 122, e1–e11.
- Tournefier, A.; Laurens, V.; Chapusot, C.; Ducoroy, P.; Padros, M.R.; Salvadori, F.; Sammut, B. Structure of MHC class I and class II cDNAs and possible immunodeficiency linked to class II expression in the Mexican axolotl. Immunol. Rev. 1998, 166, 259–277.
- Lam, S.H.; Chua, H.L.; Gong, Z.; Lam, T.J.; Sin, Y.M. Development and maturation of the immune system in zebrafish, Danio rerio: A gene expression profiling, in situ hybridization and immunological study. Dev. Comp. Immunol. 2004, 28, 9–28.
- Trede, N.S.; Langenau, D.M.; Traver, D.; Look, A.T.; Zon, L.I. The Use of Zebrafish to Understand Immunity. Immunity 2004, 20, 367–379.
- Wienholds, E.; Schulte-Merker, S.; Walderich, B.; Plasterk, R.H. Target-selected inactivation of the zebrafish rag1 gene. Science 2002, 297, 99–102.
- Petrie-Hanson, L.; Hohn, C.; Hanson, L. Characterization of rag1 mutant zebrafish leukocytes. BMC Immunol. 2009, 10, 8.





