
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Patrick Mester | + 1517 word(s) | 1517 | 2021-02-10 10:15:13 | | | |
| 2 | Vicky Zhou | Meta information modification | 1517 | 2021-02-16 16:09:09 | | |
Video Upload Options
Antimicrobial Resistance of VBNC Bacteria means the bacteria that enter the viable but non-culturable (VBNC) state, either through unfavorable environmental conditions or through potentially lethal stress, lose their ability to grow on standard enrichment media, but show a drastically increased tolerance against antimicrobials including antibiotics.
1. Introduction
Humanity′s history is a continuous battle between us and microbial pathogens and for the most part, we were on the losing side with bacterial and viral infections being among the major causes of morbidity and mortality worldwide. Thanks to the development and improvement of sanitation, hygiene practices, and especially the discovery of antibiotics and vaccinations since the early 20th century, deaths from infectious diseases have declined markedly which can be considered one of the biggest success stories in human history [1]. However, although the burden of disease has been drastically reduced, public health threats posed by microbes are still present and new challenges demand new solutions. Over the last years, an increase in infectious diseases has been observed in almost all countries, regardless of their economic development, with a rising incidence of foodborne illness and nosocomial infections [2][3][4][5]. For effective prevention and intervention measures, the assessment of microbial viability is crucial in determining the safety of food and drinking water, as well as the environmental and medical sector. Together with the rising drug resistance of pathogens against traditional antibiotic therapies, the WHO has defined this problem as one of the most serious public health challenges [6][7][8]. Figure 1 highlights the use of antibiotics in human medicine, animal husbandry, agriculture, and the subsequent circulation between these applications and the environment leading to an increase in the prevalence of antibiotic resistant bacteria in different ecological niches.
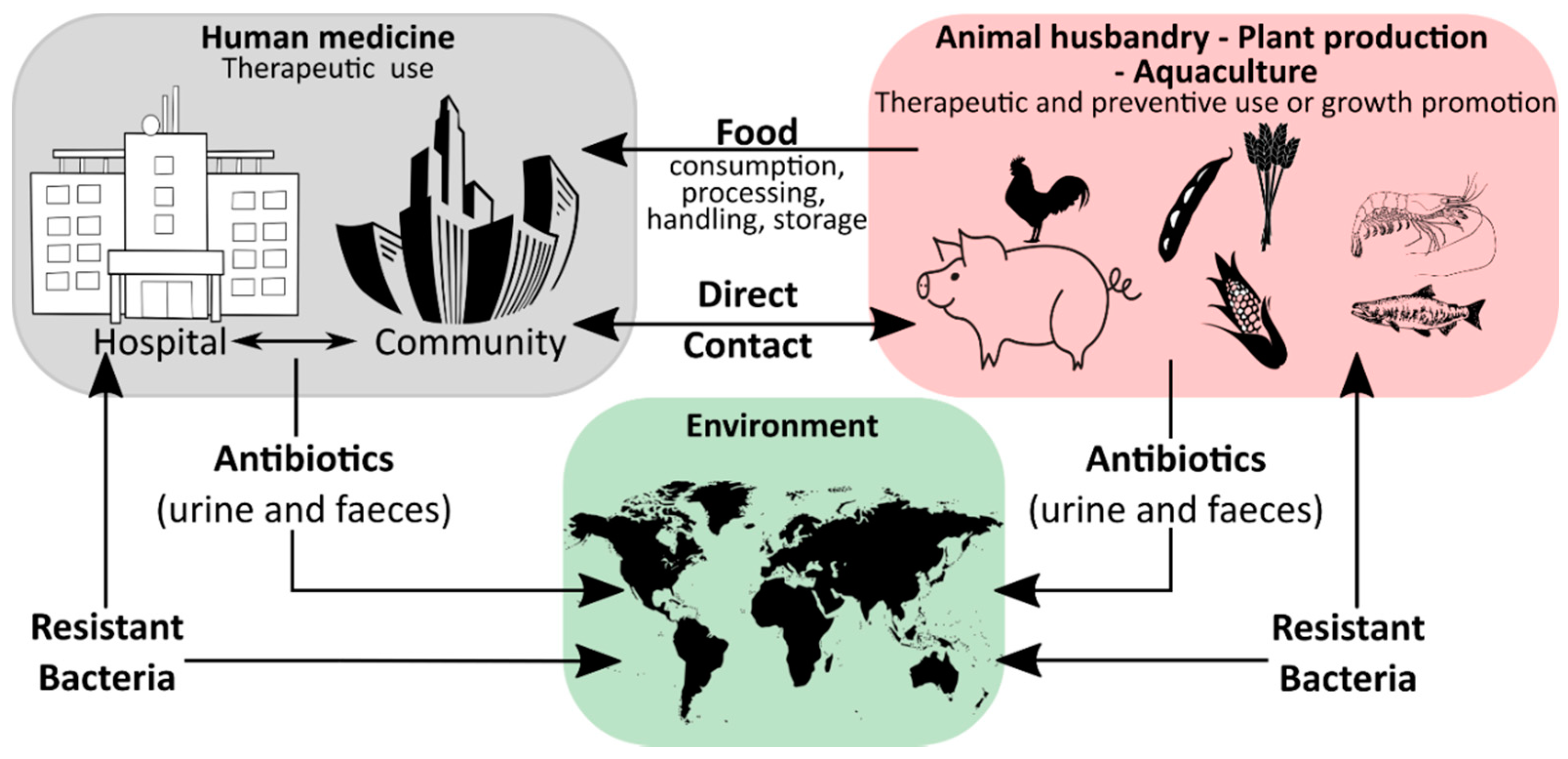
During the last decades, a plethora of antibiotic resistant bacterial genotypes have been reported and as a result, research has intensified to identify novel antibacterial agents and antibiotics [9]. While coming to the fore recently, the impact of phenotypic plasticity on antimicrobial and antibiotic resistance has been far less studied. Phenotypic plasticity describes the ability of bacteria to change their phenotype (in this case their antimicrobial resistance profile) without underlying genetic changes. While phenotypic plasticity in bacteria has been known for a long time already, the understanding of the underlying adaptive and regulatory processes, as well as their role in the infectious process has only recently been significantly expanded [6].
In order to preserve viability and adaptability in constantly changing environmental conditions, bacteria have developed various mechanisms for switching from a vegetative to a metabolically inactive, but highly resistant and persistent state. The most studied type of such cellular dormancy is sporulation of sporogenous bacteria, while in recent years, the viable but non-culturable (VBNC) state for many bacterial pathogens has gained more attention [10][11][12][13]. The VBNC state is a condition of reduced metabolic activity, induced in response to conditions of stress. Entry into the VBNC state can be facilitated by a number of environmental factors, such as a lack of nutrients, temperature shifts, or the presence of antibiotics, disinfectants, and sanitizers . When a microorganism enters the VBNC state, both the morphology and physiology of the cell changes to allow its survival under adverse conditions. Then, once the environmental conditions return to normality, the cell can get back to its culturable state, through a reactivation process, which is referred to as “resuscitation”.
This widespread, or almost universal, occurrence of bacterial cellular dormancy can lead to serious diagnostic and safety problems. Bacteria in the VBNC state represent a diagnostic challenge, due to the simple fact that the majority of current tests are either entirely culture based or need at least one cultivation step, thus failing to detect non-growing VBNC cells [12][14][15][16]. As a result, the absence of growth in culture-based methods does not always mean non-viability, but can also be interpreted as: (i) Incorrect culture medium; (ii) stress or damage of cells leading to a dormant state, (iii) low population density thus leading to no visible growth; and (iv) slow-growing cells and no visible growth. We want to emphasize that the reason for our ongoing dependency on culture-based methods is of course their proven efficiency and cost-effectiveness as diagnostic tools for many years. However, given their inability to detect dormant cells, it is important to point out that the growth and division of cells is not the only accepted parameter in assessing bacterial viability. In addition to reproduction, metabolic activity as well as an intact cell membrane are accepted general parameters for viability assessment of microorganisms, with the latter two having the advantage of being growth independent (Figure 2) [17][18]. While the issues of non-culturability, natural occurrence, induction, and resuscitation of the VBNC cell, as well as the underlying molecular mechanisms and the detection of dormant cells from environmental samples are of extreme importance, these topics have been recently reviewed [12][18][19] and therefore, will not be the main focus of this article. Rather, in this article, we want to elucidate one of the major problems associated with dormant cells, namely their high tolerance and resistance against a wide range of antimicrobials, biocides, and antibiotics . Their elevated tolerance enables cells in the VBNC state to withstand antimicrobial interventions in medical, as well as food-processing environments allowing resuscitation from the VBNC state and possible recontamination of those environments [20][21][22][23][24][25][26]. Phenotypic plasticity needs to be considered when performing antimicrobial efficacy testing. It is clear that the evaluation of antimicrobial treatments or interventions solely on cells in the logarithmic growth phase, combined with subsequent growth based quantification methods, has some significant blind spots which need to be addressed to ensure public health in the medical, as well as food production sector [14][15][27].
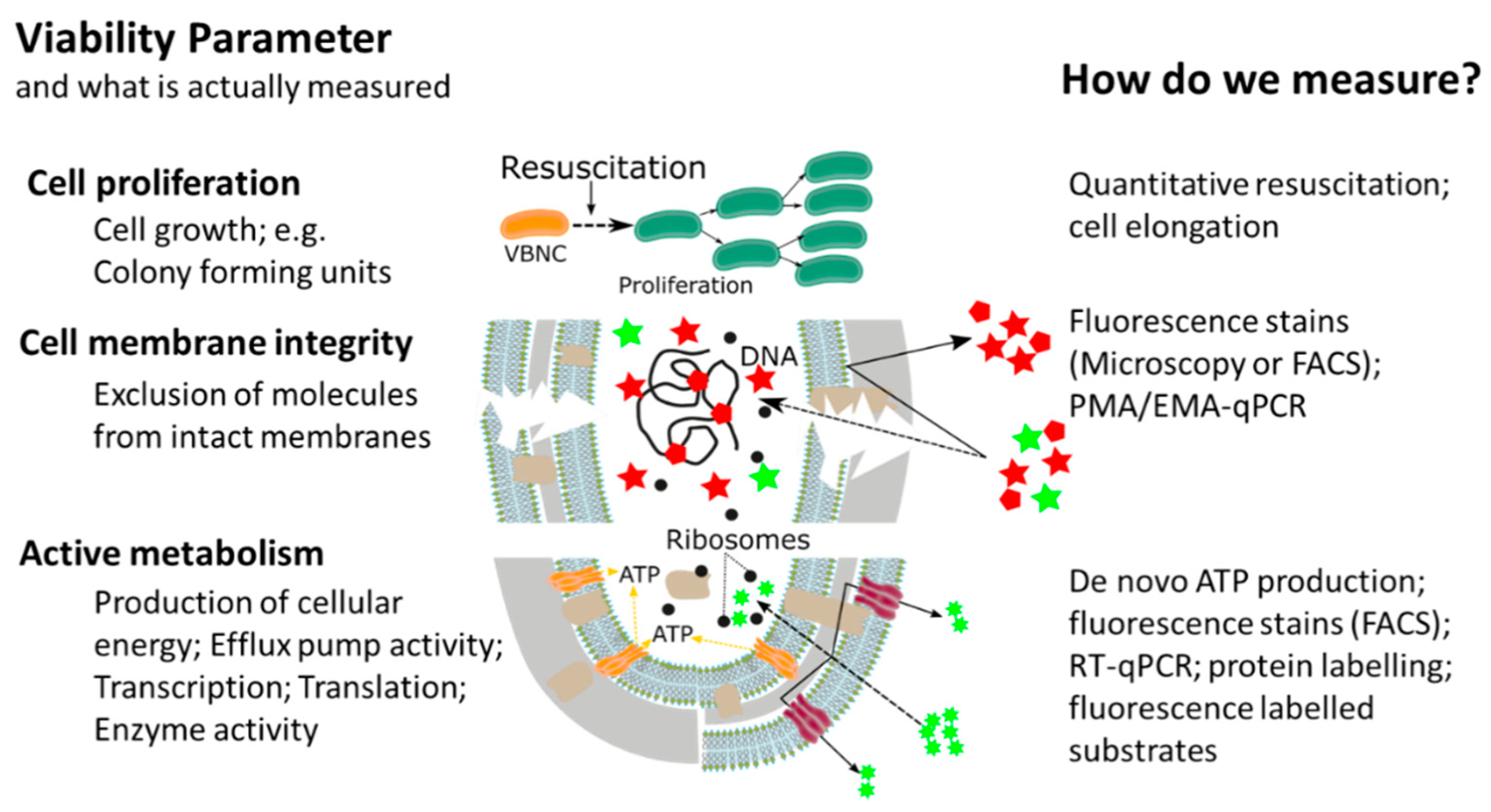
Although antimicrobial resistance is generally recognized as one of the major challenges posed by VBNC cells, the overall number of studies investigating this effect remains sparse. It is our opinion that the main reason for this is the technical and experimental difficulty of investigating antimicrobial resistance of dormant cells in a quantitative way.
2. How to Evaluate Non-Growing Cells—Current Strategies for Determining Antimicrobial Resistance of VBNC Bacteria
In recent years, the awareness about the viable but non-culturable state of bacterial pathogens has dramatically increased and the fact that bacteria in the VBNC state are not visible for routine, growth based detection methods led to the development of novel analytical methods. The majority of methodological approaches determines at least one of the generally accepted viability parameters such as resuscitation, membrane integrity, metabolic activity, or a combination thereof. Thanks to these new methodological approaches, it was discovered that, due to their low metabolic activity and cellular changes in the VBNC state, bacteria have a drastically increased tolerance against antibiotics, as well as antimicrobials and that the methodological approaches could be utilized for antimicrobial effectivity testing against VBNCs.
The majority of methodological approaches were originally developed with a clear focus on the rapid and cost-effective detection of VBNC pathogens and high-throughput methods for antimicrobial effectivity testing are still rare. However, it is our opinion that the relevance of the topic will facilitate their development. There have been several different methodological approaches in order to investigate the tolerance and resistance of VBNC cells, measuring one viability parameter (cell proliferation, cell membrane integrity, or metabolic activity). Based on their underlying experimental principles and the currently available literature, each of the methodological approaches has certain advantages, as well as limitations and for antimicrobial effectivity testing the respective method has to be chosen carefully (Table 1).
Table 1. Summary of the currently available methods for resistance screening for VBNCs.
| Method | Viability Parameter | Target and Unit | Advantage | Limitation |
|---|---|---|---|---|
| Resuscitation | Cell proliferation | Entire cell [CFU] |
|
|
| Viability PCR/qPCR | Membrane integrity | DNA [Gene copies] |
|
|
|
||||
| Fluorescence microscopy | Membrane integrity | Membrane components and DNA [Cell numbers] |
|
|
| Cellular energy |
Active metabolism | ATP [RLU] |
|
|
| Respiratory activity | Active metabolism | Reduction of fluorescencent dyes [Cell numbers] |
|
|
References
- CDC on infectious diseases in the United States: 1900-99. Popul. Dev. Rev. 1999, 25, 635–640.
- The European Union One Health 2018 Zoonoses Report. EFSA J. 2019, 17.
- Witte, A.K.; Mester, P.; Fister, S.; Süß, B.; Wagner, M.; Rossmanith, P. PCR-Stop analysis as a new tool for qPCR assay validation. Sci. Rep. 2018, 8.
- Scallan, E.; Griffin, P.M.; Angulo, F.J.; Tauxe, R.V.; Hoekstra, R.M. Foodborne illness acquired in the United states-Unspecified agents. Emerg. Infect. Dis. 2011, 17, 16–22.
- Saker, L.; Lee, K.; Cannito, B.; Gilmore, A.; Campbell-Lendrum, D.H. Globalization and Infectious Diseases: A Review of the Linkages. 2004. Available online: https://apps.who.int/iris/bitstream/handle/10665/68726/TDR_STR_SEB_ST_04.2.pdf (accessed on 22 January 2021).
- Andryukov, B.G.; Somova, L.M.; Matosova, E.V.; Lyapun, I.N. Phenotypic plasticity as a strategy of bacterial resistance and an object of advanced antimicrobial technologies. Sovrem. Tehnol. Med. 2019, 11, 164–180.
- WHO. No Time to Wait: Securing the Future from Drug-Resistant Infections; WHO: Geneva, Switzerland, 2019.
- Rice, L.B. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No ESKAPE. J. Infect. Dis. 2008, 197, 1079–1081.
- Andersson, D.I.; Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 2014, 12, 465–478.
- Zhao, X.; Zhong, J.; Wei, C.; Lin, C.-W.; Ding, T. Current perspectives on viable but non-culturable state in foodborne pathogens. Front. Microbiol. 2017, 8.
- Ayrapetyan, M.; Williams, T.C.; Oliver, J.D. Bridging the gap between viable but non-culturable and antibiotic persistent bacteria. Trends Microbiol. 2015, 23, 7–13.
- Ding, T.; Suo, Y.; Xiang, Q.; Zhao, X.; Chen, S.; Ye, X.; Liu, D. Significance of viable but nonculturable Escherichia coli: Induction, detection, and control. J. Microbiol. Biotechnol. 2017, 27, 417–428.
- Li, L.; Mendis, N.; Trigui, H.; Oliver, J.D.; Faucher, S.P. The importance of the viable but non-culturable state in human bacterial pathogens. Front. Microbiol. 2014, 5, 258.
- Bonilauri, P.; Bardasi, L.; Leonelli, R.; Ramini, M.; Luppi, A.; Giacometti, F.; Merialdi, G. Detection of food hazards in foods: Comparison of real time polymerase chain reaction and cultural methods. Ital. J. Food Saf. 2016, 5, 37–40.
- Witte, A.K.; Bromberger, B.; Mester, P.; Robben, C.; Schoder, D.; Streit, E.; Rossmanith, P. Implementation of the DEP-pooling approach for L. monocytogenes detection over 25-months by two diagnostic laboratories of an Austrian dairy company. Food Control 2020, 118.
- Mester, P.; Witte, A.K.; Robben, C.; Streit, E.; Fister, S.; Schoder, D.; Rossmanith, P. Optimization and evaluation of the qPCR-based pooling strategy DEP-pooling in dairy production for the detection of Listeria monocytogenes. Food Control 2017, 82.
- Emerson, J.B.; Adams, R.I.; Román, C.M.B.; Brooks, B.; Coil, D.A.; Dahlhausen, K.; Ganz, H.H.; Hartmann, E.M.; Hsu, T.; Justice, N.B.; et al. Schrödinger’s microbes: Tools for distinguishing the living from the dead in microbial ecosystems. Microbiome 2017, 5, 86.
- Kumar, S.S.; Ghosh, A.R. Assessment of bacterial viability: A comprehensive review on recent advances and challenges. Microbiology 2019, 165, 593–610.
- Schottroff, F.; Fröhling, A.; Zunabovic-Pichler, M.; Krottenthaler, A.; Schlüter, O.; Jäger, H. Sublethal injury and Viable but Non-culturable (VBNC) state in microorganisms during preservation of food and biological materials by non-thermal processes. Front. Microbiol. 2018, 9.
- Del Mar Lleò, M.; Benedetti, D.; Tafi, M.C.; Signoretto, C.; Canepari, P. Inhibition of the resuscitation from the viable but non-culturable state in Enterococcus faecalis. Environ. Microbiol. 2007, 9, 2313–2320.
- Ramamurthy, T.; Ghosh, A.; Pazhani, G.P.; Shinoda, S. Current perspectives on viable but non-culturable (VBNC) pathogenic bacteria. Front. Public Health 2014, 2.
- Nowakowska, J.; Oliver, J.D. Resistance to environmental stresses by Vibrio vulnificus in the viable but nonculturable state. FEMS Microbiol. Ecol. 2013, 84, 213–222.
- Lee, S.; Bae, S. Molecular viability testing of viable but non-culturable bacteria induced by antibiotic exposure. Microb. Biotechnol. 2018, 11, 1008–1016.
- Kobayashi, H.; Oethinger, M.; Tuohy, M.J.; Hall, G.S.; Bauer, T.W. Distinction between intact and antibiotic-inactivated bacteria by real-time PCR after treatment with propidium monoazide. J. Orthop. Res. 2010, 28, 1245–1251.
- Bamford, R.A.; Smith, A.; Metz, J.; Glover, G.; Titball, R.W.; Pagliara, S. Investigating the physiology of viable but non-culturable bacteria by microfluidics and time-lapse microscopy. BMC Biol. 2017, 15.
- Robben, C.; Fister, S.; Witte, A.K.; Schoder, D.; Rossmanith, P.; Mester, P. Induction of the viable but non-culturable state in bacterial pathogens by household cleaners and inorganic salts. Sci. Rep. 2018, 8.
- Ayrapetyan, M.; Oliver, J.D. The viable but non-culturable state and its relevance in food safety. Curr. Opin. Food Sci. 2016, 8, 127–133.
- Mester, P.; Robben, C.; Witte, A.K.; Kalb, R.; Ehling-Schulz, M.; Rossmanith, P.; Grunert, T. FTIR Spectroscopy Suggests a Revised Mode of Action for the Cationic Side-Chain Effect of Ionic Liquids. ACS Comb. Sci. 2019, 21, 90–97.





