| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Miroslav Přibyl | + 3372 word(s) | 3372 | 2021-01-27 04:34:35 | | | |
| 2 | Bruce Ren | -21 word(s) | 3351 | 2021-02-03 04:17:26 | | |
Video Upload Options
Among the ~22,000 human genes, very few remain that have unknown functions. One such ex-ample is suprabasin (SBSN). Originally described as a component of the cornified envelope, the function of stratified epithelia-expressed SBSN is unknown. Both the lack of knowledge about the gene role under physiological conditions and the emerging link of SBSN to various human diseas-es, including cancer, attract research interest. The association of SBSN expression with poor prognosis of patients suffering from oesophageal carcinoma, glioblastoma multiforme, and myel-odysplastic syndromes suggests that SBSN may play a role in human tumourigenesis. Three SBSN isoforms code for the secreted proteins with putative function as signalling molecules, yet with poorly described effects.
1. Introduction
Since its original description in human and mouse keratinocyte differentiation [1], suprabasin (SBSN) has been associated with multiple diseases, including cancer. SBSN isoforms are putative signalling molecules inducing cellular signalling (AKT, WNT/β-catenin, and/or p38MAPK signalling) and various cellular processes, such as migration, proliferation, neovascularization, therapy-, apoptosis- and immune-resistance. Therefore, SBSN is considered an oncogene and is a proposed biomarker in a couple of diseases, lung carcinoma and myelodysplastic syndromes (MDS). The apparent need for a deeper understating of the nature and the function of SBSN prompted us to compile all current knowledge of SBSN together with suggestions for future research work.
2. SBSN Gene Organization
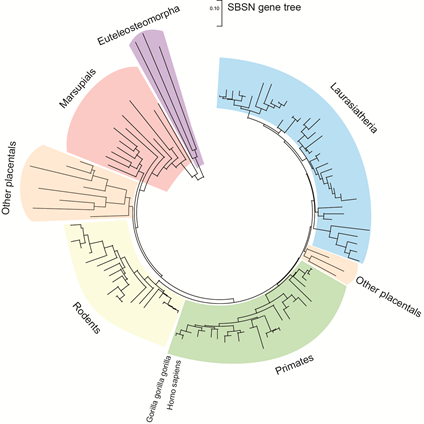
SBSN gene is located on human chromosome 19 (chr 19: 35,523,367–35,528,351 reverse strand; GRCh38:CM000681.2; band 19q13.1) close to other keratinocyte-differentiation associated genes dermokine-α/β and KDAP [1][2]. In mice it corresponds to chromosome 7 (7:30,751,471–30,756,134 forward strand; GRCm38:CM001000.2; band 7B2–7B3). The SBSN gene is a part of a coordinately expressed new stratified epithelium-related gene cluster, tentatively named stratified epithelium secreted peptides complex, SSC. SBSN was studied mostly in human and mice; however, other mammals possess SBSN homologs. Predicted SBSN peptide encoded in the genome of Gorilla gorilla gorilla possesses 97.6% amino acid sequence similarity to the human SBSN-1. Relatively high amino acid sequence identity (58.9%) between mouse and human orthologs of the largest SBSN isoform (SBSN isoform 1; SBSN-1) suggests strong gene integrity and conserved function among the species. Paralogs of SBSN were not defined, and no genes with confidently high sequence identities were identified. The Ensembl database [3] (Ensembl Genome Browser version 101, accessed on 21 August 2020) enlists 106 SBSN orthologs (26/26 primates; 30/32 rodents and related; 39/45 Laurasiatheria; 0/19 Sauropsida; 2/86 fish, and nine Monotremata and Marsupialia; Figure 1) with 96 orthologs having Gene Order Conservation Score 100 (identical four closest genes), implicating true orthology. Note, the putative SBSN fish genes show Gene Order Conservation Scores of 0 and 2.1% and 8.6% sequence identity to the human ortholog, respectively. Altogether, these observations suggest that SBSN is likely a mammalian-specific orphan gene.
Figure 1. The gene tree of suprabasin SBSN. The gene tree of SBSN generated with MEGA X software [4] utilizing Enesembl (version 101) available data . With the total sum of branch lengths (SBL) being >9.477. Distance scale = 0.10.
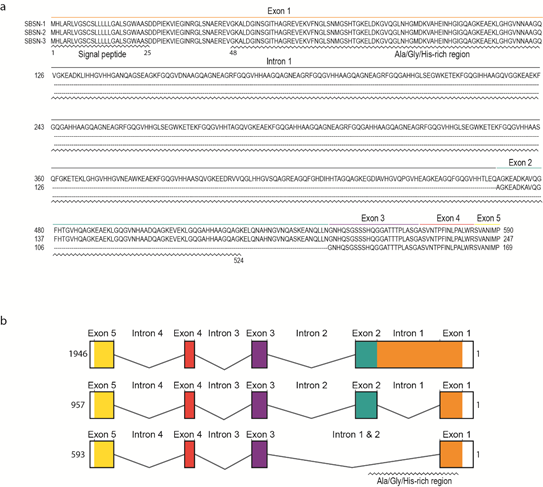
Human SBSN gene consists of five exons and four introns (Figure 2a). Human SBSN mRNA can be alternatively spliced producing three known isoforms SBSN-1 (transcript length: 1946 bp; ENST00000452271.7), SBSN-2 (957 bp; ENST00000518157.1) and SBSN-3 (593 bp; ENST00000588674.5). Importantly, SBSN-2 represents a fully spliced isoform [5], whereas SBSN-1 contains an in-frame retention of the first intron. SBSN-3 is also a fully spliced isoform but lacks exon 2. Strikingly, the mouse (Mus musculus) isoform corresponding to human SBSN-2 has not been identified. This may restrict the applicability of a mouse model for functional studies of SBSN isoforms. Interestingly, a sequence with high sequence identity (81.4%) to human exon two is present within the murine Sbsn-1 exon one, and the major difference between the human intron one exon two junction and homologous murine sequence (CACGAGGCCGGG vs. AACCAGGGTCAA) implies that in mice, the splicing site was likely not established, or was lost. The murine putative exon two is spliced together with the entire exon one in murine Sbsn-2, hence resembling human isoform three, based upon amino acids sequence identity analysis (44.5% for human SBSN-2 vs. murine Sbsn-2, and 65.5% for human SBSN-3 vs. murine Sbsn-2). Multiple other rodents lack homolog corresponding to the human SBSN-2 isoform, and interestingly, several primates (e.g., Macaca mulatta, Pan paniscus and Microcebus murinus) lack putative SBSN-2 as well. However, this may be due to the absence of its identification, rather than sequence deviation.
3. SBSN Protein Structure
The UniProtKB database [6] (The UniProt release 2020_4, accessed 21 August 2020) refers to three human SBSN isoforms (Figure 2b). SBSN-1 (Q6UWP8-1; 590 aa, predicted mass 60.541 Da) and SBSN-2 (Q6UWP8-2; 247 aa, predicted mass 25.335 Da) are well defined. However, SBSN-3 (K7ESC4) is described as a 149 aa long peptide with a predicted mass of 15.318 Da, which is incorrect due to the lacking description of putative N-terminal signal peptide (i.e., missing N-terminal sequence MHLARLVGSCSLLLLLGALS; see also sequence alignment in Figure 2a) in the database. In fact, the SBSN-3 coding sequence is 507 nts long and translates into a 169 aa long peptide. All SBSN isoforms possess a putative N-terminal signal peptide (aa 1–25) [7] granting SBSN secretory nature and extracellular localization.
Figure 2. The sequences of SBSN isoforms. (a) The amino acid sequences of human SBSN isoforms. (b) The gene organization of human SBSN isoforms.
Distinctly of the other two isoforms, SBSN-1 is alanine-(14%), glycine-(20%), and histidine-(10%) rich due to the presence of short tandem Glycine-x-Histidine-Histidine repeats (GxHH repeats, x stands for any classical amino acid) encoded by the retained intron one. Note, structural proteins often contain compositional biases of similar characteristics to SBSN-1 Ala-/Gly-/His-rich domain (aa 26—524), and based upon these observations, SBSN was proposed as a component of the cornified envelope (CE), the essential structure of corneocytes responsible for the skin protective function. Notably, no similar sequence encoding SBSN-1 Ala-/Gly-/His-rich domain is present in the human genome, indicating an evolutionarily unique and conserved role. This is further supported by a comparison of SBSN among species, showing remarkable conservation of the Ala-/Gly-/His-rich domain, as well as retention of the first intron encoding the domain. Human SBSN-2 and SBSN-3 lack most of the Ala-/Gly-/His-rich domains due to intron one splicing; however, a short repeat is located at the 3′ end of exon one. Similarly to Ala-/Gly-/His-rich domain of SBSN-1, the C-terminal sequence of all isoforms is of unknown function. As mentioned, SBSN-3 lacks exon two, but the remaining exons are present in all human SBSN isoforms.
The mouse homolog of SBSN possesses similar features. The N-terminal signal peptide was predicted (aa 1–23), and the compositional bias of Ala-/Gly-/His-rich domain (aa 97—478) in SBSN-1 (Q8CIT9; 700 aa; predicted mass 72.334 Da) is also present in mouse homolog. The current description of other murine Sbsn isoforms is rather confusing. Originally, murine Sbsn-1 isoform was identified as a 700 aa long peptide. A shorter isoform, lacking the longest isoform-specific internal repeats, is a putative Sbsn-2 (Q8CIT9-3; 164 aa; predicted mass 16.967 Da), which corresponds to human SBSN isoform-3 according to sequence analysis. Besides these two isoforms, Ensembl (version 101) identifies two other protein-coding isoforms supported by one or no EST, respectively. The UniProtKB database refers to six mouse Sbsn isoforms, including Sbsn-1 and Sbsn-2, the other isoforms are either duplications of Sbsn-1 or Sbsn-2, or unreviewed isoforms.
Secreted murine Sbsn-2 was detected using overexpression experiments, showing a mobility shift on immunoblots which indicates a post-translational modification. Indeed, murine Sbsn-1 is a substrate to tissue transglutaminase (Tgm) 2 and epithelial Tgm3 resulting in intermolecular crosslinking, and this represents the only verified post-translational modification in vitro. Its physiological relevance is still undetected. Citrullination of Sbsn was observed in the blood proteome of Parkinson’s disease in a rat model of pre-motor Parkinson’s disease [8]. Additionally, glycosylation of SBSN-1 was predicted and indeed, using mass spectrometry approach, threonine 59 of human SBSN was identified to undergo O-glycosylation in various human cell lines [9]. Three protein kinase C (PKC) phosphorylation sites in C-terminus and two casein kinase II phosphorylation sites were predicted together with 73 potential N-myristoylation sites in the mouse homolog; however, these predictions require experimental validation.
Three-dimensional structure of SBSN is still unknown. We performed Phyre2-based structure prediction [10] of all three human SBSN isoforms. In the case of SBSN-1, the predicted model predominantly consisted of disordered regions. Crystal structure of d337a mutant of Pseudomonas sp. mis38 lipase (PDB: 2ZJ6) served as a template for structure prediction with 12% sequence identity and 99.9% prediction confidence. Collagen α chain was among other possible templates with slightly lower confidence levels. The methyl-accepting chemotaxis protein of Escherichia coli (PDB: 1QU7), showing 13% sequence identity with SBSN-2, was used as a template for the structure prediction of the isoform with 98.5% confidence. Interestingly, the methyl-accepting chemotaxis protein of Thermotoga maritima (PDB: 2CH7) was suggested as a second template with 98.4% confidence. SBSN-3 isoform model was predicted upon the structure of human micelle-bound α-synuclein (PDB: 1XQ8), which showed 18% sequence identity.
4. Regulation of SBSN Expression
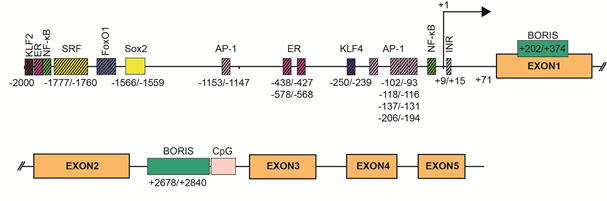
SBSN expression is tightly associated with stratified epithelia, but other expression sites have also been defined. Mechanisms of SBSN transcriptional control are not clearly described. We utilized the JASPAR algorithm [11] to predict binding sites of transcription factors in human (Supplementary Table S1) and mouse (Supplementary Table S2) SBSN proximal promoter (2 kbp upstream) and downstream coding region. Additionally, we verified putative binding sites. Based on these results, we established a model of human SBSN promoter (Figure 3). From transcription binding sites predicted within human SBSN proximal promoter region by in silico analysis, the SOX2 binding site is currently the only one experimentally validated [12] (Figure 3). Next, a Brother of the Regulator of Imprinted Sites (BORIS)-binding site within the coding region was experimentally verified [13]. Importantly, aberrant changes in expression of SBSN isoform are associated with atopic dermatitis (see below [14]), but mechanisms responsible for differential expression of isoforms under physiological and pathological conditions are not known.
SBSN is physiologically expressed in mouse stratified epithelia including the suprabasal epithelial layer of epidermis, tongue, oesophagus, palate, stomach, uterus, thyroid, trachea, lung, vagina, thymus, and urinary tract. In human, SBSN expression is associated with epidermis, thymus, uterus, tonsils, vagina, and oesophagus [15]. Originally, there was no evidence of SBSN expression in mouse embryonic and adult brain [16][17], but recent studies showed, using immunofluorescence, SBSN expression in human astrocytes, and its elevation under pathological conditions [18]. Transcriptome analysis of the human brain revealed SBSN mRNA levels in basal ganglia, but evaluation of SBSN protein presence in the human brain is needed to confirm this expression site [15]. On the contrary, the mouse brain does not show the presence of Sbsn mRNA; neither do several other mouse tissues and organs such as heart, kidney, or smooth muscle. Therefore, between mouse and human, the SBSN expression sites seem to be conserved, except for the brain.
Figure 3. SBSN promoter. Two kilobase pairs upstream of the promoter and downstream sequence of human SBSN with predicted transcription factor binding sites. SOX2 binding site is currently the only validated site together with Brother of the Regulator of Imprinted Sites (BORIS) binding sites.
Mouse Sbsn mRNA was detected on day seven of embryonic development when the expression is likely mediated by extra-embryonic tissue. Hence, the embryonic expression was detected on day 15 of the development at first, which is in coincidence with epidermal stratification. Sbsn mRNA levels then peak on day 17, and SBSN expression is associated with expression of dermokine-α/β, the components of secreted peptides complex. Similarly, SBSN mRNA was elevated in skeletal muscle cells of Alaskan sled dogs during an acute response (2 h post-exercise) after a prolonged endurance training together with dermokine and keratin 5. At the same time point, transcripts of genes involved in inflammation, oxidative stress, intermediary metabolism, immune response, and cellular compromise transcripts, e.g., S100A8, were also upregulated. The role of inflammation, immune, and stress response in SBSN expression is supported by the microarray analysis of therapy-resistant cancer cells in vitro, which showed transcript elevation of innate immune response genes and SBSN following 5-aza-2′-deoxycytidine (5-AC)-treatment or γ-radiation. Notably, activation of the ERK pathway downstream of IFN signalling emerged as a direct activator of SBSN expression [19]. Therefore, epidermal differentiation and response to inflammation provide hints at the understanding of SBSN expression inducing processes.
The composition of transcription factors and stimuli responsible for SBSN transcription is not specifically defined, but some are suggested. The Sbsn transcript elevates in differentiating mouse keratinocytes in vitro, whereas, several genes of the cornified envelope are downregulated upon SBSN knockdown. This provides additional support for the coordinated gene expression program during skin differentiation. Targets of the ERK pathway, i.e., components of the AP-1 transcription factor complex c-FOS, FRA-1, FRA-2, c-JUN, JUND, and JUNB, are differentially expressed in keratinocytes during their terminal differentiation in organotypic cultures, and AP-1 proteins are differentially expressed in the human epidermis [20]. This supports the role of ERK in SBSN expression; however, the role of ERK in keratinocyte differentiation provides contradictory results [20][21][22]. Additionally, MAL/SRF signalling also results in JUNB elevation, which plays an essential role in epidermal differentiation [21]. Inhibition of BCR-RHOA-MAL/SRF pathway resulted in the reduction of JUNB and SBSN transcripts, together with disruption of keratinocyte granulation and development of stratum corneum in an organotypic model of the human epidermis. Mouse bearing conditional knockout of Srf in basal cells, showed reduced Sbsn transcript levels [24], and indeed, the SRF binding site was predicted within the SBSN proximal promoter region of both, human and mouse (with three binding sites in mice (−1994/−1978, −1036/−1023, −789/−773 (0.83, 0.8, and 0.8 relative score) and one binding site (−1777/−1760) in human with a 0.82 relative score). Changes in the actin cytoskeleton and MAL/SRF promote physical stimuli-induced keratinocyte differentiation via JUNB [21]. Note, in dog muscle, the JUNB transcript is elevated 2 h post-exercise, while the Fos transcript is downregulated. ERK-mediated regulation of SBSN is further supported by increased expression of Sbsn in murine endothelial cells following treatment with Egf, but not bFgf [25]. Furthermore, phorbol 12-myristate 13-acetate (PMA)-mediated ERK activation enhanced SBSN expression efficiently, though this effect might be partially mediated by PKC since calcium-induced SBSN expression during differentiation of primary epidermal keratinocytes in vitro was suppressed with PKC inhibitor. Therefore, multiple pathways activated during keratinocytes differentiation may promote SBSN expression likely via, but not only, AP-1-enabled transcription. Indeed, SBSN promoter region contains multiple AP-1 binding sites. In total, a JASPAR search predicted 72 AP-1 binding sites, with extensive sequence overlaps, hence lesser number of regions is more likely. Five regions showed >0.9 relative scores (−93/−102, −112/−118, −131/−137, −194/−206, −1147/−1153).
Lower temperature (33 °C) is frequently used in biotechnology for culture/propagation of Chinese hamster ovary cells. In a recent study, a list of cold-induced genes was established using RNAseq 48 h post change of condition, which included SBSN [26]. This was further supported with ectopic expression of luciferase driven by SBSN promoter exposed to lower temperatures. S100A4 was identified to be a cold-induced gene [26], together with S100A6, which is supported by a previous study [27]. Furthermore, the JunD binding site was predicted among selected cold-regulated promoters, including SBSN, and JUND transcript levels were elevated following cold-treatment [26].
In response to hypoxic stress, high altitude acclimatization leads to lower oxygen tension and hypobaric pressure with enhanced hematopoiesis, increased blood volume, and neoangiogenesis to redistribute blood flow to vital organs, including the brain. The latter is mediated by carotid arteries. Upon acclimatization to high-altitude-associated long-term hypoxia, SBSN transcript was one of 58 significantly upregulated in carotid arteries in sheep [28]. Interestingly, most altered genes were associated with cell migration, growth and proliferation, and angiogenesis. The authors also noted that some of the regulated genes are also common targets of treatment with lipopolysaccharide (LPS), again supporting the contribution of the innate immune response to SBSN expression. Note, the upstream pathway, ERK, and a target of SBSN signalling, AKT, showed increased activation in sheep carotid arteries accompanied long-term hypoxia .
The proximal region of SBSN gene promoter was originally described as AT-rich, lacking CpG islands and containing a canonical TATA box, however, in our analysis presented here (Figure 3) we were not able to identify these regions within 250 bp upstream of +1 site of human SBSN promoter region. Conversely, we observed an initiator element at +9/+15 (Figure 3). Multiple NF-kappaB binding sites were predicted within a 2 kbp region upstream of SBSN transcription start site in both human and mouse cells [29]. We predicted thirteen NF-kappaB binding sites within this region (Figure 3). The most prominent sites depicted had a relative score > 0.92; −1921/1930, > 0.8; −48/−39). Note, both PMA and LPS are potent inducers of NF-kappaB pathway. Furthermore, several other binding sites for transcription factors, such as SP1, TF2APA, MYC, SMAD2, and FOXO1/FOXO4 were predicted within the SBSN proximal promoter [30]. Indeed, SBSN transcription in confluent human adipocyte tissue-derived stem cells (ASCs) is mediated by FOXO1 [30], since downregulation of FOXO1 with silencing RNA reduced SBSN transcript levels. We predicted four FOXO1 binding sites within 2 kbp SBSN promoter region (relative score > 0.84; −1616/1609, −1668/−1661, −1692/−1685, −1930/−1923; Figure 3). Interestingly, SBSN promotes aromatase expression, and SBSN was shown to be induced with 17beta-estradiol treatment [31]. This is supported with the prediction of estrogen receptor (ER) binding sites within the promoter. We identified thirteen ER binding sites within 2 kbp upstream region of SBSN promoter and selected three highest scoring regions to depict (relative score > 0.89; −438/−427, > 0.86; −578/−568, > 0.85; −1902/−1888; Figure 3). Altogether, dozens of transcription factors binding sites are predicted. Needless to mention, confirmation of function of the predicted binding regions require further investigation.
Multiple studies described aberrant elevation of SBSN in human malignancies [32], but the mechanisms responsible for SBSN upregulation under pathological conditions is not understood. As mentioned, SBSN expression is limited to specific tissues. Bisulfite sequencing of 11 healthy human lung tissue samples revealed methylation of SBSN promoter CpG islands. This indicates that promoter methylation, and likely associated transcriptional repression, are responsible for SBSN silencing [33]. Indeed, normal human bronchial epithelial and human small airway epithelial cells treated with demethylating agent 5-AC and histone deacetylase inhibitor trichostatin A (TSA) elevated SBSN transcript levels [33]. This is further supported by hypomethylation of SBSN promoter in approximately 50% (13/28) of primary non-small cell lung carcinoma (NSCLC) samples observed in the same study. Hypomethylation of the SBSN promoter was an effect of a dysregulated proto-oncogenic zinc finger transcription factor, CTCFL/Brother of the Regulator of Imprinted Sites (BORIS). Similarly, SBSN transcription can be induced in vitro in a salivary gland adenoid cystic carcinoma (ACC) cell line with 5-AC and trichostatin A (TSA) resulting in hypomethylation of the CpG island [34]. The SBSN transcripts are induced with 5-AC in human cancer cell lines such as DU-145, MCF-7, and HeLa. Notably, SBSN proteins were only detectable in a low-adherent subfraction of therapy-resistant 5-AC-treated cells with stem cell-like properties, indicating post-transcriptional regulation of SBSN expression. The SBSN gene promoter contains two experimentally confirmed CTCFL/BORIS binding sites in the first exon close to the transcription start site (+202/+374) and in the second intron in front of a CpG island (+2678/+2840). BORIS-mediated induction of SBSN is associated with demethylation of a CpG island in SBSN second intron and changes in histone marks comprising elevation of the active H3K4me3 and H3K14Ac, and downregulation of the repressive H3K9me3 modifications. Importantly, the elevation of SBSN transcript levels mediated by BORIS is dose-dependent. Relatively low BORIS levels were responsible for significantly higher SBSN transcript levels compared to high BORIS levels associated with re-methylation of the second intron of SBSN and increased nucleosome occupancy of SBSN transcription start site. A repressive histone mark H3K9me3 mirrored the SBSN second intron methylation pattern. Interestingly, CTCF and BORIS compete for the same binding sites of the SBSN promoter region, indicating that the epigenetic and chromatin state play essential roles in SBSN expression. Hence, methylation of regulatory sites represents the main feature responsible for the regulation (suppression) of SBSN transcription.
The connection between cell stemness and SBSN expression suggested previously is supported by the identification of SOX2, a stem cell factor commonly upregulated in cancer, as a regulator of SBSN expression in oesophageal squamous cell carcinoma (ESCC). SOX2 binding site (−1566/−1559) at the proximal region of human SBSN was confirmed by chromatin immunoprecipitation (ChIP), and remains the only determining region of SBSN promoter with the validated transcription binding factor. Furthermore, double knockout Klf2 and Klf4 mouse cardiac microvascular endothelial cells showed significantly reduced SBSN transcript levels compared to control mice [35]. We identified 25 putative KLF4 binding sites and 30 putative KLF2 binding sites in the human SBSN promoter region, two most prominent are depicted (KLF4 relative score > 0.9; −250/−239, KLF2 relative score > 0.9; −1993/−1983; Figure 3). These observations strengthen the importance of factors of stemness in SBSN expression regulation.
References
- Park, G.T.; Lim, S.E.; Jang, S.I.; Morasso, M.I. Suprabasin, a novel epidermal differentiation marker and potential cornified envelope precursor. J. Biol. Chem. 2002, 277, 45195–45202, doi:10.1074/jbc.M205380200.
- Matsui, T.; Hayashi-Kisumi, F.; Kinoshita, Y.; Katahira, S.; Morita, K.; Miyachi, Y.; Ono, Y.; Imai, T.; Tanigawa, Y.; Komiya, T.; et al. Identification of novel keratinocyte-secreted peptides dermokine-α/- β and a new stratified epithelium-secreted protein gene complex on human chromosome 19q13.1. Genomics 2004, 84, 384–397, doi:10.1016/j.ygeno.2004.03.010.
- Hunt, S.E.; McLaren, W.; Gil, L.; Thormann, A.; Schuilenburg, H.; Sheppard, D.; Parton, A.; Armean, I.M.; Trevanion, S.J.; Flicek, P.; et al. Ensembl variation resources. Database 2018, 2018, doi:10.1093/database/bay119.
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549, doi:10.1093/molbev/msy096.
- Clark, H.F. The Secreted Protein Discovery Initiative (SPDI), a Large-Scale Effort to Identify Novel Human Secreted and Transmembrane Proteins: A Bioinformatics Assessment. Genome Res. 2003, 13, 2265–2270, doi:10.1101/gr.1293003.
- Consortium, T.U. Activities at the Universal Protein Resource (UniProt). Nucleic Acids Res. 2014, 42, D191–198, doi:10.1093/nar/gkt1140.
- Moffatt, P.; Salois, P.; St-Amant, N.; Gaumond, M.H.; Lanctôt, C. Identification of a conserved cluster of skin-specific genes encoding secreted proteins. Gene 2004, 334, 123–131, doi:10.1016/j.gene.2004.03.010.
- Sancandi, M.; Uysal-Onganer, P.; Kraev, I.; Mercer, A.; Lange, S. Protein deimination signatures in plasma and plasma-evs and protein deimination in the brain vasculature in a rat model of pre-motor parkinson’s disease. Int. J. Mol. Sci. 2020, 21, 2743, doi:10.3390/ijms21082743.
- Steentoft, C.; Vakhrushev, S.Y.; Joshi, H.J.; Kong, Y.; Vester-Christensen, M.B.; Schjoldager, K.T.-B.G.; Lavrsen, K.; Dabelsteen, S.; Pedersen, N.B.; Marcos-Silva, L.; et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 2013, 32, 1478–1488, doi:10.1038/emboj.2013.79.
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Michael, J.E.S. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2016, 10, 845–858, doi:10.1038/nprot.2015-053.
- Mathelier, A.; Zhao, X.; Zhang, A.W.; Parcy, F.; Worsley-Hunt, R.; Arenillas, D.J.; Buchman, S.; Chen, C.Y.; Chou, A.; Ienasescu, H.; et al. JASPAR 2014: An extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 2014, 42, 142–147, doi:10.1093/nar/gkt997.
- Takahashi, K.; Asano, N.; Imatani, A.; Kondo, Y.; Saito, M.; Takeuchi, A.; Jin, X.; Saito, M.; Hatta, W.; Asanuma, K.; et al. Sox2 induces tumorigenesis and angiogenesis of early stage esophagealsquamous cell carcinoma through secretion of Suprabasin. Carcinogenesis 2020, 26, 1–15, doi:10.1093/carcin/bgaa014.
- Gaykalova, D.; Vatapalli, R.; Glazer, C.A.; Bhan, S.; Shao, C.; Sidransky, D.; Ha, P.K.; Califano, J.A. Dose-Dependent Activation of Putative Oncogene SBSN by BORIS. PLoS ONE 2012, 7, e40389, doi:10.1371/journal.pone.0040389.
- Li, J.; Zheng, L.; Uchiyama, A.; Bin, L.; Mauro, T.M.; Elias, P.M.; Pawelczyk, T.; Sakowicz-Burkiewicz, M.; Trzeciak, M.; Leung, D.Y.M.; et al. A data mining paradigm for identifying key factors in biological processes using gene expression data. Sci. Rep. 2018, 8, 9083, doi:10.1038/s41598-018-27258-8.
- Uhlen, M.; Fagerberg, L.; Hallstrom, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, A.; Kampf, C.; Sjostedt, E.; Asplund, A.; et al. Tissue-based map of the human proteome. Science 2015, 347, 1260419–1260419, doi:10.1126/science.1260419.
- Bazzi, H.; Fantauzzo, K.A.; Richardson, G.D.; Jahoda, C.A.B.; Christiano, A.M. Transcriptional profiling of developing mouse epidermis reveals novel patterns of coordinated gene expression. Dev. Dyn. 2007, 236, 961–970, doi:10.1002/dvdy.21099.
- Brass, E.P.; Peters, M.A.; Hinchcliff, K.W.; He, Y.D.; Ulrich, R.G. Temporal pattern of skeletal muscle gene expression following endurance exercise in Alaskan sled dogs. J. Appl. Physiol. 2009, 107, 605–612, doi:10.1152/japplphysiol.91347.2008.
- Ichinose, K.; Ohyama, K.; Furukawa, K.; Higuchi, O.; Mukaino, A.; Satoh, K.; Nakane, S.; Shimizu, T.; Umeda, M.; Fukui, S.; et al. Novel anti-suprabasin antibodies may contribute to the pathogenesis of neuropsychiatric systemic lupus erythematosus. Clin. Immunol. 2018, 193, 123–130, doi:10.1016/j.clim.2017.11.006.
- Hubackova, S.; Pribyl, M.; Kyjacova, L.; Moudra, A.; Dzijak, R.; Salovska, B.; Strnad, H.; Tambor, V.; Imrichova, T.; Svec, J.; et al. Interferon‐regulated suprabasin is essential for stress‐induced stem‐like cell conversion and therapy resistance of human malignancies. Mol. Oncol. 2019, 13, 1467–1489, doi:10.1002/1878-0261.12480.
- Mehic, D.; Bakiri, L.; Ghannadan, M.; Wagner, E.F.; Tschachler, E. Fos and Jun Proteins Are Specifically Expressed During Differentiation of Human Keratinocytes. J. Investig. Dermatol. 2005, 124, 212–220, doi:10.1111/j.0022-202X.2004.23558.x.
- Connelly, J.T.; Gautrot, J.E.; Trappmann, B.; Tan, D.W.-M.; Donati, G.; Huck, W.T.S.; Watt, F.M. Actin and serum response factor transduce physical cues from the microenvironment to regulate epidermal stem cell fate decisions. Nat. Cell Biol. 2010, 12, 711–718, doi:10.1038/ncb2074.
- Zhu, A.J.; Watt, F.M. beta-catenin signalling modulates proliferative potential of human epidermal keratinocytes independently of intercellular adhesion. Development 1999, 126, 2285–2298.
- Dubash, A.D.; Koetsier, J.L.; Amargo, E. V.; Najor, N.A.; Harmon, R.M.; Green, K.J. The GEF Bcr activates RhoA/MAL signaling to promote keratinocyte differentiation via desmoglein-1. J. Cell Biol. 2013, 202, 653–666, doi:10.1083/jcb.201304133.
- Luxenburg, C.; Pasolli, H.A.; Williams, S.E.; Fuchs, E. Developmental roles for Srf, cortical cytoskeleton and cell shape in epidermal spindle orientation. Nat. Cell Biol. 2011, 13, 203–214, doi:10.1038/ncb2163.
- Alam, M.T.; Nagao-Kitamoto, H.; Ohga, N.; Akiyama, K.; Maishi, N.; Kawamoto, T.; Shinohara, N.; Taketomi, A.; Shindoh, M.; Hida, Y.; et al. Suprabasin as a novel tumor endothelial cell marker. Cancer Sci. 2014, 105, 1533–1540, doi:10.1111/cas.12549.
- Nguyen, L.N.; Novak, N.; Baumann, M.; Koehn, J.; Borth, N. Bioinformatic Identification of Chinese Hamster Ovary (CHO) Cold-Shock Genes and Biological Evidence of their Cold-Inducible Promoters. Biotechnol. J. 2019, e1900359, doi:10.1002/biot.201900359.
- Thaisuchat, H.; Baumann, M.; Pontiller, J.; Hesse, F.; Ernst, W. Identification of a novel temperature sensitive promoter in cho cells. BMC Biotechnol. 2011, 11, 51, doi:10.1186/1472-6750-11-51.
- Goyal, R.; Longo, L.D. Acclimatization to long-term hypoxia: Gene expression in ovine carotid arteries. Physiol. Genom. 2014, 46, 725–734, doi:10.1152/physiolgenomics.00073.2014.
- Zhu, J.; Wu, G.; Li, Q.; Gong, H.; Song, J.; Cao, L.; Wu, S.; Song, L.; Jiang, L. Overexpression of Suprabasin is Associated with Proliferation and Tumorigenicity of Esophageal Squamous Cell Carcinoma. Sci. Rep. 2016, 6, 21549, doi:10.1038/srep21549.
- Ghosh, S.; Dean, A.; Walter, M.; Bao, Y.; Hu, Y.; Ruan, J.; Li, R. Cell density-dependent transcriptional activation of endocrine-related genes in human adipose tissue-derived stem cells. Exp. Cell Res. 2010, 316, 2087–2098, doi:10.1016/j.yexcr.2010.04.015.
- Stanton, A.; Mowbray, C.; Lanz, M.; Brown, K.; Hilton, P.; Tyson-Capper, A.; Pickard, R.S.; Ali, A.S.M.; Hall, J. Topical Estrogen Treatment Augments the Vaginal Response to Escherichia coli Flagellin. Sci. Rep. 2020, 10, 8473, doi:10.1038/s41598-020-64291-y.
- Pribyl, M.; Hubackova, S.; Moudra, A.; Vancurova, M.; Polackova, H.; Stopka, T.; Jonasova, A.; Bokorova, R.; Fuchs, O.; Stritesky, J.; et al. Aberrantly elevated suprabasin in the bone marrow as a candidate biomarker of advanced disease state in myelodysplastic syndromes. Mol. Oncol. 2020, 14, 2403–2419, doi:10.1002/1878-0261.12768.
- Glazer, C.A.; Smith, I.M.; Ochs, M.F.; Begum, S.; Westra, W.; Chang, S.S.; Sun, W.; Bhan, S.; Khan, Z.; Ahrendt, S.; et al. Integrative Discovery of Epigenetically Derepressed Cancer Testis Antigens in NSCLC. PLoS ONE 2009, 4, e8189, doi:10.1371/journal.pone.0008189.
- Shao, C.; Tan, M.; Bishop, J.A.; Liu, J.; Bai, W.; Gaykalova, D.A.; Ogawa, T.; Vikani, A.R.; Agrawal, Y.; Li, R.J.; et al. Suprabasin Is Hypomethylated and Associated with Metastasis in Salivary Adenoid Cystic Carcinoma. PLoS ONE 2012, 7, 1–7, doi:10.1371/journal.pone.0048582.
- Sangwung, P.; Zhou, G.; Nayak, L.; Chan, E.R.; Kumar, S.; Kang, D.-W.; Zhang, R.; Liao, X.; Lu, Y.; Sugi, K.; et al. KLF2 and KLF4 control endothelial identity and vascular integrity. JCI Insight 2017, 2, e91700, doi:10.1172/jci.insight.91700.