
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Marta Campos Campos | + 3126 word(s) | 3126 | 2020-10-08 06:15:57 | | | |
| 2 | Rita Xu | -957 word(s) | 2169 | 2021-01-27 03:09:21 | | |
Video Upload Options
The glyoxalase system was discovered over a hundred years ago and since then it has been claimed to provide the role of an indispensable enzyme system in order to protect cells from a toxic byproduct of glycolysis.
1. Introduction
The glyoxalase system, which was first described over a hundred years ago, consists of two cooperating enzymes named Glyoxalase 1 (Glo1) and Glyoxalase 2 (Glo2) [1][2]. It is a highly conserved enzyme system, which exists in every living cell from crude single prokaryotes up to complex mammalian organisms [3][4]. The initial enzyme of the glyoxalase system is glyoxalase 1 and its substrate, hemithioacetal, is formed by a spontaneous reaction of methylglyoxal (MG) and glutathione (GSH) (Figure 1). MG is highly reactive and therefore a toxic compound, but also an inevitable by-product of glycolysis and gluconeogenesis during the conversion of triose phosphate isomers (dihydroxyacetone phosphate and glyceraldehyde 3-phosphate) [5]. Interestingly, the glyoxalase system has an extremely narrow substrate specificity among dicarbonyl metabolites. In fact, it detoxifies mostly MG via the hemithioacetal, which is converted to S-D-lactoylglutathione and then hydrolyzed by Glo2 to D-lactate (Figure 2). Within this context, Glo1 represents the rate-limiting step for the detoxification of MG and is therefore of highest interest because it is believed to maintain the intracellular concentration of harmful MG in a low range. As the prevention of increased intracellular MG concentration is mandatory for the viability of a cell, in order to maintain physiological functions, it is no surprise that in eukaryotic organisms, the abundancy of Glo1 is in the top 10% of cytosolic proteins [6][7].
Glo2 is encoded by the hydroxyacylglutathione hydrolase gene and, like Glo1, it is expressed in nearly all living cells in order to hydrolyze S-D-Lactoylglutathione, the intermediate product of the glyoxalase cycle, into D-Lactate and (recycled) GSH [8][9]. Interestingly, S-D-Lactoylglutathione has been reported to be potentially cytotoxic, inducing growth arrest, and decreasing cellular viability, but further studies are needed to confirm this, because of the non-physiological doses used in this study [10]. In comparison to Glo1, Glo2 has not been located solely in the cytosol, but also in the mitochondria [8]. The amount of research conducted and therefore, available literature regarding Glo2 is far less compared to its counterpart Glo1. This may be the result of the fact that Glo1 is the rate-limiting enzyme of the glyoxalase system and seems to be more important [6]. Nevertheless, it is surprising that Glo2 is regulated by members of the tumor suppressor gene p53. In fact, Glo2 is up-regulated by p63 and p73, which are members of the p53 family, and therefore Glo2 is believed to have a role as a pro-survival factor [11]. Fortunately, recent investigations about the role of mitochondrial Glo2 have revealed new and very interesting content. Antognelli et al. suggested a pro-apoptotic role of Glo2 in non-small-cell lung cancers, and this effect is mediated by a bioactive plant compound called Oleuropein. Another study from the same group revealed that Glo2 can be viewed as a driving force of prostate tumorigenesis and therefore, may represent a novel marker of progression in prostate cancer diagnosis [12][13]. It is a nice illustration, showing that Glo2 may have been overlooked in the context of malignancies and that there is still a lot of space for further studies.
Increased levels of MG can lead to an intracellular accumulation of advanced glycation end products (AGEs) and to increased amounts of glycated DNA adducts. Both events are associated with many pathological events in humans, such as obesity and diabetes, atherosclerosis, various cancer types and neurodegenerative diseases [14][15][16][17][18][19][20]. Regarding protein glycation, the preferred targets of MG are the amino acids arginine and, to a much lesser extent, lysine. This mainly results in MG-derived hydroimidazolones called MG-H1, MG-H2 and MG-H3 (structure displayed in Figure 1). Based upon quantitative measurements, MG-H1 is the dominant type of all AGEs found within a physiological context [21][22][23]. Two other modifications should be mentioned herein, the arginine modification argpyrimidine and the most abundant lysine modification N-(1-carboxyethyl)lysine (CEL) [24][25]. Regarding the modifications of nucleic acids the most reactive nucleotide is deoxyguanosine, resulting in N2-carboxyethyl-2’-deoxyguanosine (CedG) and 3-(2’-deoxyribosyl)-6,7-dihydro-6,7-dihydroxy-6/7-methylimidazo-[2,3-b]purin-9(8)one (MG-dG) [26]. CedG has been found to be elevated in animal models and in human tissue samples, especially within the context of diabetes, but further studies are still needed to confirm its relevance (Figure 1) [27].
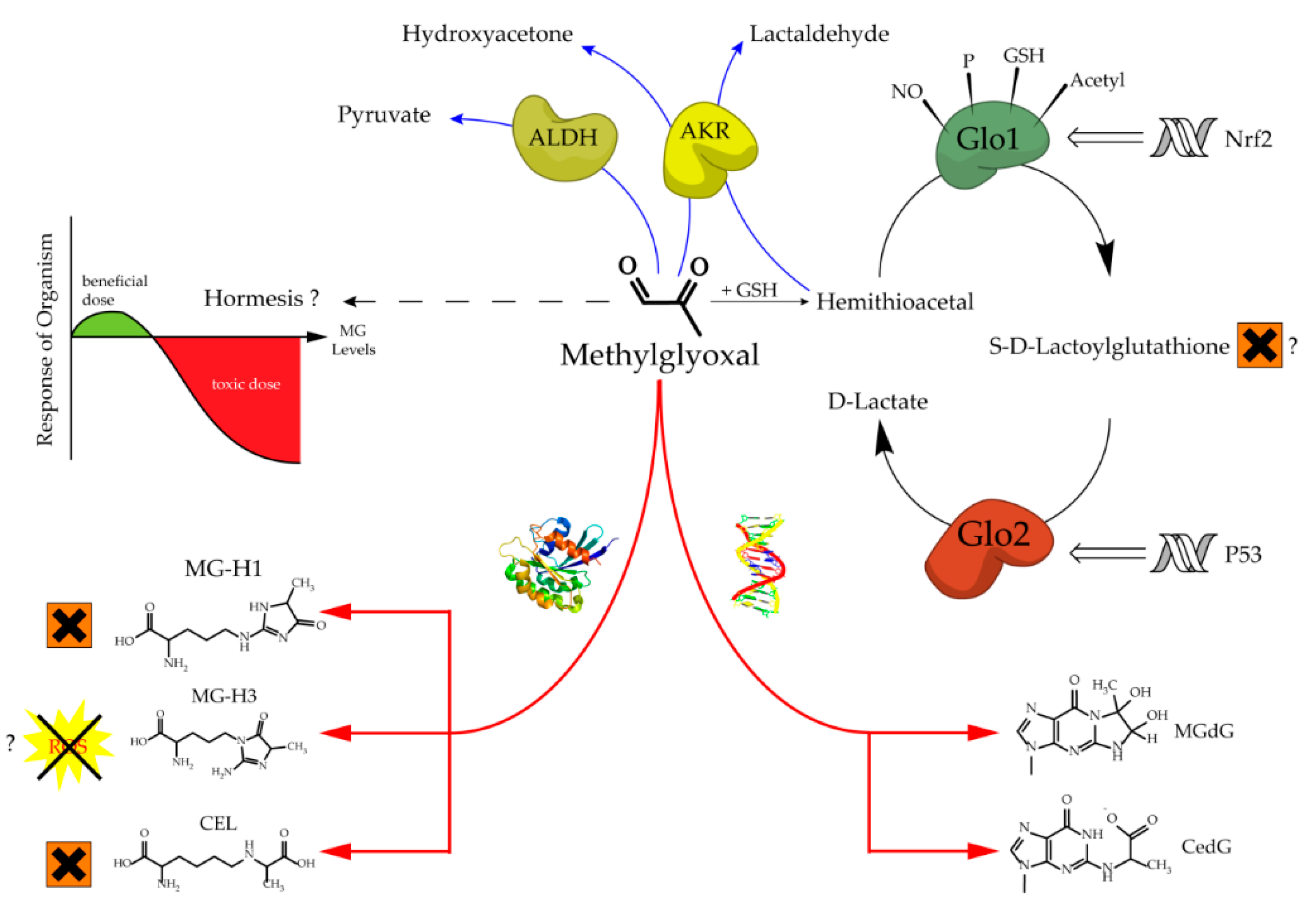
Figure 1. Aspects of methylglyoxal metabolism and its possible implications.
The regulation of Glo1 activity and expression is complex and still not well understood. However, what we know is that Glo1 has a metal responsive element, an insulin-responsive element, an antioxidant responsive element, and it is a hotspot of copy number variation [28][29][30]. Downregulation of Glo1 via hyperglycemic or hypoxic conditions induces MG stress, whereas an upregulation via Nrf2 increases Glo1 activity and therefore can alleviate intracellular MG stress [29][31]. Many experimental data suggest that the modulation of Glo1 has a high impact towards the phenotype of healthy cells exposed to cellular stress, and also of malignant cells with an increased energy demand [7][15][17][32][33][34][35]. In cruder model organisms for instance, such as Caenorhabditis elegans (C. elegans), an overexpression of Glo1 results in an increase in mean and maximum lifespan by ~40% [36]. In bovine endothelial cells, the overexpression of Glo1 reduces the intracellular accumulation of AGEs under hyperglycemic conditions [37]. Consistently, overexpression in rat models of diabetes was reported to be protective regarding ateriogenesis and renal impairment driven by microvascular alterations [35][38]. On the other hand, the partial loss of Glo1 is associated with detrimental kidney damage in a murine Glo1 knock-down model [39]. Hyperglycemic episodes and therefore diabetes related late complications, such as nephro-, retino- and neuropathy, have been frequently linked to increased MG levels due to a lower Glo1 expression in humans [14][17][18][40]. Furthermore, in a recent clinical trial, a Glo1 inducer (resveratrol-hesperetin) improved metabolic and vascular health in a small cohort of overweight and obese humans [41]. Last but not least, the naturally occurring decrease in Glo1 during aging seems to be associated with an increased risk of age-related cardiovascular diseases [42].
As already mentioned, in many types of cancer the situation is quite different. Due to the vast amounts of glucose consumption resulting from the increased proliferation rates of most tumors, Glo1 is necessary to protect the cell from harmful MG. In malignant cells with a high proliferation rate, it is undoubtedly proven that Glo1 protects those cells from the increased formation of MG due to a high glycolytic flux within the context of the Warburg effect [43][44][45]. Therefore, it is no surprise that Glo1 is generally overexpressed in numerous cancers potentially as a part of a survival strategy [15]. This is the case in urological malignancies, different breast cancer types, gastric cancer cells, bladder, colon and hepatocellular carcinomas as well as leukemia [46][47][48][49][50][51][52][53].
In prostate cancer, Glo1 is also linked to the maintenance of the metastatic phenotype by controlling the epithelial to mesenchymal transition [54]. The association of Glo1 with tumor growth could suggest its role as an oncogene, but this seems to be more an adaption to protect the tumor proteome against an increased intracellular flux of MG formation [55]. The inhibition of Glo1, therefore, may represent a potential target for anticancer therapeutics, although compounds such as bromobenzylglutathione cyclopentyl ester, which has been used in vitro, have to prove their therapeutic capacity in humans also [56].
2. Glo1 is not Indispensable for Crude and Complex Organisms
Due to its highly conserved role, the glyoxalase system has been viewed as a crucial enzyme system to maintain cellular viability [6][8][14]. This is a consequence of the necessity to detoxify the highly reactive dicarbonyl MG as fast as possible, in order to prevent glycation of proteins and DNA [7]. This has been proven in plants, crude prokaryotic microorganisms and many animals such as nematodes, arthropods, as well as chordates [4][5][36][57][58][59].
Unfortunately, the first attempts to abolish Glo1 activity failed, due to a high amount of copy number variants using the approach of gene trapping mutations [67]. The first global Glo1 KO in mice was established in 2017 [68]. Surprisingly, the effect of the complete global Glo1 KO was rather mild, at least in fish and mammals. Obviously, compensatory pathways for the glyoxalase system were underestimated in earlier studies. This is supported by the findings that in mammalian models a complete loss of Glo1 does not result in basal MG and MG-H1 elevations, even under high glucose conditions [72][74]. It has to be pointed out that most of the in vitro studies about MG-derived AGEs and DNA adducts had to use exogenously added MG in a supraphysiological range in order to achieve obvious molecular effects [64][72][75][76][77]. Certainly, Glo1 KO studies can be contradictory and provoking to the scientific dogma that silencing or inhibiting Glo1 has detrimental effects. Previously, this merged into the assumption of a cause-and-effect model between Glo1 activity and the levels of MG. If this is true, diseases associated with a complete inherited loss of Glo1 and therefore increased MG levels would have been described. As of yet, no known diseases have been attributed to Glo1 mutations in humans.
One reason for the development of late diabetic complications is the increase in MG-derived AGEs and DNA modifications, which is mainly due to the fact that Glo1 is downregulated in diabetes. Consequently, MG cannot be detoxified efficiently [14][20][21][34][77]. If this were true, then a diabetic Glo1 KO model would reflect a heavy damage of tissues which are prone to hyperglycemic conditions, such as kidneys, nerves and the eyes. However, recent data support the idea that the development of late diabetic complications due to increased AGEs and MG-derived DNA damage cannot be a result of one single downregulated enzyme. The data suggest that it is a multifactorial enzymatic system of high complexity, which is responsible for the detoxification of increased amounts of MG.
In summary, and to put the new findings into context, various in vivo studies of Glo1 KO models show that Glo1 is not an indispensable enzyme and thus, the glyoxalase system is neither. It is also a fascinating example that with increasing complexity and advanced evolution, nature may establish a backup system, which can compensate and substitute genetic mutations of highly conserved enzymes. This may not be perfectly the case in crude organisms, such as yeast and worm, but to some degree in flies and fish, and finally with perfection in a mammalian organism (Figure 2). This leads consequently to the question of why the glyoxalase system is not mandatory for a complex organism, despite it being a highly conserved and abundant system. The next part of this review tries to find answers to that essential question.
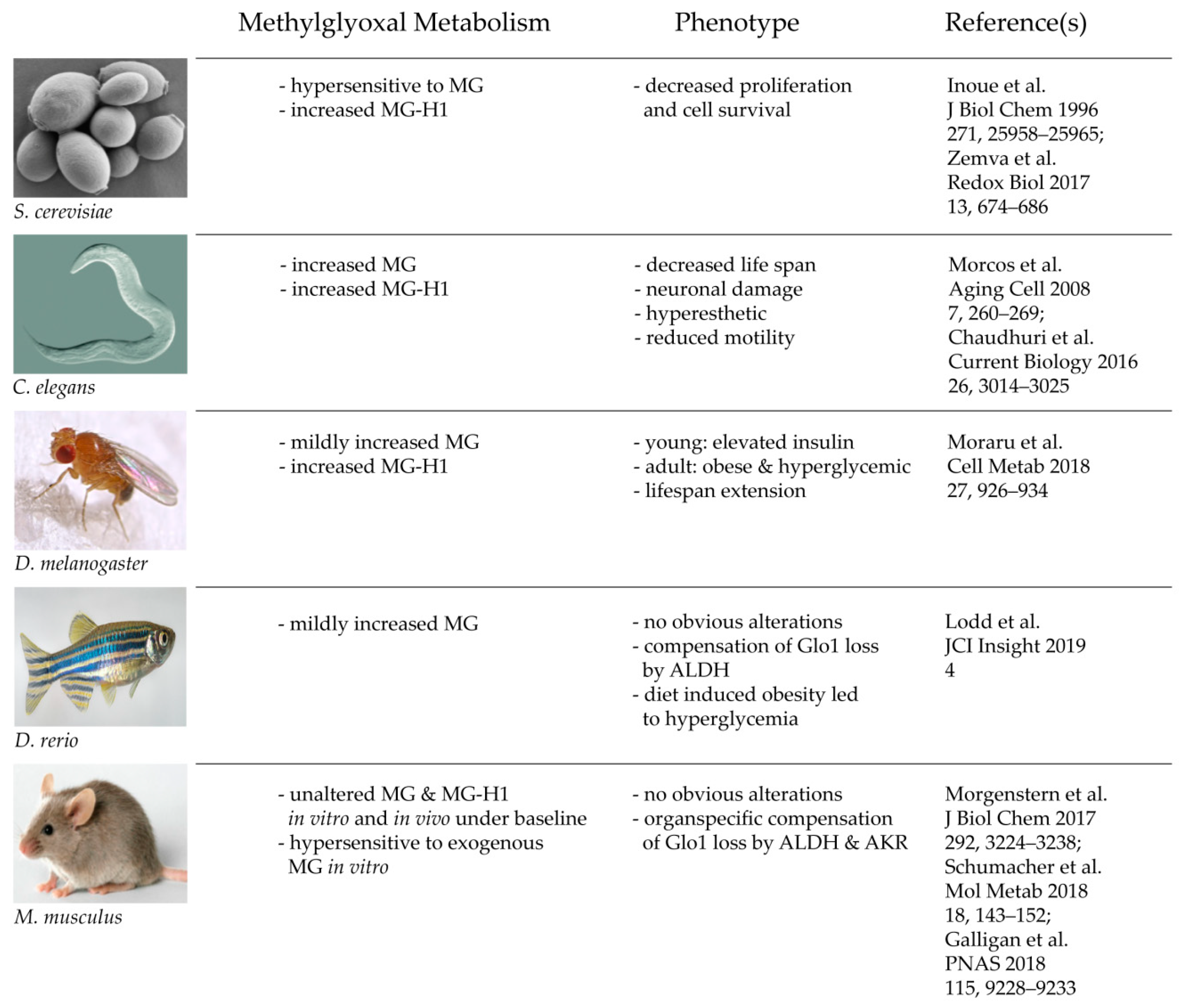
Figure 2. Consequences of a total loss of glyoxalase 1 in various model organisms.
3. Modifications of Glo1 and Other Possible Intracellular Implications
Despite the extensive research on the glyoxalase system, and in particular Glo1, investigating PTMs of Glo1 in mammals have not been a focus in the field. Studies regarding the PTMs of Glo1 are rare and the consequences of those are unfortunately not well understood [77]. However, PTMs of Glo1 have been described in yeast and plants, which suggest that such regulatory mechanisms are as highly conserved as the glyoxalase system itself.
The most studied PTM of Glo1 is the phosphorylation. There are five different putative phosphorylation sites identified for Glo1, whereas Thr107 is the only one which has been identified as the one with physiological consequences [6][84]. A recent study was able to shed more light on the molecular consequences of the phosphorylation of Thr107. It was revealed by a pharmacological and genetic approach that the driving kinase is Ca2+/Calmodulin-dependent Kinase II delta (CamKIIδ) [87]. Moreover, phosphorylated Glo1 has an elevated catalytic efficiency (lower Km; higher Vmax) and it is also protected from rapid proteasomal degradation by ubiquitination [87]. Given the background that other studies have shown that an acetylation enforces the proteasomal degradation, it is a valid hypothesis to claim that the phosphoryl group could prevent the acetylation of Glo1 in a competitive way [85]. Interestingly, the reduced state of Glo1-phosphorylation, and therefore reduced Glo1 activity in vitro and in vivo, was associated with only mild changes regarding the phenotype. Only MG-specific DNA adducts (MG-dG) were increased and this was linked to increased nuclear damage markers such as p53 and γH2Ax. This effect was more pronounced in isolated murine endothelial cells as compared to the in vivo mouse model [87]. An important finding of this recent study was the association of phosphorylated Glo1 with diabetes, ageing and malignant cells. In fact, decreased Glo1 phosphorylation has been associated with a decline in Glo1 activity in diabetes or ageing, whereas the opposite effect seems to take place in human tumor cells, in which Glo1 activity is usually upregulated [87]. This could provide an explanation for the altered Glo1 status in different intracellular environments, described by many other studies [7][15][16][17][27][28][33][34][41]. Although these recent findings regarding the driving kinase of a Glo1 phosphorylation only scratch the surface of the underlying mechanisms, it might help to clarify the precise role of the glyoxalase system and Glo1 in particular. At this point it cannot be excluded that there are other kinases too, besides CamKIIδ, which are able to phosphorylate Glo1 efficiently. For instance, it has been shown that TNFα can induce multiple phosphorylation events at Glo1 [88].
References
- Dakin, H.D.; Dudley, H.W. AN ENZYME CONCERNED WITH THE FORMATION OF HYDROXY ACIDS FROM KETONIC ALDEHYDES. J. Biol. Chem. 1913, 14, 155–157.
- Thornalley, P.J. The glyoxalase system in health and disease. Mol. Asp. Med. 1993, 14, 287–371, doi:10.1016/0098-2997(93)90002-U.
- Kaur, C.; Sharma, S.; Hasan, M.; Pareek, A.; Singla-Pareek, S.; Sopory, S. Characteristic variations and similarities in biochemical, molecular, and functional properties of glyoxalases across prokaryotes and eukaryotes. Int. J. Mol. Sci. 2017, 18, 250, doi:10.3390/ijms18040250.
- Jain, M.; Batth, R.; Kumari, S.; Mustafiz, A. Arabidopsis thaliana contains both Ni2+ and Zn2+ dependent glyoxalase I enzymes and ectopic expression of the latter contributes more towards abiotic stress tolerance in E. coli. PLoS ONE 2016, 11, e0159348, doi:10.1371/journal.pone.0159348.
- Allaman, I.; Bélanger, M.; Magistretti, P.J. Methylglyoxal, the dark side of glycolysis. Front. Neurosci. 2015, 9, 23, doi:10.3389/fnins.2015.00023.
- Thornalley, P.J. Glyoxalase I—Structure, function and a critical role in the enzymatic defence against glycation. Biochem. Soc. Trans. 2003, 31, 1343–1348, doi:10.1042/bst0311343.
- Nigro, C.; Leone, A.; Raciti, G.; Longo, M.; Mirra, P.; Formisano, P.; Beguinot, F.; Miele, C. Methylglyoxal-glyoxalase 1 balance: The root of vascular damage. Int. J. Mol. Sci. 2017, 18, 188, doi:10.3390/ijms18010188.
- Silva, M.S.; Gomes, R.; Ferreira, A.; Freire, A.P.; Cordeiro, C.A. The glyoxalase pathway: The first hundred years… and beyond. Biochem. J. 2013, 453, 1–15, doi:10.1042/BJ20121743.
- Cameron, A.D.; Ridderström, M.; Olin, B.; Mannervik, B. Crystal structure of human glyoxalase II and its complex with a glutathione thiolester substrate analogue. Structure 1999, 7, 1067–1078, doi:10.1016/s0969-2126(99)80174-9.
- Thornalley, P.J.; Tisdale, M.J. Inhibition of proliferation of human promyelocytic leukaemia HL60 cells by S-D-lactoylglutathione in vitro. Leuk. Res. 1988, 12, 897–904, doi:10.1016/0145-2126(88)90016-1.
- Xu, Y.; Chen, X. Glyoxalase II, a detoxifying enzyme of glycolysis byproduct methylglyoxal and a target of p63 and p73, is a pro-survival factor of the p53 family. J. Biol. Chem. 2006, 281, 26702–26713, doi:10.1074/jbc.M604758200.
- Antognelli, C.; Frosini, R.; Santolla, M.F.; Peirce, M.J.; Talesa, V.N. Oleuropein-induced apoptosis is mediated by mitochondrial glyoxalase 2 in NSCLC A549 cells: A mechanistic inside and a possible novel nonenzymatic role for an ancient enzyme. Oxidative Med. Cell. Longev. 2019, 2019, 8576961, doi:10.1155/2019/8576961.
- Antognelli, C.; Ferri, I.; Bellezza, G.; Siccu, P.; Love, H.D.; Talesa, V.N.; Sidoni, A. Glyoxalase 2 drives tumorigenesis in human prostate cells in a mechanism involving androgen receptor and p53-p21 axis. Mol. Carcinog. 2017, 56, 2112–2126, doi:10.1002/mc.22668.
- Hidmark, A.; Fleming, T.; Vittas, S.; Mendler, M.; Deshpande, D.; Groener, J.B.; Müller, B.P.; Reeh, P.W.; Sauer, S.K.; Pham, M.; et al. A new paradigm to understand and treat diabetic neuropathy. Exp. Clin. Endocrinol. Diabetes 2014, 122, 201–207, doi:10.1055/s-0034-1367023.
- Bellahcène, A.; Nokin, M.-J.; Castronovo, V.; Schalkwijk, C. Methylglyoxal-derived stress: An emerging biological factor involved in the onset and progression of cancer. Semin. Cancer Biol. 2018, 49, 64–74, doi:10.1016/j.semcancer.2017.05.010.
- Hanssen, N.M.J.; Stehouwer, C.D.A.; Schalkwijk, C.G. Methylglyoxal and glyoxalase I in atherosclerosis. Biochem. Soc. Trans. 2014, 42, 443–449, doi:10.1042/BST20140001.
- Maessen, D.E.M.; Stehouwer, C.D.A.; Schalkwijk, C.G. The role of methylglyoxal and the glyoxalase system in diabetes and other age-related diseases. Clin. Sci. 2015, 128, 839–861, doi:10.1042/CS20140683.
- Matafome, P.; Sena, C.; Seiça, R. Methylglyoxal, obesity, and diabetes. Endocrine 2013, 43, 472–484, doi:10.1007/s12020-012-9795-8.
- Srikanth, V.; Westcott, B.; Forbes, J.; Phan, T.G.; Beare, R.; Venn, A.; Pearson, S.; Greenaway, T.; Parameswaran, V.; Münch, G. Methylglyoxal, cognitive function and cerebral atrophy in older people. J. Gerontol. Ser. A 2013, 68, 68–73, doi:10.1093/gerona/gls100.
- Thornalley, P.J.; Langborg, A.; Minhas, H.S. Formation of glyoxal, methylglyoxal and 3-deoxyglucosone in the glycation of proteins by glucose. Biochem. J. 1999, 344, 109–116, doi:10.1042/bj3440109.
- Thornalley, P.J.; Battah, S.; Ahmed, N.; Karachalias, N.; Agalou, S.; Babaei-Jadidi, R.; Dawnay, A. Quantitative screening of advanced glycation endproducts in cellular and extracellular proteins by tandem mass spectrometry. Biochem. J. 2003, 375, 581–592, doi:10.1042/BJ20030763.
- Ahmed, N.; Thornalley, P.J.; Dawczynski, J.; Franke, S.; Strobel, J.; Stein, G.; Haik, G.M. Methylglyoxal-derived hydroimidazolone advanced glycation end-products of human lens proteins. Investig. Opthalmol. Vis. Sci. 2003, 44, 5287, doi:10.1167/iovs.03-0573.
- Ahmed, N.; Dobler, D.; Dean, M.; Thornalley, P.J. Peptide mapping identifies hotspot site of modification in human serum albumin by methylglyoxal involved in ligand binding and esterase activity. J. Biol. Chem. 2005, 280, 5724–5732, doi:10.1074/jbc.M410973200.
- Ahmed, N.; Argirov, O.K.; Minhas, H.S.; Cordeiro, C.A.A.; Thornalley, P.J. Assay of advanced glycation endproducts (AGEs): Surveying AGEs by chromatographic assay with derivatization by 6-aminoquinolyl-N-hydroxysuccinimidyl-carbamate and application to N∊-carboxymethyl-lysine- and N∊-(1-carboxyethyl)lysine-modified albumin. Biochem. J. 2002, 364, 1–14, doi:10.1042/bj3640001.
- Takahashi, K. The reactions of phenylglyoxal and related reagents with amino acids. J. Biochem. 1977, 81, 395–402, doi:10.1093/oxfordjournals.jbchem.a131471.
- Schneider, M.; Thoss, G.; Hübner-Parajsz, C.; Kientsch-Engel, R.; Stahl, P.; Pischetsrieder, M. Determination of glycated nucleobases in human urine by a new monoclonal antibody specific for N 2 -Carboxyethyl-2‘-deoxyguanosine. Chem. Res. Toxicol. 2004, 17, 1385–1390, doi:10.1021/tx049929d.
- Thornalley, P.J.; Waris, S.; Fleming, T.; Santarius, T.; Larkin, S.J.; Winklhofer-Roob, B.M.; Stratton, M.R.; Rabbani, N. Imidazopurinones are markers of physiological genomic damage linked to DNA instability and glyoxalase 1-associated tumour multidrug resistance. Nucleic Acids Res. 2010, 38, 5432–5442, doi:10.1093/nar/gkq306.
- Ranganathan, S.; Ciaccio, P.J.; Walsh, E.S.; Tew, K.D. Genomic sequence of human glyoxalase-I: Analysis of promoter activity and its regulation. Gene 1999, 240, 149–155, doi:10.1016/S0378-1119(99)00420-5.
- Xue, M.; Rabbani, N.; Momiji, H.; Imbasi, P.; Anwar, M.M.; Kitteringham, N.; Park, B.K.; Souma, T.; Moriguchi, T.; Yamamoto, M.; et al. Transcriptional control of glyoxalase 1 by Nrf2 provides a stress-responsive defence against dicarbonyl glycation. Biochem. J. 2012, 443, 213–222, doi:10.1042/BJ20111648.
- Shafie, A.; Xue, M.; Thornalley, P.J.; Rabbani, N. Copy number variation of glyoxalase I. Biochem. Soc. Trans. 2014, 42, 500–503, doi:10.1042/BST20140011.
- Zhang, H.; Li, H.; Xi, H.S.; Li, S. HIF1α is required for survival maintenance of chronic myeloid leukemia stem cells. Blood 2012, 119, 2595–2607, doi:10.1182/blood-2011-10-387381.
- Hutschenreuther, A.; Bigl, M.; Hemdan, N.; Debebe, T.; Gaunitz, F.; Birkenmeier, G. Modulation of GLO1 expression affects malignant properties of cells. Int. J. Mol. Sci. 2016, 17, 2133, doi:10.3390/ijms17122133.
- Peters, A.S.; Lercher, M.; Fleming, T.H.; Nawroth, P.P.; Bischoff, M.S.; Dihlmann, S.; Böckler, D.; Hakimi, M. Reduced glyoxalase 1 activity in carotid artery plaques of nondiabetic patients with increased hemoglobin A1c level. J. Vasc. Surg. 2016, 64, 990–994, doi:10.1016/j.jvs.2016.04.025.
- Rabbani, N.; Thornalley, P.J. Glyoxalase 1 modulation in obesity and diabetes. Antioxid. Redox Signal. 2019, 30, 354–374, doi:10.1089/ars.2017.7424.
- Brouwers, O.; Niessen, P.M.; Ferreira, I.; Miyata, T.; Scheffer, P.G.; Teerlink, T.; Schrauwen, P.; Brownlee, M.; Stehouwer, C.D.; Schalkwijk, C.G. Overexpression of glyoxalase-I reduces hyperglycemia-induced levels of advanced glycation end products and oxidative stress in diabetic rats. J. Biol. Chem. 2011, 286, 1374–1380, doi:10.1074/jbc.M110.144097.
- Morcos, M.; Du, X.; Pfisterer, F.; Hutter, H.; Sayed, A.A.R.; Thornalley, P.; Ahmed, N.; Baynes, J.; Thorpe, S.; Kukudov, G.; et al. Glyoxalase-1 prevents mitochondrial protein modification and enhances lifespan in Caenorhabditis elegans. Aging Cell 2008, 7, 260–269, doi:10.1111/j.1474-9726.2008.00371.x.
- Shinohara, M.; Thornalley, P.J.; Giardino, I.; Beisswenger, P.; Thorpe, S.R.; Onorato, J.; Brownlee, M. Overexpression of glyoxalase-I in bovine endothelial cells inhibits intracellular advanced glycation endproduct formation and prevents hyperglycemia-induced increases in macromolecular endocytosis. J. Clin. Investig. 1998, 101, 1142–1147, doi:10.1172/JCI119885.
- Brouwers, O.; Yu, L.; Niessen, P.; Slenter, J.; Jaspers, K.; Wagenaar, A.; Post, M.; Miyata, T.; Backes, W.; Stehouwer, C.; et al. Glyoxalase-1 overexpression partially prevents diabetes-induced impaired arteriogenesis in a rat hindlimb ligation model. Glycoconj. J. 2016, 33, 627–630, doi:10.1007/s10719-016-9681-3.
- Giacco, F.; Du, X.; D’Agati, V.D.; Milne, R.; Sui, G.; Geoffrion, M.; Brownlee, M. Knockdown of glyoxalase 1 mimics diabetic nephropathy in nondiabetic mice. Diabetes 2014, 63, 291–299, doi:10.2337/db13-0316.
- Rabbani, N.; Thornalley, P.J. The critical role of methylglyoxal and glyoxalase 1 in diabetic nephropathy. Diabetes 2014, 63, 50–52, doi:10.2337/db13-1606.
- Xue, M.; Weickert, M.O.; Qureshi, S.; Kandala, N.-B.; Anwar, A.; Waldron, M.; Shafie, A.; Messenger, D.; Fowler, M.; Jenkins, G.; et al. Improved glycemic control and vascular function in overweight and obese subjects by glyoxalase 1 inducer formulation. Diabetes 2016, 65, 2282–2294, doi:10.2337/db16-0153.
- Xue, M.; Rabbani, N.; Thornalley, P.J. Glyoxalase in ageing. Semin. Cell Dev. Biol. 2011, 22, 293–301, doi:10.1016/j.semcdb.2011.02.013.
- Santarius, T.; Bignell, G.R.; Greenman, C.D.; Widaa, S.; Chen, L.; Mahoney, C.L.; Butler, A.; Edkins, S.; Waris, S.; Thornalley, P.J.; et al. GLO1-A novel amplified gene in human cancer. Genes Chromosomes Cancer 2010, 49, 711–725, doi:10.1002/gcc.20784.
- Heiden, M.G.V.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science 2009, 324, 1029–1033, doi:10.1126/science.1160809.
- Ward, P.S.; Thompson, C.B. Metabolic reprogramming: A cancer hallmark even warburg did not anticipate. Cancer Cell 2012, 21, 297–308, doi:10.1016/j.ccr.2012.02.014.
- Antognelli, C.; Talesa, V.N. Glyoxalases in urological malignancies. Int. J. Mol. Sci. 2018, 19, 415, doi:10.3390/ijms19020415.
- Yang, Y.-X.; Chen, Z.-C.; Zhang, G.-Y.; Yi, H.; Xiao, Z.-Q. A subcelluar proteomic investigation into vincristine-resistant gastric cancer cell line. J. Cell. Biochem. 2008, 104, 1010–1021, doi:10.1002/jcb.21687.
- Nakamura, T.; Furukawa, Y.; Nakagawa, H.; Tsunoda, T.; Ohigashi, H.; Murata, K.; Ishikawa, O.; Ohgaki, K.; Kashimura, N.; Miyamoto, M.; et al. Genome-wide cDNA microarray analysis of gene expression profiles in pancreatic cancers using populations of tumor cells and normal ductal epithelial cells selected for purity by laser microdissection. Oncogene 2004, 23, 2385–2400, doi:10.1038/sj.onc.1207392.
- Rulli, A.; Carli, L.; Romani, R.; Baroni, T.; Giovannini, E.; Rosi, G.; Talesa, V. Expression of glyoxalase I and II in normal and breast cancer tissues. Breast Cancer Res. Treat. 2001, 66, 67–72, doi:10.1023/A:1010632919129.
- Zhang, D.; Tai, L.K.; Wong, L.L.; Chiu, L.-L.; Sethi, S.K.; Koay, E.S.C. Proteomic study reveals that proteins involved in metabolic and detoxification pathways are highly expressed in HER-2/neu -positive breast cancer*. Mol. Cell. Proteom. 2005, 4, 1686–1696, doi:10.1074/mcp.M400221-MCP200.
- Mearini, E.; Romani, R.; Mearini, L.; Antognelli, C.; Zucchi, A.; Baroni, T.; Porena, M.; Talesa, V.. Differing expression of enzymes of the glyoxalase system in superficial and invasive bladder carcinomas. Eur. J. Cancer 2002, 38, 1946–1950, doi:10.1016/S0959-8049(02)00236-8.
- Song, H.-Y.; Liu, Y.-K.; Feng, J.-T.; Cui, J.-F.; Dai, Z.; Zhang, L.-J.; Feng, J.-X.; Shen, H.-L.; Tang, Z.-Y. Proteomic analysis on metastasis-associated proteins of human hepatocellular carcinoma tissues. J. Cancer Res. Clin. Oncol. 2006, 132, 92–98, doi:10.1007/s00432-005-0044-x.
- Cui, J.-W. Proteomic analysis of human acute leukemia cells: Insight into their classification. Clin. Cancer Res. 2004, 10, 6887–6896, doi:10.1158/1078-0432.CCR-04-0307.
- Antognelli, C.; Mezzasoma, L.; Mearini, E.; Talesa, V.N. Glyoxalase 1-419C>A variant is associated with oxidative stress: Implications in prostate cancer progression. PLoS ONE 2013, 8, e74014, doi:10.1371/journal.pone.0074014.
- Rabbani, N.; Xue, M.; Weickert, M.O.; Thornalley, P.J. Multiple roles of glyoxalase 1-mediated suppression of methylglyoxal glycation in cancer biology-Involvement in tumour suppression, tumour growth, multidrug resistance and target for chemotherapy. Semin. Cancer Biol. 2018, 49, 83–93, doi:10.1016/j.semcancer.2017.05.006.
- Thornalley, P.J.; Edwards, L.G.; Kang, Y.; Wyatt, C.; Davies, N.; Ladan, M.J.; Double, J. Antitumour activity of S-p-bromobenzylglutathione cyclopentyl diester in vitro and in vivo. Inhibition of glyoxalase I and induction of apoptosis. Biochem. Pharmacol. 1996, 51, 1365–1372, doi:10.1016/0006-2952(96)00059-7.
- Jörgens, K.; Stoll, S.J.; Pohl, J.; Fleming, T.H.; Sticht, C.; Nawroth, P.P.; Hammes, H.-P.; Kroll, J. High tissue glucose alters intersomitic blood vessels in zebrafish via methylglyoxal targeting the VEGF receptor signaling cascade. Diabetes 2015, 64, 213–225, doi:10.2337/db14-0352.
- Urscher, M.; Alisch, R.; Deponte, M. The glyoxalase system of malaria parasites—Implications for cell biology and general glyoxalase research. Semin. Cell Dev. Biol. 2011, 22, 262–270, doi:10.1016/j.semcdb.2011.02.003.
- Garrido, D.; Rubin, T.; Poidevin, M.; Maroni, B.; Le Rouzic, A.; Parvy, J.-P.; Montagne, J. Fatty acid synthase cooperates with glyoxalase 1 to protect against sugar toxicity. PLoS Genet. 2015, 11, e1004995, doi:10.1371/journal.pgen.1004995.
- Ghosh, A.; Kushwaha, H.R.; Hasan, M.R.; Pareek, A.; Sopory, S.K.; Singla-Pareek, S.L. Presence of unique glyoxalase III proteins in plants indicates the existence of shorter route for methylglyoxal detoxification. Sci. Rep. 2016, 6, 18358, doi:10.1038/srep18358.
- Jagt, D.L.V.; A Hunsaker, L. Methylglyoxal metabolism and diabetic complications: Roles of aldose reductase, glyoxalase-I, betaine aldehyde dehydrogenase and 2-oxoaldehyde dehydrogenase. Chem. Biol. Interact. 2003, 143–144, 341–351.
- Jagt, D.L.V.; Hassebrook, R.K.; Hunsaker, L.A.; Brown, W.M.; Royer, R.E. Metabolism of the 2-oxoaldehyde methylglyoxal by aldose reductase and by glyoxalase-I: Roles for glutathione in both enzymes and implications for diabetic complications. Chem. Biol. Interact. 2001, 130–132, 549–562, doi:10.1016/S0009-2797(00)00298-2.
- Richarme, G.; Mihoub, M.; Dairou, J.; Bui, L.C.; Leger, T.; Lamouri, A. Parkinsonism-associated protein DJ-1/Park7 is a major protein deglycase that repairs methylglyoxal- and glyoxal-glycated cysteine, arginine, and lysine residues. J. Biol. Chem. 2015, 290, 1885–1897, doi:10.1074/jbc.M114.597815.
- Galligan, J.J.; Wepy, J.A.; Streeter, M.D.; Kingsley, P.J.; Mitchener, M.M.; Wauchope, O.R.; Beavers, W.N.; Rose, K.L.; Wang, T.; Spiegel, D.A.; et al. Methylglyoxal-derived posttranslational arginine modifications are abundant histone marks. Proc. Natl. Acad. Sci. USA 2018, 115, 9228–9233, doi:10.1073/pnas.1802901115.
- Baba, S.P.; Barski, O.A.; Ahmed, Y.; O’Toole, T.E.; Conklin, D.J.; Bhatnagar, A.; Srivastava, S. Reductive metabolism of AGE precursors: A metabolic route for preventing AGE accumulation in cardiovascular tissue. Diabetes 2009, 58, 2486–2497, doi:10.2337/db09-0375.
- Wortmann, M.; Hakimi, M.; Fleming, T.; Peters, A.S.; Sijmonsma, T.P.; Herzig, S.; Nawroth, P.P.; Böckler, D.; Dihlmann, S.A. Glyoxalase-1 knockdown does not have major short term effects on energy expenditure and atherosclerosis in mice. J. Diabetes Res. 2016, 2016, 1–8, doi:10.1155/2016/2981639.
- Shafie, A.; Xue, M.; Barker, G.; Zehnder, D.; Thornalley, P.J.; Rabbani, N. Reappraisal of putative glyoxalase 1-deficient mouse and dicarbonyl stress on embryonic stem cells in vitro. Biochem. J. 2016, 473, 4255–4270, doi:10.1042/BCJ20160691.
- Jang, S.; Kwon, D.M.; Kwon, K.; Park, C. Generation and characterization of mouse knockout for glyoxalase 1. Biochem. Biophys. Res. Commun. 2017, 490, 460–465, doi:10.1016/j.bbrc.2017.06.063.
- Ran, F.A.; Hsu, P.D.; Wright, J.; Agarwala, V.; Scott, D.A.; Zhang, F. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 2013, 8, 2281–2308, doi:10.1038/nprot.2013.143.
- Lodd, E.; Wiggenhauser, L.M.; Morgenstern, J.; Fleming, T.H.; Poschet, G.; Büttner, M.; Tabler, C.T.; Wohlfart, D.P.; Nawroth, P.P.; Kroll, J. The combination of loss of glyoxalase1 and obesity results in hyperglycemia. JCI Insight 2019, 4, doi:10.1172/jci.insight.126154.
- Moraru, A.; Wiederstein, J.; Pfaff, D.; Fleming, T.; Miller, A.K.; Nawroth, P.; Teleman, A.A. Elevated levels of the reactive metabolite methylglyoxal recapitulate progression of type 2 diabetes. Cell Metab. 2018, 27, 926–934.e8, doi:10.1016/j.cmet.2018.02.003.
- Morgenstern, J.; Fleming, T.; Schumacher, D.; Eckstein, V.; Freichel, M.; Herzig, S.; Nawroth, P. Loss of glyoxalase 1 induces compensatory mechanism to achieve dicarbonyl detoxification in mammalian schwann cells. J. Biol. Chem. 2017, 292, 3224–3238, doi:10.1074/jbc.M116.760132.
- Schemmel, K.E.; Padiyara, R.S.; D’Souza, J.J. Aldose reductase inhibitors in the treatment of diabetic peripheral neuropathy: A review. J. Diabetes Its Complicat. 2010, 24, 354–360, doi:10.1016/j.jdiacomp.2009.07.005.
- Schumacher, D.; Morgenstern, J.; Oguchi, Y.; Volk, N.; Kopf, S.; Groener, J.B.; Nawroth, P.P.; Fleming, T.; Freichel, M. Compensatory mechanisms for methylglyoxal detoxification in experimental & clinical diabetes. Mol. Metab. 2018, 18, 143–152, doi:10.1016/j.molmet.2018.09.005.
- Di Emidio, G.; Santini, S.J.; D’Alessandro, A.M.; Vetuschi, A.; Sferra, R.; Artini, P.G.; Carta, G.; Falone, S.; Amicarelli, F.; Tatone, C. SIRT1 participates in the response to methylglyoxal-dependent glycative stress in mouse oocytes and ovary. Biochim. Biophys. Acta (BBA)—Mol. Basis Dis. 2019, 1865, 1389–1401, doi:10.1016/j.bbadis.2019.02.011.
- Roy, A.; Sarker, S.; Upadhyay, P.; Pal, A.; Adhikary, A.; Jana, K.; Ray, M. Methylglyoxal at metronomic doses sensitizes breast cancer cells to doxorubicin and cisplatin causing synergistic induction of programmed cell death and inhibition of stemness. Biochem. Pharmacol. 2018, 156, 322–339, doi:10.1016/j.bcp.2018.08.041.
- Schalkwijk, C.G.; Stehouwer, C.D.A. Methylglyoxal, a highly reactive dicarbonyl compound, in diabetes, its vascular complications, and other age-related diseases. Physiol. Rev. 2020, 100, 407–461, doi:10.1152/physrev.00001.2019.
- Shin, R.; Alvarez, S.; Burch, A.Y.; Jez, J.M.; Schachtman, D.P. Phosphoproteomic identification of targets of the Arabidopsis sucrose nonfermenting-like kinase SnRK2.8 reveals a connection to metabolic processes. Proc. Natl. Acad. Sci. USA 2007, 104, 6460–6465, doi:10.1073/pnas.0610208104.
- Khan, M.M.K.; Jan, A.; Karibe, H.; Komatsu, S. Identification of phosphoproteins regulated by gibberellin in rice leaf sheath. Plant Mol. Biol. 2005, 58, 27–40, doi:10.1007/s11103-005-4013-1.
- Sankaranarayanan, S.; Jamshed, M.; Kumar, A.; Skori, L.; Scandola, S.; Wang, T.; Spiegel, D.; Samuel, M. Glyoxalase goes green: The expanding roles of glyoxalase in plants. Int. J. Mol. Sci. 2017, 18, 898, doi:10.3390/ijms18040898.
- Proietti, S.; Falconieri, G.S.; Bertini, L.; Baccelli, I.; Paccosi, E.; Belardo, A.; Timperio, A.M.; Caruso, C. GLYI4 plays A role in methylglyoxal detoxification and jasmonate-mediated stress responses in arabidopsis thaliana. Biomolecules 2019, 9, 635, doi:10.3390/biom9100635.
- Inoue, Y.; Choi, B.Y.; Murata, K.; Kimura, A. Sexual response of Saccharomyces cerevisiae: Phosphorylation of yeast glyoxalase I by a cell extract of mating factor-treated cells. J. Biochem. 1990, 108, 4–6, doi:10.1093/oxfordjournals.jbchem.a123159.
- Mitsumoto, A.; Kim, K.-R.; Oshima, G.; Kunimoto, M.; Okawa, K.; Iwamatsu, A.; Nakagawa, Y. Nitric oxide inactivates glyoxalase I in cooperation with glutathione. J. Biochem. 2000, 128, 647–654, doi:10.1093/oxfordjournals.jbchem.a022797.
- Birkenmeier, G.; Stegemann, C.; Hoffmann, R.; Günther, R.; Huse, K.; Birkemeyer, C. Posttranslational modification of human glyoxalase 1 indicates redox-dependent regulation. PLoS ONE 2010, 5, e10399, doi:10.1371/journal.pone.0010399.
- Spanos, C.; Maldonado, E.M.; Fisher, C.P.; Leenutaphong, P.; Oviedo-Orta, E.; Windridge, D.; Salguero, F.J.; Bermúdez-Fajardo, A.; Weeks, M.E.; Evans, C.; et al. Proteomic identification and characterization of hepatic glyoxalase 1 dysregulation in non-alcoholic fatty liver disease. Proteome Sci. 2018, 16, 1–16, doi:10.1186/s12953-018-0131-y.
- de Hemptinne, V.; Rondas, D.; Toepoel, M.; Vancompernolle, K. Phosphorylation on Thr-106 and NO-modification of glyoxalase I suppress the TNF-induced transcriptional activity of NF-κB. Mol. Cell. Biochem. 2009, 325, 169–178, doi:10.1007/s11010-009-0031-7.
- Morgenstern, J.; Katz, S.; Krebs-Haupenthal, J.; Chen, J.; Saadatmand, A.; Cortizo, F.G.; Moraru, A.; Zemva, J.; Campos, M.C.; Teleman, A.; et al. Phosphorylation of T107 by CamKIIδ regulates the detoxification efficiency and proteomic integrity of glyoxalase 1. Cell Rep. 2020, 32, 108160, doi:10.1016/j.celrep.2020.108160.
- de Hemptinne, V.; Rondas, D.; Vandekerckhove, J.; Vancompernolle, K. Tumour necrosis factor induces phosphorylation primarily of the nitric-oxide-responsive form of glyoxalase I. Biochem. J. 2007, 407, 121–128, doi:10.1042/BJ20070379.





