| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jesús Picazo-Aragonés | + 4116 word(s) | 4116 | 2020-12-03 10:08:01 | | | |
| 2 | Dean Liu | -1767 word(s) | 2349 | 2021-01-14 05:01:37 | | |
Video Upload Options
Volatile organic compounds (VOCs) are emitted by plants as a consequence of biotic and abiotic interaction which often change rapidly over time. Epigenetic factors, such as DNA methylation and histone modification, might trigger adaptive responses to these evolutionary pressures regulating both genes and transcription factors, as well as the rhythmic emission of VOCs through circadian clock regulation. In addition, transgenerational epigenetic effects and polyploidy could modify the generation of VOCs’ profiles of offspring, contributing to long-term evolutionary shifts.
1. Introduction
Plants synthesize an amazing diversity of secondary metabolites, which have been selected throughout their evolutionary history as a response to specific needs[1][2]. Secondary metabolites are distinguished from primary metabolites (such as nucleic acids, amino acids, carbohydrates, etc.) in that they are specialized metabolites which have a wide range of functions, mediating interaction with both biotic and abiotic environments[3][4]. Volatile organic compounds (VOCs) are one of the most important secondary metabolites produced by plants. These lipophilic compounds have a low molecular weight and high vapor pressures at ambient temperature. More than 1700 VOCs have been identified in different species from both angiosperms and gymnosperms, including a total of 90 families and 38 orders[5]. They are fundamentally released from flowers, but also fruits, leaves, stems and even roots[6].
2. Role of VOCs
VOCs define the chemical landscape of numerous ecosystems by taking part in intra- and interspecific interactions [7][8][9], being highly context dependent, and functioning in direct and indirect ways from the landscape to the intrafloral scale[10]. Plant VOCs have effects on plant–pollinator, plant–herbivore, plant–plant and other interactions and, consequently, on fitness[11]. Among all these functions, the most important and understood one is the attraction of pollinators, ensuring the plant’s reproductive success[9][12]. Floral scent promotes plant–pollinator specialization, as well as outcrossing and reproductive isolation through floral constancy[10][13]. Thus, such a sexual signal is subject to high selective pressure, being fundamental for plants’ evolution and adaptation to the environment [11].
Nevertheless, VOCs are also crucial in plant defense against herbivores and protection from pathogens[8][14]. VOCs can act as repellents, being constitutively emitted[10][15] or showing an increased emission during herbivory attack [16]. In addition, specific VOCs can perform as indirect defenses, which can drastically reduce the number of herbivores, especially insects and larvae, by attracting parasitoids and predators[7]. Furthermore, bacteria-specific VOCs are emitted by plants with a diagnostic purpose, triggering defense signaling pathways and acting as direct inhibitors of bacterial growth, therefore making plants more resistant to pathogen invasion[17].
Although pollinator attraction and plant defense may be the most important functions of VOCs, they play many other roles in the interaction of plants with their environment. VOCs are essential in plan-to-plant signaling, allowing other plants to respond to herbivores and activate defenses before being attacked[8][18] They also work as allelochemicals and neighbor detection signals, taking part in competition between plants and also having an effect on the availability of environmental factors such as light, nutrients and water[19]. Moreover, plant VOCs are also involved in other less studied interactions, like symbiotic relationships with microorganisms and fungi[6] or attraction of seed dispersers [20]. In addition, abiotic stress factors, such as temperature and high light intensity, also stimulate VOC biosynthesis and emission[21][22]. Water, salt and oxidative stresses may increase VOC emissions, but their effects are not consistent throughout the literature[23]. VOC emission, especially that of terpenes, has been proven to mitigate these stresses, allowing plants to recover rapidly from high temperature exposure[24] or alleviating oxidative stress[25] and consequently increasing plant fitness.
3. Epigenetic regulation of VOCs' biosynthesis and emission
Based on their biosynthetic origin, VOCs can be classified into three major groups: terpenoids, phenylpropanoids/benzenoids and fatty acid derivatives. Although the upstream enzymatic steps of the main biosynthetic pathways have been elucidated, knowledge about the last enzymatic steps and their regulation is scarce and is limited to model, crop and medicinal plants with high added value. However, recent progress in omics technology has led to a better understanding of plant VOC biosynthesis and the isolation of genes encoding candidate enzymes responsible for their regulation[26]. Nonetheless, the role of epigenetics in VOC biosynthesis and regulation has been neglected. Epigenetic regulation produces changes in gene function that can be mitotically and/or meiotically heritable and which do not entail a DNA sequence polymorphism[27]. The epigenetic signaling pathways altering the patterns of gene regulation and expression are cytosine DNA methylation (associated with both genes and transposable elements) and the post-transcriptional modification of histone proteins, which entails remodeling of chromatin structure[28]. Methylation of the 5′-position of cytosine residues is a reversible covalent modification of DNA, resulting in the production of 5-methyl-cytosine. Methylation changes the biophysical characteristics of DNA, causing the inhibition of DNA recognition by some proteins and allowing others to recognize it, which leads to the silencing of gene expression[29]. These proteins are known as methyl-binding proteins, transcriptional repressors that act through several mechanisms, such as the recruitment of corepressors and histone deacetylases, causing chromatin remodeling[30].
Additionally, RNA-based epigenetic mechanisms can also modify chromatin and silence transcription but are less understood. Non-coding RNAs have been known for a long time for their infrastructural role[31][32]. Furthermore, small (smRNA) and long non-coding RNAs (lncRNA) have recently emerged as key regulators of gene expression at the transcription level, RNA processing and translation[33][34]. Small RNAs can modify chromatin and silence transcription through the action of histone and DNA methyltransferases, recruited by guiding Argonaute-containing complexes to complementary nascent coding RNA or non-coding RNA scaffolds. They also act as a component of self-reinforcing positive feedback loops with an amplification component and participate in the epigenetic inheritance of histones and DNA methylation patterns. Moreover, lncRNA scaffolds recruit polycomb-group proteins and other chromatin-modifying complexes independently of smRNAs, but this mechanism remains poorly understood[35].
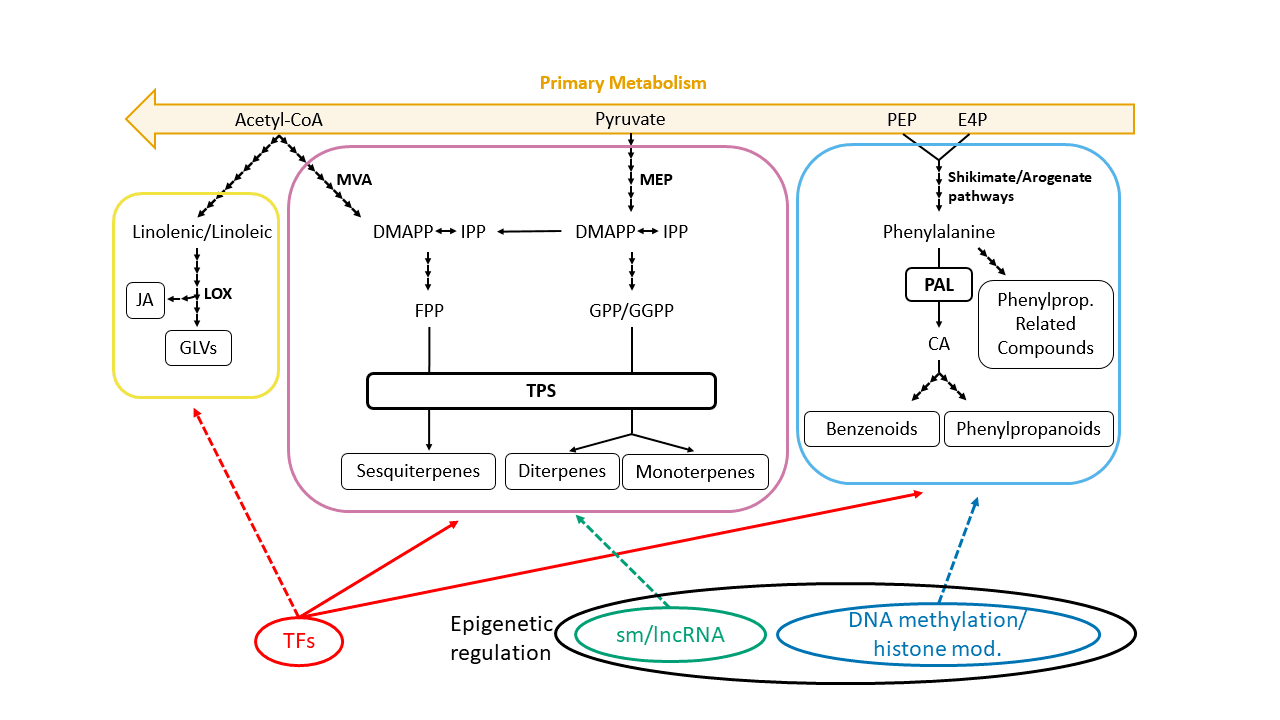
Volatile terpenes (monoterpenes, diterpenes and sesquiterpenes) are synthesized by the mevalonic acid (MVA) and methyleruthirol phosphate (MEP) pathways. Phenylpropanoid/benzenoid biosynthesis follows the shikimate and arogenate pathways. Meanwhile, fatty acid derivatives such as jasmonate (JA) and green leaf volatiles (GLVs) come from the lipoxygenase (LOX) pathway[4]. Volatile terpenoids and phenylpropanoids are transcriptionally controlled by various transcription factors and may be epigenetically controlled by small and long non-conding RNA, as well as by DNA methylation and histone modification. Volatile fatty acid derivatives biosynthesis is less understood, and also it's regulation. However it may be regulated by transcription factors (TFs) and epigenetic mechanisms.
Temporal release of floral scent is thought to be evolutionarily beneficial in order to maximize resource efficiency, attracting only truly effective pollinators and also avoiding the attraction of predators, which reduce herbivory [36][37]. These temporal patterns are accompanied by oscillations in the expression of genes in the VOC biosynthetic pathways [36][38], controlled by the circadian clock which is also epigenetically regulated. The main oscillator of the circadian clock is formed by the morning components CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) and LATE ELONGATED HYPOCOTYL (LHY) and the evening components TIMING OF CAB EXPRESSSION 1 (TOC1) and ZEITLUPE (ZTL) [39][40]. The morning and evening components regulate each other in a negative feedback loop. Light regulates ZTL expression, which ubiquitinates TOC1 for degradation[41][42]. These clock genes control the daily expression of terpene synthases and phenylpropanoid/benzenoid pathway genes. Methylation and acetylation of the H3 histone of the main clock genes causes chromatin remodeling and subsequent changes in these genes’ expression, affecting the biosynthesis and emission of VOCs[43]. Histone deacetylases (HDACs), induced by TOC1 and LHY, cause H3 deacetylation in the TOC1 promoter, repressing it. Meanwhile, the REVEILLE 8/LHY-CCA1-LIKE 5 (RVE8/LCL5) TF has an opposite role, promoting hyperacetylation of H3 in the TOC1 gene[44]. Di- and trimethylation of H3 histone also alters the expression of CCA1, TOC1 and LHY. In Arabidopsis SET DOMAIN GROUP 2/ARABIDOPSIS TRITHORAX RELATED 3 (SDG2/ATXR3) and LDL1-HDA6 alter methylation and acetylation patterns of these circadian genes.
Therefore, it seems reasonable that epigenetics would play a main role in the regulation of VOC emissions in plants. In this respect, polyploidy (whole genome duplication) could act as an important driver of VOC evolution. Polyploidy in plants has multiple effects at different scales, from the molecular (including the epigenetic landscape) to the phenotypic level, owing to advantageous evolutionary success[45]. Moreover, the persistence of epigenetic information through meiosis (i.e., transgenerational epigenetic inheritance) could also contribute to evolution of the temporal dynamic of VOCs and the plant’s biotic interactions[46]].
3. Evolution of VOCs: The Role of Polyploidy and Transgenerational Memory
Floral scent is considered an easily evolved trait, as VOCs are usually acquired and lost in related species[47][48]. Scent compounds would have mostly evolved due to gene duplication and divergence–convergence[1][2]. The large gene families, terpene synthases and O-methyl transferases, among others, would have evolved following this mechanism[1][49]. In addition, these modifications can also occur without the mediation of duplication events on single genes, changing the VOC emission throughout the evolutionary history [50]. Structural mutations modifying the enzyme catalytic sites can also cause the gaining of new functions (or the loss thereof) and hence cause changes in VOC emission. However, more interestingly, scent changes (qualitatively and quantitatively) can arise due to gene expression changes. Gene silencing or activation cannot only affect an individual VOC emission, but it can also have a cascade effect on the complete pathway[51][52]. New VOC emissions can straightforwardly arise from exogenous gene expression in new tissues. For example, in Clarkia breweri, linalool emission (its most abundant floral VOC) most likely evolved from trace emissions in stigma tissue in the ancestor C. concinna[53]. Such VOC changes could arise as a consequence of epigenetic regulation of genes, creating advantageous or disadvantageous phenotypes that are subject to natural selection. If new VOCs confer a selective advantage for the plant, the epigenetic change and the VOC production would be retained and inherited[54]. The transgenerational inheritance of floral and leaf VOCs has been recently found in Brassica rapa[55], and this could be epigenetically mediated[46]. Therefore, epigenetic change would propose a flexible evolutionary mechanism and help to explain the continuous gain and loss of specific VOCs in related plant lineages. Epigenetic variations would assist plants in surfing on an adaptative landscape in which abiotic and biotic interactions usually involve trade-offs (e.g., herbivory and pollinator attraction) [56][57].
Furthermore, all angiosperm lineages show vestiges of past rounds of whole genome duplication (WGD), highlighting the role of polyploidy as a major driving force in plant evolution[58][59][60]. Polyploidy has multiple effects at different scales, from the molecular to the phenotypic level, owing to the advantageous evolutionary success[61]. Patently, WGD (and the consequent gene duplication) is an important molecular evolutionary mechanism driving the large diversity of plant VOCs. In addition, following WGD, polyploids suffer changes in genomic structure and epigenetic remodeling[62][63]. Polyploidy usually causes an altered pattern of DNA methylation in genes, promoters and transposable elements[64][65]. These changes are enhanced in allopolyploids (WGD associated with species hybridization) because of “genomic shock”. In this case, this methylation repatterning is related to parental dominance (i.e., the higher expression of one parental homeolog) and fractionalization[45]. In addition, changes in smRNAs and chromatin have also been observed after WGD[66][67][68]. In this way, autopolyploidy (WGD without hybridization) has been recently considered as an epigenetic macromutation affecting chromatin compaction and altering contacts among chromosomes (i.e., 4D nucleome) as a side effect of the increased nucleus size[69]. This altered epigenetic landscape may promote divergent gene expression by regulating mobile element activity and silencing redundant genes[64][70]. In addition, these epigenetic alterations could create many opportunities for natural selection to act, enhancing polyploid establishment[71].
Concurrently to the epigenetic changes, WGD also exerts a cornucopia of effects on the plant phenotype, including numerous floral traits[16]. Changes in scent pattern have been shown in some orchid polyploids[72][73]]. Such changes in floral traits could impact pollinator attraction and lead to differentiation in the pollinator spectrum[50][74], causing the isolation of diploids and polyploids and facilitating polyploid establishment. Although gene duplication and later neofunctionalization[75] can promote such scent changes, rapid scent shifts could be presumably attributed to epigenetic remodeling following WGD[46][76]. Therefore, the role of the epigenetic change associated with polyploidy on scent biosynthesis and emission (and its evolutionary contribution) deserves further investigation.
In this respect, the carnation complex of Dianthus broteri is an appropriate study system to investigate the interplay between epigenetics and polyploidy on the evolutionary change of floral volatiles. This Iberian endemic complex represents the most extensive polyploid series of the Dianthus genus, with diploid (2n = 30), tetraploid (2n = 60), hexaploid (2n = 90) and dodecaploid (2n = 180) individuals[77]. Molecular and genome size studies have demonstrated that it is an autopolyploid complex[78], in which the different cytotypes form independent and monophyletic lineages and show disjunct geographic distributions with monocytotypic populations and different ecological niches within an aridity gradient, with the higher cytotypes (6× and 12×) being those that inhabit more restricted and extreme habitats[79][80]. It is also remarkable that, in this complex, polyploidy leads to an increased degree of DNA methylation[81] and of epigenetic variability, which may be crucial to phenotypic divergence (i.e., changes in VOCs), adaptation and shifts in biotic interactions[82][83][76]. Previous studies[16][84] have shown phenotypic differences in floral traits (floral size, distance to access the nectar and scent) between D. broteri cytotypes that may promote changes in pollinator patterns. The highest ploidy level (12×, D. inoxianus) involves a highly specialized pollination by a single pollinator, Hyles livornica. The floral scent of this cytotype is dominated by sesquiterpenoids and fatty acid derivatives (2-ketones), which shown a circadian emission[16]. The 2-ketones elicit responses in the antennae of the single pollinator, but also have a repellent/insecticide activity[85]. They show an explosive increase in production, triggered by experimental damage to D. inoxianus flowers, which indicates that 2-ketones may be acting like a floral filter of the alleged pollinator spectrum, leading to pollinator specialization[16]. Supporting this “filtering hypothesis”, these ketones appear in low quantities at diploid and tetraploid levels, which show a wider pollinator spectrum[86]. In addition, dominant aromatic compound changes (e.g., the disappearance of β-ocimene in 4× and 12× and the dominance of β-caryophyllene in 2× and 4×) suggest that shifts in the expression of scent-related genes may be promoting greater specialization in higher polyploids. We suggest a combination of epigenomics, biochemical and pollination biology methods to investigate whether epigenetics is involved in polyploid evolution by means of floral VOCs and pollinator shifts.
References
- Pichersky, E.; Gang, D.R. Genetics and biochemistry of secondary metabolites in plants: An evolutionary perspective. Trends Plant Sci. 2000, 5, 439–445.
- Pichersky, E.; Lewinsohn, E. Convergent Evolution in Plant Specialized Metabolism. Annu. Rev. Plant Biol. 2011, 62, 549–566.
- Pichersky, E.; Noel, J.P.; Dudareva, N. Biosynthesis of plant volatiles: Nature’s diversity and ingenuity. Science 2006, 311, 808–811.
- Dudareva, N.; Klempien, A.; Muhlemann, J.K.; Kaplan, I. Biosynthesis, function and metabolic engineering of plant volatile organic compounds. New Phytol. 2013, 198, 16–32.
- Knudsen, J.T.; Eriksson, R.; Gershenzon, J.; Ståhl, B. Diversity and distribution of floral scent. Bot. Rev. 2006, 72, 1–120.
- Farré-Armengol, G.; Filella, I.; Llusia, J.; Peñuelas, J. Bidirectional Interaction between Phyllospheric Microbiotas and Plant Volatile Emissions. Trends Plant Sci. 2016, 21, 854–860.
- Kessler, A.; Baldwin, I.T. Defensive function of herbivore-induced plant volatile emissions in nature. Science 2001, 291, 2141–2144.
- Heil, M.; Bueno, J.C.S. Within-plant signaling by volatiles leads to induction and priming of an indirect plant defense in nature. Proc. Natl. Acad. Sci. USA 2007, 104, 5467–5472.
- Farré-Armengol, G.; Filella, I.; Llusia, J.; Peñuelas, J. Floral volatile organic compounds: Between attraction and deterrence of visitors under global change. Perspect. Plant Ecol. Evol. Syst. 2013, 15, 56–67.
- Raguso, R.A. Wake Up and Smell the Roses: The Ecology and Evolution of Floral Scent. Annu. Rev. Ecol. Evol. Syst. 2008, 39, 549–569.
- Austin, A.T.; Ballaré, C.L. Plant interactions with other organisms: Molecules, ecology and evolution. New Phytol. 2014, 204, 257–260.
- Kessler, D.; Kallenbach, M.; Diezel, C.; Rothe, E.; Murdock, M.; Baldwin, I.T. How scent and nectar influence floral antagonists and mutualists. Elife 2015, 4.
- Grüter, C.; Ratnieks, F.L.W. Flower constancy in insect pollinators: Adaptive foraging behavior or cognitive limitation? Commun. Integr. Biol. 2011, 4, 633–636.
- Koornneef, A.; Pieterse, C.M.J. Cross talk in defense signaling. Plant Physiol. 2008, 146, 839–844.
- Junker, R.R.; Blüthgen, N. Floral scents repel potentially nectar-thieving ants. Evol. Ecol. Res. 2008, 10, 295–308.
- Balao, F.; Herrera, J.; Talavera, S.; Dötterl, S. Spatial and temporal patterns of floral scent emission in Dianthus inoxianus and electroantennographic responses of its hawkmoth pollinator. Phytochemistry 2011, 72, 601–609.
- Huang, M.; Sanchez-Moreiras, A.M.; Abel, C.; Sohrabi, R.; Lee, S.; Gershenzon, J.; Tholl, D. The major volatile organic compound emitted from Arabidopsis thaliana flowers, the sesquiterpene (E)-β-caryophyllene, is a defense against a bacterial pathogen. New Phytol. 2012, 193, 997–1008.
- Baldwin, I.T.; Halitschke, R.; Paschold, A.; Von Dahl, C.C.; Preston, C.A. Volatile signaling in plant-plant interactions: “Talking trees” in the genomics era. Science 2006, 311, 812–815.
- Kegge, W.; Pierik, R. Biogenic volatile organic compounds and plant competition. Trends Plant Sci. 2010, 15, 126–132.
- Rodríguez, A.; Alquézar, B.; Peña, L. Fruit aromas in mature fleshy fruits as signals of readiness for predation and seed dispersal. New Phytol. 2013, 197, 36–48.
- Hansen, U.; Seufert, G. Temperature and light dependence of β-caryophyllene emission rates. J. Geophys. Res. D Atmos. 2003, 108.
- Truong, D.H.; Delory, B.M.; Vanderplanck, M.; Brostaux, Y.; Vandereycken, A.; Heuskin, S.; Delaplace, P.; Francis, F.; Lognay, G. Temperature regimes and aphid density interactions differentially influence VOC emissions in Arabidopsis. Arthropod. Plant. Interact. 2014, 8, 317–327.
- Holopainen, J.K.; Gershenzon, J. Multiple stress factors and the emission of plant VOCs. Trends Plant Sci. 2010, 15, 176–184.
- Sharkey, T.D.; Chen, X.; Yeh, S. Isoprene increases thermotolerance of fosmidomycin-fed leaves. Plant Physiol. 2001, 125, 2001–2006.
- Vickers, C.E.; Possell, M.; Cojocariu, C.I.; Velikova, V.B.; Laothawornkitkul, J.; Ryan, A.; Mullineaux, P.M.; Nicholas Hewitt, C. Isoprene synthesis protects transgenic tobacco plants from oxidative stress. Plant Cell Environ. 2009, 32, 520–531.
- Ramya, M.; Jang, S.; An, H.-R.; Lee, S.-Y.; Park, P.-M.; Park, P.H. Volatile Organic Compounds from Orchids: From Synthesis and Function to Gene Regulation. Int. J. Mol. Sci. 2020, 21, 1160.
- Dupont, C.; Armant, D.R.; Brenner, C.A. Epigenetics: Definition, mechanisms and clinical perspective. Semin. Reprod. Med. 2009, 27, 351–357.
- Gibney, E.R.; Nolan, C.M. Epigenetics and gene expression. Heredity (Edinb.) 2010, 105, 4–13.
- Newell-Price, J.; Clark, A.J.L.; King, P. DNA methylation and silencing of gene expression. Trends Endocrinol. Metab. 2000, 11, 142–148.
- Wade, P.A. Methyl CpG binding proteins: Coupling chromatin architecture to gene regulation. Oncogene 2001, 20, 3166–3173.
- Boyko, A.; Kovalchuk, I. Genome instability and epigenetic modification-heritable responses to environmental stress? Curr. Opin. Plant Biol. 2011, 14, 260–266.
- Blomen, V.A.; Boonstra, J. Stable transmission of reversible modifications: Maintenance of epigenetic information through the cell cycle. Cell. Mol. Life Sci. 2011, 68, 27–44.
- Moazed, D. Small RNAs in transcriptional gene silencing and genome defence. Nature 2009, 457, 413–420.
- Cech, T.R.; Steitz, J.A. The noncoding RNA revolution-Trashing old rules to forge new ones. Cell 2014, 157, 77–94.
- Holoch, D.; Moazed, D. RNA-mediated epigenetic regulation of gene expression. Nat. Rev. Genet. 2015, 16, 71–84.
- Fenske, M.P.; Imaizumi, T. Circadian rhythms in floral scent emission. Front. Plant Sci. 2016, 7, 1–6.
- Zeng, L.; Wang, X.; Kang, M.; Dong, F.; Yang, Z. Regulation of the rhythmic emission of plant volatiles by the circadian clock. Int. J. Mol. Sci. 2017, 18, 2408.
- Dudareva, N.; Pichersky, E. Biochemical and molecular genetic aspects of floral scents. Plant Physiol. 2000, 122, 627–633.
- Nagel, D.H.; Kay, S.A. Complexity in the wiring and regulation of plant circadian networks. Curr. Biol. 2012, 22, R648–R657.
- McClung, C.R. Beyond Arabidopsis: The circadian clock in non-model plant species. Semin. Cell Dev. Biol. 2013, 24, 430–436.
- Alabadí, D.; Oyama, T.; Yanovsky, M.J.; Harmon, F.G.; Más, P.; Kay, S.A. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 2001, 293, 880–883.
- Yon, F.; Kessler, D.; Joo, Y.; Cortés Llorca, L.; Kim, S.G.; Baldwin, I.T. Fitness consequences of altering floral circadian oscillations for Nicotiana attenuata. J. Integr. Plant Biol. 2017, 59, 180–189.
- Du, S.; Chen, L.; Ge, L.; Huang, W. A novel loop: Mutual regulation between epigenetic modification and the circadian clock. Front. Plant Sci. 2019, 10, 1–8.
- Farinas, B.; Mas, P. Functional implication of the MYB transcription factor RVE8/LCL5 in the circadian control of histone acetylation. Plant J. 2011, 66, 318–329.
- Madlung, A.; Wendel, J.F. Genetic and epigenetic aspects of polyploid evolution in plants. Cytogenet. Genome Res. 2013, 140, 270–285.
- Kellenberger, R.T.; Schlüter, P.M.; Schiestl, F.P. Herbivore-Induced DNA Demethylation Changes Floral Signalling and Attractiveness to Pollinators in Brassica rapa. PLoS ONE 2016, 11, e0166646.
- Weiss, J.; Mühlemann, J.K.; Ruiz-Hernández, V.; Dudareva, N.; Egea-Cortines, M. Phenotypic Space and Variation of Floral Scent Profiles during Late Flower Development in Antirrhinum. Front. Plant Sci. 2016, 7, 1903.
- Prieto-Benítez, S.; Millanes, A.M.; Dötterl, S.; Giménez-Benavides, L. Comparative analyses of flower scent in Sileneae reveal a contrasting phylogenetic signal between night and day emissions. Ecol. Evol. 2016, 6, 7869–7881.
- Chen, F.; Tholl, D.; Bohlmann, J.; Pichersky, E. The family of terpene synthases in plants: A mid-size family of genes for specialized metabolism that is highly diversified throughout the kingdom. Plant J. 2011, 66, 212–229.
- Amrad, A.; Moser, M.; Mandel, T.; de Vries, M.; Schuurink, R.C.; Freitas, L.; Kuhlemeier, C. Gain and loss of floral scent croduction through changes in structural genes during pollinator-mediated Speciation. Curr. Biol. 2016, 26, 3303–3312.
- Pulido, P.; Perello, C.; Rodriguez-Concepcion, M. New Insights into Plant Isoprenoid Metabolism. Mol. Plant 2012, 5, 964–967.
- Jantzen, F.; Lynch, J.H.; Kappel, C.; Höfflin, J.; Skaliter, O.; Wozniak, N.; Sicard, A.; Sas, C.; Adebesin, F.; Ravid, J.; et al. Retracing the molecular basis and evolutionary history of the loss of benzaldehyde emission in the genus Capsella. New Phytol. 2019, 224, 1349–1360.
- Cseke, L.; Dudareva, N.; Pichersky, E. Structure and Evolution of Linalool Synthase. Mol. Biol. Evol. 1998, 15, 1491–1498.
- Quadrana, L.; Colot, V. Plant Transgenerational Epigenetics. Annu. Rev. Genet. 2016, 50, 467–491.
- Kellenberger, R.T.; Desurmont, G.A.; Schlüter, P.M.; Schiestl, F.P. Trans-generational inheritance of herbivory-induced phenotypic changes in Brassica rapa. Sci. Rep. 2018, 8, 3536.
- Farré-Armengol, G.; Fernández-Martínez, M.; Filella, I.; Junker, R.R.; Peñuelas, J. Deciphering the Biotic and Climatic Factors That Influence Floral Scents: A Systematic Review of Floral Volatile Emissions. Front. Plant Sci. 2020, 11, 1154.
- Ramos, S.E.; Schiestl, F.P. Evolution of Floral Fragrance Is Compromised by Herbivory. Front. Ecol. Evol. 2020, 8, 1–8.
- Baniaga, A.E.; Marx, H.E.; Arrigo, N.; Barker, M.S. Polyploid plants have faster rates of multivariate niche differentiation than their diploid relatives. Ecol. Lett. 2020, 23, 68–78.
- Landis, J.B.; Soltis, D.E.; Li, Z.; Marx, H.E.; Barker, M.S.; Tank, D.C.; Soltis, P.S. Impact of whole-genome duplication events on diversification rates in angiosperms. Am. J. Bot. 2018, 105, 348–363.
- Tank, D.C.; Eastman, J.M.; Pennell, M.W.; Soltis, P.S.; Soltis, D.E.; Hinchliff, C.E.; Brown, J.W.; Sessa, E.B.; Harmon, L.J. Nested radiations and the pulse of angiosperm diversification: Increased diversification rates often follow whole genome duplications. New Phytol. 2015, 207, 454–467.
- vMadlung, A.; Wendel, J.F. Genetic and epigenetic aspects of polyploid evolution in plants. Cytogenet. Genome Res. 2013, 140, 270–285.
- Doyle, J.J.; Flagel, L.E.; Paterson, A.H.; Rapp, R.A.; Soltis, D.E.; Soltis, P.S.; Wendel, J.F. Evolutionary Genetics of Genome Merger and Doubling in Plants. Annu. Rev. Genet. 2008, 42, 443–461.
- Feliner, G.N.; Casacuberta, J.; Wendel, J.F. Genomics of Evolutionary Novelty in Hybrids and Polyploids. Front. Genet. 2020, 11, 792.
- Zhang, J.; Liu, Y.; Xia, E.H.; Yao, Q.Y.; Liu, X.D.; Gao, L.Z. Autotetraploid rice methylome analysis reveals methylation variation of transposable elements and their effects on gene expression. Proc. Natl. Acad. Sci. USA 2015, 112, E7022–E7029.
- Yan, H.; Bombarely, A.; Xu, B.; Wu, B.; Frazier, T.P.; Zhang, X.; Chen, J.; Chen, P.; Sun, M.; Feng, G.; et al. Autopolyploidization in switchgrass alters phenotype and flowering time via epigenetic and transcription regulation. J. Exp. Bot. 2019, 70, 5673–5686.
- Cavé-Radet, A.; Giraud, D.; Lima, O.; El Amrani, A.; Aïnouche, M.; Salmon, A. Evolution of small RNA expression following hybridization and allopolyploidization: Insights from Spartina species (Poaceae, Chloridoideae). Plant Mol. Biol. 2020, 102, 55–72.
- Ng, D.W.K.; Lu, J.; Chen, Z.J. Big roles for small RNAs in polyploidy, hybrid vigor, and hybrid incompatibility. Curr. Opin. Plant Biol. 2012, 15, 154–161.
- Zhang, H.; Zheng, R.; Wang, Y.; Zhang, Y.; Hong, P.; Fang, Y.; Li, G.; Fang, Y. The effects of Arabidopsis genome duplication on the chromatin organization and transcriptional regulation. Nucleic Acids Res. 2019, 47, 7857–7869.
- Doyle, J.J.; Coate, J.E. Autopolyploidy: An epigenetic macromutation. Am. J. Bot. 2020, 107, 1097–1100.
- Vicient, C.M.; Casacuberta, J.M. Impact of transposable elements on polyploid plant genomes. Ann. Bot. 2017, 120, 195–207.
- Arango, J.; Beltrán, J.; Nuñez, J.; Chavarriaga, P. Evidence of epigenetic mechanisms affecting carotenoids. Subcell. Biochem. 2016, 79, 295–307.
- Gross, K.; Schiestl, F.P. Are tetraploids more successful? Floral signals, reproductive success and floral isolation in mixed-ploidy populations of a terrestrial orchid. Ann. Bot. 2015, 115, 263–273.
- Jersáková, J.; Castro, S.; Sonk, N.; Milchreit, K.; Schödelbauerová, I.; Tolasch, T.; Dötterl, S.; Jersáková, J.; Schödelbauerová, I.; Schödelbauerová, Á.I.; et al. Absence of pollinator-mediated premating barriers in mixed-ploidy populations of Gymnadenia conopsea s.l. (Orchidaceae). Evol. Ecol. 2010, 24, 1199–1218.
- Klahre, U.; Gurba, A.; Hermann, K.; Saxenhofer, M.; Bossolini, E.; Guerin, P.M.; Kuhlemeier, C. Pollinator choice in petunia depends on two major genetic loci for floral scent production. Curr. Biol. 2011, 21, 730–739.
- Barkman, T.J. Evidence for positive selection on the floral scent gene isoeugenol-O-methyltransferase. Mol. Biol. Evol. 2003, 20, 168–172. [Google Scholar] [CrossRef] [PubMed]o, L.; Wang, J.; Cheng, T.; Zhang, Q. Expansion of PmBEAT genes in the Prunus mume genome induces characteristic floral scent production. Hortic. Res. 2019, 6, 1–18.
- Mirouze, M.; Paszkowski, J. Epigenetic contribution to stress adaptation in plants. Curr. Opin. Plant Biol. 2011, 14, 267–274.
- Balao, F.; Valente, L.M.; Vargas, P.; Herrera, J.; Talavera, S. Radiative evolution of polyploid races of the Iberian carnation Dianthus broteri (Caryophyllaceae). New Phytol. 2010, 187, 542–551.
- Balao, F.; Casimiro-Soriguer, R.; Talavera, M.; Herrera, J.; Talavera, S. Distribution and diversity of cytotypes in Dianthus broteri as evidenced by genome size variations. Ann. Bot. 2009, 104, 965–973.
- López-Jurado, J.; Balao, F.; Mateos-Naranjo, E. Polyploidy-mediated divergent light-harvesting and photoprotection strategies under temperature stress in a Mediterranean carnation complex. Environ. Exp. Bot. 2020, 171, 103956.
- López-Jurado, J.; Mateos-Naranjo, E.; Balao, F. Niche divergence and limits to expansion in the high polyploid Dianthus broteri complex. New Phytol. 2019, 222, 1076–1087.
- Alonso, C.; Balao, F.; Bazaga, P.; Pérez, R. Epigenetic contribution to successful polyploidizations: Variation in global cytosine methylation along an extensive ploidy series in Dianthus broteri (Caryophyllaceae). New Phytol. 2016, 212, 571–576.
- Alonso, C.; Ramos-Cruz, D.; Becker, C. The role of plant epigenetics in biotic interactions. New Phytol. 2019, 221, 731–737.
- Balao, F.; Paun, O.; Alonso, C. Uncovering the contribution of epigenetics to plant phenotypic variation in Mediterranean ecosystems. Plant Biol. 2018, 20, 38–49.
- Herrera, J.; Balao, F. A test of phenotypic selection on petal form in the wild carnation, Dianthus inoxianus. Plant Biol. 2015, 17, 1227–1232.
- Lin, S.Y.H.; Trumble, J.T.; Kumamoto, J. Activity of volatile compounds in glandular trichomes of Lycopersicon species against two insect herbivores. J. Chem. Ecol. 1987, 13, 837–850.
- Balao, F. (University of Seville, Seville, Spain). Personal communication, 2019.