
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Andrea Balan | + 1445 word(s) | 1445 | 2020-12-08 11:59:46 | | | |
| 2 | Lily Guo | Meta information modification | 1445 | 2021-01-18 13:06:09 | | |
Video Upload Options
Mycobacterium tuberculosis is a bacterium of great medical importance because it causes tuberculosis, a disease that affects millions of people worldwide. Two important features are related to this bacterium: its ability to infect and survive inside the host, minimizing the immune response, and the burden of clinical isolates that are highly resistant to antibiotics treatment. These two phenomena are directly affected by cell envelope proteins, such as proteins from the ATP-Binding Cassette (ABC transporters) superfamily.
1. Introduction
M. tuberculosis is the agent responsible for tuberculosis, with about 10 million people infected in the world in 2018, of which 1.2 million died in 2019[1]. To make this scenario worse, many individuals have latent infections that are reactivated at some point in their lives, especially among those with other immunodeficient diseases such as co-infection with the HIV virus (human immunodeficiency virus). The bacillus has a cytoplasmic membrane, cell wall, and an external capsule formed mainly by polysaccharides that play a relevant role in the virulence of M. tuberculosis strains. The inner membrane is formed by a phospholipid bilayer with several integral membrane proteins, among which the important family of ATP-dependent transporters, the ATP-Binding Cassette (ABC) transporters, are also present. The cell wall or mycomembrane is formed by a thin layer of peptidoglycan associated with lipoarabinomananan and a layer of arabinogalactones. In addition, there are mycolic acids and other important components specific to the mycobacteria, such as different types of phosphosugars, trehalose, glycerol sulfolipids, and phenolic glycolipids [2].
Transport across the cell membrane is extremely important for the bacterium. Different molecules and substrates are required for diverse cellular processes. Species from the M. tuberculosis complex (MTBC) and strains from pathogenic groups have around 160 to 200 proteins dedicated to the ATP-dependent transport systems according to the TransportDB webserver [3] indicating the relevance of this protein family for the genus. In M. tuberculosis, 2.5% of the genome encodes components of ABC transporters, indicating their critical role in the maintenance of cellular homeostasis. These transporters, in addition to being responsible for the transport of important molecules such as sugars, amino acids, and many others, also act as efflux pumps of external agents and are thereby responsible for resistance to antibiotics[4]. Structurally, ABC transporters consist of a canonical set of two transmembrane domains (TMDs) that form a pore for the passage of the substrate (TMDs) and two nucleotide-binding domains (NDBs) that provide energy for substrate translocation through ATP hydrolysis. These components can be translated into independent polypeptides or joined in various possible combinations, forming homodimers or heterodimers[5]. In addition, ABC import systems have a third component, a periplasmic substrate-binding protein (SBP), which is responsible for the uptake and delivery of the substrate into the pore formed by the TMDs [6]. Based on the layout and architecture of the TMDs, ABC transporters can be divided into seven families, three importer families, and four exporter families[7]. Using bioinformatics analysis, structural characterization, and data mining, we have classified M. tuberculosis ABC transporter components into aforementioned families (Figure 1).
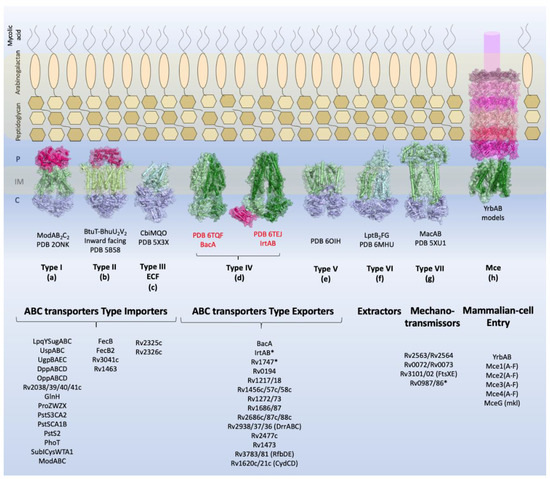
Figure 1. Overview of the types of ABC transporters and the components identified in M. tuberculosis. Schematic view of the M. tuberculosis cellular envelope is shown with the families of TMDABC transporters organized into importers (Types I–III), exporters (Types IV and V), extractors, and mechanotransmitters. Additionally, the structural representation of the Mce complex (Mammalian-cell entry proteins), which has two ABC-like TMDs, is shown. Representative structure of each transporter class is shown in the inner membrane: Type I, the molybdate transporter ModABC of Archaeoglobus fulgidus (PDB 2ONK); Type II, the vitamin transporter BtuT-BtuUV of Burkholderia cenocepacia (PDB 5B58); Type III, the cobalt energy-coupling factor transporter CbiMQO of Rhodobacter capsulatus (PDB 5 × 3X); Type IV, two ABC transporter type exporter of M. tuberculosis that had the three-dimensional structure resolved are shown: the ABC transporter of vitamin B12, BacA (gene Rv1819c) (PDB 6TQF), and the transporter of siderophores, IrtAB (PDB 6TEJ) (nomenclature in red); Type V, the transporter of O-antigen of Aquifex aeolicus VF5 (PDB 6OIH); Type VI, Cryo-EM structure of the E. coli LptB2FG transporter (PDB 6MHU); Type VII, the structure of MacAB-like efflux pump from Streptococcus pneumoniae (PDB 5XU1); Mce complex, prediction of the structural organization of Mce transporter systems (Mce1-4); the two ABC components, YrdAB, and the periplasmic domains, proteins MceA to MceF were modelled using I-Tasser server[8] based on E. coli Mla components. The structures are shown in cartoon representation with a transparent surface. The substrate-binding proteins from Type I importers and the periplasmic domains of Mce complex are shown in pink shades; the transmembrane domains are shown in dark and pale green, and the nucleotide-binding domains are shown in blue. P: periplasm, IM: inner membrane, C: cytoplasm. The ABC transporters and components of M. tuberculosis described in this review are listed below the representative structures. Transporters marked with asterisk (*) have structural organization that does not fit the functional or structural classification.
Most ABC importer systems that have been identified in M. tuberculosis so far belong to group I and include systems dedicated to the import of sugars, amino acids, peptides, and anions. However, four systems related to iron import have been classified into group II and a unique cobalt transporter into group III (Energy Coupling Factor, ECF).
M. tuberculosis ABC exporter systems fall into groups IV, V, and VII. It is interesting to observe that the secondary and or tertiary structural characteristics of some components do not fit their functional groups, as shown by experimental analysis. These components are IrtAB, BacA, Rv2563/64, Rv0072/73, Rv1747, and Rv0987/86. Functionally, IrtAB and BacA are importers, but their structures are very similar to the structures of ABC exporters. No periplasmic components were identified for these systems in the vicinity of the genome, and IrtAB also has an additional cytoplasmic domain which is thought to function in exportation. Similarly, Rv2563/64 and Rv0072/73 are structurally related to type VI transporters, although they have been associated with the import of amino acids due the presence of the NBD component. Rv2564 and Rv0073 are highly similar to the E. coli GlnQ protein, the ATPase subunit from the glutamine transporter [9]. Rv1747 and Rv0987/86, putative exporters, have a non-canonical domain architecture, with two cytoplasmic phosphorylated Fork-Head Associated (FHA) domains and two extracellular domains per TMD, respectively. This mixing of functions and structural features across families highlights how surprising and captivating the components of the M. tuberculosis inner membrane are and how little it is known about them and the proteins that belong to the bacillus envelope. Finally, we have the Mce proteins, whose function is critical for the bacillus existence. Nevertheless, their structural organization is not clear and only based on homology of the MceA-F components with the E. coli Mla proteins[10][11].
2. Identified ABC transporters in Mycobacterium tuberculosis
2.1. ABC Transporters Type Importer
ABC importers are present in prokaryotes but not in eukaryotes. They are responsible for the import of different substrates including oligopeptides, amino acids, inorganic anions, sugars, and metals. Because of such a diverse range of substrates, ABC importers are involved in many physiological processes, including transport, nutrition, virulence, and pathogenesis. Most ABC importers in M. tuberculosis are responsible for the bacillus adaptation in the human host. In this section, we describe the components and full ABC importers systems that were identified in M. tuberculosis, including their roles to keep the bacillus in full activity. We have subdivided them into five main groups according to their putative functions (Table 1).
Table 1. Genomic context and structural organization of the ATP-Binding Cassette (ABC) importers in Mycobacterium tuberculosis. Importers are presented in seven groups organized by putative function. Genes that belong to the genomic context are shown as arrows. The number of helices and structural organization predicted with TOPCONS [12] are shown in the structural organization column. Substrate-binding proteins are represented in pale pink; transmembrane domains in light and dark green; nucleotide-binding domains in purple; and the regulatory domain, when present, in pale purple. SID: siderophore interaction domain.

2.2. ABC Transporters Type Exporters
M. tuberculosis has at least 28 identified genes encoding components of ABC transporter type exporters, from which 26 form 14 complete systems subdivided into four main functional categories: (i) recycling/transport of membrane components (two systems), (ii) electron transport chain (ETC) (one system), (iii) virulence and adaptation (two systems), and (iv) drug efflux (nine systems). The genomic organization show the genes are organized in operons that also contain related genes that, depending on the function, will help in transport. Based on secondary structure predictions, domain analyses, and the prediction of the number of transmembrane helices, these transporters show a variety of structural organizations that go beyond the classical architecture of two NBDs and two TMDs. Furthermore, both transporters and components have shown a functional plasticity in M. tuberculosis and related species.
References
- WHO Global tuberculosis report 2019; Global tuberculosis report 2019. Geneva: World Health Organization; 2019. Licence: CC BY-NC-SA 3.0 IGO.
- Dulberger, C.L.; Rubin, E.J.; Boutte, C.C. The mycobacterial cell envelope — a moving target. Nat. Rev. Microbiol. 2020, 18, 47–59, doi:10.1038/s41579-019-0273-7.
- Elbourne, L.D.H.; Tetu, S.G.; Hassan, K.A.; Paulsen, I.T. TransportDB 2.0: A database for exploring membrane transporters in sequenced genomes from all domains of life. Nucleic Acids Res. 2017, 45, D320–D324, doi:10.1093/nar/gkw1068.
- Singh, R.; Dwivedi, S.P.; Gaharwar, U.S.; Meena, R.; Rajamani, P.; Prasad, T. Recent updates on drug resistance in Mycobacterium tuberculosis. J. Appl. Microbiol. 2020, 128, 1547–1567, doi:10.1111/jam.14478.
- Reis, R.; Moraes, I. Structural biology and structure–function relationships of membrane proteins. Biochem. Soc. Trans. 2018, 47, 47–61, doi:10.1042/BST20180269.
- Davidson, A.L.; Chen, J. ATP-binding cassette transporters in bacteria. Annu. Rev. Biochem. 2004, 73, 241–268, doi:10.1146/annurev.biochem.73.011303.073626.
- Thomas, C.; Tampé, R. Structural and mechanistic principles of ABC Transporters. Annu. Rev. Biochem. 2020, 89, 605–636, doi:10.1146/annurev-biochem-011520-105201.
- Yang, J.; Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 2015, 43, W174–W181, doi:10.1093/nar/gkv342.
- Braibant, M.; Gilot, P.; Content, J. The ATP binding cassette (ABC) transport systems of Mycobacterium tuberculosis. FEMS Microbiol. Rev. 2000, doi:10.1111/j.1574-6976.2000.tb00550.x.
- Ekiert, D.C.; Bhabha, G.; Isom, G.L.; Greenan, G.; Ovchinnikov, S.; Henderson, I.R.; Cox, J.S.; Vale, R.D. Architectures of lipid transport systems for the bacterial outer membrane. Cell 2017, 169, 273-285.e17, doi:10.1016/j.cell.2017.03.019.
- Kumar, A.; Chandolia, A.; Chaudhry, U.; Brahmachari, V.; Bose, M. Comparison of mammalian cell entry operons of mycobacteria: In silico analysis and expression profiling. FEMS Immunol. Med. Microbiol. 2005, 43, 185–195, doi:10.1016/j.femsim.2004.08.013.





