
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Laurence Meslet-Cladiere | + 2627 word(s) | 2627 | 2020-12-30 07:55:07 | | | |
| 2 | Vivi Li | + 336 word(s) | 2963 | 2021-01-13 05:16:49 | | |
Video Upload Options
A putative Type III Polyketide synthase (PKSIII) encoding gene was identified from a marine yeast, Naganishia uzbekistanensis strain Mo29 (UBOCC-A-208024) (formerly named as Cryptococcus sp.) isolated from deep-sea hydrothermal vents. This gene is part of a distinct phylogenetic branch compared to all known terrestrial fungal sequences. This new gene encodes a C-terminus extension of 74 amino acids compared to other known PKSIII proteins like Neurospora crassa. Full-length and reduced versions of this PKSIII were successfully cloned and overexpressed in a bacterial host, Escherichia coli BL21 (DE3). Both proteins showed the same activity, suggesting that additional amino acid residues at the C-terminus are probably not required for biochemical functions. We demonstrated by LC-ESI-MS/MS that these two recombinant PKSIII proteins could only produce tri- and tetraketide pyrones and alkylresorcinols using only long fatty acid chain from C8 to C16 acyl-CoAs as starter units, in presence of malonyl-CoA. In addition, we showed that some of these molecules exhibit cytotoxic activities against several cancer cell lines.
1. Introduction
Secondary metabolites represent an attractive and important source of natural products with a wide range of bioactivities and biotechnological applications. Organisms such as plants, algae, bacteria and fungi are able to produce bioactive compounds, many of which have been used as antitumor or antimicrobial agents, immunosuppressants, vaccine antigens or insecticides. Fungi especially are known to be one of the major producers of bioactive compounds. In fact, in 2012 the number of bioactive natural compounds characterized was 80,000–100,000, of which, 15,000 were of fungal origin [1]. Some well-known examples of bioactive fungal compounds include penicillin, developed as an antibacterial [2], lovastatin developed as a cholesterol-lowering medication [3][4], cyclosporins developed as immunosuppressive agents [5] and taxol, which has been proved to have an antitumor activity [6][7]. Up to now, most of the characterized fungal natural products are from terrestrial habitats even though fungi also colonize fresh water and marine habitats. Aquatic fungi have been less well studied compared to terrestrial fungi, but they have generated an increasing level of interest in the last decade.
Fungi from marine habitats were first studied in the 1940s when Barghoorn and Linder (1944) demonstrated their occurrence [8]. First exhaustive definition of marine fungi distinguished obligate and facultative organisms. In other words, marine fungi were divided between those “that grow and sporulate exclusively in a marine or estuarine habitat from those from freshwater or terrestrial milieus able to grow and possibly to sporulate in the marine environment” [9]. In 2015, Jones et al. provided a review of the classification of marine fungi, including the accepted names of 1112 species of characterized fungi from marine habitats [10]. Recently, there has been an increased level of interest in marine fungi, especially in defining their diversity and ecological role. Along with studies on marine filamentous fungi and yeasts, several studies have focused on the diversity of fungi in extreme environments such as the subseafloor [11][12], hydrothermal fields [13], or deep-sea hydrothermal vents [14][15]. In many cases, their activity and function has also been described, using transcriptomics to better understand their ecological role in extreme ecosystems [16]. Such studies have suggested that fungal communities from extreme ecosystems might represent an untapped reservoir of potential novel bioactive molecules, and, in recent years, secondary metabolites (also called specialized metabolites) obtained from fungi isolated from fresh water and marine habitats have gained considerable attention. According to a study by Rateb and Ebel (2011), marine fungal secondary metabolites have various biological and pharmaceuticals properties. In total, 690 novel structures of secondary metabolites have been identified from marine fungi between mid-2006 and 2010 [17]. At this time, 6222 molecules have been referenced from Dikarya fungi in MarineLit database (http://pubs.rsc.org/marinlit/). Evaluation of the biotechnological potential of marine-derived fungi has revealed that they produce mainly polyketides, terpenoids and peptides [17][18]. In addition, whole genome sequence analysis suggests that many deep-sea fungi have the potential to produce bioactive compounds. Indeed, Rédou et al. (2015) provided evidence that almost all fungal isolates in their study (96% of the 200 filamentous fungi and yeasts) had at least one gene involved in a secondary metabolite biosynthesis pathway [19], for example genes that encode enzymes involved in polyketides production (Polyketide Synthase, PKS), terpenoids (Terpene synthases, TPS) or non-ribosomal peptides (Non-Ribosomal Peptide Synthetase, NRPS). Along with this study, several studies have demonstrated antimicrobial activities of diverse marine fungi [20], deep-subseafloor fungi [21], and antimicrobial, antitumoral and antioxidant activities of sponge-associated marine fungi [22].
Polyketides represent a major and highly diverse group of natural products [17][23]. With a wide range of complex structures including macrolides, polyphenols, polyenes, and polyethers, polyketides comprise several groups of biologically important secondary metabolites such as flavonoids, phloroglucinols, resorcinols, stilbenes, pyrones, curcuminoids and chalcones [24][25]. Polyketides have numerous functions and applications as pigments, antibiotics, immunosuppressants, antioxidants, antiparasitics, cholesterol-lowering, and antitumoral agents [26]. Polyketide biosynthesis requires specific enzymes known as polyketide synthases (PKS). PKSs are a large group of enzymes, divided into three classes: type I, II or III [26][27], where type I PKSs are large multi-domain enzymes able to function in either a modular or iterative manner, type II PKSs are dissociable multi-enzyme complexes functioning in an iterative manner and type III PKSs are homodimeric enzymes, mechanistically different from the two other subgroups of PKSs, and structurally simpler [25][28]. Type III PKSs share common characteristics: they form dimers. Each monomer with a size of 40–45 kDa contains a conserved Cys-His-Asn catalytic triad within an active site cavity [25]. Polyketides are produced by iterative decarboxylative condensations of malonyl-CoA and diverse acyl-CoA thioesters (as extender and starter units), and cyclization reactions [28]. Despite their simplicity type III PKSs have an unusually wide range of substrates and play an important role in the biosynthesis of various bioactive polyketide compounds in different organisms [29]. For example, chalcones are produced by land plants [30][31]. Phlorotannins are unique to brown algae and phloroglucinols can be synthesized by diverse kind of organisms from prokaryotes (Pseudomonas) to eukaryotes (brown algae) [31][32][33][34][35]. Type III PKSs have also recently been discovered in fungi [36]. The first characterized fungal type III PKS was isolated from Neurospora crassa [37] and since then, several fungal type III PKSs have been reported. Among them, type III PKSs isolated from Aspergillus oryzae [38][39][40] and another from Aspergillus niger [41] have been characterized, as well as a type III PKS isolated from Sporotrichum laxum which appears to be involved in the production of spirolaxine, a potential drug candidate with anti-Helicobacter pylori activity [42]. A recent analysis reveals a total of 552 type III pks genes from 1193 fungal genomes (JGI Mycocosm) [43]. Only eleven type III PKSs have been biochemically characterized in fungal kingdom [43]. Polyketides produced by fungal type III PKS are grouped into triketide pyrones, tetraketide pyrones and alkylesorcinols [25]. All enzymes are isolated from terrestrial fungi. To date, no marine fungal enzymes have been described.
2. Discussion
Genes encoding type III Polyketides synthases are found in Dikarya such as Aspergillus niger, Aspergillus oryzae, Neurospora crassa, Botrytis cinerea or Sordaria macrospora [37][38][39][44][45][46], and recent genomic studies by Navarro-Munoz and Collemare revealed that there are 522 genes encoding these enzymes among 1193 fungal genomes [43]. However, currently, very few fungal pyrone synthase enzymes have been described biochemically. Indeed, there are only 11 PKSIII enzymes from fungi that have been functionally characterized and these all belong to fungal species of terrestrial origin. In this work, we have described the first biochemical characterization of PKSIII from marine fungi, a yeast, Naganishia uzbekistanensis strain Mo29 (UBOCC-A-208024) (formerly named as Cryptococcus sp.).
Marine fungi were first described in the 1980s and 1990s [9][47]. Thanks to numerous oceanographic cruises, we now know that it is possible to find many fungal species in the marine environment. Among these, the species found mainly pertained to Dikarya (Basidiomycota and Ascomycota). These species are found in different places in the marine environment such as the water column, associated with macro-organisms like algae, deep-sea sediments and hydrothermal vents [48]. The N. uzbekistanensis strain Mo29 (UBOCC-A-208024) was discovered during the MoMARDREAM-Naut oceanographic cruise (2007), on the exploration of the hydrothermal sources of the Rainbow site (-2300 m, Mid-Atlantic Ridge) [14]. Genome sequencing of this fungal species led to identify a gene encoding a PKSIII [49]. Phylogenetic analysis of the PKSIII protein sequence with enzymes from plants and bacteria, revealed that all fungal PKSIII enzymes (619 protein sequences) were grouped into a single branch. The PKSIII enzyme from N. uzbekistanensis strain Mo29 strain (UBOCC-A-208024) was not included in the fungal group/cluster. Instead, it clustered with 3 other protein sequences identified in another strain of Naganishia sp. and two other strains of Cryptococcus sp. [43]. This suggests that sequences of these three PKSIII proteins have followed an independent and early separation in the evolution compared to the other fungal sequences of PKSIII included in this study.
Interestingly, one of the features of these four enzymes is the presence of an additional 74 amino acids extension at the C-terminus. Previous studies in Sordaria macrospora and Botrytis cinerea have revealed that PKSIII may have a C-terminus extension [44][45] but the contribution of this region to the enzymatic function in these organisms is not known. Here, by expressing forms of the PKSIII Mo29 protein with and without this C-terminus extension, we show that it is not required for the enzymatic activity. Both forms are functional and produce mainly pyrones and some alkylresorcinols.
Analysis of the predicted structure of PKSIII Mo29 identified three catalytic residues: cysteine at position 171 (171C), Histidine at position 352 (352H) and Arginine at position 385 (385N). The next step is to mutate one or the three catalytic residues. This experiment could prove that these residues are involved in the enzymatic activity.
Previous studies of PKSIII have shown that another essential structure for the PKSIII protein is the “tunnel” which allows the entry of different starters such as lauroyl-CoA or stearoyl-CoA. These starters will either be cyclized or branched to the cycle synthesized by malonyl-CoA in the tunnel structure. The majority of the amino acids involved in the tunnel in PKSIII Mo29 protein are identical or similar to those found in the structure of PKSIII from N. crassa, from A. oryzae and B. cinerea (Table 1) [38][39][41][50][51][44]. Based on the structure of the PKSIII of N. crassa (3E1H) and the models of B. cinerea and A. niger, some amino acids explaining structural differences could be found. In position 144 of the PKSIII from N. uzbekistanensis strain Mo29, there is a tyrosine (144Y) whereas in N. crassa and the two other fungal proteins, there are two asparagines (PKSIIINc 125N and BcPKS 138N) and an alanine (AnPKS 140A) (amino acids not having the same chemical characteristics). Similarly, at position 212, there is a cysteine (212C), while in the three other fungal sequences, there is a non-polar amino acid (PKSIIINc 190V, BcPKS 203V and AnPKS 205L). In position 236, the PKSIII Mo29 has a serine (that is a polar amino acid) whereas in the three other fungi, there is a non-polar amino acid (PKSIIINc 206G, BcPKS 219G and AnPKS 221A). The other notable difference is in the amino acid that could bind to stearoyl-CoA. For the PKSIII Mo29, at position 355, there is a serine whereas in the other proteins, a nonpolar (309A), an aromatic (321A) or a positively charged amino acid (320R) are found, respectively, in PKSIIINc, BcPKS and AnPKS. These different amino acid changes in the structure of the PKSIII Mo29 protein could explain the fact that it does not synthetize short chain pyrones like some PKSIII proteins characterized biochemically in A. oryzae (AnCsyA and AnCsyB) [39][40].
Table 1. Comparison of residues lining the substrate-binding region (acyl-CoA starter) in different fungal and bacterial PKSIII proteins.
| Protein | Residues Near the Active Site (aa Involved in the Tunnel and Binding with Stearoyl-CoA) | |||||||||||||||||||||
| PKSIIINc | 86F | 120C | 121T | 125N | 186S | 189M | 190V | 206G | 207I | 210F | 211S | 250L | 252F | 261V | 306P | 307G | 308G | 309A | 310T | 311I | 312L | 313S |
| PKS18 | 109F | 143S | 144T | 148A | 205C | 210V | 211F | 220I | 221H | 224F | 225G | 264I | 266L | 275C | 314P | 315G | 316G | 317P | 318K | 319I | 320I | 321E |
| AoCsyA | 101F | 135C | 136T | 140N | 201C | 204F | 205F | 221A | 222M | 225F | 226G | 266I | 268F | 277P | 323P | 324G | 325G | 326Y | 327S | 328I | 329A | 330V |
| AoCsyB | 89F | 123C | 124T | 128H | 189P | 192F | 193A | 209A | 210M | 213F | 214G | 254A | 256F | 265A | 311P | 312G | 313G | 314Y | 315A | 316V | 317L | 318V |
| BcPKS | 99F | 133C | 134T | 138N | 199S | 202L | 203V | 219G | 220V | 223F | 224S | 263L | 265F | 274V | 318P | 319G | 320G | 321A | 322T | 323I | 324L | 325T |
| AnPKS | 101F | 135V | 136T | 140A | 201C | 204H | 205L | 221A | 222P | 225F | 226S | 265M | 267Y | 276A | 317P | 318G | 319G | 320R | 321A | 322V | 323I | 324Q |
| PKSIII Mo29 | 105F | 139C | 140T | 144Y | 208T | 211L | 212C | 236S | 237L | 240F | 241S | 283L | 285F | 294A | 352P | 353G | 354G | 355S | 356L | 357I | 358I | 359S |
| Protein | Residues of active site | |||||||||||||||||||||
| PKSIIINc | 152C | 305H | 338N | |||||||||||||||||||
| PKS18 | 175C | 313H | 346N | |||||||||||||||||||
| AoCsyA | 167C | 322H | 355N | |||||||||||||||||||
| AoCsyB | 155C | 310H | 343N | |||||||||||||||||||
| BcPKS | 165C | 317H | 350N | |||||||||||||||||||
| AnPKS | 167C | 316H | 349N | |||||||||||||||||||
| PKSIII Mo29 | 171C | 351H | 385N | |||||||||||||||||||
We overexpressed the entire PKSIII Mo29 protein as well as a truncated form lacking the additional 74 aa at the C-terminus. These two proteins forms are dimeric and are active. Indeed, the 74 aa extension in C-terminus does not seem to be involved in the enzymatic activity as in B. cinerea and in S. macrospora [44][45]. The two proteins are able to produce compounds in the presence of malonyl-CoA but always in the presence of an acyl-CoA starter with a carbon chain longer than 6 units. PKSIII Mo29 does not incorporate small carbon chains (acetyl-CoA, malonyl-CoA (only) and hexanoyl-CoA). No molecules are synthesized in the presence of these three starters. The known fungal pyrone synthases catalyse reactions starting from the acyl-CoA chain and ending with aldol cyclization and/or lactonization [25]. The fungal PKSIII that have been experimentally characterized are divided into two functional groups. The first group uses only long acyl-CoA chains with several malonyl-CoA. We find in this group enzymes from N. crassa, A. niger, B. cinerea and S. macrospora [50][44][45]. The second group consists of two enzymes found in A. oryzae [38][39][40]. These two enzymes can use starter molecules with chains of different lengths to produce triketide, tetraketide pyrones and alkylresorcinols. These results show that the Mo29 enzyme belongs to the first group (like the enzymes of N. crassa, A. niger, B. cinerea and S. macrospora). PKSIII Mo29 produces triketide and tetraketide pyrones in the presence of starter such as octanoyl-CoA, decanoyl-CoA, lauroyl-CoA and palmytoyl-CoA (Table 2). Another study allowed us to highlight that this protein could accommodate very long chain fatty acyl-CoA starter units produced by E. coli. Indeed, molecules synthesized by proteins overexpressed by E. coli are secreted into the culture medium. Analysis of molecules found in the culture medium show that the two types of proteins (Std and Red) are able to synthesis several molecules having 14 carbons (C14) to 28 carbons (C28). However, the majority of molecules possess 18 and 20 to 24 carbons. Among these molecules are pyrones and alkylresorcinols. The most synthesized molecules are C22H36O3 (m/z 348.2663) and C20H32O3 (m/z 320.2349) which are pyrones (triketide pyrones) and C23H38O2 (m/z 346.2869) (alkylresorcinol) (Figure 1).
Table 2. Products synthetized by PKSIII Mo29 Std and PKSIII Mo29 Red in vitro. Molecules were analyzed and detected by HPLC-Q-ToFMS. (ND: Not detected).
| Starter Substrat | Products | Rt LC/min | Parent Ion m/z | Fragment Ion m/z |
|---|---|---|---|---|
| Acetyl-CoA | ND | / | / | |
| Malonyl-CoA | ND | / | / | |
| Hexanoyl-CoA | ND | / | / | |
| Octanoyl-CoA | C12H18O3 | 17.50 | 209.1300 | ND |
| Decanoyl-CoA | C14H22O3 | 19.80 | 238.1567 | 194.1599 |
| Lauroyl-CoA | C16H26O3 | 22.05 | 266.1884 | 222.1952 |
| Palmytoyl-CoA | C20H34O3 | 26.03 | 322.2507 | 278.2554 |
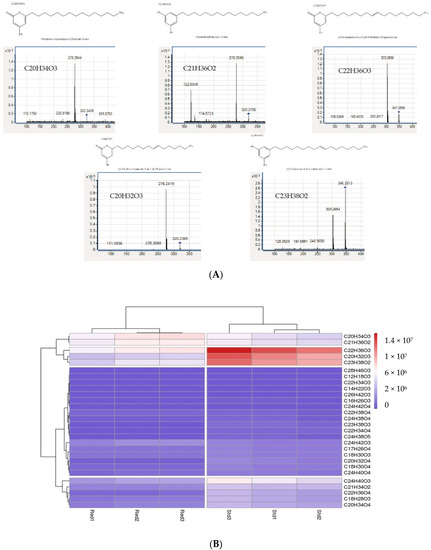
Figure 1. (A) Product identification by HPLC-Q-ToFMS. These MS/MS spectra indicate that triketide pyrones and alkylresorcinol are synthesized by PKSIII Mo29 in the presence of acyl-CoA in E. coli. (B) Heatmap. Synthesized molecules abundance from pksIII Mo29 std and pksIII Mo29 red expression in E. coli. Molecules were clustered based on the E. coli supernatants.
The PKSIII Mo29 Std and PKSIII Mo29 Red synthesize the same molecules, pyrones and alkylresorcinols. However, the truncated form seems to produce less product than the entire recombinant protein. These results were observed in In vitro experiments and in E. coli culture medium. In the E. coli culture medium, all molecules are less produced by PKSIII Mo29 Red than PKSIII Mo29 Std. These results could be explained by the absence of the C-terminus 74 aa extension. The extension could be implicated in the dimer stability or substrates binding. In this study, we have tested only classical substrates and starters such as palmitoyl-CoA; however, we did not identify natural substrates in N. uzbekistanensis strain Mo29 strain (UBOCC-A-208024) yet. So, the extension could be playing a role in the entrance of other substrates than those used in vitro.
Several α-pyrones produced by marine and terrestrial fungi have shown cytotoxic activities against several cancer cell lines [52][53][54][55]. The different molecules produced by PKSIII Mo29 In vitro have been brought into contact with different cancer cell lines, such as Caco-2 and THP1 (colon cancer cell lines and leukemic cell lines). Molecule produced by PKSIII Mo29 in the presence of malonyl-CoA and palmitoyl-CoA is a α-pyrone with a molecular formula of C20H34O3 (m/z 321.2507). When the Caco-2 cells and the THP1 are exposed for 48 h in the presence of 0.012 g/L of this molecule, the cell viability decreases. Therefore, it would mean that this α-pyrone has a cytotoxic effect on these two tumors cell lines. The molecule produced by PKSIII Mo29 in the presence of malonyl-CoA and lauroyl-CoA In vitro is also a triketide pyrone with a molecular formula of C16H26O3 (m/z 266.1884). THP1 cells were incubated for 48 h with 0.012 g/L of this molecule. Cell viability was reduced by 5%. These two molecules seem to have cytotoxic effects on cancer cell lines like Caco-2 and THP1.
3. Conclusions
The PKSIII of N. uzbekistanensis strain Mo29 (UBOCC-A-208024) is the first Polyketide Synthase of type III from a marine fungus to be described. This enzyme produces long α-pyrones and alkylresorcinols. These compounds are produced by an enzyme which have some different amino acids changed in the tunnel structure. But, to prove this hypothesis, different amino acid could be replaced by other residues using site-directed mutagenesis and mutated proteins will be tested with short starter like hexanoyl-CoA. In this study, we have discovered that some molecules produced by PKSIII Mo29 have a cytotoxic effect on two tumoral cell lines. At this time, these molecules have no antimicrobial effects on different bacteria (data not shown). We tried to discover molecules produced in N. uzbekistanensis strain Mo29 strain (UBOCC-A-208024) by PKSIII. However, at this time, we are unable to determine which natural molecules are synthetized by PKSIII Mo29. Therefore, it is important to continue the exploration of this strain to understand the biosynthetic pathway of polyketides in this marine yeast.
References
- Berdy, J. Thoughts and facts about antibiotics: Where we are now and where we are heading. J. Antibiot. 2012, 65, 385–395.
- Fleming, A. On the antibacterial action of cultures of a penicillium, with special reference to their use in the isolation of B. influenzae. Br. J. Exp. Pathol. 1929, 10, 226.
- Alberts, A.W.; Chen, J.; Kuron, G.; Hunt, V.; Huff, J.; Hoffman, C.; Rothrock, J.; Lopez, M.; Joshua, H.; Harris, E.; et al. Mevinolin: A highly potent competitive inhibitor of hydroxymethylglutaryl-coenzyme A reductase and a cholesterol-lowering agent. Proc. Natl. Acad. Sci. USA 1980, 77, 3957–3961.
- Barrios-Gonzalez, J.; Miranda, R.U. Biotechnological production and applications of statins. Appl. Microbiol. Biotechnol. 2010, 85, 869–883.
- Fliri, H.; Baumann, G.; Enz, A.; Kallen, J.; Luyten, M.; Mikol, V.; Movva, R.; Quesniaux, V.; Schreier, M.; Walkinshaw, M.; et al. Cyclosporins. Structure-activity relationships. Ann. N. Y. Acad. Sci. 1993, 696, 47–53.
- Demain, A.L.; Vaishnav, P. Natural products for cancer chemotherapy. Microb. Biotechnol. 2011, 4, 687–699.
- Stierle, A.; Strobel, G.; Stierle, D. Taxol and taxane production by Taxomyces andreanae, an endophytic fungus of Pacific yew. Science 1993, 260, 214–216.
- Barghoorn, E.S. Marine fungi: Their taxonomy and biology. Farlowia 1944, 1, 395–467.
- Kohlmeyer, J.; Kohlmeyer, E. Marine Mycology: The Higher Fungi; Press, N.Y.A., Ed.; Elsevier: Amsterdam, The Netherlands, 1979; p. 690.
- Jones, E.B.G.; Suetrong, S.; Sakayaroj, J.; Bahkali, A.; Abdel-Wahab, M.; Boekhout, T.; Pang, K.-L. Classification of marine Ascomycota, Basidiomycota, Blastocladiomycota and Chytridiomycota. Fungal Divers. 2015, 73, 1–72.
- Ciobanu, M.-C.; Burgaud, G.; Dufresne, A.; Breuker, A.; Rédou, V.; Ben Maamar, S.; Gaboyer, F.; Vandenabeele-Trambouze, O.; Lipp, J.S.; Schippers, A.; et al. Microorganisms persist at record depths in the subseafloor of the Canterbury Basin. ISME J. 2014, 8, 1370–1380.
- Orsi, W.; Biddle, J.F.; Edgcomb, V. Deep Sequencing of Subseafloor Eukaryotic rRNA Reveals Active Fungi across Marine Subsurface Provinces. PLoS ONE 2013, 8, e56335.
- Gadanho, M.; Sampaio, J.P. Occurrence and diversity of yeasts in the mid-atlantic ridge hydrothermal fields near the Azores Archipelago. Microb. Ecol. 2005, 50, 408–417.
- Burgaud, G.; Arzur, D.; Durand, L.; Cambon-Bonavita, M.-A.; Barbier, G. Marine culturable yeasts in deep-sea hydrothermal vents: Species richness and association with fauna. FEMS Microbiol. Ecol. 2010, 73, 121–133.
- Burgaud, G.; Le Calvez, T.; Arzur, D.; Vandenkoornhuyse, P.; Barbier, G. Diversity of culturable marine filamentous fungi from deep-sea hydrothermal vents. Environ. Microbiol. 2009, 11, 1588–1600.
- Pachiadaki, M.G.; Rédou, V.; Beaudoin, D.J.; Burgaud, G.; Edgcomb, V.P. Fungal and Prokaryotic Activities in the Marine Subsurface Biosphere at Peru Margin and Canterbury Basin Inferred from RNA-Based Analyses and Microscopy. Front. Microbiol. 2016, 7, 846.
- Rateb, M.E.; Ebel, R. Secondary metabolites of fungi from marine habitats. Nat. Prod. Rep. 2011, 28, 290–344.
- Ebada, S.S.; Proksch, P. Marine-derived fungal metabolites. In Springer Handbook of Marine Biotechnology; Springer: Berlin/Heidelberg, Germany, 2015; pp. 759–788.
- Rédou, V.; Navarri, M.; Meslet-Cladière, L.; Barbier, G.; Burgaud, G. Species Richness and Adaptation of Marine Fungi from Deep-Subseafloor Sediments. Appl. Environ. Microbiol. 2015, 81, 3571–3583.
- Silber, J.; Kramer, A.; Labes, A.; Tasdemir, D. From discovery to production: Biotechnology of marine fungi for the production of new antibiotics. Mar. Drugs 2016, 14, 137.
- Navarri, M.; Jegou, C.; Meslet-Cladiere, L.; Brillet, B.; Barbier, G.; Burgaud, G.; Fleury, Y. Deep Subseafloor Fungi as an Untapped Reservoir of Amphipathic Antimicrobial Compounds. Mar. Drugs 2016, 14, 50.
- Cheng, M.-M.; Tang, X.-L.; Sun, Y.-T.; Song, D.-Y.; Cheng, Y.-J.; Liu, H.; Li, P.-L.; Li, G.-Q. Biological and Chemical Diversity of Marine Sponge-Derived Microorganisms over the Last Two Decades from 1998 to 2017. Molecules 2020, 25, 853.
- Wang, X.; Zhang, Z.; Dong, X.; Feng, Y.; Liu, X.; Gao, B.; Wang, J.; Zhang, L.; Wang, J.; Shi, S.; et al. Identification and functional characterization of three type III polyketide synthases from Aquilaria sinensis calli. Biochem. Biophys. Res. Commun. 2017, 486, 1040–1047.
- Abe, I. Novel applications of plant polyketide synthases. Curr. Opin. Chem. Biol. 2012, 16, 179–185.
- Shimizu, Y.; Ogata, H.; Goto, S. TypeIII Polyketide Synthases: Functional Classification and Phylogenomics. ChemBioChem 2017, 18, 50–65.
- Staunton, J.; Weissman, K.J. Polyketide biosynthesis: A millennium review. Nat. Prod. Rep. 2001, 18, 380–416.
- Shen, B. Polyketide biosynthesis beyond the type I, II and III polyketide synthase paradigms. Curr. Opin. Chem. Biol. 2003, 7, 285–295.
- Lim, Y.P.; Go, M.K.; Yew, W.S. Exploiting the Biosynthetic Potential of Type III Polyketide Synthases. Molecules 2016, 21, 806.
- Lee, J.S. Recent advances in the synthesis of 2-pyrones. Mar. Drugs 2015, 13, 1581–1620.
- Austin, M.B.; Noel, J.P. The chalcone synthase superfamily of type III polyketide synthases. Nat. Prod. Rep. 2003, 20, 79–110.
- Ferrer, J.L.; Jez, J.M.; Bowman, M.E.; Dixon, R.A.; Noel, J.P. Structure of chalcone synthase and the molecular basis of plant polyketide biosynthesis. Nat. Struct. Biol. 1999, 6, 775–784.
- Achkar, J.; Xian, M.; Zhao, H.; Frost, J. Biosynthesis of phloroglucinol. J. Am. Chem. Soc. 2005, 127, 5332–5333.
- Funa, N.; Ohnishi, Y.; Fujii, I.; Shibuya, M.; Ebizuka, Y.; Horinouchi, S. A new pathway for polyketide synthesis in microorganisms. Nature 1999, 400, 897–899.
- Katsuyama, Y.; Ohnishi, Y. Chapter Sixteen—Type III Polyketide Synthases in Microorganisms. In Methods in Enzymology; Hopwood, D.A., Ed.; Academic Press: Cambridge, MA, USA, 2012; Volume 515, pp. 359–377.
- Meslet-Cladière, L.; Delage, L.; Leroux, C.J.-J.; Goulitquer, S.; Leblanc, C.; Creis, E.; Gall, E.A.; Stiger-Pouvreau, V.; Czjzek, M.; Potin, P. Structure/Function Analysis of a Type III Polyketide Synthase in the Brown Alga Ectocarpus siliculosus Reveals a Biochemical Pathway in Phlorotannin Monomer Biosynthesis. Plant Cell 2013, 25, 3089–3103.
- Yu, D.; Xu, F.; Zeng, J.; Zhan, J. Type III polyketide synthases in natural product biosynthesis. IUBMB Life 2012, 64, 285–295.
- Funa, N.; Awakawa, T.; Horinouchi, S. Pentaketide Resorcylic Acid Synthesis by Type III Polyketide Synthase from Neurospora crassa. J. Biol. Chem. 2007, 282, 14476–14481.
- Seshime, Y.; Juvvadi, P.R.; Kitamoto, K.; Ebizuka, Y.; Fujii, I. Identification of csypyrone B1 as the novel product of Aspergillus oryzae type III polyketide synthase CsyB. Bioorg. Med. Chem. 2010, 18, 4542–4546.
- Seshime, Y.; Juvvadi, P.R.; Kitamoto, K.; Ebizuka, Y.; Nonaka, T.; Fujii, I. Aspergillus oryzae type III polyketide synthase CsyA is involved in the biosynthesis of 3,5-dihydroxybenzoic acid. Bioorg. Med. Chem. Lett. 2010, 20, 4785–4788.
- Yu, D.; Zeng, J.; Chen, D.; Zhan, J. Characterization and reconstitution of a new fungal type III polyketide synthase from Aspergillus oryzae. Enzym. Microb. Technol. 2010, 46, 575–580.
- Li, J.; Luo, Y.; Lee, J.-K.; Zhao, H. Cloning and characterization of a type III polyketide synthase from Aspergillus niger. Bioorg. Med. Chem. Lett. 2011, 21, 6085–6089.
- Sun, L.; Wang, S.; Zhang, S.; Yu, D.; Qin, Y.; Huang, H.; Wang, W.; Zhan, J. Identification of a type III polyketide synthase involved in the biosynthesis of spirolaxine. Appl. Microbiol. Biotechnol. 2016, 100, 7103–7113.
- Navarro-Muñoz, J.C.; Collemare, J. Evolutionary Histories of Type III Polyketide Synthases in Fungi. Front. Microbiol. 2020, 10, 3018.
- Jeya, M.; Kim, T.-S.; Tiwari, M.K.; Li, J.; Zhao, H.; Lee, J.-K. The Botrytis cinerea type III polyketide synthase shows unprecedented high catalytic efficiency toward long chain acyl-CoAs. Mol. Biosyst. 2012, 8, 2864–2867.
- Ramakrishnan, D.; Tiwari, M.K.; Manoharan, G.; Sairam, T.; Thangamani, R.; Lee, J.-K.; Marimuthu, J. Molecular characterization of two alkylresorcylic acid synthases from Sordariomycetes fungi. Enzym. Microb. Technol. 2018, 115, 16–22.
- Kirimura, K.; Watanabe, S.; Kobayashi, K. Heterologous gene expression and functional analysis of a type III polyketide synthase from Aspergillus niger NRRL 328. Biochem. Biophys. Res. Commun. 2016, 473, 1106–1110.
- Rédou, V.; Vallet, M.; Meslet-Cladière, L.; Kumar, A.; Pang, K.-L.; Pouchus, Y.-F.; Barbier, G.; Grovel, O.; Bertrand, S.; Prado, S.; et al. Marine Fungi. In The Marine Microbiome: An Untapped Source of Biodiversity and Biotechnological Potential; Stal, L.J., Cretoiu, M.S., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 99–153.
- Pang, K.-L.; Overy, D.P.; Jones, E.B.G.; da Luz Calado, M.; Burgaud, G.; Walker, A.K.; Johnson, J.A.; Kerr, R.G.; Cha, H.-J.; Bills, G.F. ‘Marine fungi’ and ‘marine-derived fungi’ in natural product chemistry research: Toward a new consensual definition. Fungal Biol. Rev. 2016, 30, 163–175.
- Rédou, V.; Kumar, A.; Hainaut, M.; Henrissat, B.; Record, E.; Barbier, G.; Burgaud, G. Draft Genome Sequence of the Deep-Sea Basidiomycetous Yeast Cryptococcus sp. Strain Mo29 Reveals Its Biotechnological Potential. Genome Announc. 2016, 4.
- Goyal, A.; Saxena, P.; Rahman, A.; Singh, P.K.; Kasbekar, D.P.; Gokhale, R.S.; Sankaranarayanan, R. Structural insights into biosynthesis of resorcinolic lipids by a type III polyketide synthase in Neurospora crassa. J. Struct. Biol. 2008, 162, 411–421.
- Seshime, Y.; Juvvadi, P.R.; Fujii, I.; Kitamoto, K. Discovery of a novel superfamily of type III polyketide synthases in Aspergillus oryzae. Biochem. Biophys. Res. Commun. 2005, 331, 253–260.
- Gong, J.; Chen, C.; Mo, S.; Liu, J.; Wang, W.; Zang, Y.; Li, H.; Chai, C.; Zhu, H.; Hu, Z.; et al. Fusaresters A-E, new γ-pyrone-containing polyketides from fungus Fusarium sp. Hungcl and structure revision of fusariumin D. Org. Biomol. Chem. 2019, 17, 5526–5532.
- Han, W.; Cai, J.; Zhong, W.; Xu, G.; Wang, F.; Tian, X.; Zhou, X.; Liu, Q.; Liu, Y.; Wang, J. Protein tyrosine phosphatase 1B (PTP1B) inhibitorsfrom the deep-sea fungus Penicillium chrysogenum SCSIO 07007. Bioorg. Chem. 2020, 96, 103646.
- Keisham, S.S. Pyrone-derived Marine Natural Products: A Review on Isolation, Bio-activities and Synthesis. Curr. Org. Chem. 2020, 24, 354–401.
- Zhu, H.; Li, D.; Yan, Q.; An, Y.; Huo, X.; Zhang, T.; Zhang, M.; Wang, C.; Xia, M.; Ma, X.; et al. α-Pyrones, secondary metabolites from fungus Cephalotrichum microsporum and their bioactivities. Bioorg. Chem. 2019, 83, 129–134.





