
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | José LUIS García-Gimenez | -- | 2979 | 2024-02-07 23:26:27 | | | |
| 2 | Lindsay Dong | Meta information modification | 2979 | 2024-02-08 08:54:24 | | |
Video Upload Options
The advent of immune checkpoint inhibitors (ICIs) has represented a breakthrough in the treatment of many cancers, although a high number of patients fail to respond to ICIs, which is partially due to the ability of tumor cells to evade immune system surveillance. Non-coding microRNAs (miRNAs) have been shown to modulate the immune evasion of tumor cells, and there is thus growing interest in elucidating whether these miRNAs could be targetable or proposed as novel biomarkers for prognosis and treatment response to ICIs.
1. Introduction
1.1. miRNAs That Modulate Response to ICIs
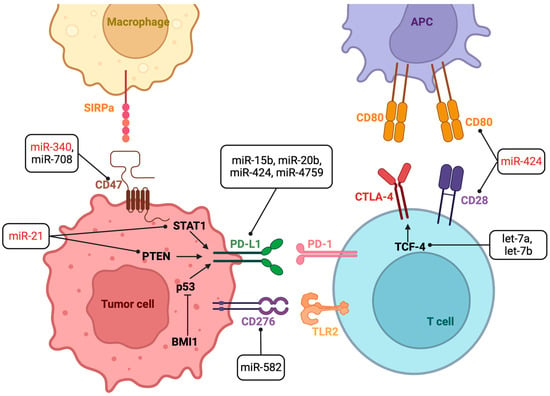
| miRNA | Cancer | miRNA Target Gene | ICI | Experimental Model | Effect on ICI Response | Refs. |
|---|---|---|---|---|---|---|
| let-7a and let-7b | Head and neck squamous cell carcinoma | TCF-4 * | anti CTLA-4 | Overexpressing let-7a/b tumor cells inoculated into mice + anti CTLA-4 | H | [13] |
| miR-15b-5p | Colorectal cancer | PD-L1 * | anti PD-1 | Overexpressing miR-15b-5p tumor cells inoculated into mice + anti PD-1. | H | [14] |
| miR-16-5p | Lung cancer | anti PD-L1 | Tumor cell + overexpressing miR-16-5p exosomes + anti PD-L1 | H | [15] | |
| miR-20b-5p | Lung cancer | PD-L1 * | anti PD-1 | Tumor cells transfected with miRNA mimic + Pembrolizumab | H | [16] |
| Breast cancer | Tumor cells transfected with miRNA mimic + Pembrolizumab | H | ||||
| miR-21 | Oral squamous cell carcinoma | PTEN | anti PD-L1 | Tumor cells inoculated into mice + miR-21 knockdown tumor-derived exosomes + anti-PD-L1 | L | [17] |
| Melanoma | STAT1 * | anti PD-1 | Tumor cells and knocked down miR-21 tumor-associated macrophages (TAM) subcutaneously injected in mice + anti PD-1 | L | [18] | |
| miR-128a | Laryngeal squamous cell carcinoma | BMI1 * | anti PD-1 | Overexpressing miR-128a tumor cells + Pembrolizumab | H | [19] |
| miR-155 | Metastatic melanoma | anti PD-1 + anti PD-L1 + anti CTLA-4 |
Tumor cells inoculated into modified mice for knockout of miR-155 in CD4/8 T cells + anti PD-1, anti PD-L1 and anti CTLA-4 | L | [20] | |
| Diffuse large B-cell lymphoma | PD-L1 * | anti PD-L1 | Overexpressing miR-155 tumor cells inoculated into mice + anti PD-L1 | L | [21] | |
| Breast cancer | SOCS1 | anti PD-L1 | Overexpressing miR-155 tumor cells inoculated into mice + anti PD-L1 | H | [22] | |
| Melanoma | PD-L1 * | anti PD-L1 | Overexpressing miR-155 tumor cells co-cultured with peripheral blood mononuclear cells + anti PD-L1 | H | [23] | |
| miR-340 | Pancreatic carcinoma | CD47 * | anti CD47 | Overexpressing miR-340 tumor cells inoculated into mice + anti CD47 | L | [24] |
| miR-424 | Colorectal cancer | CD28 and CD80 * | anti PD-1 + anti CTLA-4 | Tumor cells inoculated into miR-424 knocked mice + anti PD-1 and anti CTLA-4 | L | [25] |
| Mouse cecum orthotopic colorectal cancer + miR-424 knocked tumor cell-derived extracellular vesicles + anti PD-1 and anti CTLA-4 | L | |||||
| Hepatocellular carcinoma | PD-L1 | anti PD-L1 | Tumor cells inoculated into mice + nanobubbles carrying miR-424 mimic and anti PD-L1 | H | [26] | |
| miR-582 | B-cell precursor acute lymphoblastic leukemia | CD276 * | anti CD276 | Overexpressing miR-582 tumor cells co-cultured with NK cells + anti CD276 | H | [27] |
| miR-708 | T-acute lymphoblastic leukemia | CD47 * | anti CD47 | Overexpressing miR-708 tumor cells + anti CD47. | H | [28] |
| miR-4759 | Breast cancer | PD-L1 * | anti PD-L1 | Overexpressing miR-4759 tumor cells co-cultured with peripheral blood mononuclear cells + anti PD-L1 | H | [29] |
1.2. miRNAs Modulated after Response to ICIs
| ICI | Experimental Model | miRNA | Experimental Effect on miRNA | Refs. |
|---|---|---|---|---|
| anti PD-1 | miRNA analysis in peripheral lymphocytes from 21 good responders (complete response, partial response, or stable disease) with metastatic renal cell carcinoma before and after a 4-weeks period (2 cycles) of nivolumab administration | miR-99a, miR-708, miR-655, miR-582-3p, miR-492, miR-487a, miR-485-3p, miR-449a, miR-433, miR-431, miR-429, miR-376c, miR-342-5p, miR-340, miR-339-5p, miR-335, miR-324-5p, miR-25, miR-24, miR-22, miR-221, miR-214, miR-194, miR-18b, miR-152, miR-143, miR-100, miR-let-7ev | High levels of expression in peripheral lymphocytes after treatment compared to before treatment in good responders. | [31] |
| miRNA analysis in peripheral lymphocytes from 17 good long-responders (complete response, partial response or stable disease and progression-free survival (PFS) > 18 months) with metastatic renal cell carcinoma before and after a 4-weeks period (2 cycles) of nivolumab administration | miR-22, miR-24, miR-99a, miR-194, miR-214, miR-335, miR-339, miR-708 | High expression levels in peripheral lymphocytes after treatment compared to before treatment in good responders. | ||
| anti CTLA-4 + anti PD-1 |
Plasma from stage IV melanoma non-responders (13 patients), partial response (4 patients) and complete response (5 patients) before and after Ipilimumab and Nivolumab/Pembrolizumab treatment | miR-4649-3p and miR-615-3p | Increased levels in post- vs. pre-treatment in non-responders. No changes post- vs. pre-treatment in patients with partial response. Decreased levels post- vs. pre-treatment in patients with complete response. | [32] |
1.3. miRNAs Related to ICIs Response
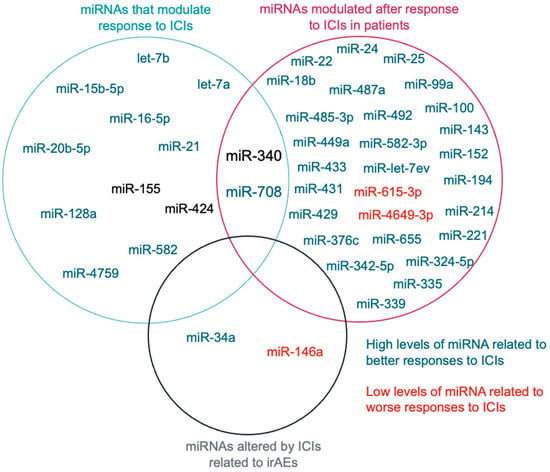
2. miRNAs Related to Immune Checkpoint Inhibitor Response
2.1. miRNAs That Modulate Response to ICIs
Several miRNAs directly or indirectly regulate PD-L1 expression (Table 1). The let-7 family (let-7a, let-7b, let-7c, let-7d and let-7e) is thought to mediate tumor suppression in cancer by inhibiting tumor cell proliferation, promoting cell death evasion or metastasis [33] but also alterations to immunity. Let-7 was significantly downregulated in tissue from head and neck squamous cell carcinoma patients compared to healthy tissue [13]. Let-7 downregulation has been observed in other types of cancer associated with reduced copy numbers, such as melanoma [34], with an upregulation of LIN28A/LIN28B, which is an RNA binding protein that inhibits Drosha or Dicer binding during let-7 biogenesis (breast cancer) [35], and with DNA hypermethylation (epithelial ovarian cancer) [36].
2.2. miRNAs Modulated after Response to ICIs
2.3. miRNAs That Regulate Immune-Related Adverse Events (irAEs)
2.4. miRNAs Related to ICIs Response
References
- Pandya, P.H.; Murray, M.E.; Pollok, K.E.; Renbarger, J.L. The Immune System in Cancer Pathogenesis: Potential Therapeutic Approaches. J. Immunol. Res. 2016, 2016, 4273943.
- Kanchan, R.K.; Doss, D.; Khan, P.; Nasser, M.W.; Mahapatra, S. To Kill a Cancer: Targeting the Immune Inhibitory Checkpoint Molecule, B7-H3. Biochim. Biophys. Acta (BBA)—Rev. Cancer 2022, 1877, 188783.
- Khan, M.; Arooj, S.; Wang, H. Soluble B7-CD28 Family Inhibitory Immune Checkpoint Proteins and Anti-Cancer Immunotherapy. Front. Immunol. 2021, 12, 651634.
- Esensten, J.H.; Helou, Y.A.; Chopra, G.; Weiss, A.; Bluestone, J.A. CD28 Costimulation: From Mechanism to Therapy. Immunity 2016, 44, 973–988.
- Saadi, W.; Fatmi, A.; Pallardó, F.V.; García-Giménez, J.L.; Mena-Molla, S. Long Non-Coding RNAs as Epigenetic Regulators of Immune Checkpoints in Cancer Immunity. Cancers 2022, 15, 184.
- Lee, Y.; Martin-Orozco, N.; Zheng, P.; Li, J.; Zhang, P.; Tan, H.; Park, H.J.; Jeong, M.; Chang, S.H.; Kim, B.-S.; et al. Inhibition of the B7-H3 Immune Checkpoint Limits Tumor Growth by Enhancing Cytotoxic Lymphocyte Function. Cell Res. 2017, 27, 1034–1045.
- Yang, H.; Xun, Y.; You, H. The Landscape Overview of CD47-Based Immunotherapy for Hematological Malignancies. Biomark. Res. 2023, 11, 15.
- Yang, F.; Shay, C.; Abousaud, M.; Tang, C.; Li, Y.; Qin, Z.; Saba, N.F.; Teng, Y. Patterns of Toxicity Burden for FDA-Approved Immune Checkpoint Inhibitors in the United States. J. Exp. Clin. Cancer Res. 2023, 42, 4.
- De Risi, I.; Sciacovelli, A.M.; Guida, M. Checkpoint Inhibitors Immunotherapy in Metastatic Melanoma: When to Stop Treatment? Biomedicines 2022, 10, 2424.
- Ratner, L.; Waldmann, T.A.; Janakiram, M.; Brammer, J.E. Rapid Progression of Adult T-Cell Leukemia–Lymphoma after PD-1 Inhibitor Therapy. N. Engl. J. Med. 2018, 378, 1947–1948.
- Ulas, E.B.; Hashemi, S.M.S.; Houda, I.; Kaynak, A.; Veltman, J.D.; Fransen, M.F.; Radonic, T.; Bahce, I. Predictive Value of Combined Positive Score and Tumor Proportion Score for Immunotherapy Response in Advanced NSCLC. JTO Clin. Res. Rep. 2023, 4, 100532.
- Jiang, P.; Gu, S.; Pan, D.; Fu, J.; Sahu, A.; Hu, X.; Li, Z.; Traugh, N.; Bu, X.; Li, B.; et al. Signatures of T Cell Dysfunction and Exclusion Predict Cancer Immunotherapy Response. Nat. Med. 2018, 24, 1550–1558.
- Yu, D.; Liu, X.; Han, G.; Liu, Y.; Zhao, X.; Wang, D.; Bian, X.; Gu, T.; Wen, L. The Let-7 Family of microRNAs Suppresses Immune Evasion in Head and Neck Squamous Cell Carcinoma by Promoting PD-L1 Degradation. Cell Commun. Signal. 2019, 17, 173.
- Liu, C.; Liu, R.; Wang, B.; Lian, J.; Yao, Y.; Sun, H.; Zhang, C.; Fang, L.; Guan, X.; Shi, J.; et al. Blocking IL-17A Enhances Tumor Response to Anti-PD-1 Immunotherapy in Microsatellite Stable Colorectal Cancer. J. Immunother. Cancer 2021, 9, e001895.
- Chen, H.-L.; Luo, Y.-P.; Lin, M.-W.; Peng, X.-X.; Liu, M.-L.; Wang, Y.-C.; Li, S.-J.; Yang, D.-H.; Yang, Z.-X. Serum Exosomal miR-16-5p Functions as a Tumor Inhibitor and a New Biomarker for PD-L1 Inhibitor-Dependent Immunotherapy in Lung Adenocarcinoma by Regulating PD-L1 Expression. Cancer Med. 2022, 11, 2627–2643.
- Jiang, K.; Zou, H. microRNA-20b-5p Overexpression Combing Pembrolizumab Potentiates Cancer Cells to Radiation Therapy via Repressing Programmed Death-Ligand 1. Bioengineered 2022, 13, 917–929.
- Li, L.; Cao, B.; Liang, X.; Lu, S.; Luo, H.; Wang, Z.; Wang, S.; Jiang, J.; Lang, J.; Zhu, G. Microenvironmental Oxygen Pressure Orchestrates an Anti- and pro-Tumoral Γδ T Cell Equilibrium via Tumor-Derived Exosomes. Oncogene 2019, 38, 2830–2843.
- Xi, J.; Huang, Q.; Wang, L.; Ma, X.; Deng, Q.; Kumar, M.; Zhou, Z.; Li, L.; Zeng, C.; Young, K.H.; et al. miR-21 Depletion in Macrophages Promotes Tumoricidal Polarization and Enhances PD-1 Immunotherapy. Oncogene 2018, 37, 3151–3165.
- Chen, H.; Guo, Y.; Huang, J.; Zhou, L. Upregulating Hsa-miR-128a Increased the Effects of Pembrolizumab on Laryngeal Cancer Cells via the P53 Pathway. Biomed. Res. Int. 2021, 2021, 2342784.
- Huffaker, T.B.; Lee, S.-H.; Tang, W.W.; Wallace, J.A.; Alexander, M.; Runtsch, M.C.; Larsen, D.K.; Thompson, J.; Ramstead, A.G.; Voth, W.P.; et al. Antitumor Immunity Is Defective in T Cell-Specific microRNA-155-Deficient Mice and Is Rescued by Immune Checkpoint Blockade. J. Biol. Chem. 2017, 292, 18530–18541.
- Zheng, Z.; Sun, R.; Zhao, H.-J.; Fu, D.; Zhong, H.-J.; Weng, X.-Q.; Qu, B.; Zhao, Y.; Wang, L.; Zhao, W.-L. MiR155 Sensitized B-Lymphoma Cells to Anti-PD-L1 Antibody via PD-1/PD-L1-Mediated Lymphoma Cell Interaction with CD8+T Cells. Mol. Cancer 2019, 18, 54.
- Wang, J.; Wang, Q.; Guan, Y.; Sun, Y.; Wang, X.; Lively, K.; Wang, Y.; Luo, M.; Kim, J.A.; Murphy, E.A.; et al. Breast Cancer Cell-Derived microRNA-155 Suppresses Tumor Progression via Enhancing Immune Cell Recruitment and Antitumor Function. J. Clin. Investig. 2022, 132, e157248.
- Vaxevanis, C.K.; Friedrich, M.; Tretbar, S.U.; Handke, D.; Wang, Y.; Blümke, J.; Dummer, R.; Massa, C.; Seliger, B. Identification and Characterization of Novel CD274 (PD-L1) Regulating microRNAs and Their Functional Relevance in Melanoma. Clin. Transl. Med. 2022, 12, e934.
- Xi, Q.; Zhang, J.; Yang, G.; Zhang, L.; Chen, Y.; Wang, C.; Zhang, Z.; Guo, X.; Zhao, J.; Xue, Z.; et al. Restoration of miR-340 Controls Pancreatic Cancer Cell CD47 Expression to Promote Macrophage Phagocytosis and Enhance Antitumor Immunity. J. Immunother. Cancer 2020, 8, e000253.
- Zhao, X.; Yuan, C.; Wangmo, D.; Subramanian, S. Tumor-Secreted Extracellular Vesicles Regulate T-Cell Costimulation and Can Be Manipulated to Induce Tumor-Specific T-Cell Responses. Gastroenterology 2021, 161, 560–574.e11.
- Liu, Y.; Xie, Q.; Ma, Y.; Lin, C.; Li, J.; Hu, B.; Liu, C.; Zhao, Y. Nanobubbles Containing PD-L1 Ab and miR-424 Mediated PD-L1 Blockade, and Its Expression Inhibition to Enable and Potentiate Hepatocellular Carcinoma Immunotherapy in Mice. Int. J. Pharm. 2022, 629, 122352.
- Li, X.; Zhang, Y.; He, F.; Gao, D.; Che, B.; Cao, X.; Huang, S.; Zheng, M.; Han, H. miR-582 Suppresses the Proliferation of B-Cell Precursor Acute Lymphoblastic Leukemia (BCP-ALL) Cells and Protects Them from Natural Killer Cell-Mediated Cytotoxicity. Front. Immunol. 2022, 13, 853094.
- Huang, W.; Wang, W.-T.; Fang, K.; Chen, Z.-H.; Sun, Y.-M.; Han, C.; Sun, L.-Y.; Luo, X.-Q.; Chen, Y.-Q. MIR-708 Promotes Phagocytosis to Eradicate T-ALL Cells by Targeting CD47. Mol. Cancer 2018, 17, 12.
- Lin, Y.-Z.; Liu, S.-H.; Wu, W.-R.; Shen, Y.-C.; Wang, Y.-L.; Liao, C.-C.; Lin, P.-L.; Chang, H.; Liu, L.-C.; Cheng, W.-C.; et al. miR-4759 Suppresses Breast Cancer through Immune Checkpoint Blockade. Comput. Struct. Biotechnol. J. 2021, 20, 241–251.
- Newman, A.M.; Liu, C.L.; Green, M.R.; Gentles, A.J.; Feng, W.; Xu, Y.; Hoang, C.D.; Diehn, M.; Alizadeh, A.A. Robust Enumeration of Cell Subsets from Tissue Expression Profiles. Nat. Methods 2015, 12, 453–457.
- Incorvaia, L.; Fanale, D.; Badalamenti, G.; Brando, C.; Bono, M.; De Luca, I.; Algeri, L.; Bonasera, A.; Corsini, L.R.; Scurria, S.; et al. A “Lymphocyte MicroRNA Signature” as Predictive Biomarker of Immunotherapy Response and Plasma PD-1/PD-L1 Expression Levels in Patients with Metastatic Renal Cell Carcinoma: Pointing towards Epigenetic Reprogramming. Cancers 2020, 12, 3396.
- Bustos, M.A.; Gross, R.; Rahimzadeh, N.; Cole, H.; Tran, L.T.; Tran, K.D.; Takeshima, L.; Stern, S.L.; O’Day, S.; Hoon, D.S.B. A Pilot Study Comparing the Efficacy of Lactate Dehydrogenase Levels Versus Circulating Cell-Free microRNAs in Monitoring Responses to Checkpoint Inhibitor Immunotherapy in Metastatic Melanoma Patients. Cancers 2020, 12, 3361.
- Zhou, J.; Xiang, H.; Cao, Z. Dual Mechanism of Let-7i in Tumor Progression. Front. Oncol. 2023, 13, 1253191.
- Zhang, L.; Huang, J.; Yang, N.; Greshock, J.; Megraw, M.S.; Giannakakis, A.; Liang, S.; Naylor, T.L.; Barchetti, A.; Ward, M.R.; et al. microRNAs Exhibit High Frequency Genomic Alterations in Human Cancer. Proc. Natl. Acad. Sci. USA 2006, 103, 9136–9141.
- Piskounova, E.; Polytarchou, C.; Thornton, J.E.; Hagan, J.P.; LaPierre, R.J.; Pothoulakis, C.; Iliopoulos, D.; Gregory, R.I. Oncogenic Lin28A and Lin28B Inhibit Let-7 microRNA Biogenesis by Distinct Mechanisms. Cell 2011, 147, 1066–1079.
- Lu, L.; Katsaros, D.; de la Longrais, I.A.R.; Sochirca, O.; Yu, H. Hypermethylation of Let-7a-3 in Epithelial Ovarian Cancer Is Associated with Low Insulin-like Growth Factor-II Expression and Favorable Prognosis. Cancer Res. 2007, 67, 10117–10122.
- Baer, C.; Squadrito, M.L.; Laoui, D.; Thompson, D.; Hansen, S.K.; Kiialainen, A.; Hoves, S.; Ries, C.H.; Ooi, C.-H.; De Palma, M. Suppression of microRNA Activity Amplifies IFN-γ-Induced Macrophage Activation and Promotes Anti-Tumour Immunity. Nat. Cell Biol. 2016, 18, 790–802.
- Hsu, J.-M.; Xia, W.; Hsu, Y.-H.; Chan, L.-C.; Yu, W.-H.; Cha, J.-H.; Chen, C.-T.; Liao, H.-W.; Kuo, C.-W.; Khoo, K.-H.; et al. STT3-Dependent PD-L1 Accumulation on Cancer Stem Cells Promotes Immune Evasion. Nat. Commun. 2018, 9, 1908.
- Chen, Y.; Xie, C.; Zheng, X.; Nie, X.; Wang, Z.; Liu, H.; Zhao, Y. LIN28/Let-7/PD-L1 Pathway as a Target for Cancer Immunotherapy. Cancer Immunol. Res. 2019, 7, 487–497.
- Palamarchuk, A.; Tsyba, L.; Tomasello, L.; Pekarsky, Y.; Croce, C.M. PDCD1 (PD-1) Is a Direct Target of miR-15a-5p and miR-16-5p. Signal Transduct. Target. Ther. 2022, 7, 12.
- Kao, S.C.; Cheng, Y.Y.; Williams, M.; Kirschner, M.B.; Madore, J.; Lum, T.; Sarun, K.H.; Linton, A.; McCaughan, B.; Klebe, S.; et al. Tumor Suppressor microRNAs Contribute to the Regulation of PD-L1 Expression in Malignant Pleural Mesothelioma. J. Thorac. Oncol. 2017, 12, 1421–1433.
- van Zandwijk, N.; Pavlakis, N.; Kao, S.C.; Linton, A.; Boyer, M.J.; Clarke, S.; Huynh, Y.; Chrzanowska, A.; Fulham, M.J.; Bailey, D.L.; et al. Safety and Activity of microRNA-Loaded Minicells in Patients with Recurrent Malignant Pleural Mesothelioma: A First-in-Man, Phase 1, Open-Label, Dose-Escalation Study. Lancet Oncol. 2017, 18, 1386–1396.
- Quagliariello, V.; Passariello, M.; Di Mauro, A.; Cipullo, C.; Paccone, A.; Barbieri, A.; Palma, G.; Luciano, A.; Buccolo, S.; Bisceglia, I.; et al. Immune Checkpoint Inhibitor Therapy Increases Systemic SDF-1, Cardiac DAMPs Fibronectin-EDA, S100/Calgranulin, Galectine-3, and NLRP3-MyD88-Chemokine Pathways. Front. Cardiovasc. Med. 2022, 9, 930797.
- Jonas, K.; Prinz, F.; Ferracin, M.; Krajina, K.; Pasculli, B.; Deutsch, A.; Madl, T.; Rinner, B.; Slaby, O.; Klec, C.; et al. MiR-4649-5p Acts as a Tumor-Suppressive microRNA in Triple Negative Breast Cancer by Direct Interaction with PIP5K1C, Thereby Potentiating Growth-Inhibitory Effects of the AKT Inhibitor Capivasertib. Breast Cancer Res. 2023, 25, 119.
- Afra, F.; Mahboobipour, A.A.; Salehi Farid, A.; Ala, M. Recent Progress in the Immunotherapy of Hepatocellular Carcinoma: Non-Coding RNA-Based Immunotherapy May Improve the Outcome. Biomed. Pharmacother. 2023, 165, 115104.
- Saini, S.; Yamamura, S.; Majid, S.; Shahryari, V.; Hirata, H.; Tanaka, Y.; Dahiya, R. MicroRNA-708 Induces Apoptosis and Suppresses Tumorigenicity in Renal Cancer Cells. Cancer Res. 2011, 71, 6208–6219.
- de Oliveira, J.C.; Scrideli, C.A.; Brassesco, M.S.; Yunes, J.A.; Brandalise, S.R.; Tone, L.G. MiR-708-5p Is Differentially Expressed in Childhood Acute Lymphoblastic Leukemia but Not Strongly Associated to Clinical Features. Pediatr. Blood Cancer 2015, 62, 177–178.
- Xia, W.; Zou, C.; Chen, H.; Xie, C.; Hou, M. Immune Checkpoint Inhibitor Induces Cardiac Injury through Polarizing Macrophages via Modulating microRNA-34a/Kruppel-like Factor 4 Signaling. Cell Death Dis. 2020, 11, 575.
- Xia, W.; Chen, H.; Chen, D.; Ye, Y.; Xie, C.; Hou, M. PD-1 Inhibitor Inducing Exosomal miR-34a-5p Expression Mediates the Cross Talk between Cardiomyocyte and Macrophage in Immune Checkpoint Inhibitor-Related Cardiac Dysfunction. J. Immunother. Cancer 2020, 8, e001293.
- Marschner, D.; Falk, M.; Javorniczky, N.R.; Hanke-Müller, K.; Rawluk, J.; Schmitt-Graeff, A.; Simonetta, F.; Haring, E.; Dicks, S.; Ku, M.; et al. MicroRNA-146a Regulates Immune-Related Adverse Events Caused by Immune Checkpoint Inhibitors. JCI Insight 2020, 5, 132334.
- Ivanova, E.; Asadullina, D.; Rakhimov, R.; Izmailov, A.; Izmailov, A.; Gilyazova, G.; Galimov, S.; Pavlov, V.; Khusnutdinova, E.; Gilyazova, I. Exosomal miRNA-146a Is Downregulated in Clear Cell Renal Cell Carcinoma Patients with Severe Immune-Related Adverse Events. Non-Coding RNA Res. 2022, 7, 159–163.
- Shimada, B.K.; Yang, Y.; Zhu, J.; Wang, S.; Suen, A.; Kronstadt, S.M.; Jeyaram, A.; Jay, S.M.; Zou, L.; Chao, W. Extracellular miR-146a-5p Induces Cardiac Innate Immune Response and Cardiomyocyte Dysfunction. Immunohorizons 2020, 4, 561–572.
- Mann, M.; Mehta, A.; Zhao, J.L.; Lee, K.; Marinov, G.K.; Garcia-Flores, Y.; Lu, L.-F.; Rudensky, A.Y.; Baltimore, D. An NF-κB-microRNA Regulatory Network Tunes Macrophage Inflammatory Responses. Nat. Commun. 2017, 8, 851.
- Li, M.; Shan, W.; Hong, B.; Zou, J.; Li, H.; Han, D.; Zhang, Y.; Li, L.; Li, D.; Lin, W. Circulating miR-92b and miR-375 for Monitoring the Chemoresistance and Prognosis of Small Cell Lung Cancer. Sci. Rep. 2020, 10, 12705.
- Ahmad, N.; Haider, S.; Jagannathan, S.; Anaissie, E.; Driscoll, J.J. MicroRNA Theragnostics for the Clinical Management of Multiple Myeloma. Leukemia 2014, 28, 732–738.
- García-Giménez, J.L.; Mena-Mollá, S.; Beltrán-García, J.; Sanchis-Gomar, F. Challenges in the Analysis of Epigenetic Biomarkers in Clinical Samples. Clin. Chem. Lab. Med. 2017, 55, 1474–1477.
- Androvic, P.; Benesova, S.; Rohlova, E.; Kubista, M.; Valihrach, L. Small RNA-Sequencing for Analysis of Circulating miRNAs: Benchmark Study. J. Mol. Diagn. 2022, 24, 386–394.





