Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Tomomitsu Hatakeyama | -- | 1885 | 2023-07-13 03:10:38 | | | |
| 2 | Jason Zhu | Meta information modification | 1885 | 2023-07-13 03:39:40 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Hatakeyama, T.; Unno, H. Lectins as Toxins from Marine Animals. Encyclopedia. Available online: https://encyclopedia.pub/entry/46719 (accessed on 26 March 2026).
Hatakeyama T, Unno H. Lectins as Toxins from Marine Animals. Encyclopedia. Available at: https://encyclopedia.pub/entry/46719. Accessed March 26, 2026.
Hatakeyama, Tomomitsu, Hideaki Unno. "Lectins as Toxins from Marine Animals" Encyclopedia, https://encyclopedia.pub/entry/46719 (accessed March 26, 2026).
Hatakeyama, T., & Unno, H. (2023, July 13). Lectins as Toxins from Marine Animals. In Encyclopedia. https://encyclopedia.pub/entry/46719
Hatakeyama, Tomomitsu and Hideaki Unno. "Lectins as Toxins from Marine Animals." Encyclopedia. Web. 13 July, 2023.
Copy Citation
Due to their remarkable structural diversity, glycans play important roles as recognition molecules on cell surfaces of living organisms. Carbohydrates exist in numerous isomeric forms and can adopt diverse structures through various branching patterns. Despite their relatively small molecular weights, they exhibit extensive structural diversity. On the other hand, lectins, also known as carbohydrate-binding proteins, not only recognize and bind to the diverse structures of glycans but also induce various biological reactions based on structural differences. Initially discovered as hemagglutinins in plant seeds, lectins have been found to play significant roles in cell recognition processes in higher vertebrates.
lectin
carbohydrate
marine animal
toxin
pore-forming protein
1. Introduction
Many plant lectins have the ability to induce cellular toxicity, likely due to their role in self-defense against invading organisms such as pathogenic bacteria or insects [1][2]. This toxicity arises from their capacity to cross-link surface carbohydrate-containing molecules on target cells, influencing the signaling pathways within those cells. Moreover, when incorporated into protein toxins, lectin domains can function as cell-binding modules to exert potent toxicity. A well-known example of a toxic lectin is ricin, which was the first discovered toxic lectin derived from the castor bean (Ricinus communis) [3]. Ricin consists of an A chain with N-glycosidase activity that inactivates eukaryotic ribosomes and a B chain that serves as a lectin subunit. The B chain assists in delivering the A chain to the cytosol by binding to cell surface glycans [4][5]. Similar toxins with enzymatic “A” subunits and receptor-binding “B” subunits (lectin subunits) are referred to as AB toxins and are also produced by various pathogenic bacteria [6]. Another type of toxic lectin includes pore-forming proteins, which form ion-permeable pores in the target cell membrane after binding to cell surface glycans through the lectin domains. These examples demonstrate the important roles that lectins often play in the actions of protein toxins. It has also been revealed that venomous marine animals produce various toxins containing lectin domains, suggesting a close association between their carbohydrate-binding activity and their toxic actions.
2. Lectin from the Venomous Sea Urchin Toxopneustes pileolus
The venom of the globiferous pedicellariae of the venomous sea urchin T. pileolus contains several toxic proteins that exert various biological effects [7]. Among these proteins, the galactose-specific lectin SUL-I exhibits various activities, such as chemotactic activity on guinea pig neutrophils and mitogenic activity on murine splenocytes [8][9]. The amino acid sequence of SUL-I indicates that it belongs to the L-rhamnose-binding lectins (RBLs), many of which have been found in fish eggs [10]. SUL-I is composed of 284 residues (30.5 kDa) with three repetitive sequence regions, each showing similarity to CRDs of rhamnose-binding lectins (RBLs) [11]. The three-dimensional structure of the rSUL-I/L-rhamnose complex was determined by X-ray crystallographic analysis with a resolution of 1.8 Å (Figure 1) [12]. The overall structure of rSUL-I consists of three distinctive CRDs, which share structural similarities with the CRDs of CSL3, the rhamnose-binding lectin from chum salmon (Oncorhynchus keta) eggs [13]. The bound L-rhamnose molecules are primarily recognized by rSUL-I through hydrogen bonds between their 2-, 3-, and 4-hydroxy groups and Asp, Asn, and Glu residues in the binding sites. When interactions of rSUL-I with oligosaccharides were examined, a higher affinity was observed for galactosylated (asialylated) N-glycans, suggesting that the potential target carbohydrate structures are galactose-terminated N-glycans. While the CRDs of rSUL-I adopt similar folds compared to those of CSL3, a significant difference was found around the variable loop in the carbohydrate-binding sites, which may be related to the difference in binding specificities for oligosaccharides between these lectins. SUL-I has carbohydrate-binding sites in its three domains, all oriented towards the same side of the protein. This structure could be advantageous for cross-linking specific membrane glycoproteins containing galactose-terminated carbohydrate chains, thereby triggering cellular responses.
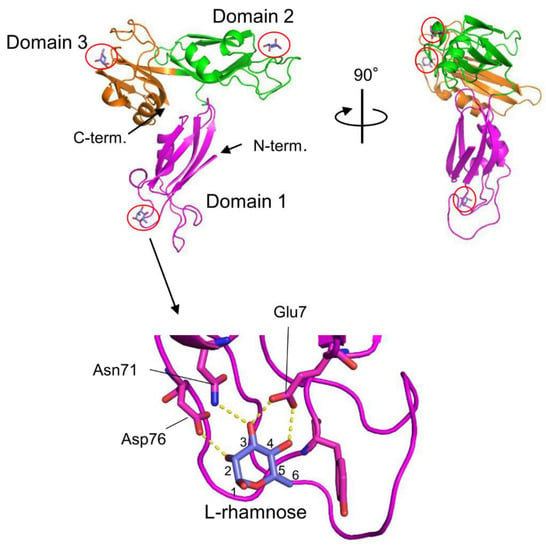
Figure 1. Crystal structures of SUL-I complexed with L-rhamnose (PDB ID: 5H4S). SUL-I is composed of three distinct domains (domains 1–3), each possessing a carbohydrate-binding site at its apex. The domains are presented in different colors. The bound L-rhamnose molecules are enclosed by red circles. In domain 1, L-rhamnose is recognized through hydrogen bonds formed between its 2-, 3-, and 4-hydroxy groups and Asp76, Asn71, and Glu7. L-Rhamnose molecules are similarly recognized in domains 2 and 3.
3. Hemolytic Lectin from the Sea Cucumber Cucumaria echinata
CEL-III is a galactose-specific lectin from C. echinata, which also contains the previously mentioned C-type lectins CEL-I and CEL-IV. Although CEL-III shows Ca2+-dependent carbohydrate binding, it belongs to the R-(ricin)-type lectin family, characterized by two β-trefoil folds [14]. Interestingly, it exhibits strong hemolytic activity and cytotoxicity [15]. Hemolysis induced by this lectin is prominent in the alkaline region above pH 7 in the presence of Ca2+ and can be inhibited by galactose and carbohydrates containing galactose at nonreducing ends. These observations suggest that the Ca2+-dependent galactose binding ability is a prerequisite for its hemolytic action. CEL-III mediates hemolysis by forming pores in the target erythrocytes, relying on its galactose-specific lectin activity. Structural analyses of CEL-III have revealed that it consists of two R-type lectin domains (domains 1 and 2) in the N-terminal two-thirds and a C-terminal one-third with a β-sheet-rich novel domain (domain 3) (Figure 2) [16][17]. Domain 1 contains two carbohydrate-binding sites, while domain 2 contains three. The binding of galactose-related carbohydrates is mediated by coordinate bonds with Ca2+ ions bound in the carbohydrate-binding sites, along with hydrogen bond networks involving nearby amino acid residues. This mechanism shares similarities with the carbohydrate-recognition mechanism of C-type CRDs, despite the lack of homology in the amino acid sequences between CEL-III and C-type CRDs.
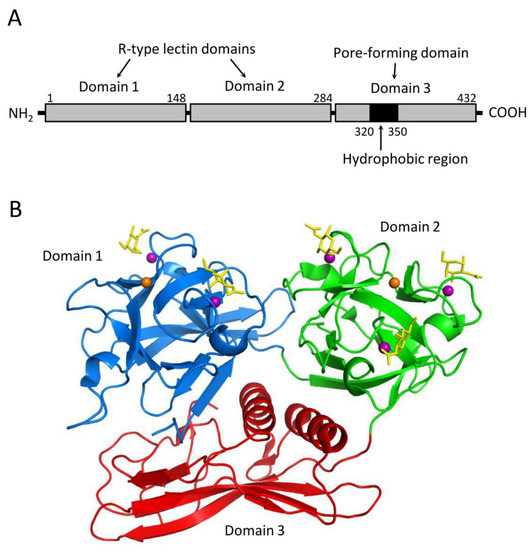
Figure 2. Structure of the hemolytic lectin CEL-III (PDB ID: 2Z48). (A) CEL-III comprises three domains. Domains 1 and 2 are R-type (β-trefoil) lectin domains, while domain 3 is a pore-forming domain that contains a hydrophobic region. (B) Crystal structure of the CEL-III/GalNAc complex. Domains 1 and 2 bind galactose-related carbohydrates by utilizing Ca2+ ions in a similar manner to C-type lectins. Domain 3 exhibits a unique structure abundant in β-strands, along with two α-helices that correspond to the hydrophobic region shown in (A). The bound GalNAc molecules are depicted as yellow stick models.
The crystal structure of the soluble oligomer of CEL-III, induced by the binding of lactulose, a galactose-containing disaccharide, revealed that CEL-III heptamerizes upon binding to galactose-containing carbohydrates (Figure 3). This binding leads to the formation of a β-barrel structure (25 Å) composed of the C-terminal β-sheet-rich domain [17]. When these conformational changes occur on the target cell surface, the β-barrel forms a membrane pore, allowing the passage of small molecules and ions across the cell membrane. This drastic change in the tertiary structure of domain 3, from a soluble form to a membrane-penetrating β-barrel, is driven by the formation of numerous hydrogen bonds within the β-barrel and hydrophobic interactions between the hydrophobic face of the β-barrel and the nonpolar interior of the membrane.
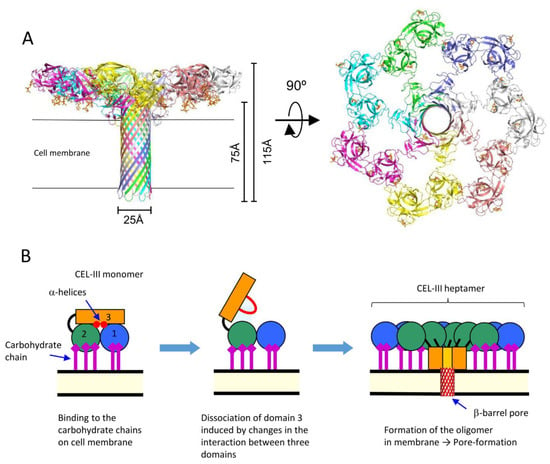
Figure 3. Oligomerization of CEL-III. (A) The crystal structure of the CEL-III heptamer (PDB ID: 3W9T). Monomer units of CEL-III are depicted in different colors. The α-helical regions in domain 3 have changed to a β-barrel structure, which is assumed to penetrate the target cell membrane. (B) Schematic drawing for presumed oligomerization processes. After binding to carbohydrate chains on the target cell surface, domain 3 may dissociate from domains 1 and 2. Concomitantly with heptamerization, the α-helical regions of domain 3 change their conformation to β-strands, forming a transmembrane pore composed of a β-barrel.
Oligomerization of CEL-III can be triggered by β-galactoside-containing disaccharides, including lactose (Galβ1-4Glc), lactulose (Galβ1-4Fru), and N-acetyllactosamine (Galβ1-4GlcNAc), but not by galactose, N-acetylgalactosamine, and melibiose (Galα1-6Glc) [18]. This suggests that heptamerization of CEL-III on the target cell surface is induced by cell surface receptors containing β-galactoside linkages, particularly glycosphingolipids such as lactosyl ceramide, which has been demonstrated to be an effective receptor for membrane pore formation in artificial lipid vesicles [19]. It appears that the drastic conformational change of CEL-III is triggered by the structural changes in the binding of the specific carbohydrate into domains 1 and 2, which are then transmitted to domain 3, leading to the opening of the interface with domain 3 [17].
The presence of β-trefoil fold domains containing QXW motifs [20] has been observed in a wide range of proteins, including lectins, toxins, carbohydrate-related enzymes, and immune cell surface receptors, which function as carbohydrate-binding modules. Structural analysis of the hemolytic lectin LSL, isolated from the mushroom Laetiporus sulphureus, has revealed that it also comprises an N-terminal carbohydrate-binding domain with a β-trefoil fold and a C-terminal pore-forming domain [21]. Notably, the pore-forming domain of this lectin exhibits remarkable structural similarity to those found in pore-forming toxins of pathogenic bacteria such as Aeromonas hydrophila and Clostridium perfringens [22]. Based on these findings, it can be inferred that LSL, similar to CEL-III, exhibits hemolytic activity by binding to cell surface glycans through its carbohydrate-binding domain and subsequently inserting a pore-forming domain into the cell membrane. Although the pore-forming domain of LSL does not share structural similarity with domain 3 of CEL-III, both domains are characterized by elongated β-sheets, which likely contribute to a favorable structure for penetrating the cell membrane.
4. Fish Spine Toxins Containing Lectin-like Domains
Some fish species possess stinging spines that contain protein toxins; among them, a toxin called natterin has been isolated from the spines of Thalassophryne nattereri [23]. Additionally, a homologous protein Ij-natterin has been found to be expressed in the spine venom of Inimicus japonicus [24]. Natterin and Ij-natterin consist of an N-terminal lectin domain, which is homologous to the oyster lectin CGL1 (DM9 domain protein) [25], and a C-terminal aerolysin-like pore-forming domain (Figure 4A). Although the specific carbohydrate-binding activity of the lectin domain has not been reported to date, its amino acid sequence similarity with CGL1 suggests that it may have mannose-specific lectin activity.
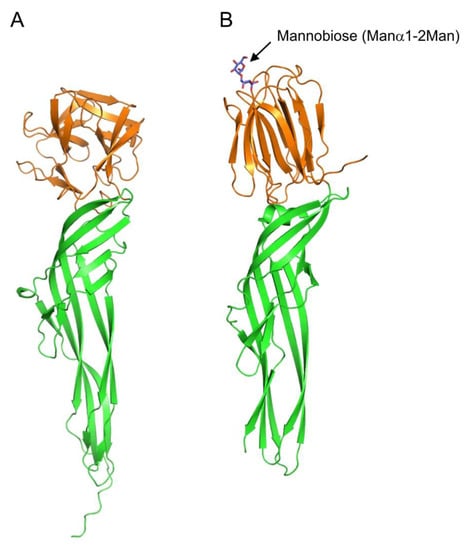
Figure 4. Comparison of tertiary structures of Ij-natterin and Dln1/mannobiose complex. (A) The tertiary structure of Ij-natterin predicted using Alphafold2 [26]. The N-terminal 50 residues are omitted due to their disordered nature. (B) The X-ray crystal structure of Dln1/mannobiose (Manα1-2Man) complex (PDB ID: 4ZNQ). The N-terminal domain demonstrates mannose-specific lectin activity. The bound mannobiose molecule is depicted as a stick model. In both proteins, the N- and C-terminal domains are colored orange and green, respectively.
Proteins that have C-terminal domains homologous to those of natterin, called natterin-like proteins, are also widely distributed in fish skin [27][28]. The crystal structure of Dln1, one of the natterin-like proteins from zebrafish (Danio rerio), has revealed that Dln1 consists of an N-terminal jacalin-like lectin domain and a C-terminal aerolysin-like domain (Figure 4B) [29]. The lectin domain was found to bind mannobiose (Manα1-2Man and Manα1-3Man), indicating that Dln1 exhibits an affinity for high-mannose glycans. Furthermore, electron microscopic analysis revealed that upon binding to high-mannose glycans, Dln1 undergoes significant conformational changes, leading to the formation of octameric pores in the lipid membrane.
Although natterin and natterin-like proteins possess C-terminal aerolysin-like domains in common, they differ in N-terminal lectin domains; the former shows a CGL1-like DM9 domain fold, while the latter has a jacalin-like domain. Considering the large conformational change during the pore-forming process of Dln1, it seems highly likely that natterin (and Ij-natterin) also induce significant conformational changes upon binding to target cell surface glycans, leading to oligomerization and pore formation. Similar large conformational changes after binding to cell surface carbohydrate chains are also observed in the course of pore formation by the hemolytic lectin CEL-III (Figure 3). The lectin domains of these pore-forming proteins not only play roles in binding to specific carbohydrate chains on the target cell but also trigger subsequent conformational changes leading to the formation of oligomers with transmembrane pores.
References
- Taylor, M.E.; Drickamer, K. Mammalian Sugar-Binding Receptors: Known Functions and Unexplored Roles. FEBS J. 2019, 286, 1800–1814.
- Taylor, M.E.; Drickamer, K. Convergent and Divergent Mechanisms of Sugar Recognition across Kingdoms. Curr. Opin. Struct. Biol. 2014, 28, 14–22.
- Olsnes, S. The History of Ricin, Abrin and Related Toxins. Toxicon 2004, 44, 361–370.
- Lord, J.M.; Roberts, L.M.; Robertus, J.D. Ricin: Structure, Mode of Action, and Some Current Applications. FASEB J. 1994, 8, 201–208.
- Hartley, M.R.; Lord, J.M. Cytotoxic Ribosome-Inactivating Lectins from Plants. Biochim. Biophys. Acta—Proteins Proteom. 2004, 1701, 1–14.
- Odumosu, O.; Nicholas, D.; Yano, H.; Langridge, W. AB Toxins: A Paradigm Switch from Deadly to Desirable. Toxins 2010, 2, 1612–1645.
- Zhang, Y.A.; Abe, J.; Siddiq, A.; Nakagawa, H.; Honda, S.; Wada, T.; Ichida, S. UT841 Purified from Sea Urchin (Toxopneustes Pileolus) Venom Inhibits Time-Dependent 45Ca2+ Uptake in Crude Synaptosome Fraction from Chick Brain. Toxicon 2001, 39, 1223–1229.
- Nakagawa, H.; Hashimoto, T.; Hayashi, H.; Shinohara, M.; Ohura, K.; Tachikawa, E. Isolation of a Novel Lectin from the Globiferous Pedicellariae of the Sea Urchin Toxopneustes Pileolus. Adv. Exp. Med. Biol. 1996, 391, 213–223.
- Takei, M.; Nakagawa, H. A Sea Urchin Lectin, SUL-1, from the Toxopneustid Sea Urchin Induces DC Maturation from Human Monocyte and Drives Th1 Polarization in Vitro. Toxicol. Appl. Pharmacol. 2006, 213, 27–36.
- Tateno, H. SUEL-Related Lectins, a Lectin Family Widely Distributed throughout Organisms. Biosci. Biotechnol. Biochem. 2010, 74, 1141–1144.
- Hatakeyama, T.; Ichise, A.; Yonekura, T.; Unno, H.; Goda, S.; Nakagawa, H. CDNA Cloning and Characterization of a Rhamnose-Binding Lectin SUL-I from the Toxopneustid Sea Urchin Toxopneustes Pileolus Venom. Toxicon 2015, 94, 8–15.
- Hatakeyama, T.; Ichise, A.; Unno, H.; Goda, S.; Oda, T.; Tateno, H.; Hirabayashi, J.; Sakai, H.; Nakagawa, H. Carbohydrate Recognition by the Rhamnose-Binding Lectin SUL-I with a Novel Three-Domain Structure Isolated from the Venom of Globiferous Pedicellariae of the Flower Sea Urchin Toxopneustes Pileolus. Protein Sci. 2017, 26, 1574–1583.
- Shirai, T.; Watanabe, Y.; Lee, M.; Ogawa, T.; Muramoto, K. Structure of Rhamnose-Binding Lectin CSL3: Unique Pseudo-Tetrameric Architecture of a Pattern Recognition Protein. J. Mol. Biol. 2009, 391, 390–403.
- Hatakeyama, T.; Unno, H.; Kouzuma, Y.; Uchida, T.; Eto, S.; Hidemura, H.; Kato, N.; Yonekura, M.; Kusunoki, M. C-Type Lectin-like Carbohydrate Recognition of the Hemolytic Lectin CEL-III Containing Ricin-Type β-Trefoil Folds. J. Biol. Chem. 2007, 282, 37826–37835.
- Hatakeyama, T.; Kohzaki, H.; Nagatomo, H.; Yamasaki, N. Purification and Characterization of Four Ca2+-Dependent Lectins from the Marine Invertebrate, Cucumaria Echinata. J. Biochem. 1994, 116, 209–214.
- Nakano, M.; Tabata, S.; Sugihara, K.; Kouzuma, Y.; Kimura, M.; Yamasaki, N. Primary Structure of Hemolytic Lectin CEL-III from Marine Invertebrate Cucumaria Echinata and Its CDNA: Structural Similarity to the B-Chain from Plant Lectin, Ricin. Biochim. Biophys. Acta—Protein Struct. Mol. Enzymol. 1999, 1435, 167–176.
- Unno, H.; Goda, S.; Hatakeyama, T. Hemolytic Lectin CEL-III Heptamerizes via a Large Structural Transition from α-Helices to a β-Barrel during the Transmembrane Pore Formation Process. J. Biol. Chem. 2014, 289, 12805–12812.
- Hatakeyama, T.; Furukawa, M.; Nagatomo, H.; Yamasaki, N.; Mori, T. Oligomerization of the Hemolytic Lectin CEL-III from the Marine Invertebrate Cucumaria Echinata Induced by the Binding of Carbohydrate Ligands. J. Biol. Chem. 1996, 271, 16915–16920.
- Hatakeyama, T.; Sato, T.; Taira, E.; Kuwahara, H.; Niidome, T.; Aoyagi, H. Characterization of the Interaction of Hemolytic Lectin CEL-III from the Marine Invertebrate, Cucumaria Echinata, with Artificial Lipid Membranes: Involvement of Neutral Sphingoglycolipids in the Pore-Forming Process. J. Biochem. 1999, 125, 277–284.
- Hazes, B. The Qxw Motif: A Flexilble Lectin Fold. Protein Sci. 1996, 5, 1490–1501.
- Mancheño, J.M.; Tateno, H.; Goldstein, I.J.; Martínez-Ripoll, M.; Hermoso, J.A. Structural Analysis of the Laetiporus Sulphureus Hemolytic Pore-Forming Lectin in Complex with Sugars. J. Biol. Chem. 2005, 280, 17251–17259.
- Podobnik, M.; Kisovec, M.; Anderluh, G. Molecular Mechanism of Pore Formation by Aerolysin-like Proteins. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372, 20160209.
- Magalhães, G.S.; Lopes-Ferreira, M.; Junqueira-De-Azevedo, I.L.M.; Spencer, P.J.; Araújo, M.S.; Portaro, F.C.V.; Ma, L.; Valente, R.H.; Juliano, L.; Fox, J.W.; et al. Natterins, a New Class of Proteins with Kininogenase Activity Characterized from Thalassophryne Nattereri Fish Venom. Biochimie 2005, 87, 687–699.
- Hatakeyama, T.; Kishigawa, A.; Unno, H. Molecular Cloning and Characterization of the Two Putative Toxins Expressed in the Venom of the Devil Stinger Inimicus Japonicus. Toxicon 2021, 201, 9–20.
- Unno, H.; Matsuyama, K.; Tsuji, Y.; Goda, S.; Hiemori, K.; Tateno, H.; Hirabayashi, J.; Hatakeyama, T. Identification, Characterization, and X-Ray Crystallographic Analysis of a Novel Type of Mannose-Specific Lectin CGL1 from the Pacific Oyster Crassostrea Gigas. Sci. Rep. 2016, 6, 29135.
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589.
- Rajan, B.; Patel, D.M.; Kitani, Y.; Viswanath, K.; Brinchmann, M.F. Novel Mannose Binding Natterin-like Protein in the Skin Mucus of Atlantic Cod (Gadus morhua). Fish Shellfish Immunol. 2017, 68, 452–457.
- Lima, C.; Disner, G.R.; Alice, M.; Falc, P.; Seni-silva, A.C.; Luis, A.; Maleski, A.; Souza, M.M.; Cristina, M.; Tonello, R.; et al. The Natterin Proteins Diversity: A Review on Phylogeny, Structure, and Immune Function. Toxins 2021, 13, 538.
- Jia, N.; Liu, N.; Cheng, W.; Jiang, Y.; Sun, H.; Chen, L.; Peng, J.; Zhang, Y.; Ding, Y.; Zhang, Z.; et al. Structural Basis for Receptor Recognition and Pore Formation of a Zebrafish Aerolysin-like Protein. EMBO Rep. 2016, 17, 235–248.
More
Information
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
633
Revisions:
2 times
(View History)
Update Date:
13 Jul 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





