
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Paolo Bergamo | -- | 3891 | 2023-06-14 09:24:09 | | | |
| 2 | Wendy Huang | -1964 word(s) | 1927 | 2023-06-14 13:56:00 | | | | |
| 3 | Wendy Huang | + 1964 word(s) | 3891 | 2023-06-16 08:26:14 | | | | |
| 4 | Wendy Huang | -1999 word(s) | 1892 | 2023-06-16 08:58:13 | | | | |
| 5 | Wendy Huang | Meta information modification | 1892 | 2023-06-16 09:01:01 | | |
Video Upload Options
Food nutrients play a key role in human metabolism and health via the modulation of multiple mechanisms, including energy metabolism, intestinal homeostasis, antioxidant homeostasis, and immune responses. The intestine is an essential organ involved in human nutrition, the metabolic activity of gut microbes is essential for maintaining host health, and alterations in its composition induce metabolic shifts that may have adverse effects. The consensus on microbiota-mediated healthy effects on the host is based on the microbe-induced biotransformation of food components into bioactive metabolites. Bioactive molecules exhibit, in combination with food components, the ability to modulate the metabolic pathways of the host or to modify the composition and metabolism of the microbiota. Studies indicated the efficacy of the carbohydrates accessible to the microbiota (MACs), polyphenols, and polyunsaturated fatty acids (PUFAs) in increasing the microbial population with the ability to yield biologically active metabolites (e.g., polyphenol metabolites, short-chain fatty acids (SCFAs)) capable of modulating redox homeostasis of the host.
1. Introduction
The beneficial effects associated with the diversity of the microbial population arise from the metabolic activities of specific microbial populations. Under eubiotic conditions, the commensal relationship between the microbiota and the host mainly consists of the capacity of bacteria to generate bioactive metabolites, starting from the ingested food, which exhibits the ability to modulate different metabolic pathways of the host [1]. For example, the production of carboxylic acids with aliphatic tails with fewer than six carbon atoms such as acetate (C2), propionate (C3), and butyrate (C4), resulting from the anaerobic fermentation of dietary plant polysaccharides, is the most relevant metabolic activity of enteric microbiota. These molecules are collectively referred to as Short-chain Fatty Acids (SCFAs) [2].
The growth of anaerobic SCFA-producing bacteria is favored by the low oxygen concentrations in the intestine where the two most abundant populations, namely, Bacteroidetes and Firmicutes, mainly produce acetate/propionate and butyrate, respectively [3]. Interestingly, due to butyrate generation during acetate metabolism, their coexistence can be consequential to mutual metabolic gain, thus resulting from the utilization of acetate produced by Bacteroidetes and Firmicutes to produce butyrate and propionate [4]. This example strongly supports the concept that the production of SCFAs is finely tuned by the balance of the bacterial species present in the gut.
| Disease | Model | Microbiota Alteration Production of SCFAs |
Ref. | |
|---|---|---|---|---|
| Diabetes | Randomized clinical trial High-fiber diet |
Type 2 diabetes ↓ SCFAs High fiber intake ↑ SCFAs ↑ SCFA-producing bacteria |
[5] | |
| Meta analysis Dietary fiber |
↑ Butyrate, propionate ↑ Bifidobacterium |
[6] | ||
| Inflammatory Bowel Disease (IBD) | 313 patients | ↓ Acetate-to-butyrate converter Firmicutes (Roseburia) ↓ Propionate ↑ Pathogens (Enterobacteriaceae, Proteobacteria) |
[7] | |
| 127 patients 87 healthy controls |
↓ Butyrate-producing bacteria (Firmicutes) ↓ SCFAs (acetate, propionate, butyrate) |
[8] | ||
| 10 inactive Crohn patients 10 healthy controls |
↓ SCFA-producing bacteria ↓ Roseburia inulinivorans, ↓ Ruminococcus torques, ↓ Clostridium lavalense, ↓ Bacteroides uniformis ↓ Faecalibacterium prausnitzii |
[9] | ||
| Nonalcoholic Fatty Liver Disease |
14 nonalcoholic fatty liver, 18 nonalcoholic steatohepatitis 27 healthy controls |
↑ SCFA levels ↑ SCFA-producing bacteria (Fusobacteriaceae, Prevotellaceae) |
[10] | |
| 25 nonalcoholic fatty liver 25 nonalcoholic steatohepatitis 25 healthy donors |
↓ Ruminococcaceae ↓ Clostridiales ↑ Bacteroidetes ↓ Firmicutes |
[11] | ||
| 30 patients F0/1 fibrosis stage 27 patients F ≥ 2 fibrosis |
↑ Bacteroidetes (F ≥ 2) ↑ Ruminococcus (F ≥ 2) ↓ Prevotella |
[12] | ||
| Neurodegeneration | Parkinson’s Disease | Nonparametric meta-analysis | ↑ Akkermansia ↓ Fecal SCFAs (acetate, propionate, butyrate) |
[13] |
| 96 patients 85 controls |
↓ Fecal SCFAs ↑ Plasma SCFAs ↑ Pro-inflammatory bacteria |
[14] | ||
| 95 patients 33 controls |
↓ Fecal SCFAs (propionic acetic, butyric) ↑ Plasma SCFA (propionic acetic) |
[15] | ||
| Alzheimer’s Disease | 25 patients | ↓ Firmicutes, Bifidobacterium ↑ Bacteroidetes |
[16] | |
| 33 dementia 22 mild cognitive impairment 120 subjective cognitive decline |
↓ SCFA-producing bacteria (Ruminococcus, Eubacterium) ↑ AD biomarkers (Amyloid-β1-42 and p-tau concentrations) |
[17] | ||
| Mouse model Sodium butyrate supplementation |
↓ Amyloid-β1-42 protein (40%) | [18] | ||
It can be assumed that butyrate, being a fundamental nutrient for colonocytes, satisfies the metabolic demands of the colon epithelium [19], and it also modulates the expression of tight junction proteins, thus preserving the intestinal barrier whose integrity is a crucial part of the overall immune response [20]. In addition, local O2 consumption during butyrate uptake and its metabolism by the intestinal epithelium stabilizes the hypoxia-inducible factor (HIF)—a transcription factor that coordinates barrier protection—which promotes the creation of an anaerobic environment. This “physiological hypoxia” stimulates the growth of SCFA-producers (anaerobic bacteria) [21], indirectly regulating the functionality of the intestinal barrier [22]. In addition, SCFAs have been shown to display an inhibitory effect on the growth of potentially pathogenic bacteria such as Salmonella typhimurium [23] or Clostridium difficile [24].
2. Effect of SCFAs on Gut Homeostasis
Acetate, propionate, and butyrate in the colon are present in the molar ratio 60:25:15, although proportions can vary depending on factors such as diet, microbiota composition, the site of fermentation, and the genotype of the host [25]. These are the predominant SCFAs present in the proximal regions of the large intestine in humans and rodents, and they are present at mM levels [26][27][28].
Acetate, propionate, and butyrate reach the highest concentrations (70–140 mM) in the proximal colon [25] where they enhance mucin secretion by increasing the expression of the MUC2 gene [29], with a concentration gradient decreasing from the lumen to the periphery [30]. When these SCFAs are absorbed into the hepatic portal circulation and the lacteal lymphatic system, they reach total concentrations ranging from 375 μM to 148 μM in the portal and hepatic blood respectively, or 79 μM in peripheral blood [25][31]. Butyrate and propionate, mostly metabolized by hepatocytes, were reported in a range of 1–15 μM in the systemic circulation, while acetate ranged between 100 and 200 μM [32][33]. However, the small amounts of SCFAs present in the bloodstream are sufficient to elicit a wide range of biological functions in different tissues.
A study on a mouse model of induced colitis demonstrated that SCFAs preserve gut homeostasis by acting on the inflammasome pathway through the upregulation of interleukin-18 [34]. Accordingly, low levels of butyrate and propionate-producing bacteria were found in patients suffering from inflammatory bowel diseases (IBD) such as ulcerative colitis or Crohn’s disease [8][9]. Several in vivo analyses have indicated that SCFAs regulate gut motility by stimulating mucosal receptors [35] or by increasing the release of the Peptide YY from gut endocrine cells, thus favoring intestine motility [36]. Other studies demonstrated that SCFAs act in preventing colonic diseases, by enhancing the absorption of minerals and decreasing the cholesterol concentration [37][38]. Experiments using germ-free animals reported that their reduced gut motility can be restored by the infusion of SCFAs [39].
Convincing evidence supports the idea that the beneficial effect of SCFAs extends beyond the colon. In fact, SCFAs participate in different physiological processes in the human body, being able to improve gut physiology, modulate the host’s glucose and lipid metabolism, and affect immune function [40][41]. In particular, upon their transport from the intestinal lumen into the blood compartment of the host, SCFAs are absorbed by the liver for gluconeogenesis or by muscle to generate energy [42]. Among SCFAs, acetate is the primary substrate for cholesterol synthesis [43], while propionate inhibits cholesterol synthesis by reducing serum lipids and has a protective effect against colon cancer [44][45]. Notably, SCFAs modulate brain functions by acting on the production of neuroactive metabolites [46]. For example, butyrate and propionate can be transferred from the gut to the brain where they act as signalling molecules through the monocarboxylate transporters that are highly expressed in the blood–brain barrier [47]. Finally, butyrate has been reported to play a protective role against carcinogenesis in colon cancer cells by enhancing the expression of cell cycle inhibitory genes [48].
3. Signaling Mechanisms Induced by SCFAs
Besides the relevant role in intestinal health, SCFAs may play their signalling role via the activation of several biochemical pathways: G-protein-coupled receptors (GPCRs), histone deacetylases (HDACs), and Nrf2 [49][50][51] (Figure 1).
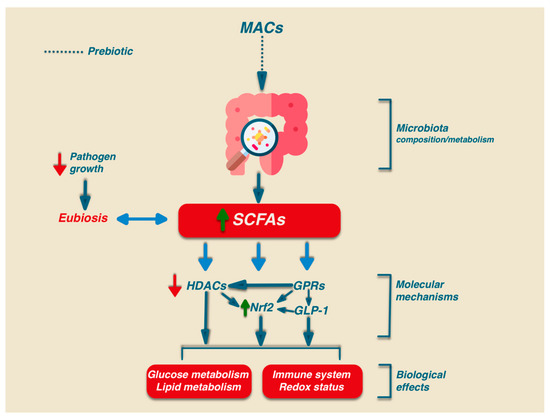
In humans, there are at least six GPRs that are sensitive to SCFAs, but among them, only GPR41, GPR43, and GPR109A are involved in SCFA-mediated signaling. GPR41 and GPR43 are the best-studied SCFA receptors [52] and are activated by acetate, propionate, and, to a lesser extent, also by butyrate. GPR41 is expressed in colon cells, in the blood vessels, and in the sympathetic nervous system, while GPR43 is mainly expressed in enteroendocrine L cells, lymphocytes, neutrophils, and monocytes [53]. GPR109A has a high affinity for niacin which can be activated by butyrate, and it is expressed only in human immune cells and colonocytes. In addition, GPR109A is highly expressed in adipocytes. The activation of this receptor in adipocytes has been linked to lipolysis and a decrease in plasma free fatty acids [54]. Activated GPCR receptors can regulate different signaling via the activation of many cellular functions such as the mitogen-activated protein kinase (MAPK) family of serine-threonine kinases, including extracellular signal-regulated kinase (ERK), c-jun N-terminal kinase (JNK), p38, and ERK5, through an intricate network of signaling. The activation of GPR43 also stimulates the phospholipase-C determining intracellular Ca2+ release and the activation of protein kinase C [55].
HDACs are a group of enzymes that affect gene transcription or alter protein activity by removing the acetyl group on the lysine ϵ-amino group of the target protein. The inhibition of HDACs is relevant for immune and inflammatory regulation by modulating either innate immunity through regulation of the Toll-Like Receptor (TLR) and Interferon (IFN) signaling pathways or by regulating antigen presentation and B and T lymphocytes to achieve adaptive immunity [56][57]. In particular, the inhibitory effect of HDACs on SCFAs, mainly due to propionate and butyrate, results in an anti-inflammatory effect through the promotion of regulatory T cell (Treg) development as well as CD4+ T cell IL-10 production [57][58].
The shared molecular pathways activated by polyphenols, PUFAs, and SCFAs support the role of Nrf2, HDACs, and GPRs in the beneficial effects elicited by dietary MACs, polyphenols, and PUFAs. Since the activation of these pathways triggers a downstream signaling cascade, this can explain why dietary bioactive molecules can exert antioxidant/beneficial effects even when present in a low plasma concentration.
Interestingly, the functional link existing between antioxidant activity and gut microbiota homeostasis has been indicated by (a) the modulatory ability of SCFAs in the Nrf2 pathway [49], (b) the age-dependent decline in the concentration of SCFAs in the gut [59], and (c) the positive association between microbiota diversity and Nrf2 efficacy [60]. In addition, the link between the production of SCFAs and the Nrf2 pathway was indicated in a recent study showing the ability of Clostridium butyricum pretreatment to increase the SCFA contents in the cecum of Enterotoxigenic Escherichia coli K88 (ETEC K88)-infected mice. In particular, the data indicated that such improvement was associated with the amelioration of the oxidative damage induced by ETEC K88 infection through the activation of the Nrf2 pathway [61]. A summary of the differential ability of microbial SCFAs in activating different receptors involved in the Nrf2 pathway is shown in Table 2.
Table 2. A brief summary of SCFAs produced by the gut microbial population and of their response to different receptors; adapted from [3][49]. Low or high affinity is denoted by + or ++, respectively.| Phylum | Family | Genus | FFAR3 (GPR41) |
FFAR2 (GPR43) |
GPR109A | |
|---|---|---|---|---|---|---|
| Firmicutes | Lachnospiraceae | Coprococcus | ACETATE | + + | + + | + + |
| Barnesiella | ||||||
| Ruminococcaceae | ||||||
| Akkermansia | ||||||
| Prevotella | ||||||
| Bifidobacterium | ||||||
| Bacteroidetes | Bacteroidaceae | Bacteroides | PROPIONATE | + | + + | + |
| Prevotellaceae | Prevotella | |||||
| Rikenellaceae | Alistipes | |||||
| Firmicutes | Eubacterium | |||||
| Blautia | ||||||
| Coprococcus | ||||||
| Veillonellaceae | Dialister | |||||
| Acidaminococcaceae | Phascolarctobacterium | |||||
| Verrucomicrobia | Verrucomicrobiaceae | Akkermansia | ||||
| Firmicutes | Lachnospiraceae | Eubacterium | BUTYRATE | + + | + + | + + |
| Roseburia | ||||||
| Clostridium | ||||||
| Eubacterium | ||||||
| Anaerostipes | ||||||
| Coprococcus | ||||||
| Ruminococcaceae | Faecalibacterium | |||||
| Subdoligranulum | ||||||
| Erysipelotrichaceae | Holdemanella |
Finally, the interplay existing between different SCFAs further strengthens the complexity of their mechanism of action [62][63] (Figure 1).
References
- Descamps, H.C.; Herrmann, B.; Wiredu, D.; Thaiss, C.A. The path toward using microbial metabolites as therapies. EBioMedicine 2019, 44, 747–754.
- Koh, A.; De Vadder, F.; Kovatcheva-Datchary, P.; Bäckhed, F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 2016, 165, 1332–1345.
- Louis, P.; Flint, H.J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 2017, 19, 29–41.
- Mahowald, M.; Rey, F.; Seedorf, H.; Turnbaugh, P.; Fulton, R.; Wollam, A.; Shah, N.; Wang, C.; Magrini, V.; Wilson, R.; et al. Characterizing a model human gut microbiota composed of members of its two dominant bacterial phyla. Proc. Natl. Acad. Sci. USA 2009, 106, 5859–5864.
- Zhao, L.; Zhang, F.; Ding, X.;Wu, G.; Lam, Y.Y.;Wang, X.; Fu, H.; Xue, X.; Lu, C.; Ma, J.; et al. Gut bacteria selectively promoted by dietary fibers alleviate type 2 diabetes. Science 2018, 359, 1151–1156.
- Ojo, O.; Feng, Q.Q.; Ojo, O.O.;Wang, X.H. The Role of Dietary Fibre in Modulating Gut Microbiota Dysbiosis in Patients with Type 2 Diabetes: A Systematic Review and Meta-Analysis of Randomised Controlled Trials. Nutrients 2020, 12, 3239. [CrossRef] Antioxidants 2023, 12, 1073 17 of 22
- Imhann, F.; Vich Vila, A.; Bonder, M.J.; Fu, J.; Gevers, D.; Visschedijk, M.C.; Spekhorst, L.M.; Alberts, R.; Franke, L.; van Dullemen, H.M.; et al. Interplay of host genetics and gut microbiota underlying the onset and clinical presentation of inflammatory bowel disease. Gut 2018, 67, 108–119.
- Machiels, K.; Joossens, M.; Sabino, J.; De Preter, V.; Arijs, I.; Eeckhaut, V.; Ballet, V.; Claes, K.; Van Immerseel, F.; Verbeke, K.; et al. A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 2014, 63, 1275–1283.
- Takahashi, K.; Nishida, A.; Fujimoto, T.; Fujii, M.; Shioya, M.; Innaeda, H.; Lnatorni, O.; Bamba, S.; Andoh, A.; Sugimoto, M. Reduced Abundance of Butyrate-Producing Bacteria Species in the Fecal Microbial Community in Crohn’s Disease. Digestion 2016, 93, 59–65.
- Rau, M.; Rehman, A.; Dittrich, M.; Groen, A.K.; Hermanns, H.M.; Seyfried, F.; Beyersdorf, N.; Dandekar, T.; Rosenstiel, P.; Geier, A. Fecal SCFAs and SCFA-producing bacteria in gut microbiome of human NAFLD as a putative link to systemic T-cell activation and advanced disease. United Eur. Gastroenterol. J. 2018, 6, 1496–1507.
- Tsai, M.C.; Liu, Y.Y.; Lin, C.C.; Wang, C.C.; Wu, Y.J.; Yong, C.C.; Chen, K.D.; Chuah, S.K.; Yao, C.C.; Huang, P.Y.; et al. Gut Microbiota Dysbiosis in Patients with Biopsy-Proven Nonalcoholic Fatty Liver Disease: A Cross-Sectional Study in Taiwan. Nutrients 2020, 12, 820.
- Boursier, J.; Mueller, O.; Barret, M.; Machado, M.; Fizanne, L.; Araujo-Perez, F.; Guy, C.D.; Seed, P.C.; Rawls, J.F.; David, L.A.; et al. The severity of nonalcoholic fatty liver disease is associated with gut dysbiosis and shift in the metabolic function of the gut microbiota. Hepatology 2016, 63, 764–775.
- Hirayama, M.; Ohno, K. Parkinson’s Disease and Gut Microbiota. Ann. Nutr. Metab. 2021, 77 (Suppl. 2), 28–35.
- Chen, S.J.; Chen, C.C.; Liao, H.Y.; Lin, Y.T.; Wu, Y.W.; Liou, J.M.; Wu, M.S.; Kuo, C.H.; Lin, C.H. Association of Fecal and Plasma Levels of Short-Chain Fatty Acids with Gut Microbiota and Clinical Severity in Patients with Parkinson Disease. Neurology 2022,98, e848–e858.
- Yang, X.; Ai, P.; He, X.; Mo, C.; Zhang, Y.; Xu, S.; Lai, Y.; Qian, Y.; Xiao, Q. Parkinson’s Disease Is Associated with ImpairedGut-Blood Barrier for Short-Chain Fatty Acids. Mov. Disord. 2022, 37, 1634–1643.
- Vogt, N.M.; Kerby, R.L.; Dill-McFarland, K.A.; Harding, S.J.; Merluzzi, A.P.; Johnson, S.C.; Carlsson, C.M.; Asthana, S.; Zetterberg, H.; Blennow, K.; et al. Gut microbiome alterations in Alzheimer’s disease. Sci. Rep. 2017, 7, 13537.
- Verhaar, B.J.H.; Hendriksen, H.M.A.; de Leeuw, F.A.; Doorduijn, A.S.; van Leeuwenstijn, M.; Teunissen, C.E.; Barkhof, F.; Scheltens, P.; Kraaij, R.; van Duijn, C.M.; et al. Gut Microbiota Composition Is Related to AD Pathology. Front. Immunol. 2021, 12, 794519.
- Fernando, W.M.A.D.; Martins, I.J.; Morici, M.; Bharadwaj, P.; Rainey-Smith, S.R.; Lim, W.L.F.; Martins, R.N. Sodium ButyrateReduces Brain Amyloid-Levels and Improves Cognitive Memory Performance in an Alzheimer’s Disease Transgenic Mouse Model at an Early Disease Stage. J. Alzheimer’s Dis. 2020, 74, 91–99.
- Barcenilla, A.; Pryde, S.; Martin, J.; Duncan, S.; Stewart, C.; Henderson, C.; Flint, H. Phylogenetic relationships of butyrate producing bacteria from the human gut. Appl. Environ. Microbiol. 2000, 66, 1654–1661.
- Ohata, A.; Usami, M.; Miyoshi, M. Short-chain fatty acids alter tight junction permeability in intestinal monolayer cells via lipoxygenase activation. Nutrition 2005, 21, 838–847.
- Rivera-Chávez, F.; Zhang, L.F.; Faber, F.; Lopez, C.A.; Byndloss, M.X.; Olsan, E.E.; Xu, G.; Velazquez, E.M.; Lebrilla, C.B.; Winter, S.E.; et al. Depletion of Butyrate-Producing Clostridia from the Gut Microbiota Drives an Aerobic Luminal Expansion of Salmonella. Cell Host Microbe 2016, 19, 443–454.
- Colgan, S.P.; Taylor, C.T. Hypoxia: An alarm signal during intestinal inflammation. Nat. Rev. Gastroenterol. Hepatol. 2010, 7, 281–287.
- Bohnhoff, M.; Miller, C.P.; Martin, W.R. Resistance of the mouse’s intestinal tract to experimental salmonella infection. II. factors responsible for its loss following streptomycin treatment. J. Exp. Med. 1964, 120, 817–828.
- Rolfe, R.D. Role of volatile fatty acids in colonization resistance to Clostridium difficile. Infect. Immun. 1984, 45, 185–191
- Cummings, J.H.; Pomare, E.W.; Branch,W.J.; Naylor, C.P.; Macfarlane, G.T. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 1987, 28, 1221–1227.
- Lin, H.V.; Frassetto, A.; Kowalik, E.J.; Nawrocki, A.R.; Lu, M.M.; Kosinski, J.R.; Hubert, J.A.; Szeto, D.; Yao, X.; Forrest, G.; et al. Butyrate and propionate protect against diet-induced obesity and regulate gut hormones via free fatty acid receptor 3-independent mechanisms. PLoS ONE 2012, 7, e35240.
- Bergman, E.N. Energy contributions of volatile fatty acids from the gastrointestinal tract in various species. Physiol. Rev. 1990, 70, 567–590.
- Macfarlane, G.T.; Macfarlane, S. Fermentation in the human large intestine: Its physiologic consequences and the potential contribution of prebiotics. J. Clin. Gastroenterol. 2011, 45, S120–S127.
- Gaudier, E.; Rival, M.; Buisine, M.P.; Robineau, I.; Hoebler, C. Butyrate enemas upregulate Muc genes expression but decrease adherent mucus thickness in mice colon. Physiol. Res. 2009, 58, 111–119.
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200
- Hamer, H.M.; Jonkers, D.; Venema, K.; Vanhoutvin, S.; Troost, F.J.; Brummer, R.J. Review article: The role of butyrate on colonic function. Aliment. Pharmacol. Ther. 2008, 27, 104–119.
- Meesters, R.; van Eijk, H.; ten Have, G.; de Graaf, A.; Venema, K.; van Rossum, B.; Deutz, N. Application of liquid chromatography mass spectrometry to measure the concentrations and study the synthesis of short chain fatty acids following stable isotope infusions. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2007, 854, 57–62.
- Moreau, N.; Goupry, S.; Antignac, J.; Monteau, F.; Le Bizec, B.; Champ, M.; Martin, L.; Dumon, H. Simultaneous measurement of plasma concentrations and C-13-enrichment of short-chain fatty acids, lactic acid and ketone bodies by gas chromatography coupled to mass spectrometry. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2003, 784, 395–403.
- Macia, L.; Tan, J.; Vieira, A.T.; Leach, K.; Stanley, D.; Luong, S.; Maruya, M.; Ian McKenzie, C.; Hijikata, A.; Wong, C.; et al. Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat. Commun. 2015, 6, 6734.
- Yajima, T. Contractile effect of short-chain fatty acids on the isolated colon of the rat. J. Physiol. 1985, 368, 667–678.
- Plaisancié, P.; Dumoulin, V.; Chayvialle, J.A.; Cuber, J.C. Luminal peptide YY-releasing factors in the isolated vascularly perfused rat colon. J. Endocrinol. 1996, 151, 421–429. [CrossRef]
- Fåk, F.; Jakobsdottir, G.; Kulcinskaja, E.; Marungruang, N.; Matziouridou, C.; Nilsson, U.; Stålbrand, H.; Nyman, M. The physico-chemical properties of dietary fibre determine metabolic responses, short-chain Fatty Acid profiles and gut microbiota composition in rats fed low- and high-fat diets. PLoS ONE 2015, 10, e0127252.
- Kilner, J.; Corfe, B.; McAuley, M.; Wilkinson, S. A deterministic oscillatory model of microtubule growth and shrinkage for differential actions of short chain fatty acids. Mol. Biosyst. 2016, 12, 93–101.
- Vincent, A.D.; Wang, X.Y.; Parsons, S.P.; Khan, W.I.; Huizinga, J.D. Abnormal absorptive colonic motor activity in germ-free mice is rectified by butyrate, an effect possibly mediated by mucosal serotonin. Am. J. Physiol. Gastrointest. Liver Physiol. 2018, 315, G896–G907.
- Gao, Z.; Yin, J.; Zhang, J.; Ward, R.E.; Martin, R.J.; Lefevre, M.; Cefalu, W.T.; Ye, J. Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes 2009, 58, 1509–1517.
- Fushimi, T.; Suruga, K.; Oshima, Y.; Fukiharu, M.; Tsukamoto, Y.; Goda, T. Dietary acetic acid reduces serum cholesterol and triacylglycerols in rats fed a cholesterol-rich diet. Br. J. Nutr. 2006, 95, 916–924.
- De Vadder, F.; Kovatcheva-Datchary, P.; Goncalves, D.; Vinera, J.; Zitoun, C.; Duchampt, A.; Backhed, F.; Mithieux, G. Microbiota-Generated Metabolites Promote Metabolic Benefits via Gut-Brain Neural Circuits. Cell 2014, 156, 84–96.
- Ukai, M.; Tomura, A.; Ito, M. Cholesterol-synthesis in germfree and conventional rats. J. Nutr. 1976, 106, 1175–1183.
- Wang, G.; Yu, Y.;Wang, Y.;Wang, J.; Guan, R.; Sun, Y.; Shi, F.; Gao, J.; Fu, X. Role of SCFAs in gut microbiome and glycolysis for colorectal cancer therapy. J. Cell. Physiol. 2019, 234, 17023–17049.
- Reigstad, C.S.; Salmonson, C.E.; Rainey, J.F.; Szurszewski, J.H.; Linden, D.R.; Sonnenburg, J.L.; Farrugia, G.; Kashyap, P.C. Gut microbes promote colonic serotonin production through an effect of short-chain fatty acids on enterochromaffin cells. FASEB J.2015, 29, 1395–1403.
- Vijay, N.; Morris, M.E. Role of monocarboxylate transporters in drug delivery to the brain. Curr. Pharm. Des. 2014, 20, 1487–1498.
- Daly, K.; Shirazi-Beechey, S. Microarray analysis of butyrate regulated genes in colonic epithelial cells. DNA Cell Biol. 2006, 25,49–62.
- Campos-Perez, W.; Martinez-Lopez, E. Effects of short chain fatty acids on metabolic and inflammatory processes in human health. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2021, 1866, 158900.
- González-Bosch, C.; Boorman, E.; Zunszain, P.A.; Mann, G.E. Short-chain fatty acids as modulators of redox signaling in health and disease. Redox Biol. 2021, 47, 102165.
- Visekruna, A.; Luu, M. The Role of Short-Chain Fatty Acids and Bile Acids in Intestinal and Liver Function, Inflammation, and Carcinogenesis. Front. Cell Dev. Biol. 2021, 9, 703218.
- Brown, A.J.; Goldsworthy, S.M.; Barnes, A.A.; Eilert, M.M.; Tcheang, L.; Daniels, D.; Muir, A.I.; Wigglesworth, M.J.; Kinghorn, I.; Fraser, N.J.; et al. The Orphan G protein-coupled receptors GPR41 and GPR43 are activated by propionate and other short chain carboxylic acids. J. Biol. Chem. 2003, 278, 11312–11319.
- Karaki, S.; Mitsui, R.; Hayashi, H.; Kato, I.; Sugiya, H.; Iwanaga, T.; Furness, J.B.; Kuwahara, A. Short-chain fatty acid receptor, GPR43, is expressed by enteroendocrine cells and mucosal mast cells in rat intestine. Cell Tissue Res. 2006, 324, 353–360.
- Ahmed, K.; Tunaru, S.; Offermanns, S. GPR109A, GPR109B and GPR81, a family of hydroxy-carboxylic acid receptors. Trends Pharmacol. Sci. 2009, 30, 557–562.
- Dorsam, R.T.; Gutkind, J.S. G-protein-coupled receptors and cancer. Nat. Rev. Cancer 2007, 7, 79–94.
- Schotterl, S.; Brennenstuhl, H.; Naumann, U. Modulation of immune responses by histone deacetylase inhibitors. Crit. Rev. Oncog. 2015, 20, 139–154.
- Weiss, U.; Möller, M.; Husseini, S.A.; Manderscheid, C.; Häusler, J.; Geisslinger, G.; Niederberger, E. Inhibition of HDAC Enzymes Contributes to Differential Expression of Pro-Inflammatory Proteins in the TLR-4 Signaling Cascade. Int. J. Mol. Sci. 2020, 21, 8943.
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly-Y, M.; Glickman, J.N.; Garrett,W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341, 569–573.
- Sun, M.;Wu,W.; Chen, L.; Yang,W.; Huang, X.; Ma, C.; Chen, F.; Xiao, Y.; Zhao, Y.; Yao, S.; et al. Microbiota-derived short-chain fatty acids promote Th1 cell IL-10 production to maintain intestinal homeostasis. Nat. Commun. 2018, 9, 3555.
- Salazar, N.; Arboleya, S.; Fernández-Navarro, T.; de Los Reyes-Gavilán, C.G.; Gonzalez, S.; Gueimonde, M. Age-Associated Changes in Gut Microbiota and Dietary Components Related with the Immune System in Adulthood and Old Age: A Cross-Sectional Study. Nutrients 2019, 11, 1765.
- Matsumaru, D.; Motohashi, H. The KEAP1-NRF2 System in Healthy Aging and Longevity. Antioxidants 2021, 10, 1929.
- Li, H.; Shang, Z.; Liu, X.; Qiao, Y.;Wang, K.; Qiao, J. Alleviates Enterotoxigenic. Front. Immunol. 2021, 12, 771826.
- Dong,W.; Jia, Y.; Liu, X.; Zhang, H.; Li, T.; Huang,W.; Chen, X.;Wang, F.; Sun,W.;Wu, H. Sodium butyrate activates NRF2 to ameliorate diabetic nephropathy possibly via inhibition of HDAC. J. Endocrinol. 2017, 232, 71–83.
- Wu, J.; Jiang, Z.; Zhang, H.; Liang, W.; Huang, W.; Li, Y.; Wang, Z.; Wang, J.; Jia, Y.; Liu, B.; et al. Sodium butyrate attenuates diabetes-induced aortic endothelial dysfunction via P300-mediated transcriptional activation of Nrf2. Free Radic. Biol. Med. 2018,124, 454–465.





