Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Takao Ishikawa | -- | 2394 | 2023-06-06 19:38:45 | | | |
| 2 | Rita Xu | Meta information modification | 2394 | 2023-06-07 07:14:33 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Buko, T.; Tuczko, N.; Ishikawa, T. DNA Data Storage. Encyclopedia. Available online: https://encyclopedia.pub/entry/45260 (accessed on 09 May 2026).
Buko T, Tuczko N, Ishikawa T. DNA Data Storage. Encyclopedia. Available at: https://encyclopedia.pub/entry/45260. Accessed May 09, 2026.
Buko, Tomasz, Nella Tuczko, Takao Ishikawa. "DNA Data Storage" Encyclopedia, https://encyclopedia.pub/entry/45260 (accessed May 09, 2026).
Buko, T., Tuczko, N., & Ishikawa, T. (2023, June 06). DNA Data Storage. In Encyclopedia. https://encyclopedia.pub/entry/45260
Buko, Tomasz, et al. "DNA Data Storage." Encyclopedia. Web. 06 June, 2023.
Copy Citation
The demand for data storage is growing at an unprecedented rate, and current methods are not sufficient to accommodate such rapid growth due to their cost, space requirements, and energy consumption. Therefore, there is a need for a new, long-lasting data storage medium with high capacity, high data density, and high durability against extreme conditions. DNA is one of the most promising next-generation data carriers, with a storage density of 10¹⁹ bits of data per cubic centimeter, and its three-dimensional structure makes it about eight orders of magnitude denser than other storage media.
bit
byte
long term data storage
next-generation information storage
1. Introduction
The demand for data storage is increasing by approximately 50% every year. In 2012, the entire world’s total information storage was 2.7 ZB [1], in 2018 it reached 33 ZB, only to rise two-fold in 2020. It is estimated that newly created data will take up about 175 ZB by 2025 [2]. This equals a 65-fold increase only in the period between 2012 and 2025.
The tremendous Global Datasphere expansion is a strong motivator for new developments in data storage. Current data storage methods, such as magnetic (e.g., hard disk), optical (e.g., Blu-ray disc), and solid-state (e.g., flash drive), are insufficient to accommodate such rapid growth [3]. The main problems with those methods are their cost, space, and energy consumption during the recording, storing, and reading of data. Moreover, their durability reaches a maximum of 50 years in perfectly optimal conditions [4]. Humidity, extreme temperatures (both high or low), magnetic fields, or mechanical failures are the main reasons why those methods are not reliable for long-term data storage.
Therefore, there is a great demand for a new, longevous data storage medium with a high capacity, high data density, and high durability against extreme conditions [1]. There are a few prototypes of next-generation data carriers that may be able to cope with the above-mentioned challenges. Among them, DNA seems to be one of the most promising. The most distinguishing features of DNA from other storage media are its density and durability against the extreme conditions.
Escherichia coli has a storage density of 1019 bits of data per cubic centimeter [5]. This means that 1.7 × 1019 bits can be stored in just 1 g of DNA. Due to its three-dimensional structure, DNA is about eight orders of magnitude denser than other storage media. Moreover, DNA replication during PCR or the cell’s proliferation enables the quick and inexpensive copying of vast amounts of data [3].
For years, a DNA specimen collected from a 700,000-year-old horse was considered to be the oldest extracted DNA. However, in 2021, this record was pushed to 1 million years. DNA extracted from mammoth teeth was successfully extracted and sequenced [6]. Additionally, scientists managed to sequence 300,000-year-old mitochondrial DNA from humans and bears [4]. These examples perfectly illustrate the longevity of DNA and proves its usefulness for archeological purposes or data storekeeping. If stored in optimal conditions and dehydrated, DNA can possibly endure for millions of years [1].
Numerous space experiments on microorganisms have proven their extraordinary durability in extreme conditions. Due to solar UV radiance, the space vacuum, and extreme temperature conditions, space is considered one of the most hostile environments [7].
UV radiance being the most deleterious parameter in space increases microorganisms’ lethality by four orders of magnitude in relation to Earth’s conditions [8]. UVB and UVC altogether cover the 200–315 nm light spectrum; these are the most hazardous to microorganisms and are responsible for their high lethality in space. This is caused by high irradiance absorption by DNA and proteins in such spectral ranges. In vegetative cells, this UV irradiance leads to DNA mutations, such as cyclobutane pyrimidine dimers and pyrimidine–pyrimidone photo products [9]. Meanwhile, in bacterial spores, thymine dimer photoproducts, so-called spore photoproducts (SP), are formed due to UV radiation. Despite this fact, all these dimers can be repaired by the direct reversal mechanism. Spores possess an additional SP-specific repair pathway that makes spores significantly more resistant to UV radiance than vegetative cells [10].
Regardless of such hostile conditions, it has been proven that spores of Bacillus subtilis shielded against UV solar radiation are able to survive in outer space for nearly 6 years. Although only 1–2% of the population recovered, the outcome was significantly increased (even to 90% of population recovered) if 5% glucose was added to the spore multilayer. It was suggested that glucose binds additional water molecules, preventing the cell from becoming completely desiccated. It also replaces water molecules, thereby stabilizing the macromolecular structure [8]. Furthermore, some microorganisms can even cope with a full space environment. For example, the lichens Rhizocarpon geographicum and Xanthoria elegans survived a 2-week exposure to outer space. After that time, the lichens completely restored their photosynthetic activity and no ultrastructural changes were revealed in most of the fungal and algal cells of lichens [11]. It is supposed that their thick cortex with UV-screening pigments (rhizocarpic and parietin phenolic acids) are responsible for their survival [12].
2. Coding Files in DNA
Encoding information in DNA is based on binary code. A specific nucleotide corresponds to a code, for example, 00 → A, 01 → C, 10 → G, and 11 → T. While binary data are “translated” into a DNA sequence, it is important to avoid long homopolymers (more than three same nucleotides in a row) and unreasonable GC content, as both might generate mistakes during the synthesis and sequencing of DNA strings. In fact, encoding a file requires converting text into a code such as ASCII (Figure 1) or Base64, and then converting the coded file into a binary system. The encoding field uses different coding algorithms, such as Huffman, to condense messages and balance code, preventing homopolymer sequences. Two examples of coding systems, their modifications, and other algorithms of a similar kind generate proper DNA strings [13][14], which are capable of long data storage.
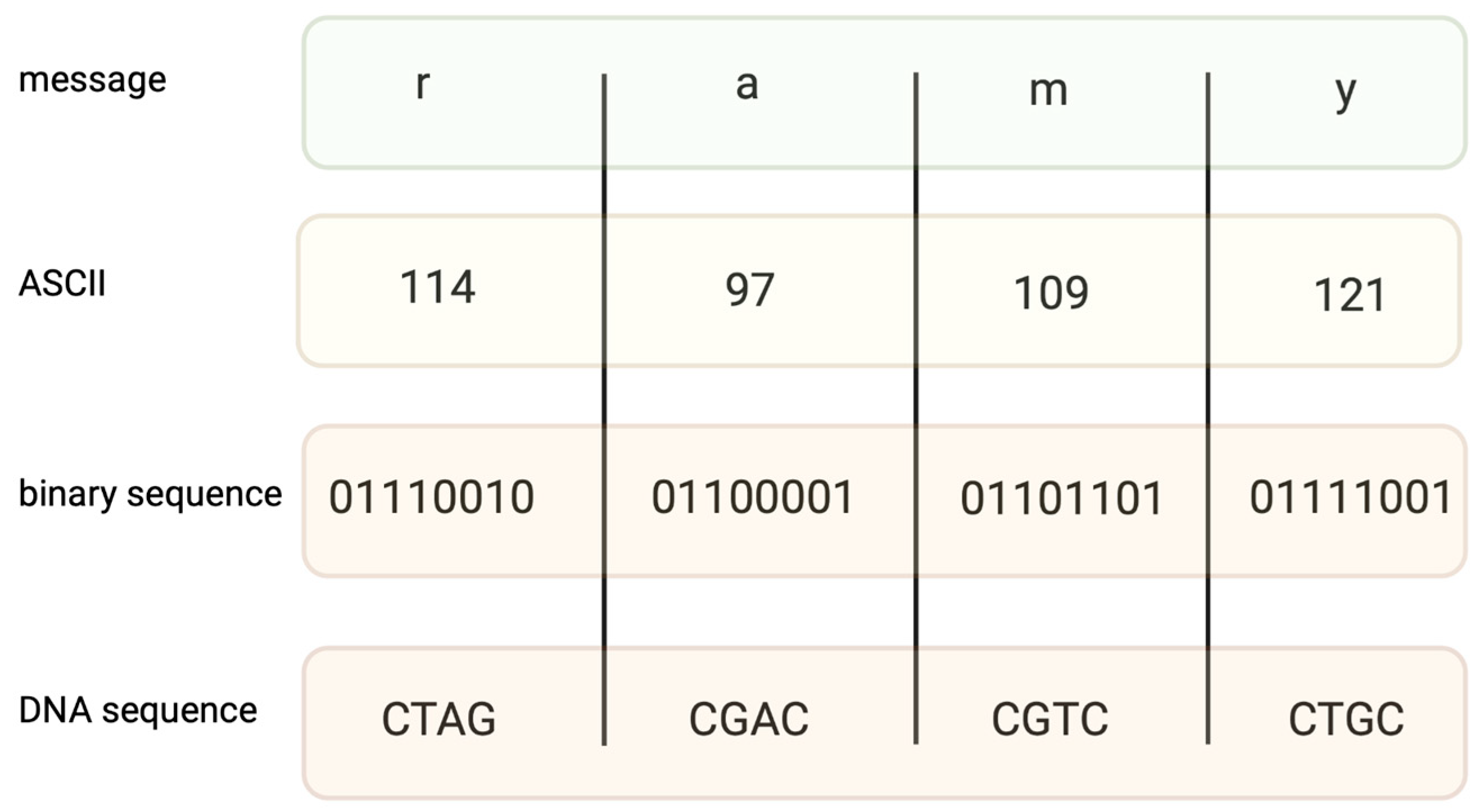
Figure 1. An example of coding the message “ramy” into an ASCII code. Converting binary data into nucleotide sequences is made by computer algorithms.
Church et al. (2012), for the first time, encoded a draft of a book, eleven JPG images and one JavaScript program in DNA [15]. For this purpose, they used a simple encoding method involving the translation of zeros into A or C and ones into T or G. As a result, the authors received 54,898 oligonucleotides, each containing three parts: 96 bases of data, 22-bases-long sequences at both ends, allowing those oligonucleotides to be parallelly amplified by PCR, and the 19-bases-long index sequence, pointing out the segment position in the original file [15]. Encoding one bit per base allowed the authors to avoid sequences that were potentially hard to write or read. Splitting information into blocks of data allowed the authors to circumvent the problems associated with the synthesis of long DNA strings. This pioneering work demonstrated the real possibility of using DNA as a data storage material, and also showed the enormous capacity of this method. An important element of the works of that time was to show the limitations of the method used. Through this work, it was noted that the information encoded in DNA is prone to sequencing errors, mainly in homopolymer regions.
One year later, Goldman et al. (2013) tried to overcome the sequencing errors occurring by encoding data with redundancy [16]. The authors encoded all 154 of Shakespeare’s sonnets, a scientific article, a medium-resolution color photograph of the European Bioinformatics Institute, and a 26 s long excerpt from Martin Luther King’s 1963 “I have a dream” speech using the Huffman algorithm to covert numeric data into a nucleotide sequence [16]. In summary, bytes of binary sequences were converted into base-3 digits (or ternary) from 0 to 2, which were then associated with three nucleotides, A, T, and C (or G if C has been used for the encoding of the previous ternary digit). DNA strings were divided into 100-nucleotide-long oligos with an overlap of 75 residues between adjacent fragments, creating four-fold redundancy (Figure 2). Alternate fragments were converted to their reverse complement, which reduces the probability of systematic failure, such as issues with DNA sequencing. Indexing sequences comprising 17 nucleotides were also encoded at the beginning and end of each fragment.
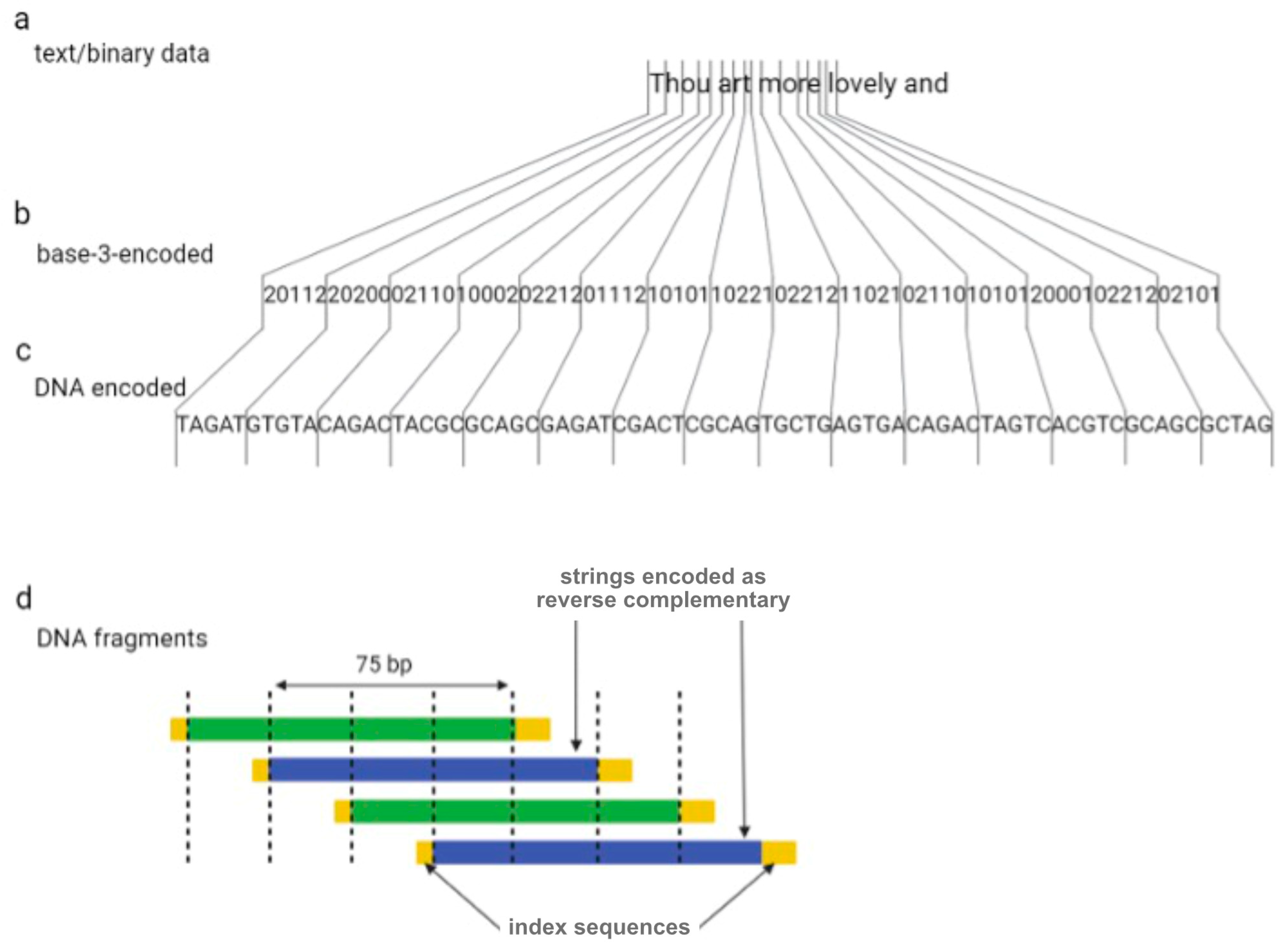
Figure 2. The coding scheme implemented by Goldman et al. Digital information (a) is converted to base-3 (b) using a Huffman code and is subsequently is converted to DNA strings (c). Dividing DNA strings as shown generated four-fold redundancy (d).
Ailenberg and Rotstein (2009) encoded text, music, and images in DNA by using modified Huffman coding (Figure 3) [17]. In their work, they constructed a plasmids library each containing 10,000 bp of information and an index plasmid that contains basic information, such as the title, author, plasmid number, and primer assignments used to read coded information [17]. The authors also constructed a separate encoding table for each type of file, which allowed the authors to encode each character from the keyboard. The authors also indicated the possibility of extending their code according to the described rules.
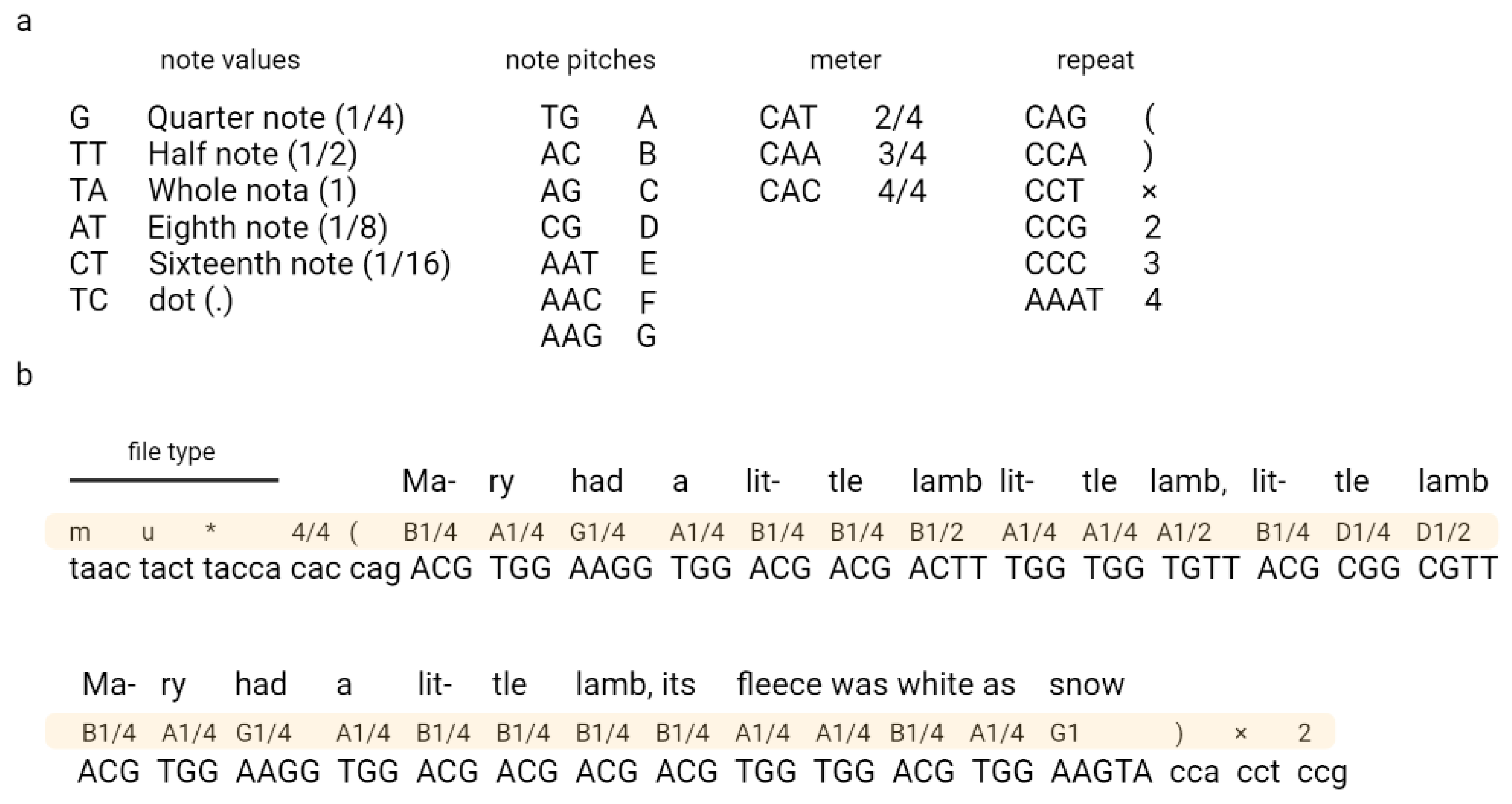
Figure 3. An example of coding music in DNA. Fragment of “Mary Had a Little Lamb” encoded using Huffman code. A nucleotide sequence corresponding to the music code is shown in (a) and the encryption part in (b). Adapted from Ailenberg and Rotstein.
The first example of the graphical file recoded in DNA was a simplified lamb drawing (Figure 4). Although this image consists of simple geometric figures, the simplicity and geometry of the image are not general requirements. Yazdi et al. (2017) managed to encode The Citizen Kane poster photograph and Smiley Face emoji (Figure 5) [18]. For this purpose, they used Base64 encoding to convert files into binary format. The DNA string length used by the authors was 1000 bp, containing 984 bp of information and 16 bp of address sequence. The purpose of the addressing method was to enable random access to codewords via highly selective PCR reactions. This approach allows the specific amplification of a pool of oligos without amplifying and reading all sequences from a given pool. This work also presented a new deletion-correcting method called homopolymer check codes. This method of correction divides DNA sequences into strings of homopolymers, e.g., {AATCCCCGA} into strings {AA, T, CCC, G, A}, which gives a homopolymer sequence of length {2,1,3,1,1}. The homopolymer length sequence contains special redundancy that protects against asymmetric substitution errors. Hypothetically, when two deletions occur in the sequence resulting in {ATCCGA}, the length of the homopolymer fragments is {1,1,2,1,1}. Recovering the original sequence is possible by correcting two bounded magnitude errors. Combining this with GC content balancing, the subsequent alignment of DNA oligonucleotides, and post-sequencing sequence sorting based on the correctness of the index sequence resulted in a new coding method.
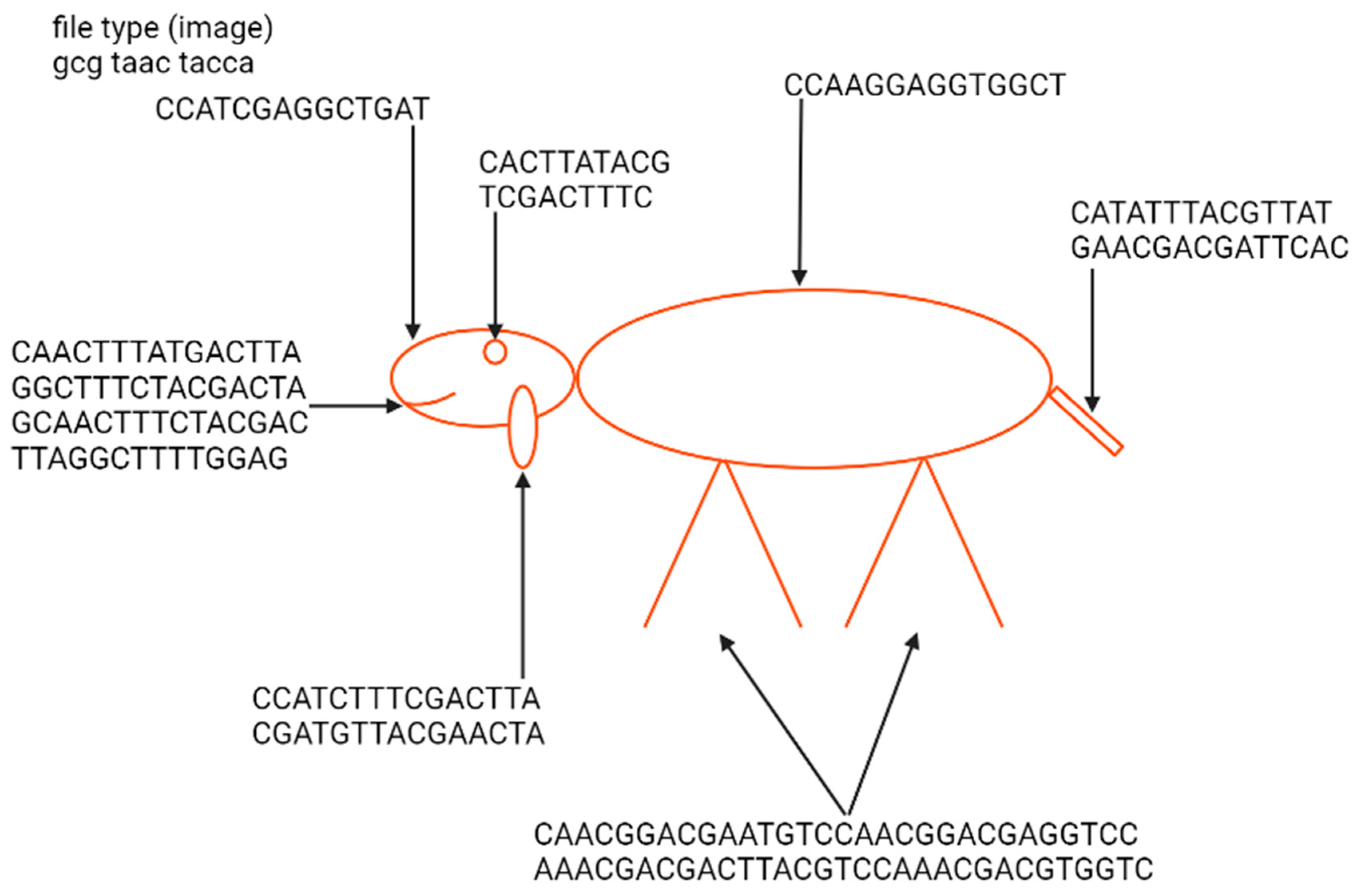
Figure 4. Indication of elements of the nucleotide sequence in which a Little Lamb was encoded and an example image presenting a lamb from the “Mary Had a Little Lamb” rhyme encoded by Ailenberg and Rotstein. The sequence of a file type defines it as an image. The geometric shape of the lamb enables the use of only 238 bp of DNA for encoding. Encoding has been performed using a template of signs indicating the type of shape and its spatial coordinates.
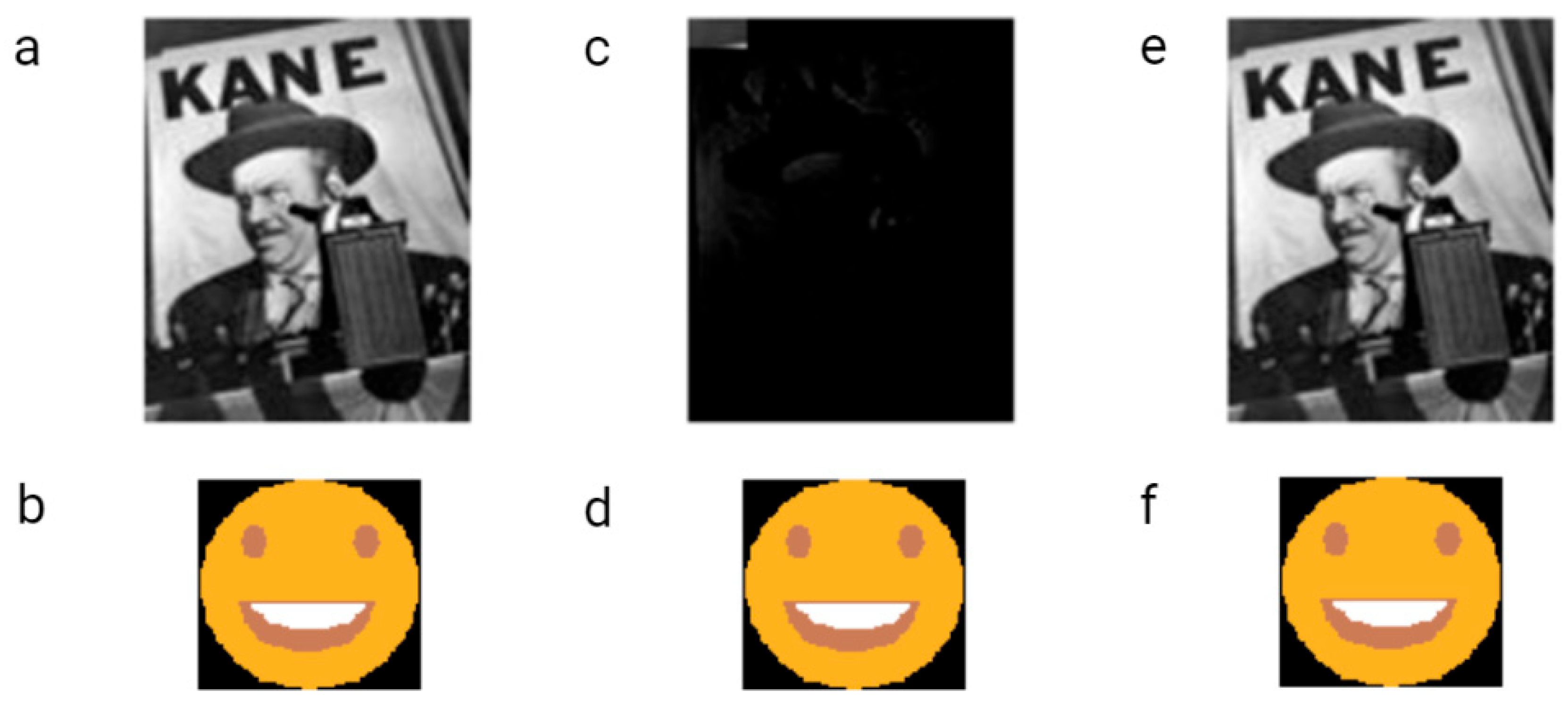

Figure 5. Smiling emoji and original Citizen Kane poster photograph encoded and decoded by Yazdi et al. The raw images were encoded and synthesized in the form of DNA strings (a,b). Images received after decoding without homopolymer check codes during processing (c,d). Images received after sequencing DNA strings when homopolymer error correction was made in order to reduce the number of errors that occurred during each encoding and decoding step (e,f). Two errors in the Citizen Kane file were sufficient to make the recovery of the image impossible. One error in the emoji did not influence the image quality.
Coding motion picture as motion GIFs and movies has also been achieved in the DNA data storage field. In 2017, Shipman et al. encoded five frames of a galloping mare from Eadweard Muybridge’s “The Human and Animal Locomotion Photographs” [19]. In their experiment, CRISPR-Cas was used to integrate an encoded short movie into the genomes of a population of living bacteria. The usage of this method does not change the overall encoding protocol. Strings of DNA are integrated into the CRISPR array thanks to appropriate integrases. Spacer sequences in the CRISPR array were used to encode barcodes defining which set of pixels was encoded in a specific part. The use of the CRISPR method for GIF encoding was of great importance because it allows the encoding of subsequent sequences without the need to additionally index them. This is because newly added sequences are almost always integrated in such a way that they push the previously integrated sequences away from the leader region. Therefore, the order of the sequence was conditioned by successive transformations in which DNA with encoded movie frames was introduced to bacterial cells.
3. Synthesis of DNA Strings
Chemical DNA synthesis has made tremendous progress since the 1970s, when fragments of about 20 nucleotides could be synthesized, to the present, when fragments of up to 500 nucleotides can be easily made. The technology commonly used for the synthesis of DNA strands enables only short 200–300 nucleotides sequences to be synthesized, which is a limitation when coding a large amount of data. Nevertheless, the technology used for DNA synthesis on microarrays seems to be more suitable for this purpose. It allows the synthesis of parallel oligonucleotides containing different sequences (Figure 6). By using it, the time and cost needed for the synthesis of large-scale DNA libraries might be greatly reduced [20]. Microarrays have enabled the high-fidelity synthesis of oligo pools of about 300 nucleotides in length [21]. Regardless of the synthesis method, long DNA fragments must be assembled from oligos. It is also necessary to add indexes to each fragment, or sequence overlapping in successive DNA fragments [3], unless—as discussed above—the CRISPR method is used to record information in the bacterial genome. In 2017, Heckel et al. considered the storage capacity using both assembly methods and have shown that an index-based coding system is optimal for data storage purposes [22].
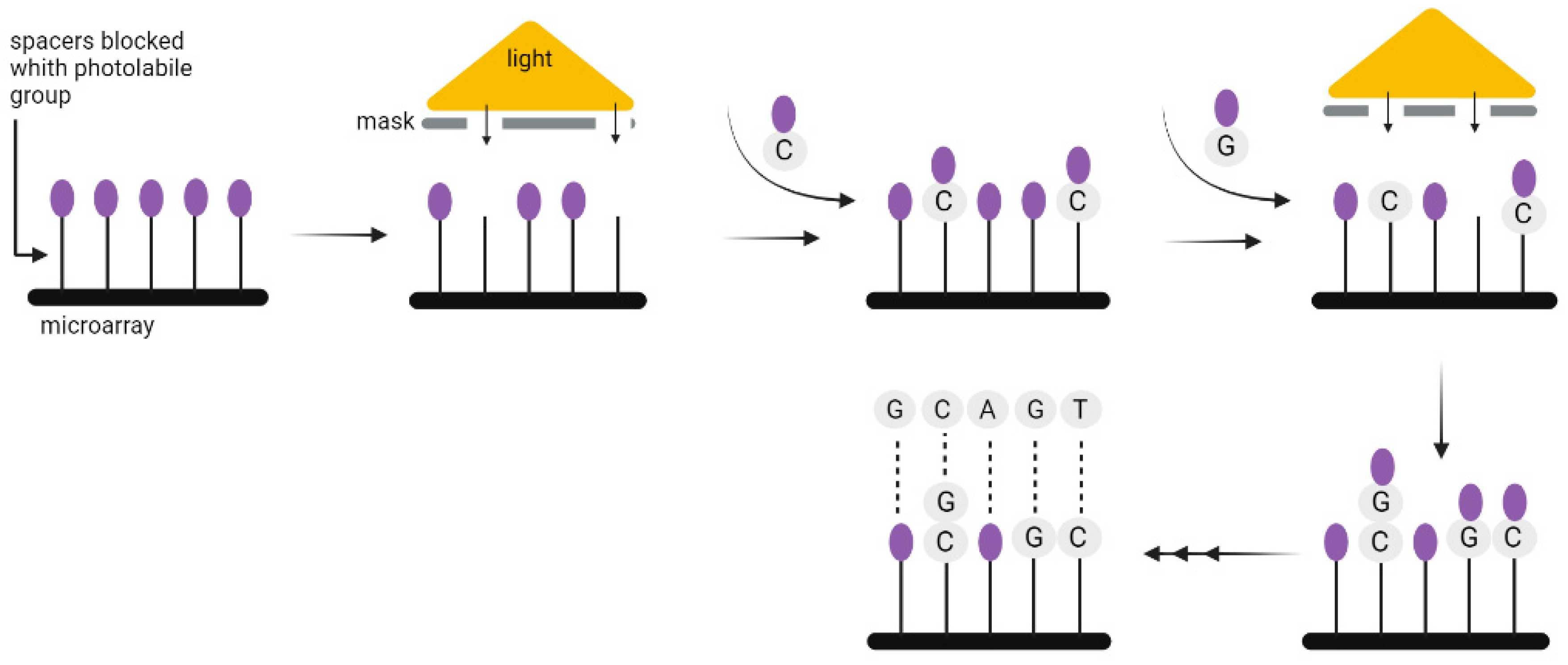
Figure 6. A solid-phase method for the synthesis of oligonucleotides using photolabile compounds. A spacer containing the photolabile group is covalently joined to the surface. Once spots on the surface are exposed to UV light through slits in the physical mask, the photolabile protecting group is removed and the synthesis of oligonucleotide begins. The subsequent appropriate phosphoramidite with the photolabile group is then applied to the entire surface of the plate. It can form covalent bonds only in the absence of the preceding photolabile group. In the subsequent steps, additional spots are exposed to radiation, and another phosphoramidite is applied where necessary. Until the final oligonucleotide is completely synthesized, the chain-extending processes are repeated.
References
- De Silva, P.Y.; Ganegoda, G.U. New Trends of Digital Data Storage in DNA. BioMed Res. Int. 2016, 2016, 8072463.
- Rydning, J.; Reinsel, D.; Gantz, J. The Digitization of the World from Edge to Core; IDC: Framingham, MA, USA, 2018.
- Ceze, L.; Nivala, J.; Strauss, K. Molecular Digital Data Storage Using DNA. Nat. Rev. Genet. 2019, 20, 456–466.
- Grass, R.N.; Heckel, R.; Puddu, M.; Paunescu, D.; Stark, W.J. Robust Chemical Preservation of Digital Information on DNA in Silica with Error-Correcting Codes. Angew. Chem. Int. Ed. Engl. 2015, 54, 2552–2555.
- Zhirnov, V.; Zadegan, R.M.; Sandhu, G.S.; Church, G.M.; Hughes, W.L. Nucleic Acid Memory. Nat. Mater. 2016, 15, 366–370.
- Van der Valk, T.; Pečnerová, P.; Díez-Del-Molino, D.; Bergström, A.; Oppenheimer, J.; Hartmann, S.; Xenikoudakis, G.; Thomas, J.A.; Dehasque, M.; Sağlıcan, E.; et al. Million-Year-Old DNA Sheds Light on the Genomic History of Mammoths. Nature 2021, 591, 265–269.
- Horneck, G.; Klaus, D.M.; Mancinelli, R.L. Space Microbiology. Microbiol. Mol. Biol. Rev. 2010, 74, 121–156.
- Horneck, G.; Bücker, H.; Reitz, G. Long-Term Survival of Bacterial Spores in Space. Adv. Space Res. 1994, 14, 41–45.
- Cadet, J.; Sage, E.; Douki, T. Ultraviolet Radiation-Mediated Damage to Cellular DNA. Mutat. Res. 2005, 571, 3–17.
- Xue, Y.; Nicholson, W.L. The Two Major Spore DNA Repair Pathways, Nucleotide Excision Repair and Spore Photoproduct Lyase, Are Sufficient for the Resistance of Bacillus Subtilis Spores to Artificial UV-C and UV-B but Not to Solar Radiation. Appl. Environ. Microbiol. 1996, 62, 2221–2227.
- Sancho, L.G.; de la Torre, R.; Horneck, G.; Ascaso, C.; de Los Rios, A.; Pintado, A.; Wierzchos, J.; Schuster, M. Lichens Survive in Space: Results from the 2005 LICHENS Experiment. Astrobiology 2007, 7, 443–454.
- Gauslaa, Y.; Solhaug, K.A. Photoinhibition in Lichens Depends on Cortical Characteristics and Hydration. Lichenologist 2004, 36, 133–143.
- Ahmed, R.K.; Mohammed, I.J. Developing a New Hybrid Cipher Algorithm Using DNA and RC4. Int. J. Adv. Comput. Sci. Appl. 2017, 8, 71.
- Zhang, Y.; Kong, L.; Wang, F.; Li, B.; Ma, C.; Chen, D.; Liu, K.; Fan, C.; Zhang, H. Information Stored in Nanoscale: Encoding Data in a Single DNA Strand with Base64. Nano Today 2020, 33, 100871.
- Church, G.M.; Gao, Y.; Kosuri, S. Next-Generation Digital Information Storage in DNA. Science 2012, 337, 1628.
- Goldman, N.; Bertone, P.; Chen, S.; Dessimoz, C.; LeProust, E.M.; Sipos, B.; Birney, E. Towards Practical, High-Capacity, Low-Maintenance Information Storage in Synthesized DNA. Nature 2013, 494, 77–80.
- Ailenberg, M.; Rotstein, O.D. An Improved Huffman Coding Method for Archiving Text, Images, and Music Characters in DNA. BioTechniques 2009, 47, 747–754.
- Yazdi, S.M.H.T.; Gabrys, R.; Milenkovic, O. Portable and Error-Free DNA-Based Data Storage. Sci. Rep. 2017, 7, 5011.
- Shipman, S.L.; Nivala, J.; Macklis, J.D.; Church, G.M. CRISPR-Cas Encoding of a Digital Movie into the Genomes of a Population of Living Bacteria. Nature 2017, 547, 345–349.
- Sinyakov, A.N.; Ryabinin, V.A.; Kostina, E.V. Application of Array-Based Oligonucleotides for Synthesis of Genetic Designs. Mol. Biol. 2021, 55, 487–500.
- Song, L.-F.; Deng, Z.-H.; Gong, Z.-Y.; Li, L.-L.; Li, B.-Z. Large-Scale de Novo Oligonucleotide Synthesis for Whole-Genome Synthesis and Data Storage: Challenges and Opportunities. Front. Bioeng. Biotechnol. 2021, 9, 689797.
- Heckel, R.; Shomorony, I.; Ramchandran, K.; Tse, D.N.C. Fundamental Limits of DNA Storage Systems. In Proceedings of the 2017 IEEE International Symposium on Information Theory (ISIT), Aachen, Germany, 25–30 June 2017; pp. 3130–3134.
More
Information
Subjects:
Others
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.1K
Revisions:
2 times
(View History)
Update Date:
07 Jun 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





