Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Janko Samardzic | -- | 3813 | 2023-05-29 08:40:48 | | | |
| 2 | Rita Xu | Meta information modification | 3813 | 2023-05-29 08:53:29 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Radosavljevic, M.; Svob Strac, D.; Jancic, J.; Samardzic, J. Pharmacogenetics of Antidepressants. Encyclopedia. Available online: https://encyclopedia.pub/entry/44945 (accessed on 21 May 2026).
Radosavljevic M, Svob Strac D, Jancic J, Samardzic J. Pharmacogenetics of Antidepressants. Encyclopedia. Available at: https://encyclopedia.pub/entry/44945. Accessed May 21, 2026.
Radosavljevic, Milica, Dubravka Svob Strac, Jasna Jancic, Janko Samardzic. "Pharmacogenetics of Antidepressants" Encyclopedia, https://encyclopedia.pub/entry/44945 (accessed May 21, 2026).
Radosavljevic, M., Svob Strac, D., Jancic, J., & Samardzic, J. (2023, May 29). Pharmacogenetics of Antidepressants. In Encyclopedia. https://encyclopedia.pub/entry/44945
Radosavljevic, Milica, et al. "Pharmacogenetics of Antidepressants." Encyclopedia. Web. 29 May, 2023.
Copy Citation
Pharmacotherapy for neuropsychiatric disorders, such as depression, has been characterized by significant inter-individual variability in drug response and the development of side effects. Pharmacogenetics, as a key part of personalized medicine, aims to optimize therapy according to a patient’s individual genetic signature by targeting genetic variations involved in pharmacokinetic or pharmacodynamic processes. Pharmacokinetic variability refers to variations in a drug’s absorption, distribution, metabolism, and elimination, whereas pharmacodynamic variability results from variable interactions of an active drug with its target molecules.
pharmacogenetics
pharmacogenomics
anxiety
depression
1. Introduction
The pharmacotherapy of neuropsychiatric disorders, such as anxiety and depression, has been characterized by significant inter-individual variability in drug response and the development of severe adverse effects, which has been recognized as a major clinical problem [1][2]. In addition to various environmental, physiological, and psychological factors, these individual differences might be largely due to genetic factors [3][4]. Therefore, various pharmacogenetic studies have been conducted to identify genetic variants that can predict patients who may optimally benefit from specific, individually tailored treatments [5][6][7][8][9] (Figure 1).
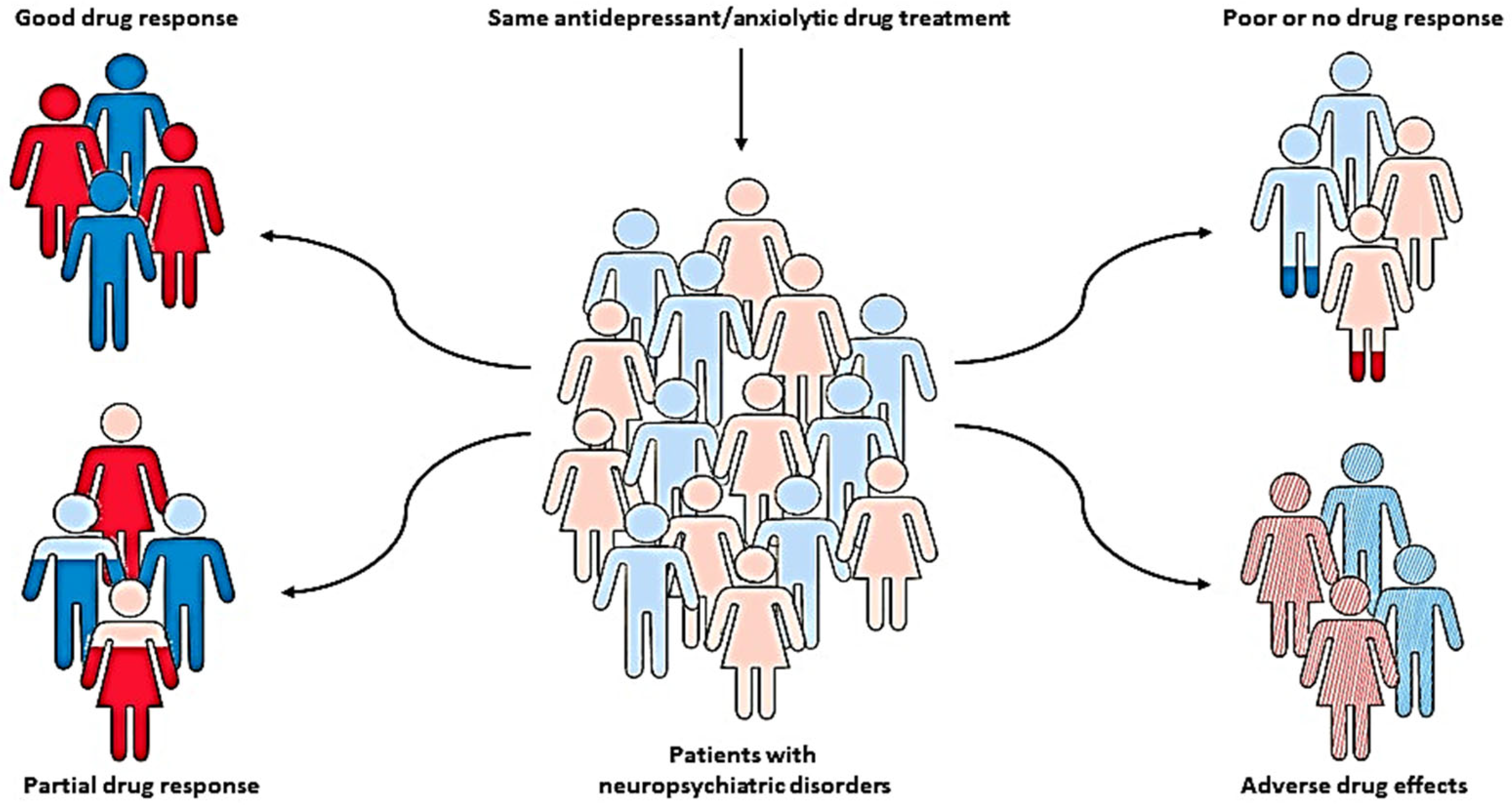
Figure 1. Pharmacogenetics, as a key part of personalized medicine, can help clinicians predict the therapeutic response and adverse drug reactions in patients with the same diagnosis and treatment, but different genotypes, and therefore identify patients who may optimally benefit from specific, individually tailored treatment.
Pharmacogenetic studies have focused primarily on candidate genes involved in drug metabolism and transport (pharmacokinetics) as well as drug action (pharmacodynamics), which can influence both treatment efficacy and the development of adverse drug effects [9][10][11]. Pharmacokinetics addresses the variability in the drug’s absorption, distribution, metabolism, and elimination (ADME), which modulates the delivery of drugs and their active metabolites or their removal from action targets.
The molecules involved in ADME processes include enzymes responsible for drug metabolism and drug transport molecules that mediate drug uptake and efflux [4]. The cytochrome P450 (CYP) and multidrug resistance (MDR) gene families have been extensively studied [6][12]. The CYP450 enzyme family in the liver is responsible for the metabolism of many psychotropic drugs [13]. For certain CYPs, the genotype affects the serum/plasma drug levels, and consequently, its efficacy and development of adverse effects [14][15]. There are four major CYP phenotypes produced by combinations of various alleles with different degrees of enzymatic activity: poor (PM), intermediate (IM), extensive (normal) (EM), and ultrarapid metabolizer (UM). PMs tend to accumulate higher drug levels in the blood and may require lower drug doses to achieve therapeutic effects, whereas UMs may require higher doses due to faster drug elimination [16][17].
In addition, the therapeutic action of psychotropic drugs depends on their effective delivery to the brain. Although some substances may diffuse passively through the brain-blood barrier (BBB), the influx and efflux of most substances are actively regulated by a complex system of transporters, influencing both pharmacokinetics and pharmacodynamics. In the case of a genetically determined decrease in functional activity or expression of transport proteins in the BBB, drug efflux from the brain into the blood is disturbed. This could lead to increased drug exposure time in the brain, its accumulation during long-term therapy, and an increased risk of developing severe adverse effects [18].
In contrast to pharmacokinetics, pharmacodynamics describes variability in drug action not dependent on variable drug concentrations but rather on the interaction of the active drug with its target molecules, including receptors, ion channels, and enzymes, and it can also influence both therapeutic responses and drug side effects.
Single nucleotide polymorphisms (SNPs) are the most commonly investigated genetic variants in both pharmacokinetic and pharmacodynamic studies [11]. Genetic polymorphism refers to the occurrence of two or more common variants (alleles) of a specific DNA sequence in a population with a frequency of more than 1% [19]. Identifying SNPs associated with variability in drug response and toxicity has been the primary focus of a significant number of pharmacogenetic and pharmacogenomic studies [7][20][21]. In these studies, two major research approaches have been used: the traditional candidate gene approach, which is hypothesis-driven and based on accumulated knowledge, and new methodologies such as genome-wide association studies (GWAS) or whole exome sequencing, which are data-driven and generate new hypotheses and knowledge [22][23].
However, pharmacogenetics is not able to explain all observed heritable variations in drug responses, and there is growing evidence that responses to drugs could be influenced by individual epigenetic states [24][25]. An emerging field of pharmacoepigenetics investigates how epigenetic mechanisms that modify gene expression without altering the genetic code might influence individual responses to drugs [24][26]. Some of the most frequently studied epigenetic modifications include DNA methylation, histone modifications, and noncoding RNA actions [25]. Various environmental factors, including drugs, nutrition, and stress, may induce epigenetic changes [27] that can be transmitted over generations [28]. Acute or chronic exposure to stressors can contribute to the development and progression of various neuropsychiatric disorders [29][30][31][32], including anxiety and depression, but it is also associated with alterations in the epigenome that may affect the expression of genes involved in drug metabolism, transport, and target molecules, and therefore impact the variability in antidepressant and anxiolytic drug responses [33][34].
2. Pharmacogenetics of Antidepressants
2.1. Pharmacokinetic Variability
2.1.1. Cytochrome P450 Family
The CYP450 family is a large group of enzymes responsible for the metabolism of various drugs and xenobiotics, including antidepressant drugs. Among the many identified CYP450 enzymes, the most important ones involved in the metabolism of a variety of psychotropic drugs are CYP2D6, CYP2C19, CYP3A4, CYP1A2, CYP2B6, CYP2C8, and CYP2C9 [9][35]. Various CYP450 enzymes are capable of metabolizing more than one drug, and a single drug can be metabolized by multiple CYP enzymes [15]. The activity of these enzymes could be influenced by genetic variations, which may result in individual differences in antidepressant drug metabolism and responses [15][16]. Most genes encoding CYPs are highly polymorphic [19]. The impact of CYP polymorphisms on drug metabolism is an important area of research in personalized medicine [19]. While more than 2000 mutations have been found in CYP genes, only specific SNPs are known to affect CYP enzymatic activity [14][15]. Depending on the genetic variants that influence enzyme activity, individuals can be classified into four main CYP phenotypes: from poor, via intermediate and extensive (normal), to ultrarapid metabolizers [17][36][37]. Due to genetic variations, PMs have little to no CYP enzyme activity, whereas IMs have reduced enzyme activity. As a result of slower antidepressant drug metabolism, both PMs and IMs may be at an increased risk for adverse drug reactions and require lower dosages or less frequent dosing regimens for antidepressant medications. On the other hand, NMs possess typical enzyme activity and a normal drug metabolism rate. However, UMs exhibit increased enzymatic activity, resulting in faster drug metabolism. Therefore, they may require higher antidepressant doses or more frequent drug administration to achieve the desired therapeutic effect in patients with depression [17][18][36][37].
CYP2D6
The CYP2D6 gene is highly polymorphic, meaning that it harbors genetic variations that can affect enzyme function [15][16][38]. CYP2D6 enzyme is involved in the metabolism of SSRIs (paroxetine, fluvoxamine, and fluoxetine), amitriptyline (TCA), and venlafaxine (SNRI) [16]. Different allele variants result in normal, decreased, and no enzyme function, characterized by extensive, intermediate, and poor metabolizer phenotypes, respectively [16][39]. CYP2D6*1, *2, *33, and *35 allele variants are associated with normal function, whereas *9, *10, *14B, *17, *29, and *41 variants reduce CYP2D6 enzymatic capacity. Furthermore, CYP2D6*3, *4, *5, *6, *7, *11, *12, *14A, *36, and *68 variants are responsible for the loss of enzyme function [16][39]. Apart from genetic variations, paroxetine and fluoxetine are potent inhibitors of CYP2D6, which may result in an “iatrogenic poor phenotype” or “phenocopy” when taken with drugs that are also metabolized by CYP2D6, such as venlafaxine. Patients with an “iatrogenic poor phenotype” may be at high risk of developing toxicity due to elevated plasma drug levels [40]. Specifically, individuals with poor CYP2D6 function may have reduced metabolism of fluoxetine and paroxetine, resulting in higher plasma drug levels and an increased risk of side effects, such as the development of suicidal ideation or antidepressant-induced mania during the initiation of treatment. As a result, the U.S. Food and Drug Administration (FDA) has issued a black box warning for SSRIs cautioning that these side effects can occur, especially in the first few weeks of treatment. Therefore, close monitoring is required for all patients starting antidepressant therapy [40][41].
Furthermore, studies have shown that poor metabolizers of fluoxetine may be at a risk of QT interval prolongation. [40]. Similar to fluoxetine, poor metabolizers of venlafaxine (genotype CYP2D6 *6/*4, *5/*4, or *6/*6) are also at risk for developing adverse effects, such as arrhythmias, gastrointestinal problems, and hyponatremia [42][43]. Ahmed et al. showed that CYP2D6 UM status contributes to venlafaxine treatment remission in patients with major depressive disorder [44]. Therefore, when prescribing SSRIs and other antidepressants, clinicians must consider a patient’s CYP2D6 genotype and adjust the dose or choose an alternative medication, if necessary, to minimize the risk of adverse effects while maximizing therapeutic benefit [6][7][20].
CYP2C19
In addition to clinically significant variants of the CYP2D6 gene, a high number of polymorphisms in the CYP2C19 gene have also been discovered [45], resulting in different metabolizer phenotypes [16][39]. Drugs that are mainly metabolized by CYP2C19 include SSRIs such as escitalopram, citalopram, and sertraline [15]. Variations in the CYP2C19 gene can result in individuals having different levels of CYP2C19 enzyme activity, affecting how drugs such as escitalopram and sertraline are metabolized [46][47]. According to previous research, the CYP2C19*1 allele variant is associated with normal function, whereas the CYP2C19*17 variant increases CYP2C19 enzymatic capacity. Furthermore, CYP2C19*2 and *3 variants are responsible for the loss of enzyme function [16][39].
For example, PMs of escitalopram are more likely to have adverse effects and psychotherapy discontinuation. However, they respond better to escitalopram, provided the therapy is tolerated. A retrospective study involving more than 2000 escitalopram-treated patients demonstrated that those with CYP2C19*1/*17 and CYP2C19*17/*17 variants (IMs) were more likely to experience treatment failure [48], whereas UMs exhibited suicidal thoughts [49]. For sertraline, recent findings demonstrated that CYP2C19 PMs had higher plasma levels in comparison to normal metabolizers [50]. Ricardo-Silgado et al. [51] suggested that CYP2C19 genotypes might be associated with an increased risk of weight gain in patients on citalopram therapy. These findings indicate the importance of CYP2C19 variants and different metabolic phenotypes in antidepressant treatment tolerability and outcome. The American Molecular Pathology published guidelines for testing CYP2C19*2, *3, and *17 variants and CYP2C19*4A-*4B, *5, *6, *7, *8, *9, *10, and *35 variants [39]. In addition, the relevance of CYP2C19 polymorphisms in inter-individual predisposition to mental diseases was also investigated. Sim et al. [52] found that individuals who were poor CYP2C19 metabolizers displayed lower levels of depressive symptoms compared to individuals who were normal metabolizers.
Moreover, a cross-sectional observational retrospective study, which included over 700 psychiatric patients with depression and anxiety, analyzed the frequencies of CYPC19*2, *4, and *17, as well as CYP2D6*2, *3, *4, *5, *6, *10, and *41 variants [35]. This study revealed that 77% of patients have at least one allele variant significantly affecting drug metabolism, with roughly half of the individuals with reduced CYP2D6 enzyme function and the majority of them being CYP2C19 rapid and ultrarapid metabolizers [35]. Therefore, due to their clinical relevance, both CYP2D6 and CYP2C19 are included in pharmacogenetics guidelines and recommendations published by the Clinical Pharmacogenetics Implementation Consortium (CPIC), Dutch Pharmacogenetics Working Group (DPWG), and regulatory agencies such as the FDA [53][54][55].
CYP2C9
The CYP2C9 gene encodes an enzyme that is involved in the metabolism of many drugs, including antidepressants, such as MAOIs, TCAs, and SSRIs [17]. The CYP2C9*1 variant is associated with normal enzyme function, whereas CYP2C9*2, *5, *8, and *11 variants reduce CYP2C9 enzymatic capacity. Furthermore, CYP2D9*3, *6, and *13 variants are responsible for the loss of enzyme function [16][39]. Different CYP2C9 variants result in normal, decreased, and no enzyme function, characterized by extensive, intermediate, and poor metabolizer phenotypes, respectively [16][39]. Moreover, variants of the CYP2C9 gene have been linked to mental disorders such as depression and psychosis [56][57].
CYP1A2
The CYP1A2 enzyme, encoded by the CYP1A2 gene, is responsible for the metabolism of approximately 24% of antidepressant drugs, including agomelatine, escitalopram, venlafaxine, duloxetine, and mirtazapine [16][58]. Several CYP1A2 variants, including *1C, *1F, and *1B, have been identified as potential indicators of the CYP1A2 phenotype and agomelatine pharmacokinetics. Notably, the CYP1A2*1C variant is associated with reduced enzyme activity, resulting in higher plasma agomelatine concentrations and an increased risk of adverse effects. On the contrary, individuals with the CYP1A2 *1F and *1B variants, who are characterized by increased enzyme activity, may metabolize agomelatine more rapidly, leading to lower plasma concentrations and potentially reduced efficacy [59][60]. Kuo et al. [61] conducted a study to investigate the association between CYP1A2 polymorphisms and the metabolism of escitalopram in patients with major depressive disorder. The study found that several SNPs in the CYP1A2 gene, such as rs2069521, rs4646425, and rs4646427 polymorphisms, were associated with altered escitalopram metabolism and an increased risk of adverse effects, such as fatigue and nausea/vomiting, particularly during the initial stages of treatment. In addition, Lin et al. [62] demonstrated that CYP1A2 SNPs, such as rs4646425, rs2472304, and rs2470890, are associated with a slower response to paroxetine treatment. Furthermore, the study of the linkage between CYP1A2 polymorphisms and the response to venlafaxine suggested that the rs2470890 polymorphism might be related to venlafaxine treatment remission [63].
2.1.2. P-glycoprotein
P-glycoprotein, commonly referred to as P-gp, is a membrane transporter within the ATP-binding cassette (ABC) transporter family. It is encoded by the ABCB1 or MDR1 gene, located on chromosome 7. P-gp functions as an efflux pump, contributing to drug absorption, distribution, and elimination [12]. P-gp has been found in various tissues throughout the body, including the liver, kidneys, intestines, and the blood-brain barrier. At the blood-brain barrier, P-gp can limit the entry of some drugs into the brain, which can affect their efficacy. The expression and activity of P-gp can vary widely between individuals, contributing to differences in drug responses and drug interactions [18]. Genetic factors play a role in determining the level of P-gp expression in different tissues and individuals. Certain genetic variants of the ABCB1 gene have been associated with altered P-gp expression and activity, which can affect the pharmacokinetics and efficacy of drugs that are substrates for P-gp [64]. P-gp has broad substrate specificity, including antidepressants such as escitalopram, fluvoxamine, paroxetine, amitriptyline, and imipramine. Therefore, when P-gp is absent or not functioning properly, these drugs can accumulate in the body, resulting in higher concentrations and potentially an increased risk of adverse effects. However, newer antidepressants (levomilnacipran, vortioxetine, and vilazodone) have been proven as poor P-gp substrates [18].
There has been interest in studying the relationship between genetic variations in the MDR1/ABCB1 gene and antidepressant treatment outcomes [65]. Several genetic variants of the MDR1/ABCB1 gene have been associated with altered P-gp activity. The three most common MDR1/ABCB1 variants, C3435T (rs1045642), C1236T (rs1128503), and G2677T (rs2032582), have been the subject of extensive research [65][66]. A recent study on experimental models suggested that the 2677G > T polymorphism in the ABCB1 gene has been associated with altered P-gp function and increased brain penetration of P-gp substrates without affecting P-gp protein expression in the blood-brain barrier [67]. In addition, genetic variants of the ABCB1 gene, including the rs2235040 and rs4148739 polymorphisms, may be associated with the onset of response to antidepressant medications rather than the response rate [8]. Furthermore, a significant link has been found between the ABCB1 rs2235015 GG genotype and better response to antidepressant treatment [8][68]. Overall, while pharmacogenetic research on MDR1/ABCB1 suggests the potential for improving outcomes of antidepressant treatment, further investigation is necessary to better comprehend the connection between genetic variations and treatment response, as well as to assess its clinical utility.
2.2. Pharmacodynamic Variability
2.2.1. Monoamine Metabolic Enzymes
Tryptophan Hydroxylase
Tryptophan hydroxylase (TPH) is an enzyme that catalyzes the conversion of the amino acid tryptophan to 5-hydroxytryptophan, which is the first step in the synthesis of serotonin, a neurotransmitter involved in the regulation of mood, appetite, sleep, and stress response [69]. There are two isoforms of the TPH enzyme, TPH1 and TPH2, encoded by separate genes. TPH1 is primarily expressed in peripheral tissues, such as the pineal gland, skin, and gut, whereas TPH2 is mainly expressed in neurons in the central nervous system (CNS). The two isoforms have similar enzymatic activities but differ in regulation, tissue distribution, and developmental expression patterns [70]. Overall, while TPH1 may be less expressed in the brain than TPH2, it appears to play an important role in the regulation of mood and stress responses and may contribute to antidepressant effects [71]. Several variants of TPH genes associated with differences in serotonin production have been identified. Some of these variants have been linked to an increased risk of developing psychiatric disorders [72]. The modulation of TPH genes has been studied as a potential approach for the treatment of various psychiatric disorders, including depression, anxiety, and addiction [73][74]. However, several studies have demonstrated controversial results regarding TPH polymorphisms and response to SSRIs [21][72][75].
Monoamine Oxidases
Monoamine oxidases (MAOs) are a family of enzymes that play critical roles in the metabolism of monoamine neurotransmitters in the CNS [76]. There are two types of MAOs, MAO-A and MAO-B, which are encoded by separate genes. MAO-A and MAO-B are found in the CNS, particularly in neurons and astroglia. MAO-A is primarily responsible for the metabolism of several important monoamine neurotransmitters, including serotonin, norepinephrine, and dopamine. These neurotransmitters play a crucial role in the regulation of mood, behavior, and cognition [77]. Several genetic variants of the MAOA gene associated with differences in enzyme activity and monoamine metabolism have been identified. In addition, some of these variants have been linked to an increased risk of developing psychiatric disorders [77]. However, the relationship between the MAOA gene variants and psychiatric disorders remains unclear. Polymorphisms in MAO genes have also been studied for their potential role in response to antidepressant medications. Some studies have suggested that certain genetic variants in MAO genes may affect the metabolism of antidepressants, potentially leading to differences in drug efficacy and side effects [78][79][80]. For example, individuals with a specific variant of the MAOA gene had a better response to fluvoxamine compared to those without this variant [79]. In addition, a recent study reported that the MAOA rs979605 polymorphism might modulate the response to antidepressant therapy in a sex-specific manner [81].
Catechol-O-Methyltransferase
Catechol-O-methyltransferase (COMT) is an enzyme that plays a crucial role in the degradation of catecholamines such as dopamine, epinephrine, and norepinephrine. The COMT gene has several polymorphisms, and the most extensively studied is the rs4680 polymorphism, also known as Val158Met [82]. This SNP causes the substitution of valine (Val) for methionine (Met) at position 158 in the COMT enzyme. The Val allele is associated with higher COMT activity, whereas the Met allele leads to lower activity. Therefore, individuals with the Val/Val genotype tend to have the highest level of COMT activity, followed by Val/Met individuals with intermediate activity, and Met/Met with the lowest activity [83]. The COMT Val158Met polymorphism has been implicated in various psychiatric disorders, including depression, anxiety, and schizophrenia [82]. Furthermore, studies have linked the COMT Val158Met polymorphism with response the to SSRIs such as fluoxetine and paroxetine [84][85]. For instance, one study discovered that the Val/Met genotype significantly affected the response to fluvoxamine [84]. Additionally, it has been suggested that the effects of the COMT genotype on antidepressant responses may depend on other factors, such as the type of medication, severity and duration of depression, and other genetic and environmental factors [86].
2.2.2. Monoamine Transporters
Serotonin Transporter
The serotonin transporter (SERT) encoded by the SLC6A4 gene plays a critical role in regulating serotonin neurotransmission in the brain. SERT is responsible for serotonin reuptake from the synaptic cleft, thereby regulating the amount of serotonin available to bind to serotonin receptors in the brain [87]. Several classes of antidepressant drugs, including SSRIs, SNRIs, and TCAs, target SERT and inhibit its activity [88].
The SLC6A4 gene was the first gene genotyped as part of the Genome-Based Therapeutic Drugs for Depression (GENDEP), and several genetic variants have been identified [89]. The most studied is the variant of the serotonin transporter-linked polymorphic region (5-HTTLPR). The 5-HTTLPR is located in the promoter region of the SLC6A4 gene and has two alleles: the short (S) and the long (L) allele. The S allele is associated with lower transcriptional efficiency and reduced expression of the SERT protein compared to the L allele. As a result, carriers of the S allele have been found to have lower serotonin reuptake efficiency, leading to higher serotonin levels. Therefore, individuals carrying the S/S genotype have been shown to have a poorer response to SSRI treatment and may experience more side effects compared to those with the L/L or L/S genotypes [21][90]. Numerous studies have investigated the relationship between the 5-HTTLPR polymorphism and response to SSRIs [91][92][93][94]. A recent case report suggested that the S/S genotype resulted in ineffective fluoxetine treatment and may have caused exacerbation of depression [95]. Maron et al. [96] investigated the association between 5-HTTLPR polymorphism and clinical response to escitalopram and found no significant association. However, they did observe a linkage between this polymorphism and a higher risk for adverse effects [96]. This finding is consistent with other studies that have reported an association between SSRIs side effects and 5-HTTLPR polymorphism [97][98][99]. Therefore, the 5-HTTLPR polymorphism may be a valuable marker for predicting antidepressant responses and tolerability in individuals with psychiatric disorders. In particular, genotyping for the 5-HTTLPR variant may help identify patients who are less likely to respond to SSRIs and may benefit from alternative treatment strategies, such as SNRIs or TCAs. Additionally, several studies have reported an association between the 5-HTTLPR S/S genotype and suicidal behavior [99][100][101], although other risk factors, such as life stressors, psychiatric disorders, and substance use, must be considered.
Norepinephrine Transporter
The norepinephrine transporter (NET) is a protein encoded by the SLC6A2 gene and plays an important role in the reuptake of norepinephrine (NE) [102]. TCAs, such as desipramine and imipramine, work by inhibiting the reuptake of NE by the NET, which increases the levels of NE in the synapse and enhances its effects on target neurons. Although antidepressants, such as SSRIs, have largely replaced TCAs, they are still used in some cases where other treatments have been ineffective. Several genetic variants have been identified in the human SLC6A2 gene, which have functional consequences and can affect the efficacy of antidepressant drugs [102]. A recent study found an association between certain SLC6A2 polymorphisms and antidepressant response variability [103]. Although, the role of SLC6A2 variants in antidepressant responses is still being studied, current evidence suggests that these associations are complex and not fully understood.
Dopamine Transporter
The dopamine transporter (DAT), encoded by the SLC6A3 gene, is a protein responsible for dopamine reuptake from the synaptic cleft into the presynaptic neuron. To date, the SLC6A3 gene has been identified with at least 502 variations [104]. However, the relationship between SLC6A3 polymorphisms and antidepressant responses is yet to be thoroughly researched. Nevertheless, some studies have suggested that specific SLC6A3 polymorphisms may influence the response to antidepressant therapy [105][106]. For example, Kirchheiner et al. [107] suggested that the SLC6A3 polymorphism increased the risk of a poorer and slower response to various antidepressants in individuals with the 9/10 and 9/9 genotypes compared to carriers of the 10/10 genotype.
References
- Willner, P.; Bergman, J.; Vanderschuren, L. The Behavioural Pharmacology of Stress-Related Disorders. Behav. Pharmacol. 2019, 30, 101–103.
- Leichsenring, F.; Steinert, C.; Rabung, S.; Ioannidis, J.P.A. The Efficacy of Psychotherapies and Pharmacotherapies for Mental Disorders in Adults: An Umbrella Review and Meta-Analytic Evaluation of Recent Meta-Analyses. World Psychiatry 2022, 21, 133–145.
- Kam, H.; Jeong, H. Pharmacogenomic Biomarkers and Their Applications in Psychiatry. Genes 2020, 11, 1445.
- Jukic, M.; Milosavljević, F.; Molden, E.; Ingelman-Sundberg, M. Pharmacogenomics in Treatment of Depression and Psychosis: An Update. Trends Pharmacol. Sci. 2022, 43, 1055–1069.
- Lunenburg, C.A.T.C.; Ishtiak-Ahmed, K.; Werge, T.; Gasse, C. Life-Time Actionable Pharmacogenetic Drug Use: A Population-Based Cohort Study in 86 040 Young People with and without Mental Disorders in Denmark. Pharmacopsychiatry 2022, 55, 95–107.
- Kee, P.S.; Maggo, S.D.S.; Kennedy, M.A.; Chin, P.K.L. The Pharmacogenetics of CYP2D6 and CYP2C19 in a Case Series of Antidepressant Responses. Front. Pharmacol. 2023, 14, 1080117.
- Xin, J.; Yuan, M.; Peng, Y.; Wang, J. Analysis of the Deleterious Single-Nucleotide Polymorphisms Associated with Antidepressant Efficacy in Major Depressive Disorder. Front. Psychiatry 2020, 11, 151.
- Geers, L.M.; Ochi, T.; Vyalova, N.M.; Losenkov, I.S.; Paderina, D.Z.; Pozhidaev, I.V.; Simutkin, G.G.; Bokhan, N.A.; Wilffert, B.; Touw, D.J.; et al. Influence of Eight ABCB1 Polymorphisms on Antidepressant Response in a Prospective Cohort of Treatment-Free Russian Patients with Moderate or Severe Depression: An Explorative Psychopharmacological Study with Naturalistic Design. Hum. Psychopharmacol. 2022, 37, e2826.
- Del Casale, A.; Pomes, L.M.; Bonanni, L.; Fiaschè, F.; Zocchi, C.; Padovano, A.; De Luca, O.; Angeletti, G.; Brugnoli, R.; Girardi, P.; et al. Pharmacogenomics-Guided Pharmacotherapy in Patients with Major Depressive Disorder or Bipolar Disorder Affected by Treatment-Resistant Depressive Episodes: A Long-Term Follow-up Study. J. Pers. Med. 2022, 12, 316.
- Shalimova, A.; Babasieva, V.; Chubarev, V.N.; Tarasov, V.V.; Schiöth, H.B.; Mwinyi, J. Therapy Response Prediction in Major Depressive Disorder: Current and Novel Genomic Markers Influencing Pharmacokinetics and Pharmacodynamics. Pharmacogenomics 2021, 22, 485–503.
- Zięba, A.; Matosiuk, D.; Kaczor, A.A. The Role of Genetics in the Development and Pharmacotherapy of Depression and Its Impact on Drug Discovery. Int. J. Mol. Sci. 2023, 24, 2946.
- Patel, K.A.; Bhatt, M.H.; Hirani, R.V.; Patel, V.A.; Patel, V.N.; Shah, G.B.; Chorawala, M.R. Assessment of Potential Drug-Drug Interactions among Outpatients in a Tertiary Care Hospital: Focusing on the Role of P-Glycoprotein and CYP3A4 (Retrospective Observational Study). Heliyon 2022, 8, e11278.
- Daniel, W.A.; Bromek, E.; Danek, P.J.; Haduch, A. The Mechanisms of Interactions of Psychotropic Drugs with Liver and Brain Cytochrome P450 and Their Significance for Drug Effect and Drug-Drug Interactions. Biochem. Pharmacol. 2022, 199, 115006.
- Mostafa, S.; Polasek, T.M.; Bousman, C.A.; Müeller, D.J.; Sheffield, L.J.; Rembach, J.; Kirkpatrick, C.M. Pharmacogenomics in Psychiatry—The Challenge of Cytochrome P450 Enzyme Phenoconversion and Solutions to Assist Precision Dosing. Pharmacogenomics 2022, 23, 857–867.
- van Westrhenen, R.; Aitchison, K.J.; Ingelman-Sundberg, M.; Jukić, M.M. Pharmacogenomics of Antidepressant and Antipsychotic Treatment: How Far Have We Got and Where Are We Going? Front. Psychiatry 2020, 11, 94.
- Alchakee, A.; Ahmed, M.; Eldohaji, L.; Alhaj, H.; Saber-Ayad, M. Pharmacogenomics in Psychiatry Practice: The Value and the Challenges. Int. J. Mol. Sci. 2022, 23, 13485.
- Shetty, P. Pharmacogenomics and Its Future Implications in Treatment-Resistant Depression. Ind. J. Priv. Psychiatry 2019, 13, 71–76.
- Wyska, E. Pharmacokinetic Considerations for Current State-of-the-Art Antidepressants. Expert Opin. Drug Metab. Toxicol. 2019, 15, 831–847.
- Samardzic, J.; Svob Strac, D.; van den Anker, J.N. The Benefit and Future of Pharmacogenetics. In Total Intravenous Anesthesia and Target Controlled Infusions; Absalom, A.R., Mason, K.P., Eds.; Springer: New York, NY, USA, 2017; pp. 697–711.
- Suzuki, Y.; Sawamura, K.; Someya, T. Polymorphisms in the 5-Hydroxytryptamine 2A Receptor and CytochromeP4502D6 Genes Synergistically Predict Fluvoxamine-Induced Side Effects in Japanese Depressed Patients. Neuropsychopharmacology 2006, 31, 825–831.
- Brunoni, A.R.; Carracedo, A.; Amigo, O.M.; Pellicer, A.L.; Talib, L.; Carvalho, A.F.; Lotufo, P.A.; Benseñor, I.M.; Gattaz, W.; Cappi, C. Association of BDNF, HTR2A, TPH1, SLC6A4, and COMT Polymorphisms with TDCS and Escitalopram Efficacy: Ancillary Analysis of a Double-Blind, Placebo-Controlled Trial. Rev. Bras. Psiquiatr. 2020, 42, 128–135.
- Motsinger-Reif, A.A.; Jorgenson, E.; Relling, M.V.; Kroetz, D.L.; Weinshilboum, R.; Cox, N.J.; Roden, D.M. Genome-Wide Association Studies in Pharmacogenomics: Successes and Lessons. Pharmacogenet. Genomics 2013, 23, 383–394.
- Li, B.; Ritchie, M.D. From GWAS to Gene: Transcriptome-Wide Association Studies and Other Methods to Functionally Understand GWAS Discoveries. Front. Genet. 2021, 12, 713230.
- Smith, D.A.; Sadler, M.C.; Altman, R.B. Promises and Challenges in Pharmacoepigenetics. Camb. Prism. Precis. Med. 2023, 1, E18.
- Cascorbi, I.; Schwab, M. Epigenetics in Drug Response. Clin. Pharmacol. Ther. 2016, 99, 468–470.
- Lauschke, V.M.; Zhou, Y.; Ingelman-Sundberg, M. Novel Genetic and Epigenetic Factors of Importance for Inter-Individual Differences in Drug Disposition, Response and Toxicity. Pharmacol. Ther. 2019, 197, 122–152.
- Kubota, T.; Miyake, K.; Hirasawa, T. Epigenetic understanding of gene-environment interactions in psychiatric disorders: A new concept of clinical genetics. Clin. Epigenetics 2012, 4, 1.
- Tuscher, J.J.; Day, J.J. Multigenerational epigenetic inheritance: One step forward, two generations back. Neurobiol. Dis. 2019, 132, 104591.
- Halbreich, U. Stress-Related Physical and Mental Disorders: A New Paradigm. BJPsych Adv. 2021, 27, 145–152.
- Abdallah, C.G.; Averill, L.A.; Akiki, T.J.; Raza, M.; Averill, C.L.; Gomaa, H.; Adikey, A.; Krystal, J.H. The Neurobiology and Pharmacotherapy of Posttraumatic Stress Disorder. Annu. Rev. Pharmacol. Toxicol. 2019, 59, 171–189.
- Miniawi, S.E.; Orgeta, V.; Stafford, J. Non-Affective Psychotic Disorders and Risk of Dementia: A Systematic Review and Meta-Analysis. Psychol. Med. 2022, 52, 1–13.
- Vafadari, B. Stress and the Role of the Gut-Brain Axis in the Pathogenesis of Schizophrenia: A Literature Review. Int. J. Mol. Sci. 2021, 22, 9747.
- Park, C.; Rosenblat, J.D.; Brietzke, E.; Pan, Z.; Lee, Y.; Cao, B.; Zuckerman, H.; Kalantarova, A.; McIntyre, R.S. Stress, epigenetics and depression: A systematic review. Neurosci. Biobehav. Rev. 2019, 102, 139–152.
- Hack, L.M.; Fries, G.R.; Eyre, H.A.; Bousman, C.A.; Singh, A.B.; Quevedo, J.; John, V.P.; Baune, B.T.; Dunlop, B.W. Moving pharmacoepigenetics tools for depression toward clinical use. J. Affect. Disord. 2019, 249, 336–346.
- Ivanov, H.Y.; Grigorova, D.; Lauschke, V.M.; Velinov, B.; Stoychev, K.; Kyosovska, G.; Shopov, P. CYP2C19 and CYP2D6 Genotypes and Metabolizer Status Distribution in a Bulgarian Psychiatric Cohort. J. Pers. Med. 2022, 12, 1187.
- van Westrhenen, R.; van Schaik, R.H.N.; van Gelder, T.; Birkenhager, T.K.; Bakker, P.R.; Houwink, E.J.F.; Bet, P.M.; Hoogendijk, W.J.G.; van Weelden-Hulshof, M.J.M. Policy and Practice Review: A First Guideline on the Use of Pharmacogenetics in Clinical Psychiatric Practice. Front. Pharmacol. 2021, 12, 640032.
- Berrou, I.; Ramsunder, A.; Palmer, R. Making the case for pharmacogenomics in the management of mental health conditions. Pharm. J. 2023, 310, 7969.
- Taylor, C.; Crosby, I.; Yip, V.; Maguire, P.; Pirmohamed, M.; Turner, R.M. A Review of the Important Role of CYP2D6 in Pharmacogenomics. Genes 2020, 11, 1295.
- Pratt, V.M.; Del Tredici, A.L.; Hachad, H.; Ji, Y.; Kalman, L.V.; Scott, S.A.; Weck, K.E. Recommendations for Clinical CYP2C19 Genotyping Allele Selection: A Report of the Association for Molecular Pathology. J. Mol. Diagn. 2018, 20, 269–276.
- Nassan, M.; Nicholson, W.T.; Elliott, M.A.; Rohrer Vitek, C.R.; Black, J.L.; Frye, M.A. Pharmacokinetic Pharmacogenetic Prescribing Guidelines for Antidepressants: A Template for Psychiatric Precision Medicine. Mayo Clin. Proc. 2016, 91, 897–907.
- Fornaro, M.; Anastasia, A.; Valchera, A.; Carano, A.; Orsolini, L.; Vellante, F.; Rapini, G.; Olivieri, L.; Di Natale, S.; Perna, G.; et al. The FDA “Black Box” Warning on Antidepressant Suicide Risk in Young Adults: More Harm than Benefits? Front. Psychiatry 2019, 10, 294.
- Suwała, J.; Machowska, M.; Wiela-Hojeńska, A. Venlafaxine Pharmacogenetics: A Comprehensive Review. Pharmacogenomics 2019, 20, 829–845.
- Chua, E.W.; Foulds, J.; Miller, A.L.; Kennedy, M.A. Novel CYP2D6 and CYP2C19 Variants Identified in a Patient with Adverse Reactions towards Venlafaxine Monotherapy and Dual Therapy with Nortriptyline and Fluoxetine. Pharmacogenet. Genom. 2013, 23, 494–497.
- Ahmed, A.T.; Biernacka, J.M.; Jenkins, G.; Rush, A.J.; Shinozaki, G.; Veldic, M.; Kung, S.; Bobo, W.V.; Hall-Flavin, D.K.; Weinshilboum, R.M.; et al. Pharmacokinetic-Pharmacodynamic Interaction Associated with Venlafaxine-XR Remission in Patients with Major Depressive Disorder with History of Citalopram / Escitalopram Treatment Failure. J. Affect. Disord. 2019, 246, 62–68.
- Bahar, M.A.; Setiawan, D.; Hak, E.; Wilffert, B. Pharmacogenetics of Drug-Drug Interaction and Drug-Drug-Gene Interaction: A Systematic Review on CYP2C9, CYP2C19 and CYP2D6. Pharmacogenomics 2017, 18, 701–739.
- Whirl-Carrillo, M.; Huddart, R.; Gong, L.; Sangkuhl, K.; Thorn, C.F.; Whaley, R.; Klein, T.E. An Evidence-Based Framework for Evaluating Pharmacogenomics Knowledge for Personalized Medicine. Clin. Pharmacol. Ther. 2021, 110, 563–572.
- Brouwer, J.M.J.L.; Nijenhuis, M.; Soree, B.; Guchelaar, H.-J.; Swen, J.J.; van Schaik, R.H.N.; Weide, J.v.d.; Rongen, G.A.P.J.M.; Buunk, A.-M.; de Boer-Veger, N.J.; et al. Dutch Pharmacogenetics Working Group (DPWG) Guideline for the Gene-Drug Interaction between CYP2C19 and CYP2D6 and SSRIs. Eur. J. Hum. Genet. 2022, 30, 1114–1120.
- Jukić, M.M.; Haslemo, T.; Molden, E.; Ingelman-Sundberg, M. Impact of CYP2C19 Genotype on Escitalopram Exposure and Therapeutic Failure: A Retrospective Study Based on 2,087 Patients. Am. J. Psychiatry 2018, 175, 463–470.
- Rahikainen, A.-L.; Vauhkonen, P.; Pett, H.; Palo, J.U.; Haukka, J.; Ojanperä, I.; Niemi, M.; Sajantila, A. Completed Suicides of Citalopram Users-the Role of CYP Genotypes and Adverse Drug Interactions. Int. J. Leg. Med. 2019, 133, 353–363.
- Hicks, J.K.; Bishop, J.R.; Gammal, R.S.; Sangkuhl, K.; Bousman, C.A.; Leeder, J.S.; Llerena, A.; Mueller, D.J.; Ramsey, L.B.; Scott, S.A.; et al. A Call for Clear and Consistent Communications Regarding the Role of Pharmacogenetics in Antidepressant Pharmacotherapy. Clin. Pharmacol. Ther. 2020, 107, 50–52.
- Ricardo-Silgado, M.L.; Singh, S.; Cifuentes, L.; Decker, P.A.; Gonzalez-Izundegui, D.; Moyer, A.M.; Hurtado, M.D.; Camilleri, M.; Bielinski, S.J.; Acosta, A. Association between CYP Metabolizer Phenotypes and Selective Serotonin Reuptake Inhibitors Induced Weight Gain: A Retrospective Cohort Study. BMC Med. 2022, 20, 261.
- Sim, S.C.; Nordin, L.; Andersson, T.M.-L.; Virding, S.; Olsson, M.; Pedersen, N.L.; Ingelman-Sundberg, M. Association between CYP2C19 Polymorphism and Depressive Symptoms. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2010, 153B, 1160–1166.
- Hicks, J.K.; Bishop, J.R.; Sangkuhl, K.; Müller, D.J.; Ji, Y.; Leckband, S.G.; Leeder, J.S.; Graham, R.L.; Chiulli, D.L.; LLerena, A.; et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline for CYP2D6 and CYP2C19 Genotypes and Dosing of Selective Serotonin Reuptake Inhibitors. Clin. Pharmacol. Ther. 2015, 98, 127–134.
- Hicks, J.K.; Sangkuhl, K.; Swen, J.J.; Ellingrod, V.L.; Müller, D.J.; Shimoda, K.; Bishop, J.R.; Kharasch, E.D.; Skaar, T.C.; Gaedigk, A.; et al. Clinical Pharmacogenetics Implementation Consortium Guideline (CPIC) for CYP2D6 and CYP2C19 Genotypes and Dosing of Tricyclic Antidepressants: 2016 Update. Clin. Pharmacol. Ther. 2017, 102, 37–44.
- Pratt, V.M.; Cavallari, L.H.; Del Tredici, A.L.; Gaedigk, A.; Hachad, H.; Ji, Y.; Kalman, L.V.; Ly, R.C.; Moyer, A.M.; Scott, S.A.; et al. Recommendations for Clinical CYP2D6 Genotyping Allele Selection: A Joint Consensus Recommendation of the Association for Molecular Pathology, College of American Pathologists, Dutch Pharmacogenetics Working Group of the Royal Dutch Pharmacists Association, and the European Society for Pharmacogenomics and Personalized Therapy. J. Mol. Diagn. 2021, 23, 1047–1064.
- LLerena, A.; Berecz, R.; Dorado, P.; González, A.P.; Peñas-LLedó, E.M.; De La Rubia, A. CYP2C9 Gene and Susceptibility to Major Depressive Disorder. Pharmacogenomics J. 2003, 3, 300–302.
- Yenilmez, E.D.; Tamam, L.; Karaytug, O.; Tuli, A. Characterization CYP1A2, CYP2C9, CYP2C19 and CYP2D6 Polymorphisms Using HRMA in Psychiatry Patients with Schizophrenia and Bipolar Disease for Personalized Medicine. Comb. Chem. High Throughput Screen. 2018, 21, 374–380.
- Cacabelos, R.; Torrellas, C. Pharmacogenomics of Antidepressants. HSOA J. Psychiatry Depress. Anxiety 2015, 1, 001.
- Saiz-Rodríguez, M.; Ochoa, D.; Belmonte, C.; Román, M.; Vieira de Lara, D.; Zubiaur, P.; Koller, D.; Mejía, G.; Abad-Santos, F. Polymorphisms in CYP1A2, CYP2C9 and ABCB1 Affect Agomelatine Pharmacokinetics. J. Psychopharmacol. 2019, 33, 522–531.
- Song, L.; Du, Q.; Jiang, X.; Wang, L. Effect of CYP1A2 Polymorphism on the Pharmacokinetics of Agomelatine in Chinese Healthy Male Volunteers. J. Clin. Pharm. Ther. 2014, 39, 204–209.
- Kuo, H.-W.; Liu, S.C.; Tsou, H.-H.; Liu, S.-W.; Lin, K.-M.; Lu, S.-C.; Hsiao, M.-C.; Hsiao, C.-F.; Liu, C.-Y.; Chen, C.-H.; et al. CYP1A2 Genetic Polymorphisms Are Associated with Early Antidepressant Escitalopram Metabolism and Adverse Reactions. Pharmacogenomics 2013, 14, 1191–1201.
- Lin, K.-M.; Tsou, H.-H.; Tsai, I.-J.; Hsiao, M.-C.; Hsiao, C.-F.; Liu, C.-Y.; Shen, W.W.; Tang, H.-S.; Fang, C.-K.; Wu, C.-S.; et al. CYP1A2 Genetic Polymorphisms Are Associated with Treatment Response to the Antidepressant Paroxetine. Pharmacogenomics 2010, 11, 1535–1543.
- Zhu, Y.; Zhang, N.; Ren, D.; Bi, Y.; Xu, F.; Niu, W.; Sun, Q.; Guo, Z.; Yuan, R.; Yuan, F.; et al. CYP1A2 Genetic Polymorphism Is Associated with Treatment Remission to Antidepressant Venlafaxine in Han Chinese Population. Clin. Neuropharmacol. 2019, 42, 32–36.
- Lazarowski, A.; Czornyj, L. Potential Role of Multidrug Resistant Proteins in Refractory Epilepsy and Antiepileptic Drugs Interactions. Drug Metabol. Drug Interact. 2011, 26, 21–26.
- Magarbeh, L.; Hassel, C.; Choi, M.; Islam, F.; Marshe, V.S.; Zai, C.C.; Zuberi, R.; Gammal, R.S.; Men, X.; Scherf-Clavel, M.; et al. ABCB1 Gene Variants and Antidepressant Treatment Outcomes: A Systematic Review and Meta-Analysis Including Results from the CAN-BIND-1 Study. Clin. Pharmacol. Ther. 2023, 6, 2854.
- Sarginson, J.E.; Lazzeroni, L.C.; Ryan, H.S.; Ershoff, B.D.; Schatzberg, A.F.; Murphy, G.M., Jr. ABCB1 (MDR1) Polymorphisms and Antidepressant Response in Geriatric Depression. Pharmacogenet. Genom. 2010, 20, 467–475.
- Yamasaki, Y.; Moriwaki, T.; Ogata, S.; Ito, S.; Ohtsuki, S.; Minegishi, G.; Abe, S.; Ohta, Y.; Kazuki, K.; Kobayashi, K.; et al. Influence of MDR1 Gene Polymorphism (2677G>T) on Expression and Function of P-Glycoprotein at the Blood-Brain Barrier: Utilizing Novel P-Glycoprotein Humanized Mice with Mutation. Pharmacogenet. Genom. 2022, 32, 288–292.
- Shan, X.-X.; Qiu, Y.; Xie, W.-W.; Wu, R.-R.; Yu, Y.; Wu, H.-S.; Li, L.-H. ABCB1 Gene Is Associated with Clinical Response to SNRIs in a Local Chinese Han Population. Front. Pharmacol. 2019, 10, 761.
- Nakamura, K.; Hasegawa, H. Developmental Role of Tryptophan Hydroxylase in the Nervous System. Mol. Neurobiol. 2007, 35, 45–54.
- Crisafulli, C.; Fabbri, C.; Porcelli, S.; Drago, A.; Spina, E.; De Ronchi, D.; Serretti, A. Pharmacogenetics of Antidepressants. Front. Pharmacol. 2011, 2, 6.
- Kato, M.; Serretti, A. Review and Meta-Analysis of Antidepressant Pharmacogenetic Findings in Major Depressive Disorder. Mol. Psychiatry 2010, 15, 473–500.
- Moskaleva, P.V.; Shnayder, N.A.; Dmitrenko, D.V.; Shilkina, O.S.; Neznanov, N.G.; Nasyrova, R.F. Association of TPH1 and TPH2 Gene Polymorphisms with the Risk of Developing Psychoneurological Disorders. Neurosci. Behav. Physiol. 2022, 52, 462–469.
- Matthes, S.; Mosienko, V.; Bashammakh, S.; Alenina, N.; Bader, M. Tryptophan Hydroxylase as Novel Target for the Treatment of Depressive Disorders. Pharmacology 2010, 85, 95–109.
- Du, J.; Zhang, Z.; Li, W.; He, L.; Xu, J.; Shi, Y. Association Study of the TPH2 Gene with Major Depressive Disorder in the Han Chinese Population. Eur. J. Psychiatry 2016, 30, 131–140.
- Secher, A.; Bukh, J.; Bock, C.; Koefoed, P.; Rasmussen, H.B.; Werge, T.; Kessing, L.V.; Mellerup, E. Antidepressive-Drug-Induced Bodyweight Gain Is Associated with Polymorphisms in Genes Coding for COMT and TPH1. Int. Clin. Psychopharmacol. 2009, 24, 199–203.
- Yeung, A.W.K.; Georgieva, M.G.; Atanasov, A.G.; Tzvetkov, N.T. Monoamine Oxidases (MAOs) as Privileged Molecular Targets in Neuroscience: Research Literature Analysis. Front. Mol. Neurosci. 2019, 12, 143.
- Jones, D.N.; Raghanti, M.A. The Role of Monoamine Oxidase Enzymes in the Pathophysiology of Neurological Disorders. J. Chem. Neuroanat. 2021, 114, 101957.
- Bi, Y.; Ren, D.; Guo, Z.; Ma, G.; Xu, F.; Chen, Z.; An, L.; Zhang, N.; Ji, L.; Yuan, F.; et al. Influence and Interaction of Genetic, Cognitive, Neuroendocrine and Personalistic Markers to Antidepressant Response in Chinese Patients with Major Depression. Prog. Neuropsychopharmacol. Biol. Psychiatry 2021, 104, 110036.
- Yoshida, K.; Naito, S.; Takahashi, H.; Sato, K.; Ito, K.; Kamata, M.; Higuchi, H.; Shimizu, T.; Itoh, K.; Inoue, K.; et al. Monoamine Oxidase: A Gene Polymorphism, Tryptophan Hydroxylase Gene Polymorphism and Antidepressant Response to Fluvoxamine in Japanese Patients with Major Depressive Disorder. Prog. Neuropsychopharmacol. Biol. Psychiatry 2002, 26, 1279–1283.
- Clark, S.L.; Adkins, D.E.; Aberg, K.; Hettema, J.M.; McClay, J.L.; Souza, R.P.; van den Oord, E.J.C.G. Pharmacogenomic Study of Side-Effects for Antidepressant Treatment Options in STAR*D. Psychol. Med. 2012, 42, 1151–1162.
- Chappell, K.; Colle, R.; Bouligand, J.; Trabado, S.; Fève, B.; Becquemont, L.; Corruble, E.; Verstuyft, C. The MAOA Rs979605 Genetic Polymorphism Is Differentially Associated with Clinical Improvement Following Antidepressant Treatment between Male and Female Depressed Patients. Int. J. Mol. Sci. 2022, 24, 497.
- Srivastava, K.; Ochuba, O.; Sandhu, J.K.; Alkayyali, T.; Ruo, S.W.; Waqar, A.; Jain, A.; Joseph, C.; Poudel, S. Effect of Catechol-O-Methyltransferase Genotype Polymorphism on Neurological and Psychiatric Disorders: Progressing towards Personalized Medicine. Cureus 2021, 13, e18311.
- Hall, K.T.; Loscalzo, J.; Kaptchuk, T.J. Systems Pharmacogenomics—Gene, Disease, Drug and Placebo Interactions: A Case Study in COMT. Pharmacogenomics 2019, 20, 529–551.
- Benedetti, F.; Dallaspezia, S.; Colombo, C.; Lorenzi, C.; Pirovano, A.; Smeraldi, E. Effect of Catechol-O-Methyltransferase Val(108/158)Met Polymorphism on Antidepressant Efficacy of Fluvoxamine. Eur. Psychiatry 2010, 25, 476–478.
- He, Q.; Shen, Z.; Ren, L.; Wang, X.; Qian, M.; Zhu, J.; Shen, X. The Association of Catechol-O-Methyltransferase (COMT) Rs4680 Polymorphisms and Generalized Anxiety Disorder in the Chinese Han Population. Int. J. Clin. Exp. Pathol. 2020, 13, 1712–1719.
- Gonda, X.; Petschner, P.; Eszlari, N.; Baksa, D.; Edes, A.; Antal, P.; Juhasz, G.; Bagdy, G. Genetic Variants in Major Depressive Disorder: From Pathophysiology to Therapy. Pharmacol. Ther. 2019, 194, 22–43.
- Yang, D.; Gouaux, E. Illumination of Serotonin Transporter Mechanism and Role of the Allosteric Site. Sci. Adv. 2021, 7, eabl3857.
- Krout, D.; Rodriquez, M.; Brose, S.A.; Golovko, M.Y.; Henry, L.K.; Thompson, B.J. Inhibition of the Serotonin Transporter Is Altered by Metabolites of Selective Serotonin and Norepinephrine Reuptake Inhibitors and Represents a Caution to Acute or Chronic Treatment Paradigms. ACS Chem. Neurosci. 2017, 8, 1011–1018.
- Uher, R.; Perroud, N.; Ng, M.Y.M.; Hauser, J.; Henigsberg, N.; Maier, W.; Mors, O.; Placentino, A.; Rietschel, M.; Souery, D.; et al. Genome-Wide Pharmacogenetics of Antidepressant Response in the GENDEP Project. Am. J. Psychiatry 2010, 167, 555–564.
- Stein, K.; Maruf, A.A.; Müller, D.J.; Bishop, J.R.; Bousman, C.A. Serotonin Transporter Genetic Variation and Antidepressant Response and Tolerability: A Systematic Review and Meta-Analysis. J. Pers. Med. 2021, 11, 1334.
- Mrazek, D.A.; Rush, A.J.; Biernacka, J.M.; O’Kane, D.J.; Cunningham, J.M.; Wieben, E.D.; Schaid, D.J.; Drews, M.S.; Courson, V.L.; Snyder, K.A.; et al. SLC6A4 Variation and Citalopram Response. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2009, 150B, 341–351.
- Huezo-Diaz, P.; Uher, R.; Smith, R.; Rietschel, M.; Henigsberg, N.; Marušič, A.; Mors, O.; Maier, W.; Hauser, J.; Souery, D.; et al. Moderation of Antidepressant Response by the Serotonin Transporter Gene. Br. J. Psychiatry 2009, 195, 30–38.
- Porcelli, S.; Fabbri, C.; Serretti, A. Meta-Analysis of Serotonin Transporter Gene Promoter Polymorphism (5-HTTLPR) Association with Antidepressant Efficacy. Eur. Neuropsychopharmacol. 2012, 22, 239–258.
- Zou, Z.; Huang, Y.; Wang, J.; Min, W.; Zhou, B. The Association between Serotonin-Related Gene Polymorphisms and Susceptibility and Early Sertraline Response in Patients with Panic Disorder. BMC Psychiatry 2020, 20, 388.
- Stäuble, C.K.; Meier, R.; Lampert, M.L.; Mikoteit, T.; Hatzinger, M.; Allemann, S.S.; Hersberger, K.E.; Meyer Zu Schwabedissen, H.E. Case Report: Non-Response to Fluoxetine in a Homozygous 5-HTTLPR S-Allele Carrier of the Serotonin Transporter Gene. Front. Psychiatry 2022, 13, 942268.
- Maron, E.; Tammiste, A.; Kallassalu, K.; Eller, T.; Vasar, V.; Nutt, D.J.; Metspalu, A. Serotonin Transporter Promoter Region Polymorphisms Do Not Influence Treatment Response to Escitalopram in Patients with Major Depression. Eur. Neuropsychopharmacol. 2009, 19, 451–456.
- Kato, M.; Fukuda, T.; Wakeno, M.; Fukuda, K.; Okugawa, G.; Ikenaga, Y.; Yamashita, M.; Takekita, Y.; Nobuhara, K.; Azuma, J.; et al. Effects of the Serotonin Type 2A, 3A and 3B Receptor and the Serotonin Transporter Genes on Paroxetine and Fluvoxamine Efficacy and Adverse Drug Reactions in Depressed Japanese Patients. Neuropsychobiology 2006, 53, 186–195.
- Zhu, J.; Klein-Fedyshin, M.; Stevenson, J.M. Serotonin Transporter Gene Polymorphisms and Selective Serotonin Reuptake Inhibitor Tolerability: Review of Pharmacogenetic Evidence. Pharmacotherapy 2017, 37, 1089–1104.
- Sarmiento-Hernández, E.I.; Ulloa-Flores, R.E.; Camarena-Medellín, B.; Sanabrais-Jiménez, M.A.; Aguilar-García, A.; Hernández-Muñoz, S. Association between 5-HTTLPR Polymorphism, Suicide Attempt and Comorbidity in Mexican Adolescents with Major Depressive Disorder. Actas Esp. Psiquiatr. 2019, 47, 1–6.
- Gonda, X.; Fountoulakis, K.N.; Harro, J.; Pompili, M.; Akiskal, H.S.; Bagdy, G.; Rihmer, Z. The Possible Contributory Role of the S Allele of 5-HTTLPR in the Emergence of Suicidality. J. Psychopharmacol. 2011, 25, 857–866.
- Antypa, N.; Serretti, A.; Rujescu, D. Serotonergic Genes and Suicide: A Systematic Review. Eur. Neuropsychopharmacol. 2013, 23, 1125–1142.
- Nemoda, Z.; Angyal, N.; Tarnok, Z.; Birkas, E.; Bognar, E.; Sasvari-Szekely, M.; Gervai, J.; Lakatos, K. Differential Genetic Effect of the Norepinephrine Transporter Promoter Polymorphisms on Attention Problems in Clinical and Non-Clinical Samples. Front. Neurosci. 2018, 12, 1051.
- Ochi, T.; Vyalova, N.M.; Losenkov, I.S.; Paderina, D.Z.; Pozhidaev, I.V.; Loonen, A.J.M.; Simutkin, G.G.; Bokhan, N.A.; Wilffert, B.; Ivanova, S.A. Polymorphisms in the Adrenergic Neurotransmission Pathway Impact Antidepressant Response in Depressed Patients. Neurosci. Appl. 2023, 2, 101016.
- Porcelli, S.; Drago, A.; Fabbri, C.; Gibiino, S.; Calati, R.; Serretti, A. Pharmacogenetics of Antidepressant Response. J. Psychiatry Neurosci. 2011, 36, 87–113.
- Zhou, Z.; Zhen, J.; Karpowich, N.K.; Goetz, R.M.; Law, C.J.; Reith, M.E.A.; Wang, D.-N. LeuT-Desipramine Structure Reveals How Antidepressants Block Neurotransmitter Reuptake. Science 2007, 317, 1390–1393.
- Lavretsky, H.; Siddarth, P.; Kumar, A.; Reynolds, C.F., 3rd. The Effects of the Dopamine and Serotonin Transporter Polymorphisms on Clinical Features and Treatment Response in Geriatric Depression: A Pilot Study. Int. J. Geriatr. Psychiatry 2008, 23, 55–59.
- Kirchheiner, J.; Nickchen, K.; Sasse, J.; Bauer, M.; Roots, I.; Brockmöller, J. A 40-Basepair VNTR Polymorphism in the Dopamine Transporter (DAT1) Gene and the Rapid Response to Antidepressant Treatment. Pharmacogenomics J. 2007, 7, 48–55.
More
Information
Subjects:
Pharmacology & Pharmacy
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
660
Revisions:
2 times
(View History)
Update Date:
29 May 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





