Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Jan van Turnhout | -- | 1940 | 2023-05-08 12:47:51 | | | |
| 2 | Sirius Huang | Meta information modification | 1940 | 2023-05-09 09:45:44 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Mantri, D.; Wymenga, L.; Van Turnhout, J.; Van Zeijl, H.; Zhang, G. SARS-CoV-2 Virus Manipulation Using Electric Fields. Encyclopedia. Available online: https://encyclopedia.pub/entry/43978 (accessed on 02 April 2026).
Mantri D, Wymenga L, Van Turnhout J, Van Zeijl H, Zhang G. SARS-CoV-2 Virus Manipulation Using Electric Fields. Encyclopedia. Available at: https://encyclopedia.pub/entry/43978. Accessed April 02, 2026.
Mantri, Devashish, Luutzen Wymenga, Jan Van Turnhout, Henk Van Zeijl, Guoqi Zhang. "SARS-CoV-2 Virus Manipulation Using Electric Fields" Encyclopedia, https://encyclopedia.pub/entry/43978 (accessed April 02, 2026).
Mantri, D., Wymenga, L., Van Turnhout, J., Van Zeijl, H., & Zhang, G. (2023, May 08). SARS-CoV-2 Virus Manipulation Using Electric Fields. In Encyclopedia. https://encyclopedia.pub/entry/43978
Mantri, Devashish, et al. "SARS-CoV-2 Virus Manipulation Using Electric Fields." Encyclopedia. Web. 08 May, 2023.
Copy Citation
Studying the mechanisms of virus deactivation could have huge benefits for battling pandemics such as COVID-19. Using electric fields as a tool may allow us to integrate the trapping, sampling and inactivation of viruses on a single microchip platform.
micro-electrodes
virus in-activation
virus sampling
virus concentration
SARS-CoV-2
dielectrophoresis
1. Concentration of Moderately Large Viruses (>200 nm)
A polarizable particle suspended in a liquid medium exposed to a nonuniform electric field experiences a force known as the dielectrophoretic force (DEP). When the medium is more polarizable than the particle, the particle will be repelled by the electrode, resulting in a negative DEP force (nDEP). Whereas when the particle is more polarizable, than the medium, the particle, will be attracted to the electrode, resulting in a positive DEP force (pDEP). Dielectrophoretic manipulation has been used to successfully trap and concentrate bacteria and viruses. Concentration of the viral particulates facilitates the formation of localized clusters that can be easily examined and inactivated. Morgan et al. successfully demonstrated the use of DEP in concentrating viruses [1]. They concentrated Tobacco Mosaic Virus (TMV) (ϕ 280 nm) with a pDEP using saw-tooth electrodes with 2–6 μm pitch spacing. The virus was found to be highly polarizable, which was attributed to the absence of an insulating membrane. The group also showed that TMV could be separated and filtered out from a mixture of TMV and Herpes Simplex Virus (HSV, ϕ 150–240 nm) by exploiting the difference in the Clausius–Mossotti (CM) factor. The CM factor depends on the virus’s composition and structure [2]. Finally, the group explored the prospects of DEP on HSV. The HSV was concentrated by a pDEP force with an electric field (freq. 4.5 MHz, 5 Vpp, E = 106 V/m) [3]. The manipulation of the Vaccinia virus (ϕ 240 nm) is also well documented. A study reported the use of interdigitated Ti/Pt electrodes to trap Vaccinia virus using a pDEP with 7 Vpp and 1 MHz [4].
Recently, the use of nanofibers for trapping viruses was investigated [5]. Although, the nanofibers are particularly good at trapping the virus from a flowing sample, they cannot concentrate the virus. Similar inferences can be gathered from studies that suggests the use of electrostatics to trap the corona virus [6][7]. The virus can only be trapped, but not concentrated. The viruses described above are moderately large DNA viruses with an inner surface area high enough to amass enough charges to facilitate its manipulation. As particle dimensions shrink, the controlled manipulation of particles becomes increasingly difficult as induced dipole moments scale with the third power of the particle’s radius [8]. This could be partly compensated by increasing the electric field strength through electrode geometry optimization and/or increasing the source voltage. However, the latter likely results in the electrolysis of the medium.
2. Concentration of Smaller Viruses
Smaller viruses were also shown to respond to the dielectrophoretic effect. Influenza (ϕ 80 nm) and Hepatitis virus (ϕ 32 nm) were manipulated and concentrated using negative dielectrophoresis (nDEP) on planar electrode arrays. Planar platforms and 3D structures that make use of quadrupole and octupole electrode cages establish points of confluence, where the virus can be trapped effectively [9]. Cowpea Mosaic Virus (CPMV) (30 nm), a spherical virus, was successfully trapped using castellated electrodes with a pitch size of 2 μm [10]. Despite having a small size, CPMV shows a higher polarizability than that of the medium, because of its non-enveloped nature. The same is true for the Adenovirus and Rotavirus, which despite having a small size, were easily trapped and concentrated using a pDEP [11]. The pDEP trapping of viruses was observed in media with higher solution conductivities than that of bacteria. The prime reason for this might be attributed to the smaller size of viruses compared to that of bacteria.
A challenging problem is that with the increase in electrolyte conductivity, the ionic strength increases. This may result in more electrochemical reactions. Usually, virus samples require a relatively high ionic strength for their storage. Another challenge is that smaller viruses (<100 nm) show a random Brownian motion. Brownian motion is characterized by random movements that could hinder the trapping of viruses [11][12]. These problems can be resolved by attaching the virus to a larger molecule, e.g., Hepatitis A virus (27 nm) bonded to a streptavidin-coated particle [13].
Insulator-based DEP (iDEP) is a technique that uses insulating structures to concentrate the electric field. The technique seems to become more common for concentrating smaller viruses as well as larger viruses in recent years. For example, influenza virus (90 nm) [14], Sindbis virus (130 nm) [15], bacteriophages such as T4 and SPN3UP (90 nm) [16] and other larger viruses such as TMV [17] have been concentrated by iDEP. A study using circular and oval-shaped electrodes employed this method and was devised to trap three different strains of the same bacteriophage, demonstrating the high specificity of this technique [16]. However, no study has established a better efficiency of iDEP over DEP in terms of trapping and concentrating the smaller virus particles. Moreover, iDEP devices are more susceptible to Joule heating and electrolysis of the medium [18].
A summary of all the different aspects of the practical studies that concentrated viruses using DEP have been tabulated in Table 1 below.
Table 1. Comparison of the practical studies that concentrated viruses using DEP.
| Name | Size | Type | Capsulation | Trapping | Electrodes |
|---|---|---|---|---|---|
| Tobacco Mosaic | 280 nm | RNA virus | Non-enveloped | pDEP | Sawtooth (6 µm) [1] |
| Herpes simplex | 240 nm | DNA virus | Enveloped | pDEP | Quadrupole (6 µm) [3] |
| Vaccinia | 360 × 270 × 250 nm | DNA virus | Enveloped | pDEP | Interdigitated (10 µm) [4] |
| Influenza | 90 nm | RNA virus | Enveloped | nDEP | Quadrupole, Interdigitated (6 µm), (40 µm) [9] |
| Hepatitis A | 27 nm | RNA virus | Non-enveloped | nDEP/pDEP | Quadrupole Octrupole (2 µm) [13] |
| Cowpea Mosaic | 30 nm | RNA virus | Non-enveloped | pDEP | Castellated (2 µm) [10] |
| Adeno | 90 nm | DNA virus | Non-enveloped | iDEP | Castellated/Interdigitated (10 µm) [11] |
| Sindbis | 130 nm | RNA virus | Enveloped | iDEP | Sawtooth gradient (0–700 V) [15] |
| T4 bacteriophage | 90 nm | DNA virus | Non-enveloped | iDEP | Circular and oval (80 µm) [16] |
3. Trapping of SARS-CoV-2: Theoretical Approach
The spherical SARS-CoV-2 virus is characterized by a lipid insulating membrane, and hence, the particle may not be easily polarizable. Tuning properties of the medium becomes instrumental if we desire to attract and concentrate the virus particles with pDEP. The Clausius–Mossotti factor of SARS-CoV-2 can be modelled by using a core-shell model, as depicted in Figure 1. The inner layer represents the core and the outer layer represents the lipid membrane. The DEP force on a neutral particle can be described by:
where εp* is the permittivity of the particles, εs* that of the suspending solvent, R the radius of the particle, E the applied field, and ∇ the divergence. According to Equation (1), the force can be attractive or repulsive depending on the values for εp* and εs*. The ratio of permittivities in Equation (1) equals the CM-factor, so:
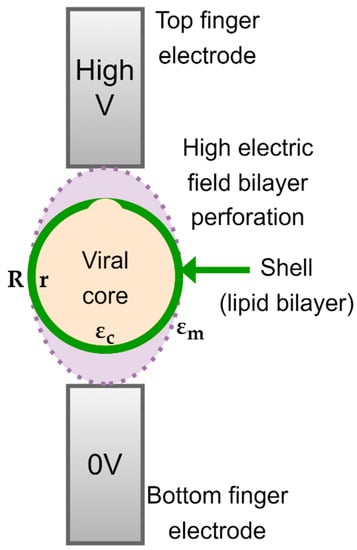
Figure 1. Core-shell model of SARS-CoV-2.
Note that the permittivities are complex quantities, the real and imaginary part of which usually will change with the applied radial frequency ω. As a result, F will depend on ω as well. The fact that εp* and εs* and, thus, CM* are complex is indicated in Equations (1) and (2) by the asterisk in superscript. The symbol εs′ means the real part of εs*. The imaginary part is denoted by εs″. We may thus write:
If εs″ is caused by conduction losses, then ε′′s(ω)=γs/(εoω) where γs is the conductivity of the solvent and εo the permittivity of vacuum εo = 8.854 × 10−12 F/m. In addition to ionic conduction, dipole relaxations may also occur in a dielectric medium. Assume that these are not strong in the present case and so they are neglected. This implies that εs′ will remain constant. make the same assumptions for the complex permittivities of the core and the membrane of the virus, εc* and εm*. This means that:
The change in the CM-factor with frequency f, f = ω/(2π), will evidently derive from the conductivity and permittivity of the solvent, as well as from the conductivity and permittivity of the core and membrane of the particle.
For a layered sphere, the equation for εp* obeys:
which may also be written as:
The core-shell structure of the virus is visualized in Figure 1, in which the permittivity of the RNA-core εc* and that of the membrane proteins εm* are complex, frequency dependent quantities. Both are assumed to obey the expressions given in Equation (4). The radius of the core is r and that of the outer layer is R. When substitute Equation (5) in the CM factor given by Equation (2), we obtain:
As indicated, the CM* given by Equation (7) will depend on the frequency. The same applies to the DEP force given by Equation (1), as have already mentioned above.
It should be realized that εs* from the suspending solvent will differ from the permittivity of the solvent proper. The solvent with virus particles forms a suspension, a mixture. For the calculation of the permittivity of a mixture, many formulae are available. A convenient one of Maxwell-Garnett, based on the mean field approximation, gives an explicit formula for the permittivity of a suspension of spherical particles:
Note that εsp* may also written as:
where v is the volume fraction of the virus particles. It is this εsp* which we should substitute for εs* in Equations (2) and (7). This leads to:
We can compute CM from Equations (5), (8) and (9) with a few lines of code using Matlab, Maple or Mathematica. The plots shown in Figure 2 are obtained with a Mathematica script.
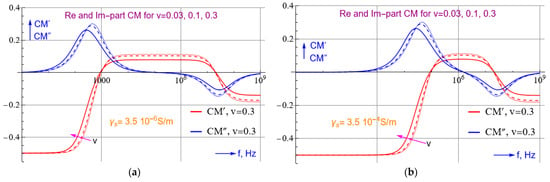
Figure 2. The real and imaginary CM-spectra calculated for two buffer medium conductivities of γs = 3.5 × 10−6 (a) and 3.5 × 10−4 S/m (b) and three volume fractions v of the virus. CM′ becomes positive only in a certain frequency region. This region narrows if the conductivity of the buffer increases. The CM″ spectra can be approximated roughly by dCM′/dlnω. They thus show a peak when the slope of CM′ changes the most. The position of these peaks corresponds roughly to those in the dielectric loss spectra. The spectra given hold for εs′ = 78, εc′ = 70, εm′ = 12, γs = 3.5 × 10−4 S/m, γc = 0.2 S/m, γm = 10−9 S/m, r = 54 nm and R = 60 nm. The modelling was performed with Mathematica.
Selecting a non-aqueous medium, such as that of γs < γp, a pDEP force on the corona virus can be produced at lower frequencies to effectively concentrate the virus. Unfortunately, studies have not delineated the electrical parameters of SARS-CoV-2. Nonetheless, for a KCl solution with a low conductivity, and as long as ε′s>ε′c,ε′s>ε′m, the graph in Figure 2 shows that the virus will be attracted to the electrodes at acceptable frequencies, for which the real part of CM becomes positive. Figure 2 also suggests that with a lower medium conductivity, pDEP can be brought about by lower frequency electric fields. This calculation does not consider the negative charges that are already present on the RNA molecule inside the core of the virus. This assumption is supported by previous studies that show that dielectrophoretic mobility is not affected by the interior charges in Cowpea Chlorotic Mottle virus [19] as well as the corona virus [6]. If ε′s>ε′c and a medium conductivity of <0.005 S/m is maintained, there is a high chance that corona virus can be concentrated with moderately high frequencies and electric fields.
References
- Morgan, H.; Green, N.G. Dielectrophoretic manipulation of rod-shaped viral particles. J. Electrost. 1997, 42, 279–293.
- Morgan, H.; Hughes, M.P.; Green, N.G. Separation of Submicron Bioparticles by Dielectrophoresis. Biophys. J. 1999, 77, 516–525.
- Hughes, M.; Morgan, H.; Rixon, F.J.; Burt, J.P.; Pethig, R. Manipulation of herpes simplex virus type 1 by dielectrophoresis. Biochim. Biophys. Acta 1998, 1425, 119–126.
- Akin, D.; Li, H.; Bashir, R. Real-Time Virus Trapping and Fluorescent Imaging in Microfluidic Devices. Nano Lett. 2004, 4, 257–259.
- Madiyar, F.R.; Syed, L.U.; Culbertson, C.T.; Li, J. Manipulation of bacteriophages with dielectrophoresis on carbon nanofiber nanoelectrode array. Electrophoresis 2013, 34, 1123–1130.
- Javidpour, L.; Božič, A.; Naji, A.; Podgornik, R. Electrostatic interactions between the SARS-CoV-2 virus and a charged electret fibre. Soft Matter 2021, 17, 4296–4303.
- Leung, W.W.F.; Sun, Q. Electrostatic charged nanofiber filter for filtering airborne novel coronavirus (COVID-19) and nano-aerosols. Sep. Purif. Technol. 2020, 250, 116886.
- Lapizco-Encinas, B.H.; Rito-Palomares, M. Dielectrophoresis for the manipulation of nanobioparticles. Electrophoresis 2007, 28, 4521–4538.
- Grom, F.; Kentsch, J.; Müller, T.; Schnelle, T.; Stelzle, M. Accumulation and trapping of hepatitis A virus particles by electrohydrodynamic flow and dielectrophoresis. Electrophoresis 2006, 27, 1386–1393.
- Ermolina, I.; Milner, J.; Morgan, H. Dielectrophoretic investigation of plant virus particles: Cow Pea Mosaic Virus and Tobacco Mosaic Virus. Electrophoresis 2006, 27, 3939–3948.
- Nakano, M.; Ding, Z.; Suehiro, J. Dielectrophoresis and dielectrophoretic impedance detection of adenovirus and rotavirus. Jpn. J. Appl. Phys. 2016, 55, 017001.
- Al Ahmad, M.; Mustafa, F.; Ali, L.M.; Karakkat, J.V.; Rizvi, T.A. Label-Free Capacitance-Based Identification of Viruses. Sci. Rep. 2015, 5, 9809.
- Kentsch, J.; Dürr, M.; Schnelle, T.; Gradl, G.; Müller, T.; Jager, M.; Normann, A.; Stelzle, M. Microdevices for separation, accumulation, and analysis of biological micro- and nanoparticles. IEE Proc.-Nanobiotechnol. 2003, 150, 82–89.
- Masuda, T.; Maruyama, H.; Honda, A.; Arai, F. Virus Enrichment for Single Virus Infection by Using 3D Insulator Based Dielectrophoresis. PLoS ONE 2014, 9, e94083.
- Ding, J.; Lawrence, R.M.; Jones, P.V.; Hogue, B.G.; Hayes, M.A. Concentration of Sindbis virus with optimized gradient insulator-based dielectrophoresis. Analyst 2016, 141, 1997–2008.
- De Peña, A.C.; Redzuan, N.H.M.; Abajorga, M.K.; Hill, N.; Thomas, J.A.; Lapizco-Encinas, B.H. Analysis of Bacteriophages with Insulator-Based Dielectrophoresis. Micromachines 2019, 10, 450.
- Lapizco-Encinas, B.; Davalos, R.V.; Simmons, B.; Cummings, E.B.; Fintschenko, Y. An insulator-based (electrodeless) dielectrophoretic concentrator for microbes in water. J. Microbiol. Methods 2005, 62, 317–326.
- Gallo-Villanueva, R.C.; Perez-Gonzalez, V.H.; Cardenas-Benitez, B.; Jind, B.; Martinez-Chapa, S.O.; Lapizco-Encinas, B.H. Joule heating effects in optimized insulator-based dielectrophoretic devices: An interplay between post geometry and temperature rise. Electrophoresis 2019, 40, 1408–1416.
- Johnson, M.W.; Wagner, G.W.; Bancroft, J.B. A Titrimetric and Electrophoretic Study of Cowpea Chlorotic Mottle Virus and its Protein. J. Gen. Virol. 1973, 19, 263–273.
More
Information
Subjects:
Biotechnology & Applied Microbiology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
520
Revisions:
2 times
(View History)
Update Date:
09 May 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





