Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Makoto Sahara | -- | 3945 | 2023-04-07 17:13:27 | | | |
| 2 | Catherine Yang | Meta information modification | 3945 | 2023-04-10 03:08:50 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Sahara, M. Recent Advances in Cardiac Organoids. Encyclopedia. Available online: https://encyclopedia.pub/entry/42876 (accessed on 29 March 2026).
Sahara M. Recent Advances in Cardiac Organoids. Encyclopedia. Available at: https://encyclopedia.pub/entry/42876. Accessed March 29, 2026.
Sahara, Makoto. "Recent Advances in Cardiac Organoids" Encyclopedia, https://encyclopedia.pub/entry/42876 (accessed March 29, 2026).
Sahara, M. (2023, April 07). Recent Advances in Cardiac Organoids. In Encyclopedia. https://encyclopedia.pub/entry/42876
Sahara, Makoto. "Recent Advances in Cardiac Organoids." Encyclopedia. Web. 07 April, 2023.
Copy Citation
Cardiac organoids are in vitro self-organizing and three-dimensional structures composed of multiple cardiac cells (i.e., cardiomyocytes, endothelial cells, cardiac fibroblasts, etc.) with or without biological scaffolds. Since cardiac organoids recapitulate structural and functional characteristics of the native heart to a higher degree compared to the conventional two-dimensional culture systems, their applications, in combination with pluripotent stem cell technologies, are being widely expanded for the investigation of cardiogenesis, cardiac disease modeling, drug screening and development, and regenerative medicine.
cardiac organoid
cardiac tissue engineering
cardiogenesis
regenerative medicine
biotechnology
1. Various Types of Cardiac Organoids
The approaches for generation of in vitro cardiac organoids are classified into scaffold-based and scaffold-free methods. While biomaterials such as (natural or synthetic) hydrogels or decellularized bioscaffolds are used in the former, the approach inducing spherical aggregation of cultured cells on an anti-adhesive environment are usually adopted in the latter [1][2]. The cell types used in the processes for constructing cardiac organoids are categorized into PSC aggregates such as embryoid bodies (EBs) and differentiated cardiac cells such as CMs, ECs, and CFBs, which are derived from either PSC differentiation or primary cells.
1.1. Cardiac Organoids in an Early Era
In 2017, utilizing the EHT technologies, the early-type cardiac organoids have been reported (Table 1) [3][4]. Mills et al. developed a 96-well high-throughput device, called the Heart Dynamometer, each well of which has a culture insert composed of an elliptical cell seeding area and two elastomeric posts [3]. Using human PSC (hPSC)-derived cardiac cells (~70% hPSC-CMs with the rest stromal cells on day 15 in Wnt-modulated hPSC-CM differentiation) embedded with ECM (collagen I and Matrigel) on this device, the team generated human cardiac organoids, and successfully conducted both a functional screening of the optimal metabolic and mechanical loading conditions for CM function and maturation [3]. They also conducted a drug screening of 105 small molecules for identifying pro-proliferative compounds under this system [5]. Similarly, Voges et al. constructed human circular cardiac organoids via fabrication of human ESC (hESC)-derived cardiac cells in the mold with ECM (collagen I), and developed a cardiac disease modeling platform such as an injury model [4]. Interestingly, following cryoinjury, the cardiac organoids exhibited an endogenous regenerative response with fully functional recovery 2 weeks after injury, indicating the previously unrecognized regenerative capabilities of immature human heart tissues.
Table 1. Overview of cardiac organoid models in an early era (2017–2020).
| Authors & Years | CO Models/Platform | Sp. | Cells | Scaffold | Chemicals | Features | Chambers/Cavities | Applications | Refs |
|---|---|---|---|---|---|---|---|---|---|
| Mills et al., 2017 Mills et al., 2019 |
Heart Dynamometer-engineered CO | H | hPSC-derived cardiac cells | Collagen I Matrigel |
CHIR, BMP4, Activin A, FGF2, IWP4 | - A 96-well device for high-throughput functional screening of hiPSC-derived CO to facilitate testing for maturation conditions - Identification of two pro-proliferative molecules without side effects on cardiac function, acting via the Mevalonate pathway |
(−) | Maturation & drug screening (pro-proliferation) |
[3][5] |
| Voges et al., 2017 | Circular CO engineered in the mold | H | hPSC-derived cardiac cells | Collagen I | CHIR, BMP4, Activin A, FGF2, IWP4 | - Human CO exhibiting an endogenous and full regenerative response 2 weeks after acute injury | (−) | Disease modeling (cryoinjury) | [4] |
| Hoang et al., 2018 Hoang et al., 2021 |
Spacially-patterned CO | H | hPSCs | PDMS stencils with aligned holes | CHIR, IWP4 | - Biomaterial-based cell patterning combined with stem cell organoid engineering - Optimization of CO geometries for efficient CO production, reflecting high consistency and large morphology - Quantification of the embryotoxic potential of 9 pharmaceutical compounds |
(−) | Cardiogenesis modeling & drug screening (embryo toxicity) |
[6][7] |
| Lee et al., 2020 | Murine CO | M | mESC-EB | Laminin Entactin |
FGF4, BIO, BMP4 | - Innovative approach to generate CO with chamber formation (i.e., both atria- and ventricle-like parts) from mESC EBs via FGF4 and ECM | (+) | Cardiogenesis modeling | [8] |
| Richards et al., 2017 Richards et al., 2020 |
Multicellular CO | H | hiPSC-CM, hCFB, HUVEC, hADSC | Agarose | Not applicable * | - Generation of human CO that resembled the lumenized vascular network in the developing heart - Modeling of human heart structure after MI by oxygen diffusion gradient and NA stimulation |
(−) | Cardiogenesis & disease modeling (ischemia) |
[9][10] |
| Buono et al., 2020 | Multicellular CO | H | hiPSC-CM, HCMEC, hCFB | CHIR, C59 | - CO generation with a tri-culture approach - Clear differences in structures and beating behavior in between HCM Pt. (MYH7-mutant) hiPSC-derived and control COs |
(−) | Disease modeling (HCM) | [11] |
CO, cardiac organoid; EB, embryoid body; ECM, extracellular matrix; H, human; hADSC, human adipose-derived stem cell; hCFB, human cardiac fibroblast; HCM, hypertrophic cardiomyopathy; HCMEC, human cardiac microvascular endothelial cell; hiPSC, human induced pluripotent stem cell; hPSC, human pluripotent stem cell; HUVEC, human umbilical vein endothelial cell; M, mouse; mESC, mouse embryonic stem cell; MI, myocardial infarction; NA, noradrenaline; Pt., patient; Sp, species. In the “Chambers/Cavities” column, “(+)” means that the CO generates clear heart chambers and cavities in the structure, while “(−)” means that the CO does not exhibit them. * This CO model uses the primary cells.
One of the critical issues in organoid formation is the heterogeneity of the generated organoids in regards to the random positioning of specific cell types and the organoids’ size. To tackle this issue, Hoang et al. introduced biomaterial-based spatial micropatterning in hPSC organoid engineering [6]. Geometric confinement was given to human iPSCs (hiPSCs) with a poly(ethylene glycol)-based micropatterned substrate, created by oxygen plasma etching and a polydimethylsiloxane stencil with holes aligned in a grid. In the result, the team generated spatially organized early-developing cardiac organoids with contracting CMs in the center, surrounded by stromal cells and with a certain size. Further, using the optimized conditions for organoid production such as micropatterning in 600 μm diameter circles, Hoang et al. conducted a drug screening to detect developmental cardiotoxicity and quantified the embryotoxic potential of nine pharmaceutical compounds [7].
In many cases, cardiac organoids are engineered from PSC aggregates, i.e., EBs. By combining mouse ESC-derived EBs with the optimized ECM environment, composed of the laminin–entactin complex supplemented with fibroblast growth factor 4, Lee et al. developed functional murine heart organoids that possessed atrium- and ventricle-like chambers, which were similar to those of in vivo embryonic hearts [8]. It will be interesting and important to confirm whether this promising system can be also applied to a human PSC-derived cardiac organoid model. In contrast, in some cases, (terminally) differentiated cardiac cells (i.e., CMs, ECs, CFBs, etc.) are used to form cardiac organoids as multicellular strategies [9][10][11]. Using defined cell types and ratios, i.e., 50% hiPSC-CMs and 50% non-myocytes (at a 4:2:1 ratio of human CFBs, human umbilical vein ECs [HUVECs], and human adipose-derived stem cells, all of which were derived from the primary cells) in non-adhesive agarose hydrogel molds, Richards et al. developed human cardiac organoids that structurally and functionally resembled a lumenized vascular network in the embryonic heart [9]. Importantly, the team modeled the structure of the human myocardium after myocardial infarction (MI) by providing these organoids with an oxygen-diffusion gradient (10% oxygen) and stimulation with noradrenaline, both of which induced hallmarks of MI, including fibrosis, pathological metabolic shifts, and impaired calcium handling at the transcriptomic, structural and functional levels [10]. Similarly, Buono et al. generated human cardiac organoids with a triculture approach using hiPSC-CMs (from healthy subjects and hypertrophic cardiomyopathy patients carrying the MYH7 gene mutation), human cardiac microvascular ECs and human CFBs at a 3:5:2 ratio [11]. The team successfully showed clear differences in structural and electrophysiological properties between healthy and cardiomyopathic organoids, the latter of which exhibited an arrhythmic phenotype. Overall, these multicellular strategies-based cardiac organoids were proven to be a useful tool for modeling a genetic or non-genetic cardiac disease.
1.2. Latest Cardiac Organoids since 2021
Since 2021, an increased number of developed self-organizing cardiac organoids with more advanced structures and functions have been reported (Table 2). Rossi and colleagues captured early heart organogenesis with an in vivo-like spatiotemporal repeatability using murine embryonic organoids, termed “gastruloids”, which were formed by mouse ESC-derived EBs [12]. Using axially patterned murine gastruloids treated with a Wnt activator (CHIR99021), and cardiogenic factors such as basic fibroblast growth factor (bFGF), vascular endothelial growth factor-A (VEGF-A), and ascorbic acid in ultra-low attachment microplates under shaking, the team successfully modeled the earliest stages of heart development, such as the formation of an in vivo cardiac crescent-like structure composed of specified first heart field (FHF) and second heart field (SHF) progenitor cells. Further, the cardiac crescent-like structure developed into an early cardiac tube-like beating tissue that was adjacent to a co-developed primitive gut-like structure, mutually separated by a networked endocardial-like layer. Although these cardiac structures did not further follow the in vivo developing stages such as looping and the formation of four chambers, their system sheds light on key aspects in organogenesis through the coordinated development of multiple tissues and organs [12]. Similarly, Silva et al. generated cooperative cardiac and gut tissues originating from different germ lineages within a single organoid model, formed by hiPSC-derived mesendoderm progenitors’ spheroids [13]. These multilineage organoids showed distinct structural features and extensive tissue growth of the developing human heart and gut, containing epithelial endoderm, complexed and segregated CM subtypes (e.g., MLC2v+ ventricular CMs), and a TNNT2−TBX18+ epicardial layer. With this unique system, the team demonstrated that co-emergence of the two defined tissues and their mutual crosstalk promoted physiological maturation of cardiac tissue, especially of atrial/nodal CMs, although the cardiac parts of these organoids were not morphologically similar to the structure of the in vivo developing heart [13].
Biological scaffolds offer a better environment to cells in in vitro 3D culture. Drakhlis and colleagues generated highly structured heart-forming organoids (HFOs) by embedding hPSC aggregates in Matrigel, followed by directed CM differentiation simply via biphasic Wnt signaling modulation [14]. After 10 days in differentiation, the HFOs exhibited self-assembly with distinct layers, consisting of an inner layer containing endodermal foregut-like and endothelial cells, an endocardial cell layer at the interface between the inner and middle layers, a middle layer containing mostly CMs with epicardial cells, and an outer layer containing mesenchymal and liver cells. All of these structures resembled aspects of early native heart development, supported by a crosstalk with foregut endoderm development. Importantly, the team applied their organoid model to investigate a cardiac genetic disease, and demonstrated that NKX2-5-knockout (KO) hESC-derived HFOs showed decreased CM adhesion and hypertrophy with reduced tissue compaction, which were reminiscent of cardiac malformations previously observed in NKX2-5-KO mice [15], highlighting the utility of the HFO system for the in vitro modeling of gene KO phenotypes [14]. One of the main issues to be addressed in cardiac organoid formation is that most of the existing models do not recapitulate in vivo self-organizing cardiac architecture, such as four cardiac chambers with formation of inner endocardial cavities. To tackle this point, Hofbauer et al. created cavity-forming cardiac organoids, termed “cardioids”, using hPSC aggregates sequentially treated with a multitude of signaling modifiers and cardiogenic factors, including CHIR99021, bone morphogenic protein 4 (BMP4), bFGF, Acitivin A, LY294002 (a PI3K inhibitor), IWP2 (a Wnt inhibitor), insulin, VEGF-A, and retinoic acid (RA) in ultra-low-attachment 96-well plates pre-coated with vitronectin or laminin [16]. While the inner lining of endothelial/endocardial cells in a formed cardiac chamber-like structure has not been observed in previous cardiac organoid models, the team revealed that cavity morphogenesis and cardiac specification was differentially controlled by a mesodermal Wnt-BMP signaling axis, respectively, and that the cavity formation required a Wnt-BMP signaling’s downstream transcription factor HAND1. Cardioids contained three major cardiac cell types, i.e., CMs, endocardial cells and epicardial cells, resembling self-organizing principles of human cardiogenesis such as chamber-like morphogenesis. Further, upon cryoinjury, cardioids exhibited ECM accumulation, an early hallmark of both heart regenerative and pathological responses, together with rapid recruitment of ECs and epicardial fibroblast-like cells. This implies that human cardioids represent an excellent platform for cardiac disease modeling and future translational research [16]. Another study by Lewis-Israeli et al. also reported interesting cavity-forming cardiac organoids [17]. The team adopted a three-step Wnt signaling modulation strategy (activation/inhibition/re-activation) at specific time points on suspension hPSC-EBs for the induction of cardiac mesoderm and epicardial cells. In combination with cardiogenic growth factors such as BMP4 and Activin A, their relatively simple approach generated self-assembling human cardiac organoids that contained internal chambers with multi-lineage cardiac cell types including CMs, epicardial cells/CFBs, and endothelial/endocardial cells. Their cardiac organoids recapitulated heart field formation, developed a vasculature, and displayed well-organized sarcomeres in CMs with robust beating and normal electrophysiological properties. Using this platform, the team also modeled pregestational diabetes-induced congenital heart defects, and showed that high glucose and insulin treatment resulted in decreased oxygen consumption, increased glycolysis, arrhythmia, and irregular mitochondria distribution and lipid droplets in the organoids, indicating successful disease modeling [17].
From 2022 onwards, the cardiac organoid’s studies have been continuously and increasingly reported (Table 2). Ormsted et al. developed a human multi-lineage gastruloid model by optimizing the conditions for the in vitro development of interconnected neuro-cardiac lineages in a single organoid [18]. Using their uniquely developed elongating multi-lineage organized (EMLO) gastruloids [19], which were derived from hiPSC aggregates and contained co-developing central and peripheral neurons and trunk mesendoderm, the team modified the procedure of the EMLO gastruloid-derived organoid development for facilitating human cardiogenesis. Following the initial gastruloid induction phase in a 2D environment supplemented with CHIR99021 and bFGF for 2 days, aggregation of dissociated cells was obtained in an anti-adherent 3D shaking culture supplemented with bFGF, hepatocyte growth factor (HGF), and insulin-like growth factor-1 (IGF-1), the last two of which were replaced into the cardiogenic growth factors such as VEGF-A and ascorbic acid after 48 h. EMLO gastruloid-derived cardiac structures (EMLOCs) emerged on day 5 onwards, and the contractile EMLOCs resembled the features of early developing hearts, including heart tube formation, chamber-like structures, and formation of a putative outflow tract. Of particular interest, the EMLOCs were surrounded by neurons co-developed in a spatially organized pattern, mimicking the innervated heart. Thus, this human EMLOC model appears to be an attractive tool to dissect the mechanisms underlying concomitant neurogenesis and cardiogenesis [18]. Another study by Branco et al. focused on the in vitro formation of pro-epicardium (PE) and foregut/liver bud, which provided crucial support to the development of cardiac organoids containing an inner myocardium-like core and an outer epicardium-like layer [20]. With the modulation of Wnt, BMP, and RA signaling in in vitro hPSC aggregates’ differentiation, the team obtained uniquely CM aggregates and PE septum transversum mesenchyme (STM)-posterior foregut/hepatic diverticulum (PFH) organoids separately. After their dissociation, the singularized cells were reaggregated at a 9:1 ratio of CMs and PE-STM-PFH cells. In the end, this co-culture approach in a 3D environment generated an epicardium–myocardium heart organoid, comprising a WT1+ epicardial-like layer that completely surrounds a TNNT2+ myocardium-like tissue, properly recapitulating the in vivo activities and functions of the PE/epicardial cells to promote heart development and CM proliferation and maturation [20]. Lee et al. developed a human chamber-forming cardiac organoid with a simple, optimized approach using hiPSC aggregates embedded into Matrigel (10%) in ultra-low attachment dishes that were rotated on a shaker and a Wnt-modulated CM differentiation protocol [21]. Importantly, the team demonstrated that key functions and morphological features of the contractile human cardiac organoids were maintained through vascularization after in vivo subcutaneous transplantation into nude mice, paving a path to a future therapeutic transplantation of a cardiac organoid model for heart disease.
Although some of the previously reported cardiac organoid models exhibited chamber formation in the structures, their chamber identities (i.e., atrium or ventricle, right or left) have not been cautiously examined. To address this, Feng et al. established the two in vitro 3D CM differentiation protocols, and using these, produced atrial and ventricular cardiac organoids, respectively [22]. The team found that in hiPSC differentiation in a 2D monolayer or 3D environment, following Wnt signaling modulation (activation/inhibition), additional treatment with RA (1 μM) at the cardiac mesoderm stage induced heart cells and organoids with predominantly atrial lineage identity in both 2D and 3D conditions, while no addition of RA treatment induced with predominantly ventricular lineage identity. Using these chamber-specific cardiac organoids, they further modeled a congenital heart defect such as Ebstein’s anomaly carrying a point mutation (c.673 C > A) in the human NKX2-5 gene. They showed that the (CRISPR-mediated) NKX2-5 mutant hiPSC line-derived organoids generated on a ventricular differentiation condition had consistently atrial CM-like phenotypes, including higher beating rates and electrophysiological and transcriptomic features, indicating successful recapitulation of the disease’s atrialized ventricular defects [22]. Next, it will be interesting and important to investigate whether these atrial and ventricular organoids can be further combined via atrioventricular connection in vitro and generate coordinated atrium–ventricle organoids. Most recently, Meier and colleagues have reported a hPSC aggregate-derived unique cardiac organoid model, termed “epicardioid”, which displayed self-organization of ventricular myocardium and epicardium through a bunch of signaling moderators and growth factors, featuring RA signaling activation [23]. Importantly, the epicardioids captured the main functions of the embryonic epicardium, such as providing several cardiac progenitor sources as well as paracrine mediators driving myocardial compaction and maturation, all of which are essential for developed self-organization and higher maturation of myocardium but have not been achieved in previous cardiac organoid models. Further, the team investigated the multicellular pathogenesis by successfully modeling human congenital (Noonan syndrome carrying a PTPN11N308S/+ mutation) or stress-induced cardiac hypertrophy and fibrotic remodeling. As such, they have further expanded the role and significance of human cardiac organoids as an excellent tool to address fundamental questions in cardiogenesis, heart disease, and drug discovery [23].
Table 2. Overview of the latest cardiac organoid models since 2021.
| Authors & Years | CO Models/Platform | Sp. | Cells | Scaffold | Chemicals | Features | Chambers/Cavities | Applications | Refs |
|---|---|---|---|---|---|---|---|---|---|
| Rossi et al., 2021 | Gatruloid-derived CO | M | mESC aggregates | CHIR, FGF2, Ascorbic Acid, VEGF-A | - Gastruloids containing the three germ layer derivatives - Induction of cardiac crescent-like FHF/SHF structure - Formation of primitive gut-like structures with a codeveloped CM heart tube with a vascular/endocardium-like network |
(−) | Cardiogenesis modeling | [12] | |
| Silva et al., 2021 | Multilineage CO | H | hiPSC-derived mes-endoderm progenitors’ aggregates | CHIR, IWP2, Ascorbic Acid, | - Reconstitution (force aggregation) of hiPSC-derived mesendoderm progenitors - Co-emergence of cardiac core and gut-like tube cells with epicardial lining, promoting CM compaction and maturation |
(−) | Cardiogenesis modeling | [13] | |
| Drakhlis et al., 2021 | Heart-Forming Organoid | H | hPSC aggregates | Matrigel | CHIR, IWP2 | - Formation of three-layered self-assembly: (inner) endothelial/endocardial/foregut cells; (middle) myocardial/epicardial cells; (outer) mesenchyme/liver cells - Recapitulation of early heart and foregut development |
(+) | Cardiogenesis & disease modeling (non-compact HCM by NKX2-5 KO) |
[14] |
| Hofbauer et al., 2021 | Cardioid | H | hPSC aggregates | Vitronectin Laminin |
CHIR, BMP4, FGF2, Activin A, LY294002, Insulin, IWP2, VEGF-A, RA | - Identification of mesodermal Wnt-BMP signaling axis (with HAND1)-modulated cavity formation principles, assembled by epicardium and myocardium and lined by a layer of ECs | (+) | Cardiogenesis & disease modeling (cryoinjury & CHD by NKX2-5 or HAND1 KO) |
[16] |
| Lewis-Israeli et al., 2021 | Scaffold-free self-assembling CO | H | hPSC aggregates | CHIR (On/Off/On), BMP4, Activin A, C59 | - Three-step Wnt signaling modulation for induction of cardiac mesoderm and epicardial cells - Recapitulation of internal chambers formed by multi-lineage cardiac cell types with well-organized sarcomeres in CMs and developed vasculature |
(+) | Cardiogenesis & disease modeling (pregestational diabetes-induced cardiomyopathy) |
[17] | |
| Ormsted et al., 2022 | EMLOC-induced CO | H | hPSC aggregates | CHIR, FGF2, HGF, IGF-1, VEGF-A, Ascorbic Acid | - Interconnected neuro-cardiac lineages in a single gastruloid model - Induction of heart tube formation, chamber-like structures, formation of a putative OFT, and innervated heart-like structure populated by neurons |
(+) | Cardiogenesis modeling | [18] | |
| Branco et al., 2022 | Epicardium-myocardium organoid (EMO) | H | hPSC aggregate-derived CM aggregates and PE/STM/PFH organoids | CHIR, BMP4, RA, Ascorbic Acid | - Wnt/BMP4/RA-mediated hPSC-PE/STM/PFH organoids - EMO generated by reaggregating hPSC-derived CM aggregates and PE/STM/PFH-dissociated cells - EMO comprising an epicardium layer fully surrounding a myocardium layer |
(−) | Cardiogenesis modeling | [20] | |
| Lee et al., 2022 | Chamber-forming CO | H | hiPSC aggregates | Matrigel | CHIR, C59 | - Manufacturing of chamber-forming hiPSC-derived COs based on Matrigel (10%) in anti-adherent dishes with dynamic culture | (+) | Cardiogenesis modeling & in vivo transplantation |
[21] |
| Feng et al., 2022 | Chamber (atrium/ventricle)-specific CO | H | hiPSC aggregates | CHIR, C59, RA (+ or −) | - Generation of atrial-lineage and ventricular-lineage COs with or without RA treatment, enabling the study of heart disease with a specific chamber defect | (+) | Cardiogenesis & disease modeling (Ebstein’s anomaly by NKX2-5 mutant) |
[22] | |
| Meier et al., 2023 | Epicardioid | H | hPSC aggregates | Collagen I | CHIR, BMP4, FGF2, Activin A, LY294002, Insulin, IWP2, RA | - Generation of self-organizing COs displaying morphological and functional patterning of the epicardium and myocardium typical of the left ventricular wall - Elucidation of fundamental roles and cellular heterogeneity of epicardial cells during ventricular development |
(−) | Cardiogenesis & disease modeling (CHD using Noonan syndrome Pt. hiPSCs & ET1-induced HCM) |
[23] |
CHD, congenital heart defect; CO, cardiac organoid; EMLOC, elongated multi-lineage organized gastruloid-derived cardiogenesis; ET1, endothelin 1; H, human; HCM, hypertrophic cardiomyopathy; hiPSC, human induced pluripotent stem cell; hPSC, human pluripotent stem cell; KO, knockout; M, mouse; mESC, mouse embryonic stem cell; PE, pro-epicardium; PFH, posterior foregut and hepatic diverticulum; Pt., patient; RA, retinoic acid; Sp, species; STM, septum transversum mesenchyme. In the “Chambers/Cavities” column, “(+)” means that the CO generates clear heart chambers and cavities in the structure, while “(−)” means that the CO does not exhibit them.
2. Applications of Cardiac Organoids
As noted above, to date, modern cardiac organoid models have been applied to various platforms, including modeling cardiogenesis, genetic and non-genetic heart disease modeling, drug screening/testing, and transplantation/regenerative medicine (Figure 1). In addition to each of the cases described in the 2nd section, the following interesting examples have been further reported.
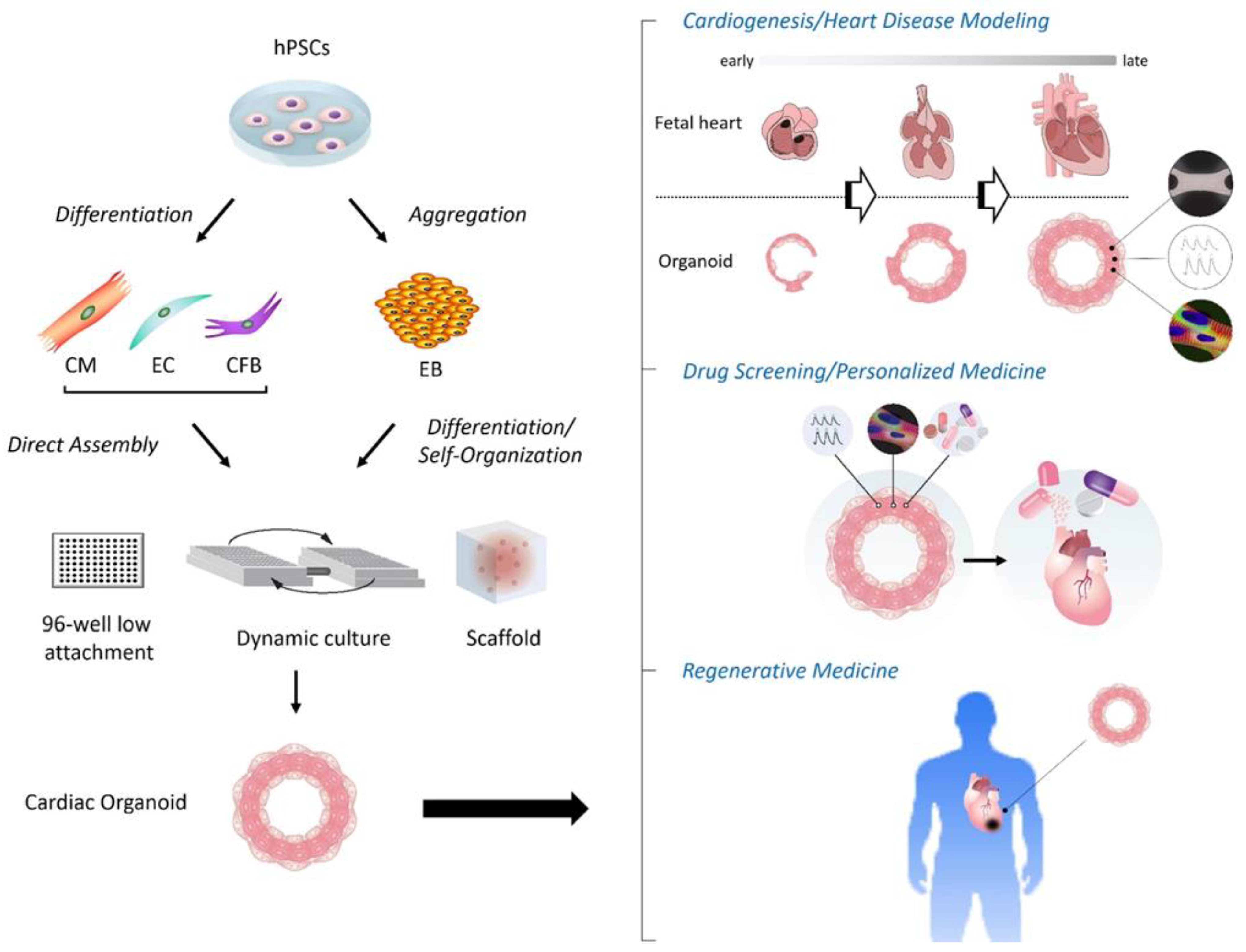
Figure 1. Schema of construction and applications of human cardiac organoids. (Left) Following aggregation or directed differentiation of hPSCs, cardiac organoids are constructed by differentiation and self-organization of aggregated hPSCs (i.e., EBs), or direct assembly of hPSC-derived cardiac cells such as CMs, ECs and CFBs under the 3D environment with anti-adherent culture plates, dynamic culture (shaking), and/or biological scaffolds. (Right) The applications of cardiac organoids involve the modeling of cardiogenesis and/or heart diseases, drug screening and development with or without targeting personalized medicine, and regenerative medicine. CFB, cardiac fibroblast; CM, cardiomyocyte; EB, embryoid body; EC, endothelial cell; hPSC, human pluripotent stem cell.
Varzideh et al. generated human cardiac organoids by co-culture with hPSC-derived cardiac progenitor cells, hPSC-derived mesenchymal stem cells, and HUVECs on a Matrigel bed for three days, and transplanted them intraperitonially into immunodeficient mice [24]. After 4 weeks, the team found that the transplanted cardiac organoids induced neovascularization with chimeric connection to host vasculature, and further promoted CM maturation as indicated by a more developed ultrastructure, transcriptomic profile, and electronic excitability patterns, compared to in vitro cardiac organoids and in vivo CM transplants. This indicated the successful transplantation of cardiac organoids for the maturation of CMs towards an adult-like phenotype [24]. In another case, although not a typical cardiac organoid, Long et al. established a disease modeling platform for Duchenne muscular dystrophy (DMD) by generating engineered heart muscles via coculture with DMD patients’ hiPSC-derived CMs and human fibroblasts in bovine collagen [25]. More critically, after correction of DMD mutation in the human dystrophin gene by CRISPR/Cas9 technology, the 3D engineered heart muscles exhibited restored dystrophin expression and improved mechanical force of contraction. This study provided clear evidence that in vitro 3D cardiac models, combined with hPSCs and genome editing, are a powerful approach to explore genetic pathogenesis and to obtain mechanistic insights into heart disease. More recently, Mills et al. utilized their unique cardiac organoid models [3][5], fabricated with a minor modification such as the use of 80% hPSC-derived cardiac cells (at a 7:3 ratio of CMs and fibroblasts) and 20% hPSC-derived ECs in mixed culture, for the screening of cardio-protective drugs against cardiac injury and dysfunction in a SARS-CoV-2 infection setting [26]. The team identified that a cocktail of inflammatory cytokines (“cytokine storm”) induced diastolic dysfunction in the cardiac organoids. They also demonstrated that bromodomain and extra-terminal family (BET) inhibition was a promising therapeutic candidate to prevent COVID-19-induced heart damage, as BET inhibitors recovered dysfunction in cardiac organoids treated with the cytokine storm or COVID-19 patient-derived serum, and rescued cardiac dysfunction and death in SARS-CoV-2-infected K18-hACE2 mice. Consequently, this study further supported the notion that compared to conventional 2D culture methods, human in vivo 3D cardiac organoid models are relevant for drug testing and discovery against heart disease.
References
- Moroni, L.; Burdick, J.A.; Highley, C.; Lee, S.J.; Morimoto, Y.; Takeuchi, S.; Yoo, J.J. Biofabrication strategies for 3D in vitro models and regenerative medicine. Nat. Rev. Mater. 2018, 3, 21–37.
- Zuppinger, C. 3D Cardiac Cell Culture: A Critical Review of Current Technologies and Applications. Front. Cardiovasc. Med. 2019, 6, 87.
- Mills, R.J.; Titmarsh, D.M.; Koenig, X.; Parker, B.L.; Ryall, J.G.; Quaife-Ryan, G.A.; Voges, H.K.; Hodson, M.P.; Ferguson, C.; Drowley, L.; et al. Functional screening in human cardiac organoids reveals a metabolic mechanism for cardiomyocyte cell cycle arrest. Proc. Natl. Acad. Sci. USA 2017, 114, E8372–E8381.
- Voges, H.K.; Mills, R.J.; Elliott, D.A.; Parton, R.G.; Porrello, E.R.; Hudson, J.E. Development of a human cardiac organoid injury model reveals innate regenerative potential. Development 2017, 144, 1118–1127.
- Mills, R.J.; Parker, B.L.; Quaife-Ryan, G.A.; Voges, H.K.; Needham, E.J.; Bornot, A.; Ding, M.; Andersson, H.; Polla, M.; Elliott, D.A.; et al. Drug Screening in Human PSC-Cardiac Organoids Identifies Pro-proliferative Compounds Acting via the Mevalonate Pathway. Cell Stem Cell 2019, 24, 895–907.
- Hoang, P.; Wang, J.; Conklin, B.R.; Healy, K.E.; Ma, Z. Generation of spatial-patterned early-developing cardiac organoids using human pluripotent stem cells. Nat. Protoc. 2018, 13, 723–737.
- Hoang, P.; Kowalczewski, A.; Sun, S.; Winston, T.S.; Archilla, A.M.; Lemus, S.M.; Ercan-Sencicek, A.G.; Gupta, A.R.; Liu, W.; Kontaridis, M.I.; et al. Engineering spatial-organized cardiac organoids for developmental toxicity testing. Stem Cell Rep. 2021, 16, 1228–1244.
- Lee, J.; Sutani, A.; Kaneko, R.; Takeuchi, J.; Sasano, T.; Kohda, T.; Ihara, K.; Takahashi, K.; Yamazoe, M.; Morio, T.; et al. In vitro generation of functional murine heart organoids via FGF4 and extracellular matrix. Nat. Commun. 2020, 11, 4283.
- Richards, D.J.; Coyle, R.C.; Tan, Y.; Jia, J.; Wong, K.; Toomer, K.; Menick, D.R.; Mei, Y. Inspiration from heart development: Biomimetic development of functional human cardiac organoids. Biomaterials 2017, 142, 112–123.
- Richards, D.J.; Li, Y.; Kerr, C.M.; Yao, J.; Beeson, G.C.; Coyle, R.C.; Chen, X.; Jia, J.; Damon, B.; Wilson, R.; et al. Human cardiac organoids for the modelling of myocardial infarction and drug cardiotoxicity. Nat. Biomed. Eng. 2020, 4, 446–462.
- Filippo Buono, M.; von Boehmer, L.; Strang, J.; Hoerstrup, S.P.; Emmert, M.Y.; Nugraha, B. Human Cardiac Organoids for Modeling Genetic Cardiomyopathy. Cells 2020, 9, 1733.
- Rossi, G.; Broguiere, N.; Miyamoto, M.; Boni, A.; Guiet, R.; Girgin, M.; Kelly, R.G.; Kwon, C.; Lutolf, M.P. Capturing Cardiogenesis in Gastruloids. Cell Stem Cell 2021, 28, 230–240.
- Silva, A.C.; Matthys, O.B.; Joy, D.A.; Kauss, M.A.; Natarajan, V.; Lai, M.H.; Turaga, D.; Blair, A.P.; Alexanian, M.; Bruneau, B.G.; et al. Co-emergence of cardiac and gut tissues promotes cardiomyocyte maturation within human iPSC-derived organoids. Cell Stem Cell 2021, 28, 2137–2152.
- Drakhlis, L.; Biswanath, S.; Farr, C.M.; Lupanow, V.; Teske, J.; Ritzenhoff, K.; Franke, A.; Manstein, F.; Bolesani, E.; Kempf, H.; et al. Human heart-forming organoids recapitulate early heart and foregut development. Nat. Biotechnol. 2021, 39, 737–746.
- Pashmforoush, M.; Lu, J.T.; Chen, H.; Amand, T.S.; Kondo, R.; Pradervand, S.; Evans, S.M.; Clark, B.; Feramisco, J.R.; Giles, W.; et al. Nkx2-5 pathways and congenital heart disease; loss of ventricular myocyte lineage specification leads to progressive cardiomyopathy and complete heart block. Cell 2004, 117, 373–386.
- Hofbauer, P.; Jahnel, S.M.; Papai, N.; Giesshammer, M.; Deyett, A.; Schmidt, C.; Penc, M.; Tavernini, K.; Grdseloff, N.; Meledeth, C.; et al. Cardioids reveal self-organizing principles of human cardiogenesis. Cell 2021, 184, 3299–3317.
- Lewis-Israeli, Y.R.; Wasserman, A.H.; Gabalski, M.A.; Volmert, B.D.; Ming, Y.; Ball, K.A.; Yang, W.; Zou, J.; Ni, G.; Pajares, N.; et al. Self-assembling human heart organoids for the modeling of cardiac development and congenital heart disease. Nat. Commun. 2021, 12, 5142.
- Olmsted, Z.T.; Paluh, J.L. A combined human gastruloid model of cardiogenesis and neurogenesis. iScience 2022, 25, 104486.
- Olmsted, Z.T.; Paluh, J.L. Co-development of central and peripheral neurons with trunk mesendoderm in human elongating multi-lineage organized gastruloids. Nat. Commun. 2021, 12, 3020.
- Branco, M.A.; Dias, T.P.; Cabral, J.M.S.; Pinto-do-Ó, P.; Diogo, M.M. Human multilineage pro-epicardium/foregut organoids support the development of an epicardium/myocardium organoid. Nat. Commun. 2022, 13, 6981.
- Lee, S.G.; Kim, Y.J.; Son, M.Y.; Oh, M.S.; Kim, J.; Ryu, B.; Kang, K.R.; Baek, J.; Chung, G.; Woo, D.H.; et al. Generation of human iPSCs derived heart organoids structurally and functionally similar to heart. Biomaterials 2022, 290, 121860.
- Feng, W.; Schriever, H.; Jiang, S.; Bais, A.; Wu, H.; Kostka, D.; Li, G. Computational profiling of hiPSC-derived heart organoids reveals chamber defects associated with NKX2-5 deficiency. Commun. Biol. 2022, 5, 399.
- Meier, A.B.; Zawada, D.; de Angelis, M.T.; Martens, L.D.; Santamaria, G.; Zengerle, S.; Nowak-Imialek, M.; Kornherr, J.; Zhang, F.; Tian, Q.; et al. Epicardioid single-cell genomics uncover principles of human epicardium biology in heart development and disease. Nat. Biotechnol. 2023, in press.
- Varzideh, F.; Pahlavan, S.; Ansari, H.; Halvaei, M.; Kostin, S.; Feiz, M.S.; Latifi, H.; Aghdami, N.; Braun, T.; Baharvand, H. Human cardiomyocytes undergo enhanced maturation in embryonic stem cell-derived organoid transplants. Biomaterials 2019, 192, 537–550.
- Long, C.; Li, H.; Tiburcy, M.; Rodriguez-Caycedo, C.; Kyrychenko, V.; Zhou, H.; Zhang, Y.; Min, Y.L.; Shelton, J.M.; Mammen, P.P.A.; et al. Correction of diverse muscular dystrophy mutations in human engineered heart muscle by single-site genome editing. Sci. Adv. 2018, 4, eaap9004.
- Mills, R.J.; Humphrey, S.J.; Fortuna, P.R.J.; Lor, M.; Foster, S.R.; Quaife-Ryan, G.A.; Johnston, R.L.; Dumenil, T.; Bishop, C.; Rudraraju, R.; et al. BET inhibition blocks inflammation-induced cardiac dysfunction and SARS-CoV-2 infection. Cell 2021, 184, 2167–2182.
More
Information
Subjects:
Cell & Tissue Engineering
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
832
Revisions:
2 times
(View History)
Update Date:
10 Apr 2023
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





