1. Introduction
The detection of miRNAs (short, non-coding RNA sequences of approximately 19 to 25 nucleotides) has tremendous potential for early diagnosis of varying disorders, most notably life-threatening and unpredictable diseases such as cancer. Their function as circulating biomarkers is however hindered by their sparseness in body fluids, small size, and minor differences between the types of miRNAs. Thus, their detection in a short time with high accuracy in biological samples remains a challenge. Body fluids affect large spectra of detecting systems due to their high viscosity, the pH influence, the presence of other interfering biomolecules, and as previously mentioned, low amounts of target miRNA
[1].
The miRNA expression spectrum in respective body fluids can differ not only depending on the disease type and pathological conditions but also on other aspects related to the patient (e.g., medication, diet, age, etc.) The body fluids in which miRNAs are detected can be obtained by either invasive or non-invasive procedures. Currently, the research is more often focused on the detection in body fluids obtained by non-invasive or weakly invasive means, such as urine, whole blood, and serum. For the miRNA capture, the body fluid is typically mixed with nanoprobe suspension
[1][2].
Methods and approaches of miRNA detection based on nanotechnologies offer an alternative path to a faster and less complicated detection process, nevertheless also with specific challenges, including reproducibility, standardization, optimization, normalization, and data processing. The techniques vary in utilized material (organic, inorganic, or hybrid), nanostructures (nanorods, nanowires, nanosheets, etc.), and the strategy of the complementary DNA application (hairpin conjugates, molecular beacons, catalytic self-assembly, spherical nucleic acids, etc.)
[3][4][5].
As such, hybrid NPs are particularly interesting since they combine the unique physical properties of the inorganic core (especially useful for their detection/quantification) with the biocompatibility/biospecificity of the organic shell.
After the selection of the nanomaterial as the substrate, for example, for the nanoprobe, the next step is the functionalization via an appropriate conjugation strategy, generally in the form of the attachment of an oligonucleotide sequence. The cornerstone of miRNA targeting approaches is antisense technology based on hybridizing the target miRNA with a complementary oligonucleotide sequence, leading to the formation of a duplex with higher stability. Usually, a complementary miRNA of the same length as the target miRNA is used, since such molecules are widely commercially available along with a wide range of modifications. Other possibilities, such as single- or double-stranded DNA, various hairpin probes, and molecular beacons, depend on the chosen approach and subsequent detection technique. The form of the target miRNA depends on the chosen media of miRNA detection—for validation of a method in model solutions, it is possible to use commercially produced sequences mimicking the naturally occurring miRNA, whose detection in biofluids can be hindered by other biomolecules and proteins present
[6].
The preparation of the functionalized material can be, at times, less scrutinized on the road to miRNA detection, the reports often understandably focused on achieving the lowest detection limits
[7]. Regarding miRNA detection methods analyze new developments and novel experimental techniques
[5][8] or are devoted to some particular nanomaterial
[9] or functionalized ligands
[10]. An attempt to gain a deeper insight in the conjugation strategies and methods (
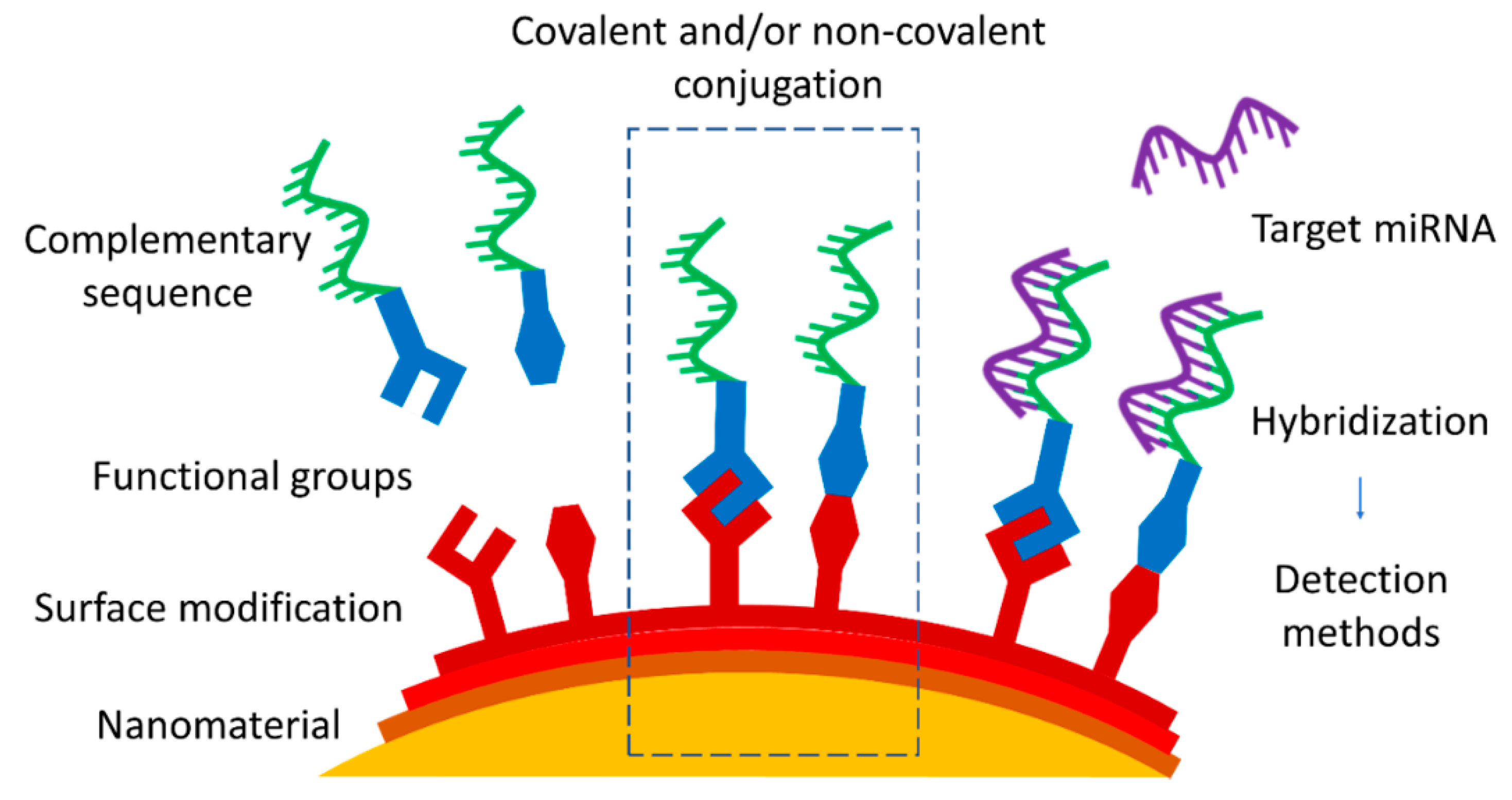
Figure 1).
Figure 1. Approaches of nanomaterial functionalization by covalent or non-covalent conjugation with complementary miRNA or DNA sequence as a part of the strategy of miRNA nanoprobe development.
2. Nanomaterials Used as Substrates
All nanomaterials mentioned in this brief overview have high surface energy and a high surface-to-volume ratio, leading to a higher catalytical activity. Their more specific properties vary depending on the nanomaterial, which oftentimes leads to a combination of two or more types of nanomaterials to take better advantage of their strengths (Table 1).
2.1. Gold Nanoparticles
Gold nanoparticles (AuNPs) have excellent catalytic and electrical properties and unique optical properties, which are extensively studied for signal amplification with the aim of reaching highly sensitive biosensing. Their optical properties are rooted in the localized surface plasmon resonance (LSPR) phenomenon, and additionally, the plasmonic surface can quench the fluorescence. With the increase of their size and/or upon aggregation of the AuNPs in aqueous suspensions (colloids), their LSPR band shifts are detected by means of a UV-Vis spectrometer or seen with a naked eye as a color change
[11].
AuNPs are biocompatible due to their chemical stability in biological fluids. Possible toxicity concerns are connected to their long-term retention in the body. On the other hand, AuNPs can be easily modified with miRNA molecules, particularly due to the gold surface affinity to the thiol group
[12]. Used as electrochemical or optical sensors in fluorescence, surface-enhanced Raman scattering (SERS), surface plasmon resonance (SPR), or colorimetry-based detection approaches, AuNPs can either be dispersed in the aqueous media
[13] or immobilized on solid support
[14]. The enhancement of AuNPs is widely used in the applications of biosensors in miRNA detection, achieving higher selectivity and sensitivity starting from sub-nanomolar levels
[11].
2.2. Silver Nanoparticles
Compared with AuNPs, the LSPR band of silver nanoparticles (AgNPs) has a shorter position (approximately 400 nm instead of 535 nm). Nevertheless, both the monodispersity and reproducibility are more difficult to control with AgNPs compared with AuNPs. This leads to a frequent combination of AgNPs with other metals, such as gold, for example, in the form of core–shell nanoparticles
[15][16][17].
AgNPs have found application as sensors in electrochemical and optical detection due to their high extinction coefficient, high scattering-to-extinction ratio, and high field enhancement. AgNPs show distinct amplified signals
[18] along with strong Raman and fluorescence enhancement. Successful approaches are based on plasmonic properties of AgNPs implemented in platforms applying SERS
[19], LSPR
[20], or fluorescence readouts for detecting miRNA
[21]. Similar to other metal nanoparticles, AgNPs also need a compatible organic coating before their use in biological systems to positively influence their stability and possible cytotoxicity
[22][23].
2.3. Magnetic Nanoparticles
The term “magnetic nanoparticles” (MNPs) generally encompasses metal (e.g., iron, nickel platinum, and cobalt), metal oxide (e.g., iron oxides Fe
3O
4, γ-Fe
2O
3, and ferrites), and metal alloy NPs (e.g., FePt and FeCo). MNPs present several advantages enabling their application in biomedicine, since after proper modification, MNPs are non-toxic and biocompatible and can be utilized as nanovectors for specific targeting. Their magnetic properties allow the use of magnetic separation for the binding and detection of biomolecules
[24][25]. MNPs are often combined with other nanomaterial types to unite the specific advantages
[26].
Considering that miRNAs are generally present in very low concentrations in body fluids, the possibility to reconcentrate the miRNAs captured by MNPs via magnetic separation or extraction presents an enticing enhancement in the detection process. The miRNA extraction from a larger volume sample also represents insight into a larger population of the molecules and provides simpler handling procedures. However, the magnetic separation can be hampered if the viscous drag of body fluids overwhelms the magnetophoretic force
[27].
2.4. Quantum Dots
Quantum dots (QDs) are luminescent semiconductor nanocrystals with the possibility to tune the maximum wavelength of their light emission spectra. Stable QDs can be obtained by an easy one-pot and/or one-step synthesis directly in a water medium
[28]. Aqueous suspensions of QDs have a high photoluminescent quantum yield and resistance to photobleaching
[29]. These qualities prompted their advancing application as optical, electrochemical, and chemiluminescent biosensors
[4].
Due to high surface reactivity, QDs can serve as a nanoscale scaffold for further functionalization with common conjugation chemistry.
All in all, QDs are useful in miRNA detection due to their strong fluorescence provided by high quantum yield, narrow emission, and broad absorption spectra which provide multicolor labels with one light source and a strongly active surface for conjugations. Interactions on the surface of QDs lead to the activation or quenching of the fluorescence signal, allowing miRNA detection in complex media, such as body fluids.
2.5. Carbon Nanomaterials
Various carbon nanomaterials offer different characteristics (higher surface area, biocompatibility, and non-toxicity) useful for biosensing. Generally, carbon nanoparticles have strong and adjustable photoluminescence
[30].
Carbon nanomaterials for miRNA detection can be in the form of carbon nanoparticles
[31], carbon nanotubes
[32], nanofibers
[33], quantum dots
[34], fullerenes
[35], graphene nanosheets
[36], and graphene oxide
[37].
The good electrical conductivity and sensitivity of carbon nanomaterials are applied for the design and construction of electrochemical biosensors, particularly for electrode surface modification and for the preparation of modified electrodes
[30][38][39][40].
Table 1. Types of nanomaterials used as substrates for miRNA nanoprobes.
2.6. Characterization of Properties of the Nanomaterial before and after Conjugations
The determination of the physico-chemical properties of the chosen nanomaterial goes hand in hand with the need for monitoring their changes, both before and after the successful conjugation of a biomolecule. Some characterization techniques are more popular than others; however, as a rule, there is usually a combination of two or more methods used to cover possible inaccuracies and weak spots of the measurements.
Dynamic light scattering (DLS) measurements remain crucial in the confirmation of the size distribution of nanoparticles in aqueous dispersion and their colloidal stability
[43]. The determination of the hydrodynamic diameter (DH) is usually joined by the evaluation of the zeta potential representative of the surface charges since the change of this parameter is often used to confirm the surface modification of the nanomaterial
[48][49]. However, with small biomolecules such as miRNA, the changes in DH and zeta potential are generally too small to be reliable proof of conjugation. Although DLS and zetametry measurements are very common and can be found in the majority of performed studies, they are also necessarily joined by other complementary techniques which allow for more accurate confirmations
[50].
Gel electrophoresis (of both the agarose and the polyacrylamide types) can be used to determine whether the conjugation between the nanomaterial and the ligand, or between two ligands, occurred. The migration of the unattached ligands is noticeably different compared with the migration of the conjugated ones, allowing for the separation of both. As a control, samples of NPs conjugated to a nucleotide sequence complementary to target miRNA, the non-conjugated NPs
[44][51], and the solutions of free sequences are often used
[52][53]. However, when it comes to confirmation of the binding of a large number of biomolecules, it is necessary to have a reliable purification technique to eliminate the excess of free ligands.
Structural and elemental analyses with X-ray diffraction (XRD) and X-ray photoelectron spectroscopy are often combined
[38] and are generally used as a precise way to study the composition and bulk properties of a nanomaterial, including the presence of biomolecules on its surface. XRD provides information about the structural properties of a nanomaterial
[44], including crystallinity and phase, while it can also give a rough idea of the average size of the NPs. XPS is the most sensitive spatially resolved technique enabling the determination of the bonding nature and elemental ratio in the nanomaterial, data from which the composition of the layers can be deduced
[35]. Both techniques are less efficient with very small NPs, with smaller precision in structural measurements, and with amorphous NPs, where different atomic lengths can affect the measurement.
Thermogravimetric analysis (TGA) is centered on the weight loss of a sample as a function of increased temperature. Due to the organic content, the characteristic weight loss profile before and after the conjugation is investigated
[12][14]. Generally, TGA is useful to confirm the presence of organic molecules conjugated to the inorganic nanomaterial, as is also the case with spectroscopic techniques.
Fourier transform infrared (FT-IR) spectroscopy is based on the detection of certain functional groups present on the surface of the nanomaterial in the event of successful conjugation and is considered acceptable for the detection of biological ligands onto inorganic nanomaterial
[34][54][55]. Nevertheless, FT-IR is not always sensitive enough. Additionally, non-reacted components need to be removed with the appropriate purification method prior to FT-IR measurements.
Fluorescence spectroscopy is quite favored for confirmation of both the successful functionalization of the nanomaterial and the subsequent hybridization of target miRNA. The strategies are based on either the quenching of the fluorescence signal
[56] or its increase after the binding event
[34]. It can be the nanomaterial itself that has fluorescent properties, such as QDs, or a complementary sequence to the target miRNA modified with fluorescent dye. Therefore, in the case of using fluorescent components, the characterization of the nanoprobe and the detection of the target miRNA capture are closely entwined. Notably, it is important to have an efficient strategy of sample purification or a reliable way to distinguish the signals, as the non-conjugated fluorescence components can easily interfere with the accuracy of the measurements.