Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Christian Barbato | -- | 2382 | 2022-11-17 13:02:07 | | | |
| 2 | Sirius Huang | Meta information modification | 2382 | 2022-11-18 02:07:14 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Riccardi, G.; Bellizzi, M.G.; Fatuzzo, I.; Zoccali, F.; Cavalcanti, L.; Greco, A.; Vincentiis, M.D.; Ralli, M.; Fiore, M.; Petrella, C.; et al. Salivary Proteomic Approaches and Methods. Encyclopedia. Available online: https://encyclopedia.pub/entry/35116 (accessed on 06 June 2026).
Riccardi G, Bellizzi MG, Fatuzzo I, Zoccali F, Cavalcanti L, Greco A, et al. Salivary Proteomic Approaches and Methods. Encyclopedia. Available at: https://encyclopedia.pub/entry/35116. Accessed June 06, 2026.
Riccardi, Gabriele, Mario Giuseppe Bellizzi, Irene Fatuzzo, Federica Zoccali, Luca Cavalcanti, Antonio Greco, Marco De Vincentiis, Massimo Ralli, Marco Fiore, Carla Petrella, et al. "Salivary Proteomic Approaches and Methods" Encyclopedia, https://encyclopedia.pub/entry/35116 (accessed June 06, 2026).
Riccardi, G., Bellizzi, M.G., Fatuzzo, I., Zoccali, F., Cavalcanti, L., Greco, A., Vincentiis, M.D., Ralli, M., Fiore, M., Petrella, C., Minni, A., & Barbato, C. (2022, November 17). Salivary Proteomic Approaches and Methods. In Encyclopedia. https://encyclopedia.pub/entry/35116
Riccardi, Gabriele, et al. "Salivary Proteomic Approaches and Methods." Encyclopedia. Web. 17 November, 2022.
Copy Citation
Oral squamous cell carcinoma (OSCC) is one of the most frequent cancers worldwide. Endoscopic methods may be useful in the evaluation of oral injuries even though the diagnostic gold standard is a biopsy. Targeted screenings could be considered the best way to prevent the occurrence of oral cancer. Aimed to elucidate the potential identification of specific biomarkers of OSCC, the use of saliva is convenient and noninvasive. Many studies reported more than a hundred putative saliva biomarkers for OSCC, and proteogenomic approaches were fundamental to disclosing this issue.
oral squamous cell carcinoma (OSCC)
salivary biomarkers
head neck cancer
proteomics
otolaryngology
1. Introduction
The most important etiological factors of oral cancer are alcohol, tobacco, and herpes papilloma virus infections [1][2]. Some premalignant lesions are associated with OSCC, and the most common is leukoplakia [3]. In the evaluation of oral injuries, the main diagnostic gold standard is biopsy, in addition to endoscopy. Targeted screenings could be considered the best way to prevent the occurrence of oral cancer and improve public health status, as is performed for breast and cervical cancer [4]. The World Health Organization (WHO) indicates that wide screening is the only effective strategy to lower the morbidity of each cancer, as well as for OSCC. Moreover, with the aim of increasing survival rates of patients with oral cancer, the identification of approved biomarkers as early outcomes is essential in the diagnostic algorithm of OSCC.
In this regard, saliva samples could be considered a useful tool for the study of general clinical conditions due to their numerous molecular contents. Thus, the search for salivary biomarkers could be employed in preventing oral cancer and diagnosing OSCC at an early stage. Saliva is a product of the salivary glands and it contains various proteins, microorganisms, skin cells, serum, and blood derivatives [5]. Cancer-related proteins released by malignant cells and/or other proteins related to various systemic diseases have been discovered in salivary sampling [6][7]. Knowledge of the salivary proteome can help us find potential biomarkers for early oral cancer screening [8].
Saliva proteomics can serve as a reliable source for biomarker validation for OSCC (Figure 1). Saliva is a promising biofluid that could mirror the physiological and pathological state of the organism [9]. From oral cavity cells and salivary glands, a plethora of proteins are released into the saliva, and their profiling in OSCC is a promising approach. In addition, the liquid biopsy performed on the patient’s saliva is a non-invasive analysis.
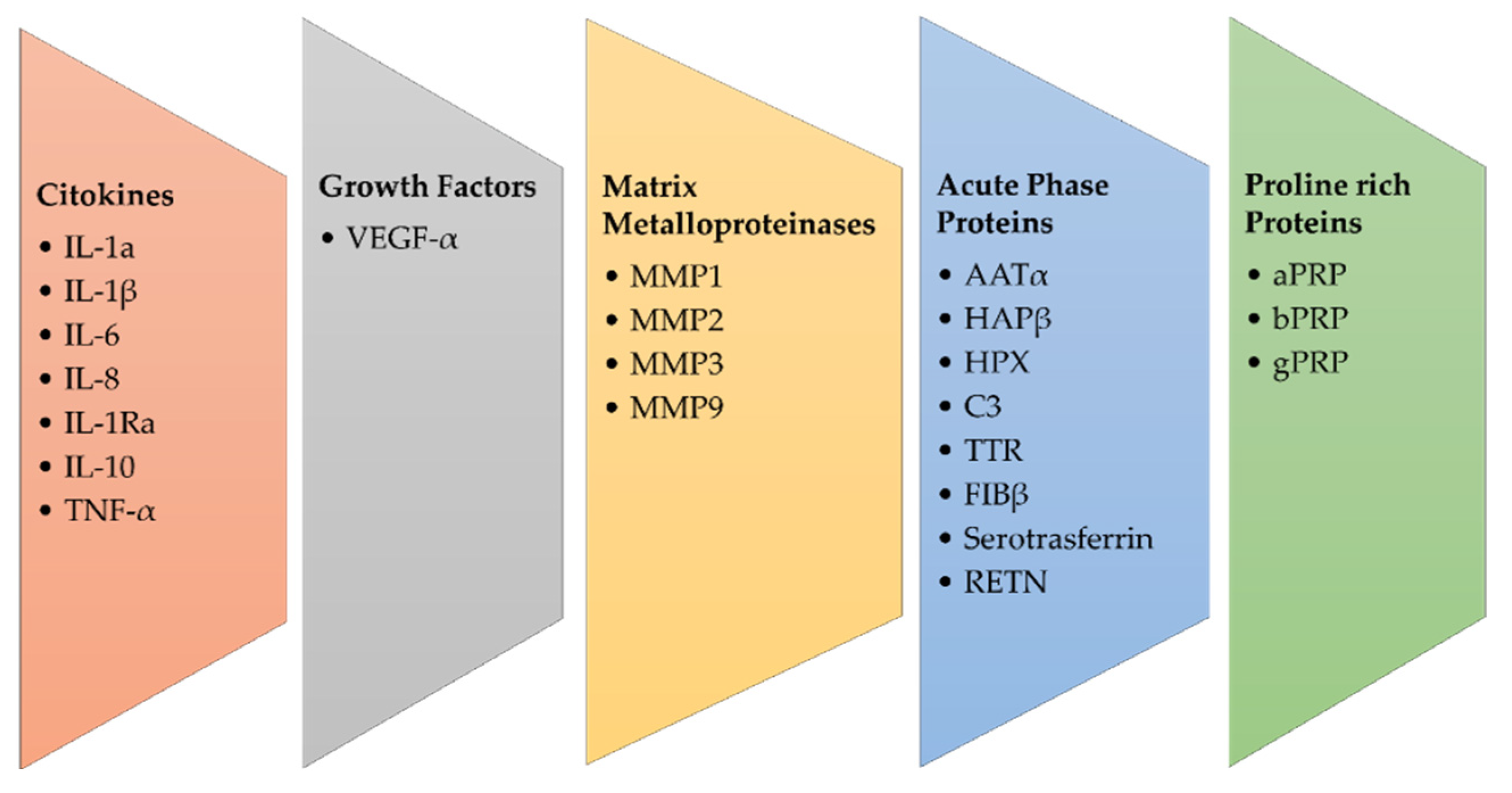
Figure 1. Main groups of salivary biomarkers in OSCC.
Physiologically, the proteins most present in saliva are alfa-amylase, cystatins, pro-line-rich peptides, mucins, and serum albumin. Because more than 50% of cases of OSCC are diagnosed at late stages of disease progression, an early saliva protein content analysis might be a useful strategy to reveal cancer oral disease.
In this regard, the time window of saliva protein analysis could be kept in mind for several studies. In addition, the normal saliva composition with respect to sex, age, and the circadian cycle is a fundamental point. The following reports show a partial view with respect to these mentioned points, but reinforce the opportunity for saliva proteomics.
Cytokeratin has been shown as a potential diagnostic biomarker in head and neck cancer [10]. Salivary levels of Interleukin-1α (IL-1 α), IL-6, IL-8, Vascular-Endothelial Growth Factor- α (VEGF- α) and Tumor Necrosis Factor- α (TNF- α) can support analysis of the progression of premalignant lesions of the tongue and could be used for cancer screening and early diagnosis [11]. Salivary matrix metalloproteinase (MMP) concentrations may be useful for detecting and monitoring OSCC; in fact, as reported by Stott-Miller et al., salivary concentrations of MMP1 and MMP3 were 6–15 times higher in patients with OSCC than in healthy people, with a tendency to increase with higher stage disease [12]. The expression of MMP-9 in saliva has been associated with OSCC and is necessary to obtain an early diagnosis [13].
Shiptzer et al. reported that expression of cyclin D1, Ki67, LDH, and matrix metalloproteinase 9 (MMP-9) were elevated, while measurement of DNA 8-oxoquanine glycosylase (OGG1) and Maspin was lowered in oral cancer saliva patients [14]. MMP2 and MMP-9 are associated with tumor invasion and metastasis, and the concentration of these biomarkers differs significantly in the saliva of healthy individuals, patients with premalignant diseases of the oral cavity, and OSCC patients, suggesting their diagnostic and prognostic role [15][16][17]. In fact, the main protein families involved in salivary proteomics are proline-rich proteins (PRP), including acidic PRP (aPRP), basic PRP (bPRP), and glycosylated PRP (gPRP), α-amylase, mucins, salivary (type S) cystatins, histatins, and statins [18]. Krapfenbauer et al. identified 25 proteins specific to OSCC, suggesting their measurement as biomarkers for oral cancer. Among these, 12 new proteins identified are the proteins galectin-7, cofilin, precursor of CRP, creatine kinase, fatty acid binding protein of the m-chain, type II keratin, myosin light chains 2 and 3, plakoglobin, retinoic acid binding protein II, nucleoside diphosphate kinase A and phosphoglycerate mutase 1 [19].
Starting from the observation that chronic inflammation and oral cancer are connected, acute phase response proteins (APPs) have been detected in the saliva of patients with OSCC, and in particular the APP Haptoglobinβ (HAPβ), α-antitrypsin (AATα), complement-C3 (C3), hemopexin (HPX), serotransferrin, transthyretin (TTR) and fibrinogen β (FIBβ) [19]. In addition, it was suggested that increased levels of HAPβ, AATα, C3, HPX, TF, TTR, FIBβ, and ABG were suitable biomarkers for the early diagnosis of OSCC [19][20]. Yu et al. supported this hypothesis by assembling an inflammatory protein panel for the initial screening of OSCC [21].
Finally, they described another biomarker for the early diagnosis of OSCC, named resistin (RETN), which is a cysteine-rich adipose-derived peptide hormone. Initially, it was considered an endocrine molecule, and after it was associated with type II diabetes mellitus, inflammation, and cardiology disease. RETN has been shown to be highly correlated with the advanced grade of OSCC and metastases. This result highlighted that RETN is a putative salivary biomarker for early diagnosis and is mainly a possible negative prognostic factor of OSCC.
However, the authors are convinced that a standardized system for the collection and analysis of saliva is necessary for it to be used as an early diagnosis target, given the vulnerability of salivary proteins to proteolytic enzymes, oral microorganisms, and the circadian cycle.
2. Salivary Proteomic Approaches and Methods
An important starting point is the perspective of combining salivary proteomic analysis with standard oral examinations in order to demonstrate the role of salivary proteomic analysis for diagnosis in OSCC (Table 1).
Table 1. Selected studies on Proteomic and OSCC.
| Author | Country | Institutional Affiliation | Method | Purpose |
|---|---|---|---|---|
| Aro et al. [22] | Finland | University of Helsinki | Genomics; Proteomics and transcriptomics; Lipidomics; Metabolomics; Microbiomics | Review of the recent advancements in the field of salivary diagnostics in oral cancer |
| Li et al. [23] | China | Sichuan University | Salivary proteomics for oral cancer | Putative salivary proteomic biomarkers in oral cancer screening |
| Chattopadhyay et al. [24] | India | Central University of Tamil Nadu | GSTT1 Polymorphism in Salivary DNA; Role of circulating tumor DNA (ctDNA); Salivary methylome; Salivary transcriptome; Salivary Proteomics, Salivary Microbiome; Salivary Metabolomics | Saliva biomarkers as diagnostic and prognostic tools |
| Ni et al. [25] | China | Nanjing University | Protein biomarker | Early detection by tissues, salivary and serum |
| Kaczor-Urbanowicz et al. [26] | USA | University of California at Los Angeles | Poc diagnostics; RNA sequencing; Liquid biopsy; Electromagnetic field-based techniques; Electric field stimulates release and measurement method | Introduce population-based screening programs |
| Khurshid et al. [27] | Saudi Arabia | King Faisal University | Whole-Mouth Saliva Biomarker | Non-invasive diagnosis |
| Chakraborty et al. [28] | India | Vellore Institute of Technology | Microfluidics systems Salivary biomarkers (Telomerase, HPV16 DNA, Reactive nitrogen species and antioxidants, Salivary vitamin E and C) | Early stage of diagnosis |
| Manzano-Moreno et al. [29] | Spain | University of Granada | Saliva, MicroRNA and Cytokines | Diagnosis and prognosis of OSCC |
Salivary proteins have been observed to be vulnerable to external factors, and therefore, saliva samples must be collected, refrigerated at 4 °C, and analyzed at a cold temperature to avoid bacterial contamination within 3–6 h. High-speed and high-sensitivity mass spectrometry (MS) allows the investigation of salivary proteomes in a thorough and scrupulous way, also from the point of view of gene expression and post-translational modifications. This approach is usually combined with surface enhanced laser desorption ionization (SELDI), matrix-assisted laser desorption ionization (MALDI) or time-of-flight (TOF) with the aim of measuring intact proteins or peptides (SELDI-TOF-MS) [30].
A biomarker can be defined as a measurable and quantifiable biological parameter that has the ability to play a role in health or environmental exposure and drug responses to a therapeutic protocol. Prognostic biomarkers are used to monitor a benign or malignant condition and give us information about the future development of the health condition, while diagnostic biomarkers indicate the presence or absence of a pathological state, such as cancer or chronic diseases.
Kaczor-Urbanowicz et al. suggest that an integrated electrochemical multiplexing saliva-based platform for oral cancer detection has emerged [26]. This platform is very useful in detecting salivary proteins and nucleic acids such as DNA and RNA, analyzing up to eight different biomarkers in a single session in a few minutes. This saliva test was applied in an Indian cohort of saliva samples in OSCC patients and achieved 90% sensitivity and 90% specificity for both interleukin 8 (IL-8) and IL-8 mRNA. These promising data indicate that this method could be useful for screening and risk assessment for oral cancer and for selecting patients who may need a biopsy [31].
OSCC is common worldwide and, despite improvements in treatment, the 5-year survival rate is disastrous. Factors associated with this failure appear to be late diagnosis and the development of relapses of primary tumors. Therefore, the identification of an OSCC protein biomarker during the initiation and progression of cancer would aid in its diagnosis and treatment by a salivary proteomic approach.
Technologies such as differential gel electrophoresis, two-dimensional polyacrylamide gel electrophoresis, and multidimensional protein identification technology can be used for profiling. Specifically, the use of a small amount of unfractionated serum sample added to a “protein chip”, which is subsequently analyzed by surface-enhanced laser desorption-ionization time-of-flight mass spectrometry (SELDI-TOF-MS) to generate a proteomic signature of the serum, represents an advanced technology [32][33].
Currently, OSCC is diagnosed after a thorough physical examination of the oral cavity looking for signs and symptoms of the disease. When the clinical examination shows an abnormal area in the oral cavity, a tissue biopsy is performed with resection of the same and the verification of the presence of malignant tumor cells by a pathologist. Ishikawa et al. verified that 12 hours after dinner is the best time to collect saliva samples because they report that saliva collected in this time frame expressed significantly different concentrations of metabolites in OSCC patients compared to healthy patients. The study, therefore, suggested a longer fasting period if saliva should be used for diagnosis to improve the discriminatory capacity of any method of analysis [27]. To identify putative protein biomarkers suitable for OSCC detection, all proteins detected in whole saliva samples from patients with OSCC were analyzed. Saliva samples were profiled using rifle proteomics based on C4 reverse phase liquid chromatography for pre-fractionation reverse phase capillary liquid chromatography with quadruple time-of-flight mass spectrometry.
Results demonstrated that the five markers, Mac-2 Binding protein (M2BP), Migration inhibitory factor-related protein 14 (MRP14), CD59 Glycoprotein, catalase, and profilin, provide 90% sensitivity and 83% specificity for OSCC detection [34]. With the aim of identifying biomarkers of cancer development with respect to precancerous lesions, salivary proteins from 12 patients with OSCC and 12 healthy subjects were taken and separated by two-dimensional gel electrophoresis (2DE) and mass spectrometry (MS), and the solid association of α-1-antitrypsin and haptoglobin with OSCC was further enhanced by immunohistochemical staining of tumor tissues [35]. In another similar study, a total of 41 OSCC patients and 30 OSCC-free control subjects were recruited and 2DE and MS methods were used. The sensitivity of the transferrin-based ELISA for oral cancer prediction was 100% for patients with stage T1 oral cancer, 86.6% for stage T2 and 100% for stage T3/T4. This suggests that salivary transferrin may be a biomarker for the detection of the early stage of oral cancer [36]. As for malignant lesions of the oral cavity, a differential proteomic profiling was performed on saliva from dysplastic leukoplakia patients. Initially, 93 proteins were found, and up to 30 overexpressed were selected, and CD44, S100A7, and S100P were significantly associated as putative candidates for the early phase of tumor progression [37]. Aiming to differentiate the early to the late phase of OSCC patients, more recently, Jain et al., individuated from saliva samples several proteins by LC-MS in oral squamocellular cell cancer and thereafter twelve were validated using targeted proteomics [38]. Among these, Alpha 2 Glycoprotein 1 (AZGP1), Alpha-2-Heremans-Schmid Glycoprotein (AHSG), Keratin 6C (KRT6C) and BPI fold-containing family B member 2 (BPIFB2) showed high sensitivity and specificity, suggesting theirs as a potential salivary biomarker of different stages of OSCC [39].
Chattopadhyay et al. summarize the potential of omics studies on salivary biomarkers as diagnostic and prognostic approaches to the detection of oral cancer [24]. The salivary proteome is defined as the complete protein content present in human saliva. Up to two thousand proteins are released into the oral cavity by the acinar cells of all the salivary glands. The salivary proteome has been useful in identifying possible biomarkers for OSCC using advanced proteomics technologies such as mass spectrometry, liquid chromatography, and protein labeling. Endothelin-1 represents a potential biomarker for the development of OSCC in patients with oral lichen planus and IL-8, IL-1β, glycoprotein M2BP (Mac-2 binding protein), CD59, myeloid protein 14 (MRP14) and catalase as salivary biomarkers of oral cancer [40].
Chakraborty et al. summarize all the advances in oral cancer detection and thus also on salivary proteins for OSCC screening and detection [28]. In addition to being non-invasive and inexpensive, the use of saliva has the advantage that sample collection is simple and easy to learn. Ease of storage and transport, sample availability, and ease of repeated collection of salivary samples can improve their feasibility for diagnostic applications [41]. In this article, they are mentioned as molecules for clinical diagnosis in patients with OSCC Cyfra-21 and CA-125, but also mRNA, microRNA, and reactive nitrogen species [42].
Hu et al. analyzed the role of salivary biomarkers in the detection of oral cancer [43]. As well, other authors have shown that saliva contains a collection of analytes, such as proteins, DNA, and mRNA, as well as several metabolites that can be potential biomarkers for clinical and translational applications [44]. Therefore, salivary analysis is an effective option for prevention, monitoring, diagnosis, and prognosis and this biological liquid could even become the first choice for the screening and identification of biomarkers given the constant contact with cancer cells inside the oral cavity [45].
References
- Ferraguti, G.; Terracina, S.; Petrella, C.; Greco, A.; Minni, A.; Lucarelli, M.; Agostinelli, E.; Ralli, M.; de Vincentiis, M.; Raponi, G.; et al. Alcohol and Head and Neck Cancer: Updates on the Role of Oxidative Stress, Genetic, Epigenetics, Oral Microbiota, Antioxidants, and Alkylating Agents. Antioxidants 2022, 11, 145.
- Galati, L.; Chiocca, S.; Duca, D.; Tagliabue, M.; Simoens, C.; Gheit, T.; Arbyn, M.; Tommasino, M. HPV and head and neck cancers: Towards early diagnosis and prevention. Tumour Virus Res. 2022, 14, 200245.
- Salcedo-Bellido, I.; Requena, P.; Mateos, R.; Ortega-Rico, C.; Olmedo-Requena, R.; Lozano-Lorca, M.; Arrebola, J.P.; Barrios-Rodríguez, R. Factors associated with the development of second primary tumours in head and neck cancer patients. Eur. J. Cancer Care 2022, 18, e13699.
- Chen, H.; Shu, H.-M.; Chang, Z.-L.; Wang, Z.-F.; Yao, H.-H.; Zhu, H.-M.; Lu, T.-M.; Ma, Q.-Y.; Yang, B.-L. Efficacy of Pap test in combination with ThinPrep cytological test in screening for cervical cancer. Asian Pac. J. Cancer Prev. 2012, 13, 1651–1655.
- Loo, J.A.; Yan, W.; Ramachandran, P.; Wong, D.T. Comparative Human Salivary and Plasma Proteomes. J. Dent. Res. 2010, 89, 1016–1023.
- Wong, D.T. Salivaomics. J. Amer. Dent. Assoc. 2012, 143, 19S–24S.
- Wang, X.; Kaczor-Urbanowicz, K.E.; Wong, D.T. Salivary biomarkers in cancer detection. Med. Oncol. 2017, 34, 7.
- Cristaldi, M.; Mauceri, R.; Di Fede, O.; Giuliana, G.; Campisi, G.; Panzarella, V. Salivary Biomarkers for Oral Squamous Cell Carcinoma Diagnosis and Follow-Up: Current Status and Perspectives. Front. Physiol. 2019, 10, 1476.
- Sugimoto, M. Salivary Metabolomics for Cancer Detection. Expert Rev. Proteom. 2020, 17, 639–648.
- Tang, K.D.; Kenny, L.; Perry, C.; Frazer, I.; Punyadeera, C. The overexpression of salivary cytokeratins as potential diagnostic biomarkers in head and neck squamous cell carcinomas. Oncotarget 2017, 8, 72272–72280.
- Korostoff, A.; Reder, L.; Masood, R.; Sinha, U.K. The role of salivary cytokine biomarkers in tongue cancer invasion and mortality. Oral Oncol. 2011, 47, 282–287.
- Stott-Miller, M.; Houck, J.R.; Lohavanichbutr, P.; Méndez, E.; Upton, M.P.; Futran, N.D.; Schwartz, S.M.; Chen, C. Tumor and salivary matrix metalloproteinase levels are strong diagnostic markers of oral squamous cell carcinoma. Cancer Epidemiol. Biomark. Prev. 2011, 20, 2628–2636.
- Smriti, K.; Ray, M.; Chatterjee, T.; Shenoy, R.P.; Gadicherla, S.; Pentapati, K.C.; Rustaqi, N. Salivary MMP-9 as a Biomarker for the Diagnosis of Oral Potentially Malignant Disorders and Oral Squamous Cell Carcinoma. Asian Pac. J. Cancer Prev. 2020, 21, 233–238.
- Shiptzer, T.; Hamzany, Y.; Bahar, G.; Feinmesser, R.; Savulescu, D.; Borovoi, I.; Gavish, M.; Nagler, R.M. Salivary analysis of oral cancer biomarkers. Br. J. Cancer 2009, 101, 1194–1198.
- Valavanidis, A.; Vlachogianni, T.; Fiotakis, K.; Loridas, S. Pulmonary Oxidative Stress, Inflammation and Cancer: Respirable Particulate Matter, Fibrous Dusts and Ozone as Major Causes of Lung Carcinogenesis through Reactive Oxygen Species Mechanisms. Int. J. Environ. Res. Public Health 2013, 10, 3886–3907.
- Roszkowski, K.; Jozwicki, W.; Blaszczyk, P.; Mucha-Malecka, A.; Siomek, A. Oxidative damage DNA: 8-oxoGua and 8-oxodG as molecular markers of cancer. Med. Sci. Monit. 2011, 17, CR329–CR333.
- Agha-Hosseini, F.; Mirzaii-Dizgah, I.; Farmanbar, N.; Abdollahi, M. Oxidative stress status and DNA damage in saliva of human subjects with oral lichen planus and oral squamous cell carcinoma. J. Oral Pathol. Med. 2012, 41, 736–740.
- Korkmaz, K.S.; Butuner, B.D.; Roggenbuck, D. Detection of 8-OHdG as a diagnostic biomarker. J. Lab. Precis. Med. 2018, 3, 95.
- Krapfenbauer, K. Identification of tumour related proteins as potential screening markers by proteome analysis-protein profiles of human saliva as a predictive and prognostic tool. EPMA 2014, J5, 1–15.
- Castagnola, M.; Cabras, T.; Iavarone, F.; Fanali, C.; Nemolato, S.; Peluso, G.; Bosello, S.L.; Faa, G.; Ferraccioli, G.; Messana, I. The human salivary proteome: A critical overview of the results obtained by different proteomic platforms. Expert Rev. Proteom. 2012, 9, 33–46.
- Yu, J.S.; Chen, Y.T.; Chiang, W.F.; Hsiao, Y.C.; Chu, L.J.; See, L.C.; Wu, C.S.; Tu, H.T.; Chen, H.W.; Chen, C.C. Saliva protein biomarkers to detect oral squamous cell carcinoma in a highrisk population in Taiwan. Proc. Natl. Acad. Sci. USA 2016, 113, 11549.
- Aro, K.; Kaczor-Urbanowicz, K.; Carreras-Presas, C.M. Salivaomics in oral cancer. Curr. Opin. Otolaryngol. Head Neck Surg. 2019, 27, 91–97.
- Li, Q.; Ouyang, X.; Chen, J.; Zhang, P.; Feng, Y. A Review on Salivary Proteomics for Oral Cancer Screening. Curr. Issues Mol. Biol. 2020, 37, 47–56.
- Chattopadhyay, I.; Panda, M. Recent trends of saliva omics biomarkers for the diagnosis and treatment of oral cancer. J. Oral Biosci. 2019, 61, 84–94.
- Ni, Y.H.; Ding, L.; Hu, Q.G.; Hua, Z.C. Potential biomarkers for oral squamous cell carcinoma: Proteomics discovery and clinical validation. Proteom. Clin. Appl. 2015, 9, 86–97.
- Kaczor-Urbanowicz, K.E.; Martín Carreras-Presas, C.; Kaczor, T.; Tu, M.; Wei, F.; Garcia-Godoy, F.; Wong, D.T. Emerging technologies for salivaomics in cancer detection. J. Cell Mol. Med. 2017, 21, 640–647.
- Khurshid, Z.; Zafar, M.S.; Khan, R.S.; Najeeb, S.; Slowey, P.D.; Rehman, I.U. Role of Salivary Biomarkers in Oral Cancer Detection. Adv. Clin. Chem. 2018, 86, 23–70.
- Chakraborty, D.; Natarajan, C.; Mukherjee, A. Advances in oral cancer detection. Adv. Clin Chem. 2019, 91, 181–200.
- Manzano-Moreno, F.J.; Costela-Ruiz, V.J.; García-Recio, E.; Olmedo-Gaya, M.V.; Ruiz, C.; Reyes-Botella, C. Role of Salivary MicroRNA and Cytokines in the Diagnosis and Prognosis of Oral Squamous Cell Carcinoma. Int. J. Mol. Sci. 2021, 22, 12215.
- Domon, B.; Aebersold, R. Mass spectrometry and protein analysis. Science 2006, 312, 212–217.
- Wu, C.C.; Chu, H.W.; Hsu, C.W.; Chang, K.P.; Liu, H.P. Saliva proteome profiling reveals potential salivary biomarkers for detection of oral cavity squamous cell carcinoma. Proteomics 2015, 15, 3394–3404.
- Kawas, S.A.; Rahim, Z.H.A.; Ferguson, D.B. Potential uses of human salivary protein and peptide analysis in the diagnosis of disease. Arch. Oral Biol. 2012, 57, 1–9.
- Wei, F.; Patel, P.; Liao, W.; Chaudhry, K.; Zhang, L.; Arellano-Garcia, M.; Hu, S.; Elashoff, D.; Zhou, H.; Shukla, S.; et al. Electrochemical sensor for multiplex biomarkers detection. Clin. Cancer Res. 2009, 15, 4446–4452.
- Nunes, L.A.S.; Brenzikofer, R.; Macedo, D.V. Reference intervals for saliva. analytes collected by a standardized method in a physically active population. Clin. Biochem. 2011, 44, 1440–1444.
- Jessie, K.; Jayapalan, J.J.; Ong, K.-C.; Rahim, Z.H.A.; Zain, R.M.; Wong, K.-T.; Hashim, O.H. Aberrant proteins in the saliva of patients with oral squamous cell carcinoma. Electrophoresis 2013, 34, 2495–2502.
- Sivadasan, P.; Gupta, M.K.; Sathe, G.; Sudheendra, H.V.; Sunny, S.P.; Renu, D.; Hari, P.S.; Gowda, H.; Suresh, A.; Kuriakose, M.A.; et al. Salivary proteins from dysplastic leukoplakia and oral squamous cell carcinoma and their potential for early detection. J. Proteom. 2020, 212, 103574.
- de Jong, E.P.; Xie, H.; Onsongo, G.; Stone, M.D.; Chen, X.B.; Kooren, J.A.; Refsland, E.W.; Griffin, R.J.; Ondrey, F.G.; Wu, B.; et al. Quantitative proteomics reveals myosin, actin as promising saliva biomarkers for distinguishing pre-malignant, malignant oral lesions. PLoS ONE 2010, 5, e11148.
- Jain, A.; Kotimoole, C.N.; Ghoshal, S.; Bakshi, J.; Chatterjee, A.; Prasad, T.S.K.; Pal, A. Identification of potential salivary biomarker panels for oral squamous cell carcinoma. Sci. Rep. 2021, 11, 3365.
- Jou, Y.-U.; Lin, C.-H.; Lai, C.-H.; Chen, C.-H.; Kao, J.-U.; Chen, S.-H.; Tsai, M.-I.; Huang, S.-U.; Lin, C.-H. Proteomic identification of salivary transferrin as a biomarker for early detection of oral cancer. Anal. Chim. Acta 2010, 681, 41–48.
- Lau, W.W.; Hardt, M.; Zhang, Y.H.; Freire, M.; Ruhl, S. The Human Salivary Proteome Wiki: A Community-Driven Research Platform. J. Dent Res. 2021, 100, 1510–1519.
- Wong, D.T. Salivary diagnostics powered by nanotechnologies, proteomics and genomics. J. Am. Dent. Assoc. 2006, 137, 313–321.
- Balan, J.J.; Rao, R.S.; Premalatha, B.R.; Patil, S. Analysis of tumor marker CA 125 in saliva of normal and oral squamous cell carcinoma patients: A comparative study. J. Contemp. Dent. Pract. 2012, 13, 671–675.
- Hu, S.; Arellano, M.; Boontheung, P.; Wang, J.; Zhou, H.; Jiang, J.; Elashoff, D.; Wei, R.; Loo, J.A.; Wong, D.T. Salivary proteomics for oral cancer biomarker discovery. Clin. Cancer Res. 2008, 14, 6246–6252.
- Ishikawa, S.; Sugimoto, M.; Konta, T.; Kitabatake, K.; Ueda, S.; Edamatsu, K.; Okuyama, N.; Yusa, K.; Iino, M. Salivary Metabolomics for Prognosis of Oral Squamous Cell Carcinoma. Front. Oncol. 2022, 11, 789248.
- Elmahgoub, F. Could salivary biomarkers be useful in the early detection of oral cancer and oral potentially malignant disorders, and is there a relationship between these biomarkers and risk factors? Evid. Based Dent. 2022, 23, 30–31.
More
Information
Subjects:
Pathology
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.1K
Revisions:
2 times
(View History)
Update Date:
18 Nov 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





