
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Camila Xu | -- | 2898 | 2022-11-17 01:41:14 |
Video Upload Options
Acetylation (or in IUPAC nomenclature ethanoylation) describes a reaction that introduces an acetyl functional group into a chemical compound. Deacetylation is the removal of an acetyl group. Acetylation refers to the process of introducing an acetyl group (resulting in an acetoxy group) into a compound, namely the substitution of an acetyl group for an active hydrogen atom. A reaction involving the replacement of the hydrogen atom of a hydroxyl group with an acetyl group (CH3CO) yields a specific ester, the acetate. Acetic anhydride is commonly used as an acetylating agent reacting with free hydroxyl groups. For example, it is used in the synthesis of aspirin, heroin, and THC-O-acetate.
1. Protein Acetylation
Acetylation is an important modification of proteins in cell biology; and proteomics studies have identified thousands of acetylated mammalian proteins.[1][2][3] Acetylation occurs as a co-translational and post-translational modification of proteins, for example, histones, p53, and tubulins. Among these proteins, chromatin proteins and metabolic enzymes are highly represented, indicating that acetylation has a considerable impact on gene expression and metabolism. In bacteria, 90% of proteins involved in central metabolism of Salmonella enterica are acetylated.[4][5]
1.1. N-Terminal Acetylation
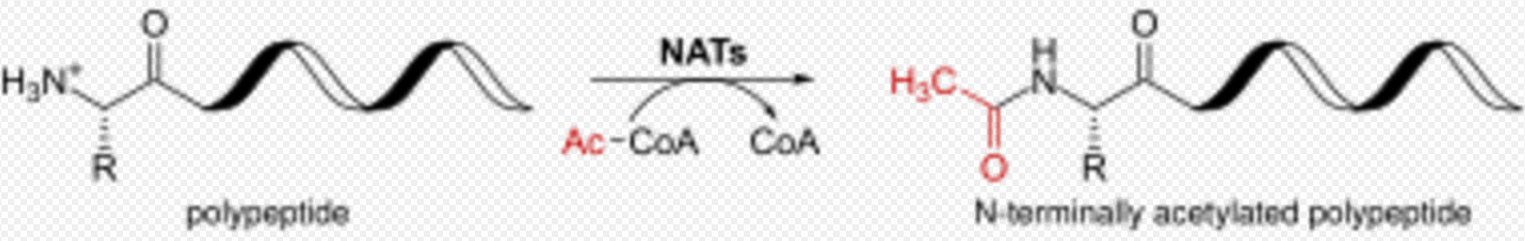
N-terminal acetylation is one of the most common co-translational covalent modifications of proteins in eukaryotes, and it is crucial for the regulation and function of different proteins. N-terminal acetylation plays an important role in the synthesis, stability and localization of proteins. About 85% of all human proteins and 68% in yeast are acetylated at their Nα-terminus.[6] Several proteins from prokaryotes and archaea are also modified by N-terminal acetylation.
N-terminal Acetylation is catalyzed by a set of enzyme complexes, the N-terminal acetyltransferases (NATs). NATs transfer an acetyl group from acetyl-coenzyme A (Ac-CoA) to the α-amino group of the first amino acid residue of the protein. Different NATs are responsible for the acetylation of nascent protein N-terminal, and the acetylation was found to be irreversible so far.[7]
N-terminal acetyltransferases
To date, seven different NATs have been found in humans - NatA, NatB, NatC, NatD, NatE, NatF and NatH. Each of these different enzyme complexes is specific for different amino acids or amino acid sequences which is shown in the following table.
Table 1. The Composition and Substrate specificity of NATs.
| NAT | Subunits (catalytic subunits are in bold.) | Substrates |
|---|---|---|
| NatA | Naa10 (Ard1) Naa15 (Nat1) | Ser-, Ala-, Gly-, Thr-, Val-, Cys- N-termini |
| NatB | Naa20 (Nat3) Naa25 (Mdm20) | Met-Glu-, Met-Asp-, Met-Asn-, Met-Gln- N-termini |
| NatC | Naa30 (Mak3) Naa35 (Mak10) Naa38 (Mak31) | Met-Leu-, Met-Ile-, Met-Trp-, Met-Phe- N-termini |
| NatD | Naa40 (Nat4) | Ser-Gly-Gly-, Ser-Gly-Arg- N-termini |
| NatE | Naa50 (Nat5) Naa10 (Ard1) Naa15 (Nat1) | Met-Leu-, Met-Ala-, Met-Lys-, Met-Met- N-termini |
| NatF | Naa60 | Met-Lys-, Met-Leu-, Met-Ile-, Met-Trp-, Met-Phe- N-termini |
| NatH | Naa80 | Actin- N-termini |
NatA
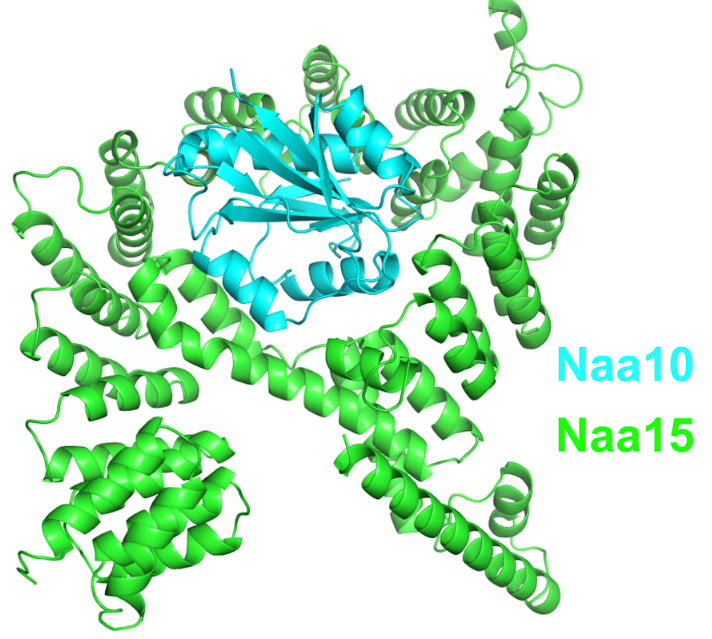
NatA is composed of two subunits, the catalytic subunit Naa10 and the auxiliary subunit Naa15. NatA subunits are more complex in higher eukaryotes than in lower eukaryotes. In addition to the genes NAA10 and NAA15, the mammal-specific genes NAA11 and NAA16, make functional gene products, which form different active NatA complexes. Four possible hNatA catalytic-auxiliary dimers are formed by these four proteins. However, Naa10/Naa15 is the most abundant NatA.[9]
NatA acetylates Ser, Ala-, Gly-, Thr-, Val- and Cys N-termini after the initiator methionine is removed by methionine amino-peptidases. These amino acids are more frequently expressed in the N-terminal of proteins in eukaryotes, so NatA is the major NAT corresponding to the whole number of its potential substrates.[10]
Several different interaction partners are involved in the N-terminal acetylation by NatA. Huntingtin interacting protein K (HYPK) interacts with hNatA on the ribosome to affect the N-terminal acetylation of a subset of NatA substrates. Subunits hNaa10 and hNaa15 will increase the tendency for aggregation of Huntingtin if HYPK is depleted. Hypoxia-inducible factor (HIF)-1α has also been found to interact with hNaa10 to inhibit hNaa10-mediated activation of β-catenin transcriptional activity.[11]
NatB
NatB complexes are composed of the catalytic subunit Naa20p and the auxiliary subunit Naa25p, which are both found in yeast and humans. In yeast, all the NatB subunits are ribosome-associated; but in humans, NatB subunits are both found to be ribosome-associated and non-ribosomal form. NatB acetylates the N-terminal methionine of substrates starting with Met-Glu-, Met-Asp-, Met-Asn- or Met-Gln- N termini.
NatC
NatC complex consists of one catalytic subunit Naa30p and two auxiliary subunits Naa35p and Naa38p. All three subunits are found on the ribosome in yeast, but they are also found in non-ribosomal NAT forms like Nat2. NatC complex acetylates the N-terminal methionine of substrates Met-Leu-, Met-Ile-, Met-Trp- or Met-Phe N-termini.
NatD
NatD is only composed with the catalytic unit Naa40p and Naa40p and it is conceptually different form the other NATs. At first, only two substrates, H2A and H4 have been identified in yeast and humans. Secondly, the substrate specificity of Naa40p lies within the first 30-50 residues which are quite larger than the substrate specificity of other NATs. The acetylation of histones by NatD is partially associate with ribosomes and the amino acids substrates are the very N-terminal residues, which makes it different from lysine N-acetyltransferases (KATs).[12]
NatE
NatE complex consists with subunit Naa50p and two NatA subunits, Naa10p and Naa15p. The N terminus of Naa50p substrates is different from those acetylated by the NatA activity of Naa10p.[13]
NatF
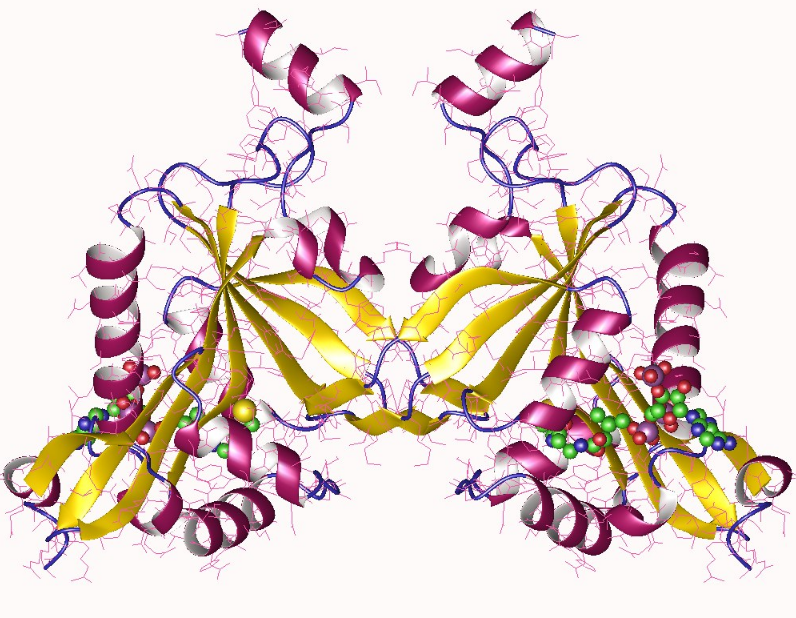
NatF is a NAT that is composed of the Naa60 enzyme. Initially, it was thought that NatF was only found in higher eukaryotes, since it was absent from yeast.[14] However, it was later found that Naa60 is found throughout the eukaryotic domain, but was secondarily lost in the fungi lineage.[15] Compared to yeast, NatF contributes to the higher abundance of N-terminal acetylation in humans. NatF complex acetylates the N-terminal methionine of substrates Met-Lys-, Met-Leu-, Met-Ile-, Met-Trp- and Met-Phe N termini which are partly overlapping with NatC and NatE.[6] NatF has been shown to have an organellar localization and acetylates cytosolic N-termini of transmembrane proteins.[16] The organellar localization of Naa60 is mediated by its unique C-terminus, which consists of two alpha helices that peripherally associate with the membrane and mediate interactions with PI(4)P.[17]
NAA80/NatH
NAA80/NatH is an N-terminal acetyltransferase that specifically acetylates the N-terminus of actin. [18]
N-terminal acetylation function
Protein stability
N-terminal acetylation of proteins can affect protein stability, but the results and mechanism were not very clear until now.[19] It was believed that N-terminal acetylation protects proteins from being degraded as Nα-acetylation N-termini were supposed to block N-terminal ubiquitination and subsequent protein degradation.[20] However, several studies have shown that the N-terminal acetylated protein have a similar degradation rate as proteins with a non-blocked N-terminus.[21]
Protein localization
N-terminal acetylation has been shown that it can steer the localization of proteins. Arl3p is one of the ‘Arf-like’ (Arl) GTPases, which is crucial for the organization of membrane traffic.[22] It requires its Nα-acetyl group for its targeting to the Golgi membrane by the interaction with Golgi membrane-residing protein Sys1p. If the Phe or Tyr is replaced by an Ala at the N-terminal of Arl3p, it can no longer localized to the Golgi membrane, indicating that Arl3p needs its natural N-terminal residues which could be acetylated for proper localization.[23]
Metabolism and apoptosis
Protein N-terminal acetylation has also been proved to relate with cell cycle regulation and apoptosis with protein knockdown experiments. Knockdown of the NatA or the NatC complex leads to the induction of p53-dependent apoptosis, which may indicate that the anti-apoptotic proteins were less or no longer functional because of reduced protein N-terminal acetylation.[24] But in contrast, the caspase-2, which is acetylated by NatA, can interact with the adaptor protein RIP associated Ich-1/Ced-3 homologous protein with a death domain (RAIDD). This could activate caspase-2 and induce cell apoptosis.[25]
Protein synthesis
Ribosome proteins play an important role in the protein synthesis, which could also be N-terminal acetylated. The N-terminal acetylation of the ribosome proteins may have an effect on protein synthesis. A decrease of 27% and 23% in the protein synthesis rate was observed with NatA and NatB deletion strains. A reduction of translation fidelity was observed in the NatA deletion strain and a defect in ribosome was noticed in the NatB deletion strain.[26]
Cancer
NATs have been suggested to act as both onco-proteins and tumor suppressors in human cancers, and NAT expression may be increased and decreased in cancer cells. Ectopic expression of hNaa10p increased cell proliferation and up regulation of gene involved in cell survival proliferation and metabolism. Overexpression of hNaa10p was in the urinary bladder cancer, breast cancer and cervical carcinoma.[27] But a high level expression of hNaa10p could also suppress tumor growth and a reduced level of expressed hNaa10p is associated with a poor prognosis, large tumors and more lymph node metastases.
Table 2. Overview of the expression of NatA subunits in various cancer tissues[28]
| Nat subunits | Cancer tissue | Expression pattern |
|---|---|---|
| hNaa10 | lung cancer, breast cancer, colorectal cancer, hepatocellular carcinoma | high in tumors |
| hNaa10 | lung cancer, breast cancer, pancreatic cancer, ovarian cancer | loss of heterozygosity in tumors |
| hNaa10 | breast cancer, gastric cancer, lung cancer | high in primary tumors, but low with lymph node metastases |
| hNaa10 | Non-small cell lung cancer | low in tumors |
| hNaa15 | papillary thyroid carcinoma, gastric cancer | high in tumors |
| hNaa15 | neuroblastoma | high in advanced stage tumors |
| hNaa11 | hepatocellular carcinoma | loss of heterozygosity in tumors |
1.2. Lysine Acetylation and Deacetylation
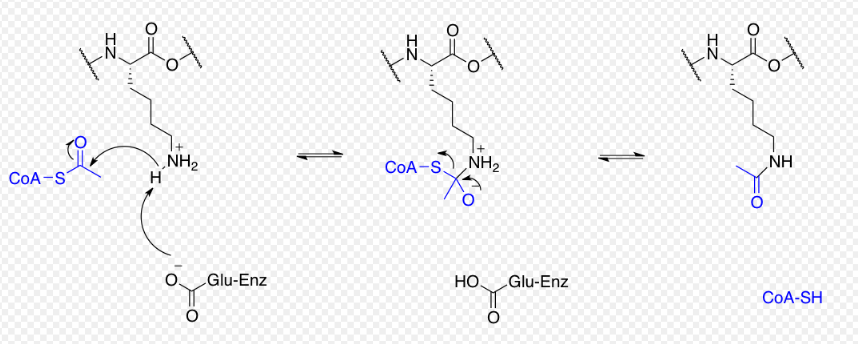
Proteins are typically acetylated on lysine residues and this reaction relies on acetyl-coenzyme A as the acetyl group donor. In histone acetylation and deacetylation, histone proteins are acetylated and deacetylated on lysine residues in the N-terminal tail as part of gene regulation. Typically, these reactions are catalyzed by enzymes with histone acetyltransferase (HAT) or histone deacetylase (HDAC) activity, although HATs and HDACs can modify the acetylation status of non-histone proteins as well.[29]
The regulation of transcription factors, effector proteins, molecular chaperones, and cytoskeletal proteins by acetylation and deacetylation is a significant post-translational regulatory mechanism[30] These regulatory mechanisms are analogous to phosphorylation and dephosphorylation by the action of kinases and phosphatases. Not only can the acetylation state of a protein modify its activity but there has been recent suggestion that this post-translational modification may also crosstalk with phosphorylation, methylation, ubiquitination, sumoylation, and others for dynamic control of cellular signaling.[31] The regulation of tubulin protein is an example of this in mouse neurons and astroglia.[32][33] A tubulin acetyltransferase is located in the axoneme, and acetylates the α-tubulin subunit in an assembled microtubule. Once disassembled, this acetylation is removed by another specific deacetylase in the cell cytosol. Thus axonemal microtubules, which have a long half-life, carry a "signature acetylation," which is absent from cytosolic microtubules that have a shorter half-life.
In the field of epigenetics, histone acetylation (and deacetylation) have been shown to be important mechanisms in the regulation of gene transcription. Histones, however, are not the only proteins regulated by posttranslational acetylation. The following are examples of various other proteins with roles in regulating signal transduction, whose activities are also affected by acetylation and deacetylation.
p53
The p53 protein is a tumor suppressor that plays an important role in the signal transactions in cells, especially in maintaining the stability of the genome by preventing mutation. Therefore, it is also known as “the guardian of the genome". It also regulates the cell cycle and arrests cell growth by activating a regulator of the cell cycle, p21. Under severe DNA damage, it also initiates programmed cell death.The function of p53 is negatively regulated by oncoprotein Mdm2. Studies suggested that Mdm2 will form a complex with p53 and prevent it from binding to specific p53-responsive genes.[34][35]
Acetylation of p53
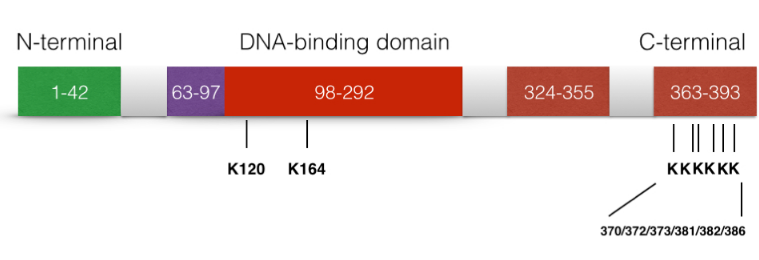
The acetylation of p53 is indispensable for its activation. It has been reported that the acetylation level of p53 will increase significantly when cell undergo stress. There are three major acetylation sites on p53: K164, K120 and C terminus.[36] If only one of the acetylation sites is defected, the activation of p21 is still observed. However, if all three activation sites are blocked, the activation of p21 and the suppression of cell growth caused by p53 will be completely lost. In addition, the acetylation of p53 prevents its binding to the repressor Mdm2 on DNA.[37] In addition, it is suggested that the p53 acetylation is crucial for its transcription-independent proapoptotic functions.[38]
Implications for cancer therapy
Since the major function of p53 is tumor suppressor, the idea that activation of p53 is an appealing strategy for cancer treatment. Nutlin-3[39] is a small molecule designed to target p53 and Mdm2 interaction that kept p53 from deactivation.[40] Reports also shown that the cancer cell under the Nutilin-3a treatment, acetylation of lys 382 was observed in the c-terminal of p53.[41][42]
Microtubule
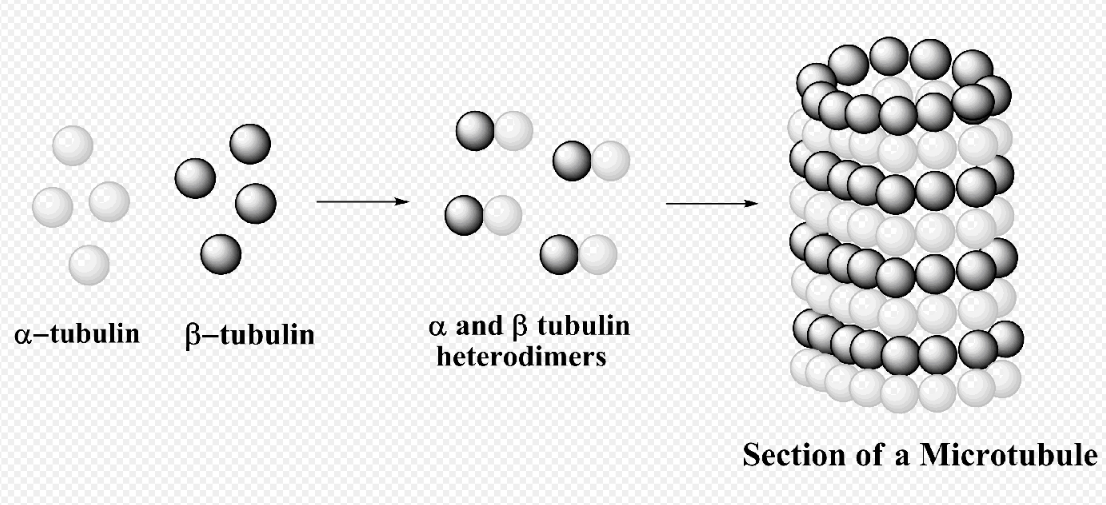
The structure of microtubules is long, hollow cylinder dynamically assembled from α/β-tubulin dimers. They play an essential role in maintaining the structure of the cell as well as cell processes, for example, movement of organelles.[43] In addition, microtubule is responsible of forming mitotic spindle in eukaryotic cells to transport chromosomes in cell division.[44][45]
Acetylation of tubulin
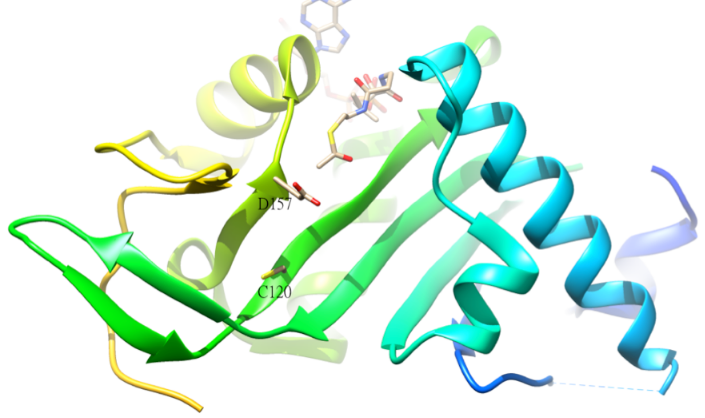
The acetylated residue of α-tubulin is K40, which is catalyzed by α-tubulin acetyl-transferase (α-TAT) in human. The acetylation of K40 on α-tubulin is a hallmark of stable microtubules. The active site residues D157 and C120 of α-TAT1 are responsible for the catalysis because of the shape complementary to α-Tubulin. In addition, some unique structural features such as β4-β5 hairpin, C-terminal loop, and α1-α2 loop regions are important for specific α-Tubulin molecular recognition.[46] The reverse reaction of the acetylation is catalyzed by histone deacetylase 6.[47]
Implications for cancer therapy
Since microtubules play an important role in cell division, especially in the G2/M phase of the cell cycle, attempts have been made to impede microtubule function using small molecule inhibitors, which have been successfully used in clinics as cancer therapies.[48] For example, the vinca alkaloids and taxanes selectively bind and inhibit microtubules, leading to cell cycle arrest.[49] The identification of the crystal structure of acetylation of α-tubulin acetyl-transferase (α-TAT) also sheds a light on the discovery of small molecule that could modulate the stability or de-polymerization of tubulin. In other words, by targeting α-TAT, it is possible to prevent the tubulin from acetylation and result in the destabilization of tubulin, which is a similar mechanism for tubulin destabilizing agents.[46]
STAT3
Signal transducer and activator of transcription 3 (STAT3) is a transcription factor that is phosphorylated by receptor associated kinases, for example, Janus-family tyrosine kinases, and translocate to nucleus. STAT3 regulates several genes in response to growth factors and cytokines and play an important role in cell growth. Therefore, STAT3 facilitates oncogenesis in a variety of cell growth related pathways. On the other hand, it also play a role in the tumor suppressor.[50]
Acetylation of STAT3
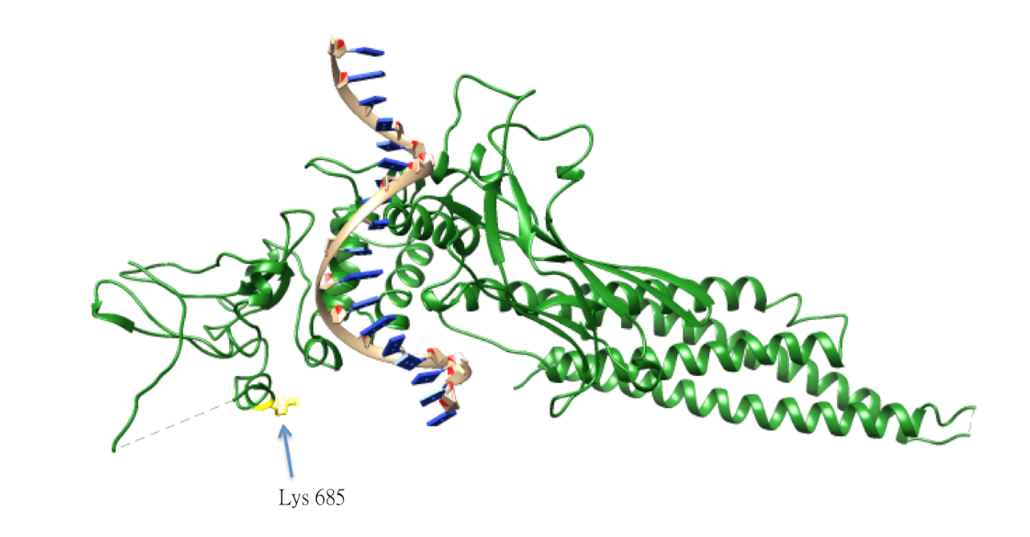
The acetylation of Lys685 of STAT3 is important for STAT3 homo-dimerization, which is essential for the DNA-binding and the transcriptional activation of oncogenes. The acetylation of STAT3 is catalyzed by histone acetyltransferase p300, and reversed by type 1 histone deacetylase. The lysine acetylation of STAT3 is also elevated in cancer cells.[51]
Therapeutic implications for cancer therapy
Since the acetylation of STAT3 is important for its oncogenic activity and the fact that the level of acetylated STAT3 is high in cancer cells, it is implied that targeting acetylated STAT3 for chemoprevention and chemotherapy is a promising strategy. This strategy is supported by treating resveratrol, an inhibitor of acetylation of STAT3, in cancer cell line reverses aberrant CpG island methylation.[52]
2. Acetylation of Wood
Since the beginning of the 20th century, acetylation of wood was researched as a method to upgrade the durability of wood in resistance against rotting processes and molds. Secondary benefits include the improvement of dimensional stability, improved surface hardness, and no decrease in mechanical properties due to the treatment.
The physical properties of any material are determined by its chemical structure. Wood contains an abundance of chemical groups called free hydroxyls. Free hydroxyl groups adsorb and release water according to changes in the climatic conditions to which the wood is exposed. This is an explanation as to why wood swells and shrinks. It is also believed that the digestion of wood by enzymes initiates at the free hydroxyl sites – which is one of the principal reasons why wood is prone to decay.
Acetylation changes the free hydroxyls within the wood into acetyl groups. This is done by reacting the wood with acetic anhydride, which comes from acetic acid (known as vinegar when in its dilute form). When the free hydroxyl group is transformed to an acetyl group, the ability of the wood to absorb water is greatly reduced, rendering the wood more dimensionally stable and, because it is no longer digestible, extremely durable.
In 2007, Titan Wood, a London-based company, with production facilities in The Netherlands, achieved cost-effective commercialization and began large-scale production of acetylated wood under the trade name "Accoya".
References
- "Lysine acetylation targets protein complexes and co-regulates major cellular functions". Science 325 (5942): 834–840. 2009. doi:10.1126/science.1175371. PMID 19608861. Bibcode: 2009Sci...325..834C. https://dx.doi.org/10.1126%2Fscience.1175371
- "Mitochondrial acetylome analysis in a mouse model of alcohol-induced liver injury utilizing SIRT3 knockout mice". J. Proteome Res. 11 (3): 1633–1643. 2012. doi:10.1021/pr2008384. PMID 22309199. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3324946
- Brook, Tom. "Protein Acetylation: Much More than Histone Acetylation". Cayman Chemical. Archived from the original on 2014-02-28. https://web.archive.org/web/20140228141824/https://www.caymanchem.com/app/template/Article.vm/article/2152.
- "Regulation of cellular metabolism by protein lysine acetylation". Science 327 (5968): 1000–1004. 2010. doi:10.1126/science.1179689. PMID 20167786. Bibcode: 2010Sci...327.1000Z. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3232675
- Wang, Qijun; Zhang, Yakun; Yang, Chen; Xiong, Hui; Lin, Yan; Yao, Jun; Li, Hong; Xie, Lu et al. (2010-02-19). "Acetylation of Metabolic Enzymes Coordinates Carbon Source Utilization and Metabolic Flux" (in en). Science 327 (5968): 1004–1007. doi:10.1126/science.1179687. ISSN 0036-8075. PMID 20167787. Bibcode: 2010Sci...327.1004W. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4183141
- "NatF contributes to an evolutionary shift in protein N-terminal acetylation and is important for normal chromosome segregation". PLOS Genet. 7 (7): e1002169. 2011. doi:10.1371/journal.pgen.1002169. PMID 21750686. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3131286
- "Protein N-terminal acetyltransferases: when the start matters". Trends Biochem. Sci. 37 (4): 152–161. 2012. doi:10.1016/j.tibs.2012.02.003. PMID 22405572. https://dx.doi.org/10.1016%2Fj.tibs.2012.02.003
- "Molecular basis for N-terminal acetylation by the heterodimeric NatA complex". Nat. Struct. Mol. Biol. 20 (9): 1098–105. 2013. doi:10.1038/nsmb.2636. PMID 23912279. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3766382
- "Composition and biological significance of the human Nalpha-terminal acetyltransferases". BMC Proceedings 3 Suppl 6 (Suppl 6): S3. 2009. doi:10.1186/1753-6561-3-s6-s3. PMID 19660096. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2722096
- "Proteomics analyses reveal the evolutionary conservation and divergence of N-terminal acetyltransferases from yeast and humans". Proc. Natl. Acad. Sci. U.S.A. 106 (20): 8157–8162. 2009. doi:10.1073/pnas.0901931106. PMID 19420222. Bibcode: 2009PNAS..106.8157A. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2688859
- "The chaperone-like protein HYPK acts together with NatA in cotranslational N-terminal acetylation and prevention of Huntingtin aggregation". Mol. Cell. Biol. 30 (8): 1898–1909. 2010. doi:10.1128/mcb.01199-09. PMID 20154145. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2849469
- "The human N-alpha-acetyltransferase 40 (hNaa40p/hNatD) is conserved from yeast and N-terminally acetylates histones H2A and H4". PLOS ONE 6 (9): e24713. 2011. doi:10.1371/journal.pone.0024713. PMID 21935442. Bibcode: 2011PLoSO...624713H. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3174195
- "The yeast N(alpha)-acetyltransferase NatA is quantitatively anchored to the ribosome and interacts with nascent polypeptides". Mol. Cell. Biol. 23 (20): 7403–7414. 2003. doi:10.1128/mcb.23.20.7403-7414.2003. PMID 14517307. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=230319
- "NatF contributes to an evolutionary shift in protein N-terminal acetylation and is important for normal chromosome segregation.". PLOS Genet. 7 (7): e1002169. July 2011. doi:10.1371/journal.pgen.1002169. PMID 21750686. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3131286
- Rathore, Om Singh; Faustino, Alexandra; Prudêncio, Pedro; Van Damme, Petra; Cox, Cymon J.; Martinho, Rui Gonçalo (2016). "Absence of N-terminal acetyltransferase diversification during evolution of eukaryotic organisms.". Scientific Reports 6: 21304. doi:10.1038/srep21304. PMID 26861501. Bibcode: 2016NatSR...621304R. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4748286
- Aksnes (March 3, 2015). "An Organellar Nα-Acetyltransferase, Naa60, Acetylates Cytosolic N Termini of Transmembrane Proteins and Maintains Golgi Integrity". Cell Reports 10 (8): 1362–74. doi:10.1016/j.celrep.2015.01.053. PMID 25732826. https://dx.doi.org/10.1016%2Fj.celrep.2015.01.053
- Aksnes, Henriette; Goris, Marianne; Strømland, Øyvind; Drazic, Adrian; Waheed, Qaiser; Reuter, Nathalie; Arnesen, Thomas (2017). "Molecular determinants of the N-terminal acetyltransferase Naa60 anchoring to the Golgi membrane.". Journal of Biological Chemistry 292 (16): 6821–6837. doi:10.1074/jbc.M116.770362. PMID 28196861. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5399128
- "NAA80 is actin's N-terminal acetyltransferase and regulates cytoskeleton assembly and cell motility". Proc Natl Acad Sci U S A 115 (17): 4399–4404. 2018. doi:10.1073/pnas.1718336115. PMID 29581253. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5924898
- "N-terminal acetylation and other functions of Nα-acetyltransferases". Biol. Chem. 393 (4): 291–8. 2012. doi:10.1515/hsz-2011-0228. PMID 22718636. https://dx.doi.org/10.1515%2Fhsz-2011-0228
- "Role of the alpha-amino group of protein in ubiquitin-mediated protein breakdown". Proc. Natl. Acad. Sci. U.S.A. 81 (22): 7021–5. 1984. doi:10.1073/pnas.81.22.7021. PMID 6095265. Bibcode: 1984PNAS...81.7021H. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=392068
- "N-terminal acetylation of cellular proteins creates specific degradation signals". Science 327 (5968): 973–977. 2010. doi:10.1126/science.1183147. PMID 20110468. Bibcode: 2010Sci...327..973H. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4259118
- "Targeting of the Arf-like GTPase Arl3p to the Golgi requires N-terminal acetylation and the membrane protein Sys1p". Nat. Cell Biol. 6 (5): 405–413. 2004. doi:10.1038/ncb1120. PMID 15077113. https://dx.doi.org/10.1038%2Fncb1120
- "Knockdown of human N alpha-terminal acetyltransferase complex C leads to p53-dependent apoptosis and aberrant human Arl8b localization". Mol. Cell. Biol. 29 (13): 3569–3581. 2009. doi:10.1128/mcb.01909-08. PMID 19398576. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2698767
- "Depletion of the human Nα-terminal acetyltransferase A induces p53-dependent apoptosis and p53-independent growth inhibition". Int. J. Cancer 127 (12): 2777–2789. 2010. doi:10.1002/ijc.25275. PMID 21351257. https://dx.doi.org/10.1002%2Fijc.25275
- "Metabolic regulation of protein N-alpha-acetylation by Bcl-xL promotes cell survival". Cell 146 (4): 607–620. 2011. doi:10.1016/j.cell.2011.06.050. PMID 21854985. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3182480
- "N(α)-Acetylation of yeast ribosomal proteins and its effect on protein synthesis". J Proteomics 74 (4): 431–441. 2011. doi:10.1016/j.jprot.2010.12.007. PMID 21184851. https://dx.doi.org/10.1016%2Fj.jprot.2010.12.007
- "Immunohistochemical analysis of human arrest-defective-1 expressed in cancers in vivo". Oncol. Rep. 21 (4): 909–15. 2009. doi:10.3892/or_00000303. PMID 19287988. https://dx.doi.org/10.3892%2For_00000303
- "Protein N-terminal acetyltransferases in cancer". Oncogene 32 (3): 269–276. 2013. doi:10.1038/onc.2012.82. PMID 22391571. https://dx.doi.org/10.1038%2Fonc.2012.82
- "Regulation of protein turnover by acetyltransferases and deacetylases". Biochimie 90 (2): 306–12. 2008. doi:10.1016/j.biochi.2007.06.009. PMID 17681659. https://dx.doi.org/10.1016%2Fj.biochi.2007.06.009
- "Acetylation and deacetylation of non-histone proteins". Gene 363: 15–23. 2005. doi:10.1016/j.gene.2005.09.010. PMID 16289629. https://dx.doi.org/10.1016%2Fj.gene.2005.09.010
- "Lysine acetylation: codified crosstalk with other posttranslational modifications". Mol. Cell 31 (4): 449–61. 2008. doi:10.1016/j.molcel.2008.07.002. PMID 18722172. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2551738
- "Posttranslational modifications of tubulin in cultured mouse brain neurons and astroglia". Biol. Cell 65 (2): 109–117. 1989. doi:10.1016/0248-4900(89)90018-x. PMID 2736326. https://dx.doi.org/10.1016%2F0248-4900%2889%2990018-x
- "The acetylation of alpha-tubulin and its relationship to the assembly and disassembly of microtubules". J. Cell Biol. 103 (2): 571–579. 1986. doi:10.1083/jcb.103.2.571. PMID 3733880. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2113826
- Alberts, Bruce (March 2002). Molecular Biology of the Cell. Garland Science. ISBN 0815332181.
- Weinberg, Robert A. (2013). Biology of cancer. (2. ed.). [S.l.]: Garland Science. ISBN 978-0815342205.
- "The impact of acetylation and deacetylation on the p53 pathway". Protein Cell 2 (6): 456–462. 2011. doi:10.1007/s13238-011-1063-9. PMID 21748595. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3690542
- "Acetylation is indispensable for p53 activation". Cell 133 (4): 612–626. 2008. doi:10.1016/j.cell.2008.03.025. PMID 18485870. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2914560
- "p53 acetylation is crucial for its transcription-independent proapoptotic functions". J. Biol. Chem. 284 (17): 11171–11183. 2009. doi:10.1074/jbc.M809268200. PMID 19265193. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2670122
- "In vivo activation of the p53 pathway by small-molecule antagonists of MDM2". Science 303 (5659): 844–848. 2004. doi:10.1126/science.1092472. PMID 14704432. Bibcode: 2004Sci...303..844V. https://dx.doi.org/10.1126%2Fscience.1092472
- "Small-molecule inhibitors of the MDM2-p53 protein-protein interaction to reactivate p53 function: a novel approach for cancer therapy". Annu. Rev. Pharmacol. Toxicol. 49 (1): 223–241. 2009. doi:10.1146/annurev.pharmtox.48.113006.094723. PMID 18834305. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2676449
- "Nutlin-3a, an MDM2 antagonist and p53 activator, helps to preserve the replicative potential of cancer cells treated with a genotoxic dose of resveratrol". Mol. Biol. Rep. 40 (8): 5013–5026. 2013. doi:10.1007/s11033-013-2602-7. PMID 23666059. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3723979
- "Nutlin-3a activates p53 to both down-regulate inhibitor of growth 2 and up-regulate mir-34a, mir-34b, and mir-34c expression, and induce senescence". Cancer Res. 68 (9): 3193–3203. 2008. doi:10.1158/0008-5472.CAN-07-2780. PMID 18451145. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2440635
- Kreis, ed. by Thomas; Vale, Ronald (1999). Guidebook to the cytoskeletal and motor proteins (2. ed.). Oxford [u.a.]: Oxford Univ. Press. ISBN 0198599560.
- al.], Harvey Lodish ... [et (2013). Molecular cell biology (7th ed.). New York: W.H. Freeman and Co.. ISBN 978-1429234139.
- Fojo, edited by Tito (2008). The role of microtubules in cell biology, neurobiology, and oncology ([Online-Ausg.] ed.). Totowa, N. J.: Humana Press. ISBN 978-1588292940. https://archive.org/details/microtubuletarge00anto.
- "Structure of the α-tubulin acetyltransferase, αTAT1, and implications for tubulin-specific acetylation". Proc. Natl. Acad. Sci. U.S.A. 109 (48): 19655–19660. 2012. doi:10.1073/pnas.1209357109. PMID 23071314. Bibcode: 2012PNAS..10919655F. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3511727
- "HDAC6 is a microtubule-associated deacetylase". Nature 417 (6887): 455–458. 2002. doi:10.1038/417455a. PMID 12024216. Bibcode: 2002Natur.417..455H. https://dx.doi.org/10.1038%2F417455a
- Teresa Carlomagno, ed (2009). Tubulin-binding agents : synthetic, structural, and mechanistic insights. contributions by K.-H. Altmann. Berlin: Springer. ISBN 978-3540690368.
- Zito, edited by Thomas L. Lemke, David A. Williams; associate editors, Victoria F. Roche, S. William (2013). Foye's principles of medicinal chemistry (7th ed.). Philadelphia: Wolters Kluwer Health/Lippincott Williams & Wilkins. ISBN 978-1609133450.
- Müller-Decker, Friedrich Marks, Ursula Klingmüller, Karin (2009). Cellular signal processing : an introduction to the molecular mechanisms of signal transduction. New York: Garland Science. ISBN 978-0815342151.
- "Stat3 dimerization regulated by reversible acetylation of a single lysine residue". Science 307 (5707): 269–273. 2005. doi:10.1126/science.1105166. PMID 15653507. Bibcode: 2005Sci...307..269Y. https://dx.doi.org/10.1126%2Fscience.1105166
- "Acetylated STAT3 is crucial for methylation of tumor-suppressor gene promoters and inhibition by resveratrol results in demethylation". Proc. Natl. Acad. Sci. U.S.A. 109 (20): 7765–7769. 2012. doi:10.1073/pnas.1205132109. PMID 22547799. Bibcode: 2012PNAS..109.7765L. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3356652





