
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Beatrix Zheng | -- | 3941 | 2022-11-16 01:43:13 | | | |
| 2 | Beatrix Zheng | -50 word(s) | 3891 | 2022-11-16 04:07:51 | | |
Video Upload Options
The Planctomycetes are a phylum of widely distributed bacteria, occurring in both aquatic and terrestrial habitats. They play a considerable role in global carbon and nitrogen cycles, with many species of this phylum capable of anaerobic ammonium oxidation, also known as anammox. Many planctomycetes occur in relatively high abundance as biofilms, often associating with other organisms such as macroalgae and marine sponges. Planctomycetes are included in the PVC superphylum along with Verrucomicrobia, Chlamydiae, Lentisphaerae, Kiritimatiellaeota, and Candidatus Omnitrophica. The phylum Planctomycete is composed of the classes Planctomycetia and Phycisphaerae. First described in 1924, members of the Planctomycetes were identified as eukaryotes and were only later described as bacteria in 1972. Early examination of members of the Planctomycetes suggested a cell plan differing considerably from other bacteria, although they are now confirmed as Gram-negative bacteria, but with many unique characteristics. Bacteria in the Planctomycetes are often small, spherical cells, but a large amount of morphological variation is seen. Members of the Planctomycetes also display distinct reproductive habits, with many species dividing by budding, in contrast to all other free-living bacteria, which divide by binary fission. Interest is growing in the Planctomycetes regarding biotechnology and human applications, mainly as a source of bioactive molecules. In addition, some Planctomycetes were recently described as human pathogens. The species Gemmata obscuriglobus has been identified specifically as comprising bacteria with unique characteristics among the Planctomycetes, such as their ability to synthesize sterols.
1. Structure and Morphology
1.1. Cell shape and Appendages

The distinct morphological characteristics of bacteria in the Planctomycetes have been discussed extensively.[2] The common morphology is often spherical cells roughly 2 μm in diameter, as observed in the species Aquisphaera giovannonii. However, the diversity in cell shape often varies greatly in them. Ovoid and pear-shaped cells have been described in some species, and often occur in rosettes of three to 10 cells.[3] Gemmata obscuriglobus is a well studied species in the Planctomycetes with spherical cells. In contrast, bacteria in the species Planctopirus limnophila have ovoid cells.[4]
Many Planctomycetes species display structures and appendages on the outer surface of the cell. Flagella, common in most bacteria, have also been observed in the species P. limnophila.[3][5][6] Many Planctomycetes also have a holdfast, or stalk, which attaches the cell to a surface or substrate.[5][6] Members of some species, though, such as Isosphaera pallida lack a holdfast.[5]
Unique appendages known as crateriform structures have been observed[3][5][6] in species of Planctomycetes belonging to the class Planctomycetia.[7] The outer surface of cells in the species P. limnophila display both large and small crateriform structures. Large crateriform structures often cover the cell surface, while small crateriform structures are often only at the end of the cell. Light microscopy demonstrated fibers of both stalk and pili type in P. limnophila and G. obscuriglobus. The pili fibers in both these species were often associated with large crateriform structures; in contrast, the stalk fibers were associated with small crateriform structures.[6]
1.2. Cell Wall Composition
Early examination of the Planctomycetes suggested that their cell plan differed considerably from both Gram-positive and Gram-negative bacteria.[5] Until recently, bacteria in the Planctomycetes were thought to lack peptidoglycans in their cell walls, and were instead suggested to have proteinaceous cell walls. Peptidoglycan is an essential polymer of glycans, present in all free-living bacteria, and its rigidity helps maintain integrity of the cell. Peptidoglycan synthesis is also essential during cell division. Recently, those in the species G. obscuriglobus were found to have peptidoglycan in their cell walls.[5][6]
1.3. Internal Cell Composition
Planctomycetes were once thought to display distinct compartmentalization within the cytosol.[5][6] Three-dimensional electron tomography reconstruction of G. obscuriglobus displayed varying interpretations of this suggested compartmentalization.[1] The cytosol was suggested to be separated into compartments, both the paryphoplasm and pirellulosome, by an intracytoplasmic membrane. This interpretation has since been demonstrated to be incorrect. In fact, the intracytoplasmic membrane is well known to be the cytoplasmic membrane which displays unique invaginations, giving the appearance of compartmentalization within the cytosol.[1][5][6] Planctomycetes therefore display the two compartments typical of Gram-negative bacteria, the cytoplasm and periplasm.
The excess membrane observed in G. obscuriglobus triples the surface area of the cell relative to its volume, which is suggested to be associated with sterol synthesis.[1]
1.4. Pigments
Many Planctomycetes spp. display pink or orange coloring, suggested to result from the production of carotenoid pigments. Carotenoids are produced by plants and fungi, and by some heterotrophic bacteria to protect against oxidative stress. Three different carotenoid pigments have been identified in two different strains of the Planctomycetes.[8]
In marine environments, Planctomycetes are often suspended in the water column or present as biofilms on the surface of macroalgae, and are often exposed to harmful ultraviolet radiation. More highly pigmented species of the Planctomycetes are more resistant to ultraviolet radiation, although this is not yet well understood.[9]
1.5. Unique Characteristics of Anammox Cells
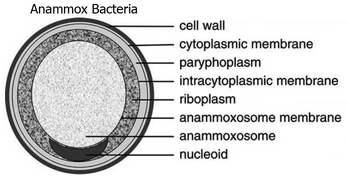
Bacteria in the Planctomycetes that are anammox-capable form the order Brocadiales.[10] The cells of anammox bacteria are often coccoid with a diameter of about 0.8 μm,[11] and are suggested to contain three compartments, each surrounded by a membrane. The outer membrane encloses the cell and the protoplasm and the innermost membrane surrounds the anammoxosome, the central structure of anammox bacteria.[6][12] Anammox membrane composition resembles that of all other living organisms, and is composed of glycolipid.[12]
2. Life History and Reproduction
2.1. Growth
Planctomycetes spp. grow slowly, when compared to other bacteria,[5][11][13][14] often forming rosette structures of 3-5 cells.[5][14] Planctomycetes in the species P. limnophila is suggested to be relatively fast growing,[5][15] with a doubling time of roughly 6-14 days. In contrast, some other planctomycetes have doubling times of around 30 days.[15] Their high abundance in many ecosystems is surprising, given their slow growth rates.[11][13]
2.2. Lifecycle
Planctomycetes often perform a lifestyle switch between both a sessile stalked stage and a free-swimming stage.[14] Members of the species P. limnophila perform a lifestyle switch that is often associated with cell division. The sessile mother cell produces a free-swimming daughter cell. The daughter cell must then attach to a surface before starting the cycle over again. However, not all of the Planctomycetes have a motile stage, and the lifestyle switch observed in many species may not be common among all Planctomycetes.[5]
2.3. Reproduction
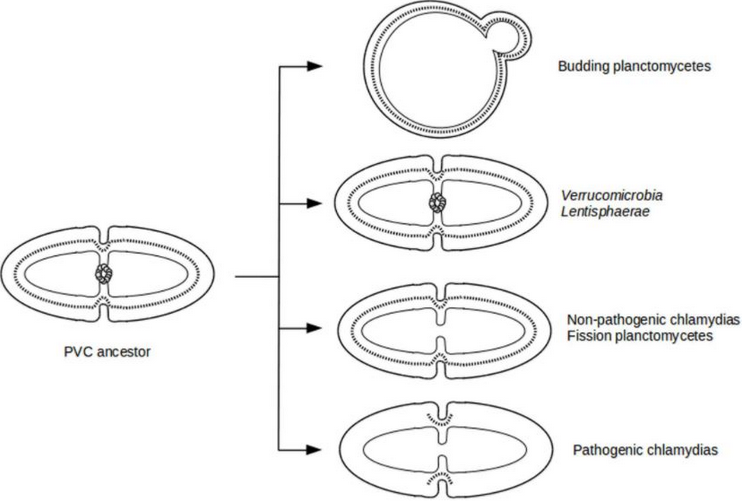
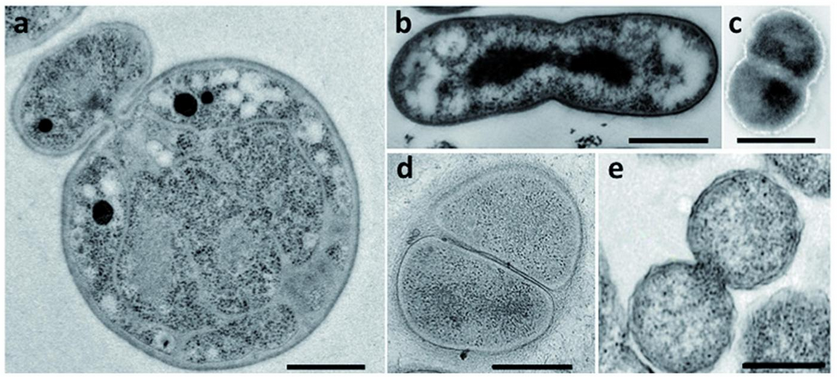
The current understanding of bacterial cell division is based on model organisms such as Escherichia coli.[4] The dominant form of reproduction observed in almost all bacteria is cell division by binary fission, which involves the synthesis of both peptidoglycans and proteins known as FtsZ.[4][16] In contrast, many bacteria in the Planctomycetes divide by budding.[5][7][17]
FtsZ proteins are suggested to be similar in structure to that of tubulin, the protein present in eukaryotes,[18] and is essential for septal formation during cell division.[2][5] The lack of FtsZ proteins is often lethal.[5] Peptidoglycan also play a considerable role in cell division by binary fission.[16]
Planctomycetes is one of the only known phyla whose members lack FtsZ proteins.[5][16][18] Bacteria in the Chlamydiales, also a member of the PVC superphylum, also lack FtsZ.[18] Although bacteria in the Planctomycetes lack FtsZ, two distinct modes of cell division have been observed.[5] Most Planctomycetes divide by binary fission, mainly species of the class Phycisphaerae. In contrast, species of the class Planctomycetia divide by budding.[5][7][17]
The mechanisms involved in budding have been described extensively for yeast cells. However, bacterial budding observed in planctomycetes is still poorly understood.[4] Budding has been observed in both radial symmetric cells, such as bacteria in the species P. limnophila, and axially symmetric cells.[7] During cell division in members of P. limnophila, the daughter cells originate from the region opposite to the pole with the holdfast or stalk.
Considerable diversity has been observed in cell division among bacteria in the Planctomycetes.[7][17] During cell division in Fuerstia marisgermanicae, a tubular structure is connected from the bud to the mother cell.[5][10] The species Kolteria novifilia forms a distinct clade of Planctomycetes, and is the only known species to divide by lateral budding at the middle of the cell. Lastly, members of the clade Saltatorellus are capable of switching between both binary fission and budding.[7][17]
3. Genetic Characteristics
3.1. Molecular Signatures
Planctomycetes are known for their unusual cellular characteristics, and their distinctness from all other bacteria is additionally supported by the shared presence of two conserved signature indels (CSIs).[19] These CSIs demarcate the group from neighboring phyla within the PVC group.[20] An additional CSI has been found that is shared by all Planctomycetes species, with the exception of Kuenenia stuttgartiensis. This supports the idea that K. stuttgartiensis forms a deep branch within the Planctomycete phylum.
A CSI has also been found to be shared by the entire PVC group, including the Planctomycetes.[19][20] Planctomycetes also contain an important conserved signature protein that has been characterized to play an important housekeeping function that is exclusive to members belonging to the PVC superphylum.[21]
3.2. General Characteristics
The genome size of Rhodopirellula baltica has been estimated to be over 7 million bases, making it one of the largest prokaryotic genomes sequenced. Extensive genome duplication takes up about 25% of the genome sequence.[2] This may be a way for the organism to adapt to mutations, allowing for redundancy if a part of the genome is damaged. The polymerase chain reaction primer used often mismatches with the genes, creating difficulty when sequencing the genome.[22]
When comparing under a microscope, a defining characteristic for some Planctomycetes is that a single unlinked rRNA operon can be identified near the origin. The changes of genetic material is through internal chromosomal inversion, and not through lateral gene transfer. This creates a way of diversification in the Planctomycetes variants as multiple transposon genes in these regions have reverse orientation that transfers to rearrangements.
Some Planctomycetes thrive in regions containing highly concentrated nitrate,[2] and have genes that are required for heterotactic acid fermentation. The enzyme lactate dehydrogenase plays a key role in this process. The genetic process also has ultraviolet radiation protection response, and is associated with the genes recA, lexA, uvrA, uvrB, and uvrC, in addition to a photolyase gene that is expressed when the environment offers excessive ultraviolet radiation stress. Other stress responses include the decomposition of hydrogen peroxide and oxidation.
Many Planctomycetes also express sulfatase genes. The genome of Pirellula sp. strain 1 incorporates 110 genes that contribute to encoding proteins that produce sulfatase enzymes. In comparison with a different species of prokaryotic, Pseudomonas aeruginosa, only 6 sulfatases occur and the genes that express these proteins are contained as two to five pairs, usually clustered in 22 groups.[2]
3.3. Molecular Evolution
Planctomycetes originate from within the Bacteria and these similarities between proteins in Planctomycetales and eukaryotes reflect convergent evolution. Gained protein families in Gemmataceae, a subgroup within Planctomycetes, have low sequence similarity to eukaryotic proteins; however, they show highest sequence similarity to other Gemmataceae protein families.[23]
There is massive emergence of novel protein families within the Gemmataceae. More than one thousand protein families were acquired by duplications and domain rearrangements. The new paralogs function in signal transduction, regulatory systems, and protein interaction pathways. They are related to the functional organisation of the cell, which can be interpreted as an adaptation to a more complex lifestyle.[23] The protein length is longer in the Gemmataceae than in most other bacteria and the genes have linkers. There is an overlap between the longest proteins in Planctomycetales and the shortest proteins in eukaryotes. In the terms of gene paralogy, protein length, and protein domain structures, prokaryotes and eukaryotes do not have sharp boundaries.[23]
4. Phylogeny
Originally classified as a eukaryote due to morphology, the advent of genetic sequencing allowed researchers to agree that the Planctomycetes belong to the domain Bacteria.[5] Within that domain, Planctomycetes are classified as their own phylum, however, other researchers have argued they could also be categorized as part of a larger superphylum entitled PVC, which would encompass the phyla Verrucomicrobia, Chlamydiae and Lentisphaerae, and the candidate phylum Candidatus Omnitrophica.[22] Within this superphylum, its members have been found to be closely related through the creation of 16S rRNA trees. Both the Planctomycetes and Chlamydiae encode proteins for nucleotide transporters, and the Verrucomicrobia have also been found to have features common among eukaryotic cells. Thus, a common ancestor of this superphylum may have been the start of the eukaryotic lineage.[22] While this is one possible explanation, because PVC is not the start of the bacterial tree,[24] the existence of eukaryotic traits and genes is more likely explained through lateral gene transfer, and not a more recent eukaryotic ancestor.[22]
The phylogeny based on the work of the All-Species Living Tree Project:[25]

5. Ecology
5.1. Distribution and Abundance
Members of the Planctomycetes are found in a diverse range of environments, both geographically and ecologically,[26] and occur in both aquatic and terrestrial habitats.[5] In aquatic environments, they are found in both freshwater and marine systems.[26] Planctomycetes were originally believed to exist exclusively in aquatic environments, but they are noe known to be also abundant in soils[27] and hypersaline environments.[28] They are widespread on five continents, including Antarctica and Australia .[26][27]
Fluorescence in situ hybridization was used to detect Planctomycetes in various environments, and Planctomycetes are found in abundance in pphagnum bogs. Some Planctomycetes were found in the digestive systems of marine lifeforms, while others tend to live among eukaryotes.[22]
5.2. Environmental Influences on Distribution
Planctomycetes account for roughly 11% of prokaryotic communities in marine systems, and their vast distribution demonstrates their ability to inhabit many different environments. They can also adapt to both aerobic and anaerobic conditions. Many factors can affect their distribution, such as humidity, oxygen levels, and pH levels. Planctomycete diversity and abundance are strongly associated with relative humidity. The effects of oxygen levels demonstrate the energy needs of the individual. Many species of Planctomycetes are chemoheterotrophic, including G. obscuriglobus. Thermostilla marina, a thermophilic anaerobic species occupying hydrothermal vent regions, can use elemental sulfur to generate sulfide and respire with nitrate. Planctomycetes can also inhabit regions with ranges in pH levels from 4.2 to 11.6.[29]
5.3. Ecological Impacts and Global Carbon Cycle
Planctomycetes have a significant impact on global biogeochemistry and climate, with their ability to mineralize and break down detritus particles in the water column.[2][9]
Planctomycetes play a considerable role in the global carbon cycle.[2][5][7][30] As both obligate and facultative aerobic chemoheterotrophs, the primary source of carbon used by Planctomycetes is from carbohydrates. Many Planctomycetes have the ability to breakdown extremely complex carbohydrates, making these nutrients available to other organisms. This ability to recycle carbon has been linked to specific C1 metabolism genes observed in many Planctomycetes and are suggested to play a significant role, but this area of research is still poorly understood.
Planctomycetes also display many sulfatase enzymes, which are capable of breaking down sulfated heteropolysaccharides, which are produced by many groups of macroalgae. The breakdown of these sulfated heteropolysaccharides by Planctomycetes are then used as an energy source. Some Planctomycetes are suggested to be capable of breaking down carrageenan.[30]

5.4. Association with Other Organisms
Planctomycetes have often been observed in association with many organisms, including, macroalgae, microalgae, marine sponges, and plants such as lichens and bryophytes.[29] They have also been observed inhabiting deep-sea cold seeps, where they are dominant organisms living on tube worms.[5]
5.5. Macroalgae

Planctomycetes are often associated with marine surfaces high in nutrients. They occur as biofilms on algal surfaces in relatively high abundance.[11] Macroalgae such as the kelps Laminaria hyperborea and Ecklonia radiata are suggested to be an important habitat for Planctomycetes.[5][31] Roughly 70% of the bacterial community on Ecklonia radiata were Planctomycetes.[5][13] Almost 150 Planctomycetes species have been isolated from the biofilms of macroalgae, and these communities associated with macroalgae are mainly independent of changes in geographical distribution. This would suggest a symbiotic relationship.[29]
Kelp forests dominate the rocky coastlines of temperature regions, and provide habitat, shelter, and food for many organisms, including the Planctomycetes.[5] Given the considerable role of kelp forests in coastal primary productivity, the association of the Planctomycetes with kelp could indicate their significant role in coastal habitats.[32] Planctomycetes also play an important role as components of detritus in the water column, also known as marine snow,[5][32] given their ability to attach to surfaces.[33]
As the climate continues to warm, the abundance of Planctomycetes associated with macroalgae might increase. The seaweed Caulerpa taxifolia was incubated under higher CO2 conditions, and the abundance of Planctomycetes increased substantially, as much as 10 times in some species.[5]
5.6. Microalgae and Diatom Blooms
While macroalgae are well known substrates for Planctomycetes communities, their abundance has also been known to correlate with blooms of microalgae such as diatoms.[5][32] Blooms of cyanobacteria, diatoms, and dinoflagellates provide nutrients for Planctomycetes, which could explain the association.[29]
5.7. Marine Sponges
Planctomycetes spp. are often associated with the surfaces of marine sponges.[29][33] They interact with sponges either by attachment with a holdfast, or through a symbiotic relationship. A high diversity of Planctomycetes is present as biofilms on sponges. The symbiotic relationship among sponges and Planctomycetes contributes to the health of the sponge, and the sponge often provides suitable habitat and nutrients to the Planctomycetes.[29]
5.8. Lichen Communities and Sphagnum Bogs
Planctomycetes were found to be highly abundant in lichen communities throughout northwestern Siberia and displayed extremely high diversity. Planctomycetes have also been associated with lichen communities and Sphagnum wetlands. Sphagnum wetlands store large amounts of carbon, contributing to the global carbon cycle. Planctomycetes play a considerable role in the degradation of sphagnum, accounting for roughly 15% of the bacterial community.[29]
5.9. Other Bacterial Communities
Planctomycetes display associations with other bacterial communities, mainly Alphaproteobacteria, Bacteroidetes, Gemmatimonadetes, and Verrucomicrobia. The growth of many Planctomycetes is often supported by the essential nutrients provided by other bacteria within the community, and some Planctomycetes rely strongly on symbiotic relationships with other bacteria.[29]
6. Physiology
6.1. Endocytosis
The existence of membrane coat proteins near the intracytoplasmic membrane could be used for an endocytosis-like uptake system, which would be the first instance this function has been found outside of the eukaryotic domain. However, now that the existence of a rigid peptidoglycan cell wall has been confirmed, these vesicles to be able to pass through this cell wall seems unlikely. Additionally, deletion of one of these membrane coat proteins within P. limnophila found no decrease in macromolecule uptake.[6] In addition, with the use of cryoelectron tomography-based three-dimensional reconstruction of Planctomycetes has found that what were originally thought to be vesicles being held in the periplasm are actually just folds in the cytoplasmic membrane.[5] Yet it has been demonstrated that the Planctomycetes can survive on high-molecular-weight polysaccharides as their only source of carbon, meaningthey must have the ability to incorporate complex carbon substrates into their cytoplasm. Three hypotheses have been put forth: First, the Planctomycetes excrete an enzyme which, outside of the cell wall, degrades the complex substrates into smaller monosaccharides, which can more easily be transported through the different membranes. Second, the complex substrates become anchored to the outside of Planctomycetes, which are then able to slowly break down these substrates into oligosaccharides, which are able to be transported into the periplasm of Planctomycetes by specialized proteins. The third hypothesis involves the crateriform structures found on the outside of Planctomycete cell walls. These structures have fibers lining their pits that may be able to absorb whole polysaccharides into the periplasm, where they would then be digested.[6]
6.2. Osmotic Regulation
Almost all bacteria have a cytosol following the outer shape of their peptidoglycan cell wall. Eukaryotes are different in that they have their cytosol divided into multiple compartments to create organelles such as a nucleus. Planctomycetes are unique in that they have large invaginations of their cytoplasmic membrane, pulling away from the peptidoglycan cell wall and leaving room for the periplasm. Traditionally, the cytoplasmic membrane has been thought to be responsible for controlling the osmotic pressure of bacterial cells. Yet due to the folds in the cytoplasmic membrane, and the existence of large spaces of periplasm within Planctomycetes, their peptidoglycan acts as an osmotic barrier with the periplasm being isotonic to the cytosol.[5]
6.3. Anaerobic Ammonium Oxidation (Anammox)

Anammox is the process of oxidizing ammonium where nitrite acts as the electron acceptor. This process creates energy for the organism performing the reaction in the same way humans gain energy from oxidizing glucose.[34] In a marine environment, this ultimately removes nitrogen from the water, as N2 gas cannot be used by phytoplankton and is released into the atmosphere. Up to 67% of dinitrogen gas production in the ocean can be attributed to anammox[35] and about 50% of the nitrogen gas in the atmosphere is thought to be produced from anammox.[36] Planctomycetes are the most dominant phylum of bacteria capable of performing anammox, thus the Planctomycetes capable of performing anammox play an important role in the global cycling of nitrogen.[37]
6.4. Sterol Synthesis
The synthesis of sterols, often observed in eukaryotes and uncommon among bacteria, has been observed very rarely in Planctomycetes.[4][5] The synthesis of sterols such as lanosterol has been observed in G. obscuriglobus. Lanosterol is common in eukaryotes and two other groups of bacteria, both methylotrophic proteobacteria and myxobacteria. The synthesis of sterols observed in G. obscuriglobus is unique within Planctomycetes. Sterol synthesis is suggested to be associated with regulation of membrane fluidity in Planctomycetes,[4] and has been described as essential to the proper growth and reproduction of G. obscuriglobus.[38]
7. Biotechnology and Human Applications
Recently, interest has arisen in examining the Planctomycetes regarding their potential roles in biotechnology, mainly as a source of bioactive molecules,[29][39] of interest mainly to the pharmaceutical industry. Bioactive compounds are mainly present as secondary metabolites,[39] although little is known about Planctomycetes secondary metabolites.[40] This is unexpected, as the Planctomycetes have several key features as other known producers of bioactive molecules, such as the Myxobacteria.[40] However, a number of ongoing studies serve as various first steps in including Planctomycetes in small-molecule drug development for humans.
Planctomycetes spp. are worthwhile considerations in challenging the current models for the origin of the nucleus, along with other aspects of origin and evolution of the eukaryotic endomembrane system.[28]
7.1. Climate Change
The impacts of research on Planctomycetes and their uses might be of global significance with regards to nutrient cycling processes and assist in furthering understanding for global marine biogeochemistry. However, with Planctomycetes' growing influences on metabolic processes involving water and air, it may also have a role in interchanges between oceans and atmosphere, potentially affecting climate change.[28]
7.2. Planctomycetes as Pathogens
Planctomycetes spp. were recently identified as being an opportunistic human pathogen, but a lack of culture media limits studies on the bacteria in the Planctomycetes as pathogens of humans.[29]
References
- "Three-dimensional reconstruction of bacteria with a complex endomembrane system". PLOS Biology 11 (5): e1001565. 2013-05-21. doi:10.1371/journal.pbio.1001565. PMID 23700385. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3660258
- "Complete genome sequence of the marine planctomycete Pirellula sp. strain 1". Proceedings of the National Academy of Sciences of the United States of America 100 (14): 8298–303. July 2003. doi:10.1073/pnas.1431443100. PMID 12835416. Bibcode: 2003PNAS..100.8298G. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=166223
- "Insights into the ultrastructural morphology of novel Planctomycetes". Antonie van Leeuwenhoek 104 (4): 467–76. October 2013. doi:10.1007/s10482-013-9969-2. PMID 23857394. https://dx.doi.org/10.1007%2Fs10482-013-9969-2
- "Non-essentiality of canonical cell division genes in the planctomycete Planctopirus limnophila". Scientific Reports 10 (1): 66. January 2020. doi:10.1038/s41598-019-56978-8. PMID 31919386. Bibcode: 2020NatSR..10...66R. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=6952346
- "On the maverick Planctomycetes". FEMS Microbiology Reviews 42 (6): 739–760. November 2018. doi:10.1093/femsre/fuy029. PMID 30052954. https://dx.doi.org/10.1093%2Ffemsre%2Ffuy029
- "Determining the bacterial cell biology of Planctomycetes". Nature Communications 8 (1): 14853. April 2017. doi:10.1038/ncomms14853. PMID 28393831. Bibcode: 2017NatCo...814853B. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5394234
- "Cultivation and functional characterization of 79 planctomycetes uncovers their unique biology". Nature Microbiology 5 (1): 126–140. January 2020. doi:10.1038/s41564-019-0588-1. PMID 31740763. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=7286433
- "Pink- and orange-pigmented planctomycetes produce saproxanthin-type carotenoids including a rare C45 carotenoid". Environmental Microbiology Reports 11 (6): 741–748. December 2019. doi:10.1111/1758-2229.12796. PMID 31600855. http://plymsea.ac.uk/id/eprint/8248/1/1758-2229.12796.pdf.
- "High ultraviolet C resistance of marine Planctomycetes". Antonie van Leeuwenhoek 104 (4): 585–95. October 2013. doi:10.1007/s10482-013-0027-x. PMID 24052365. https://dx.doi.org/10.1007%2Fs10482-013-0027-x
- "Fuerstia marisgermanicae gen. nov., sp. nov., an Unusual Member of the Phylum Planctomycetes from the German Wadden Sea" (in English). Frontiers in Microbiology 7: 2079. 2016. doi:10.3389/fmicb.2016.02079. PMID 28066393. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5177795
- "Alienimonas californiensis gen. nov. sp. nov., a novel Planctomycete isolated from the kelp forest in Monterey Bay". Antonie van Leeuwenhoek 113 (12): 1751–1766. December 2020. doi:10.1007/s10482-019-01367-4. PMID 31802338. https://dx.doi.org/10.1007%2Fs10482-019-01367-4
- Kartal, Boran; de Almeida, Naomi M.; Maalcke, Wouter J.; Op den Camp, Huub J.M.; Jetten, Mike S.M.; Keltjens, Jan T. (2013-05-01). "How to make a living from anaerobic ammonium oxidation". FEMS Microbiology Reviews 37 (3): 428–461. doi:10.1111/1574-6976.12014. ISSN 0168-6445. PMID 23210799. https://dx.doi.org/10.1111%2F1574-6976.12014
- Kohn, Timo; Rast, Patrick; Kallscheuer, Nicolai; Wiegand, Sandra; Boedeker, Christian; Jetten, Mike S. M.; Jeske, Olga; Vollmers, John et al. (2020). "The Microbiome of Posidonia oceanica Seagrass Leaves Can Be Dominated by Planctomycetes" (in English). Frontiers in Microbiology 11: 1458. doi:10.3389/fmicb.2020.01458. ISSN 1664-302X. PMID 32754127. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=7366357
- "Three marine strains constitute the novel genus and species Crateriforma conspicua in the phylum Planctomycetes". Antonie van Leeuwenhoek 113 (12): 1797–1809. December 2020. doi:10.1007/s10482-019-01375-4. PMID 31894495. https://dx.doi.org/10.1007%2Fs10482-019-01375-4
- "Characterization of Planctomyces limnophilus and development of genetic tools for its manipulation establish it as a model species for the phylum Planctomycetes". Applied and Environmental Microbiology 77 (16): 5826–9. August 2011. doi:10.1128/AEM.05132-11. PMID 21724885. Bibcode: 2011ApEnM..77.5826J. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3165242
- "Evolutionary Cell Biology of Division Mode in the Bacterial Planctomycetes-Verrucomicrobia- Chlamydiae Superphylum" (in English). Frontiers in Microbiology 7: 1964. 2016. doi:10.3389/fmicb.2016.01964. PMID 28018303. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=5147048
- ""Candidatus Laterigemmans baculatus" gen. nov. sp. nov., the first representative of rod shaped planctomycetes with lateral budding in the family Pirellulaceae". Systematic and Applied Microbiology 44 (2): 126188. April 2021. doi:10.1016/j.syapm.2021.126188. PMID 33647766. https://dx.doi.org/10.1016%2Fj.syapm.2021.126188
- "FtsZ-less cell division in archaea and bacteria". Current Opinion in Microbiology. Growth and development: eukaryotes/prokaryotes 13 (6): 747–52. December 2010. doi:10.1016/j.mib.2010.10.005. PMID 21050804. https://dx.doi.org/10.1016%2Fj.mib.2010.10.005
- "Molecular Signatures for the PVC Clade (Planctomycetes, Verrucomicrobia, Chlamydiae, and Lentisphaerae) of Bacteria Provide Insights into Their Evolutionary Relationships". Frontiers in Microbiology 3: 327. 2012. doi:10.3389/fmicb.2012.00327. PMID 23060863. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3444138
- "Impact of genomics on the understanding of microbial evolution and classification: the importance of Darwin's views on classification". FEMS Microbiology Reviews 40 (4): 520–53. July 2016. doi:10.1093/femsre/fuw011. PMID 27279642. https://dx.doi.org/10.1093%2Ffemsre%2Ffuw011
- "Signature protein of the PVC superphylum". Applied and Environmental Microbiology 80 (2): 440–5. January 2014. doi:10.1128/AEM.02655-13. PMID 24185849. Bibcode: 2014ApEnM..80..440L. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3911108
- Wagner, Michael; Horn, Matthias (2006-06-01). "The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance" (in en). Current Opinion in Biotechnology. Environmental biotechnology/Energy biotechnology 17 (3): 241–249. doi:10.1016/j.copbio.2006.05.005. ISSN 0958-1669. PMID 16704931. https://www.sciencedirect.com/science/article/pii/S0958166906000644.
- Mahajan, Mayank; Yee, Benjamin; Hägglund, Emil; Guy, Lionel; Fuerst, John A; Andersson, Siv G E (2019-12-11). "Paralogization and New Protein Architectures in Planctomycetes Bacteria with Complex Cell Structures". Molecular Biology and Evolution 37 (4): 1020–1040. doi:10.1093/molbev/msz287. ISSN 0737-4038. http://dx.doi.org/10.1093/molbev/msz287.
- "Deciphering the evolution and metabolism of an anammox bacterium from a community genome". Nature 440 (7085): 790–794. 6 April 2006. doi:10.1038/nature04647. PMID 16598256. Bibcode: 2006Natur.440..790S. https://dx.doi.org/10.1038%2Fnature04647
- "16S rRNA-based LTP release 123 (full tree)". Silva Comprehensive Ribosomal RNA Database. http://www.arb-silva.de/fileadmin/silva_databases/living_tree/LTP_release_123/LTPs123_SSU_tree.pdf.
- Fuerst, John (2004). "Planctomycetes: a phylum of emerging interest for microbial evolution and ecology.". World Federation for Culture Collections Newsletter 38. http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.538.2883&rep=rep1&type=pdf.
- Buckley, Daniel H.; Huangyutitham, Varisa; Nelson, Tyrrell A.; Rumberger, Angelika; Thies, Janice E. (2006-07-01). "Diversity of Planctomycetes in Soil in Relation to Soil History and Environmental Heterogeneity" (in en). Applied and Environmental Microbiology 72 (7): 4522–4531. doi:10.1128/AEM.00149-06. ISSN 0099-2240. PMID 16820439. Bibcode: 2006ApEnM..72.4522B. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1489350
- Fuerst, J. A. (1995-07-01). "The planctomycetes: emerging models for microbial ecology, evolution and cell biology". Microbiology 141 (7): 1493–1506. doi:10.1099/13500872-141-7-1493. ISSN 1350-0872. PMID 7551018. http://dx.doi.org/10.1099/13500872-141-7-1493.
- "Planctomycetes as Host-Associated Bacteria: A Perspective That Holds Promise for Their Future Isolations, by Mimicking Their Native Environmental Niches in Clinical Microbiology Laboratories" (in English). Frontiers in Cellular and Infection Microbiology 10: 519301. 2020. doi:10.3389/fcimb.2020.519301. PMID 33330115. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=7734314
- "Fosmids of novel marine Planctomycetes from the Namibian and Oregon coast upwelling systems and their cross-comparison with planctomycete genomes". The ISME Journal 1 (5): 419–35. September 2007. doi:10.1038/ismej.2007.63. PMID 18043661. https://dx.doi.org/10.1038%2Fismej.2007.63
- "Community composition of the Planctomycetes associated with different macroalgae". FEMS Microbiology Ecology 88 (3): 445–56. June 2014. doi:10.1111/1574-6941.12258. PMID 24266389. https://dx.doi.org/10.1111%2F1574-6941.12258
- "Intracellular compartmentation in planctomycetes". Annual Review of Microbiology 59 (1): 299–328. October 2005. doi:10.1146/annurev.micro.59.030804.121258. PMID 15910279. https://dx.doi.org/10.1146%2Fannurev.micro.59.030804.121258
- "Isolation and diversity of planctomycetes from the sponge Niphates sp., seawater, and sediment of Moreton Bay, Australia". Antonie van Leeuwenhoek 104 (4): 533–46. October 2013. doi:10.1007/s10482-013-0003-5. PMID 23959164. https://dx.doi.org/10.1007%2Fs10482-013-0003-5
- Kartal, Boran (2011). Cultivation, Detection, andEcophysiology of AnaerobicAmmonium-Oxidizing Bacteria. San Diego CA: Academic Press. pp. 89–108. ISBN 978-0-12-381294-0.
- Qian (2018). "Diversity and distribution of anammox bacteria in water column and sediments of the Eastern Indian Ocean". International Biodeterioration & Biodegradation 133: 52–62. doi:10.1016/j.ibiod.2018.05.015. https://dx.doi.org/10.1016%2Fj.ibiod.2018.05.015
- Teeseling (2015). "Anammox Planctomycetes have a peptidoglycan cell wall". Nature Communications 6: 6878. doi:10.1038/ncomms7878. PMID 25962786. Bibcode: 2015NatCo...6.6878V. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4432595
- Jing (2015). "A snapshot on spatial and vertical distribution of bacterial communities in the eastern Indian Ocean". Acta Oceanologica Sinica 35 (6): 85–93. doi:10.1007/s13131-016-0871-4. https://dx.doi.org/10.1007%2Fs13131-016-0871-4
- Rivas-Marin, Elena; Stettner, Sean; Gottshall, Ekaterina Y.; Santana-Molina, Carlos; Helling, Mitch; Basile, Franco; Ward, Naomi L.; Devos, Damien P. (2019-07-02). "Essentiality of sterol synthesis genes in the planctomycete bacterium Gemmata obscuriglobus" (in en). Nature Communications 10 (1): 2916. doi:10.1038/s41467-019-10983-7. ISSN 2041-1723. PMID 31266954. Bibcode: 2019NatCo..10.2916R. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=6606645
- "Planctomycetes as Novel Source of Bioactive Molecules" (in English). Frontiers in Microbiology 7: 1241. 2016. doi:10.3389/fmicb.2016.01241. PMID 27570520. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=4982196
- Jeske, Olga; Jogler, Mareike; Petersen, Jörn; Sikorski, Johannes; Jogler, Christian (2013-10-01). "From genome mining to phenotypic microarrays: Planctomycetes as source for novel bioactive molecules" (in en). Antonie van Leeuwenhoek 104 (4): 551–567. doi:10.1007/s10482-013-0007-1. ISSN 1572-9699. PMID 23982431. https://doi.org/10.1007/s10482-013-0007-1.





