Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Mohamed Abouzid | -- | 2781 | 2022-11-11 18:46:57 | | | |
| 2 | Conner Chen | + 13 word(s) | 2794 | 2022-11-15 07:46:33 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Ahmed, A.A.; Abouzid, M.; Kaczmarek, E. Deep Learning Applications in Tumor Pathology. Encyclopedia. Available online: https://encyclopedia.pub/entry/34215 (accessed on 06 June 2026).
Ahmed AA, Abouzid M, Kaczmarek E. Deep Learning Applications in Tumor Pathology. Encyclopedia. Available at: https://encyclopedia.pub/entry/34215. Accessed June 06, 2026.
Ahmed, Alhassan Ali, Mohamed Abouzid, Elżbieta Kaczmarek. "Deep Learning Applications in Tumor Pathology" Encyclopedia, https://encyclopedia.pub/entry/34215 (accessed June 06, 2026).
Ahmed, A.A., Abouzid, M., & Kaczmarek, E. (2022, November 11). Deep Learning Applications in Tumor Pathology. In Encyclopedia. https://encyclopedia.pub/entry/34215
Ahmed, Alhassan Ali, et al. "Deep Learning Applications in Tumor Pathology." Encyclopedia. Web. 11 November, 2022.
Copy Citation
The revolution of artificial intelligence and its impacts on our daily life has led to tremendous interest in the field and its related subtypes: machine learning and deep learning. Scientists and developers have designed machine learning- and deep learning-based algorithms to perform various tasks related to tumor pathologies, such as tumor detection, classification, grading with variant stages, diagnostic forecasting, recognition of pathological attributes, pathogenesis, and genomic mutations. Pathologists are interested in artificial intelligence to improve the diagnosis precision impartiality and to minimize the workload combined with the time consumed, which affects the accuracy of the decision taken.
artificial intelligence
image analysis
deep learning
machine learning
1. Diagnosis of Tumor
Pathologists must differentiate cancer from healthy cells and malignant from benign tumors, and these distinctions may significantly impact clinical decisions for various therapeutic approaches. Researchers have been able to develop artificial intelligence (AI) algorithms for that purpose; for instance, convolutional neural network (CNN)-based AI algorithms have been designed by Bardou et al. [1] (Figure 1).
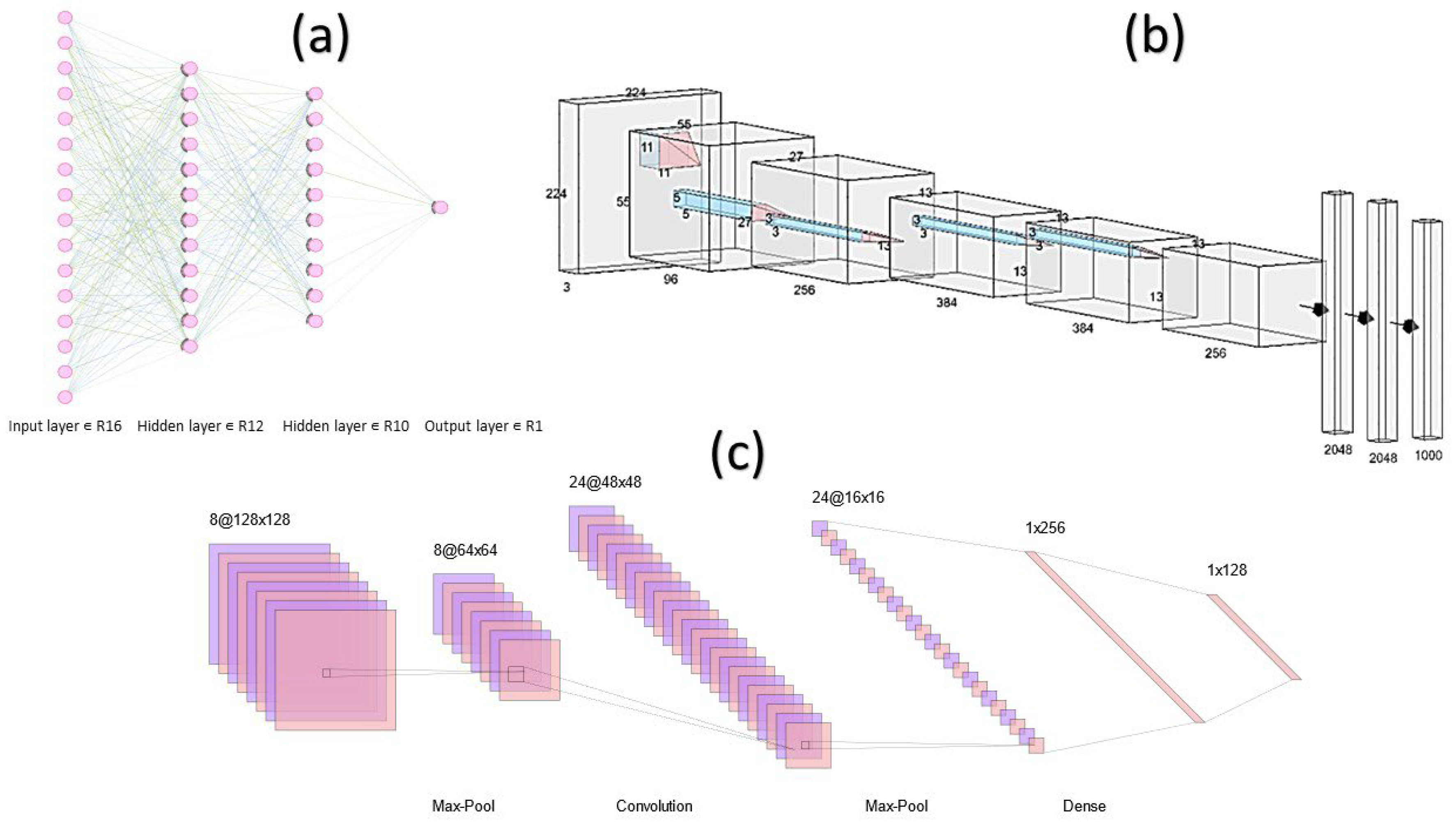
To distinguish the whole-slide images (WSIs) of breast cancer into two groups (cancer and non-cancer) with a precision level of 83.3% and categorize the result into four groups (healthy tissue, benign lesions, cancer in situ, and invasive cancer) with 77.8% precision, a stacked CNN was first trained to identify relatively lower attributes and then used as an input dataset to build a higher level of the stacked network. This program was developed by Bejnordi et al. [5]. They could differentiate breast malignancy from typical lesions with a 0.962 of the regions under the recipient operating curve (AUC or AUROC) and characterize invasive ductal cancer, ductal cancer in situ, and benign lesions with a precision of 81.3% using this CNN model. Bejnordi et al. [6] developed an algorithm based on the CNN system to integrate known stroma attributes to differentiate benign lesions from breast cancer, taking into account the impact of stroma on tumors. Skilled pathologists and deep learning (DL)-based AI algorithms were able to distinguish between malignant and benign tissues of colorectal tumors [7][8], as well as skin cancer from nevi (the plural of nevus) [9]. Mercan et al. [10] categorized breast tumors as proliferative, non-proliferative, atypical hyperplasia, cancer in situ, and invasive cancer based on breast biopsy WSIs with an 81% accuracy. This was made by using weakly supervised DL models that significantly decreased the burden of labeling. With an 86.5% precision, Wang et al. [11] categorized lesions of gastric tissues into normal, dysplasia, and cancer, while Tomita et al. [12] classified esophageal tissue as cancer, dysplasia, and Barrett esophagus with an 83% precision. Pathologists should conduct cytology analysis parallel with biopsy and excision specimens as part of their regular work. In the images diagnosed based on liquid and smear samples for the cervical cytology, AI could identify cells as normal or abnormal with a precision of 98.3% and 98.6%, respectively [13]. Based on the attributes of the cell [14] or WSI level features [15], AI-based algorithms have the power to distinguish high-grade urothelial carcinoma and its suspected cases from other urine cytology. According to the cytological images, AI also demonstrated promising potential in the comparative diagnosis of thyroid tumors [16].
2. Classification of Tumor
Different subtypes of cancer have different therapeutic approaches. Images from biopsy samples, frosted tissues, and formalin-fixed paraffin-embedded (FFPE) tissues showed a high AUC (0.83–0.97) in a study that used a CNN-based algorithm to directly separate non-small cell lung cancer (NSCLC) into squamous cell carcinoma, large cell carcinoma, adenocarcinoma, and normal lung tissue [17]. Bearing in mind the divergent patterns of lung adenocarcinoma cell growth that have been linked to patient clinical results, the CNN model designed by Gertych et al. [18] and Wei et al. [19] was used to classify every single image tile considering the pattern of growth for each individual and produce a likelihood map for the WSI, making it easier for pathologists to describe the principal and malignant elements of lung adenocarcinoma, including papillary, micropapillary, solid, and acinar components, quantitatively. Cervical squamous cell carcinoma, colorectal polyp [20], thyroid tumor [21], ovarian cancer [22], and breast tumor [23] were all multi-classified using a DL-based AI. This ability allowed the AI-based models to identify the different lung cancer histological subtypes with a precision of 60% to 89% based on cytological images [24].
3. Grading of Tumor
Pathologists evaluate tumor grades mainly depending on the tumor cell variation, cell division, necrosis, glandular structure, and other contextual factors affecting treatment decisions and clinical surveillance. To determine the grade of gliomas, Ertosun and Rubin [25] designed two different CNNs: one was able to correctly classify the patients with low-grade glioma or with glioblastoma multiforme with a 96% accuracy, while the other was able to distinguish the grade II glioma from grade III with a 71% accuracy. A CNN-based algorithm correctly identified medium-, moderate-, and high-grade breast cancers in 69% of breast biopsy images [26]. With a 91% precision, pathologists have used DL-based methods effectively to distinguish between the grades of colorectal adenocarcinoma into normal tissue, low-grade, and high-grade [8]. In the prostate cancer area, AI and machine learning (ML) algorithms have shown accurate and promising models in the grading process of prostate cancer. Several studies found that these models can achieve pathologist-level performance. One of the famous prostate cancer competitions is the PANDA challenge, which stands for Prostate cANcer graDe Assessment using the Gleason grading system [27]. The PANDA challenge involved 12,625 whole-slide images (WSIs) of prostate biopsies from 6 different areas and engaged 1010 groups from more than 60 countries, making it the most significant histopathology competition. The challenge system proved efficient, resulting in the first team achieving pathologist-level grading performance in only ten days. The PANDA challenge, hosted on the Kaggle platform in April–July 2020, rigorously validated the top-performing algorithms across international patient cohorts. Perincheri et al. developed a model from 118 cases to detect high-grade prostatic intraepithelial neoplasia with a 97.7% sensitivity and 99.3% specificity [28]. By using 549 slides for training and 2501 slides for testing, Pantanowitz et al. developed a model with 99.7% accuracy to detect atypical small acinar proliferation (ASAP) and perineural invasion (PNI) [29]. Moreover, Ström et al. created a model for prostate cancer detection and Gleason score using 6953 biopsies for training and 1718 biopsy for testing, resulting in a model with an AUC of 0.997 [30].
4. Staging of Tumor
Pathologists should have as many details as possible about excision samples for tumor node metastases (TNM) staging to achieve the proper treatment decisions. The developed CNN-based algorithm was able to identify three categories of the region of interest (ROI) in osteosarcomas, such as a tumor, non-tumor, and necrotic portion (e.g., cartilage, bones), on the patch level (around 64,000 patches from 82 WSIs) with a precision of 92.4% [31]. Additionally, it is possible to measure the rate of necrosis, a variable element in prognosis. For that purpose, numerous DL-based models were established to identify breast cancer tumor areas [10][32][33]. Pathologists must evaluate lymph node metastasis as part of tumor staging, but unfortunately, this process consumes time, and there is a possibility of false outcomes. Two AI models outperformed the pathologists’ findings in the Cancer Metastases in Lymph Nodes Challenge (CAMELYON16). The challenge aimed to compare the performance of AI systems and human pathologists in evaluating novel algorithms that detect the metastasis of cancer cells to lymph nodes in breast cancer. In slide-level diagnosis (recognizing whether cancer metastasis has existed), the best model achieved an AUC of 0.994.
Moreover, another two algorithms surpassed pathologists’ skill in detecting the level of lesions (identifying all metastases without discrete tumor cells) with the best mean accuracy obtained over six false-positive rates of 0.807 [34]. Furthermore, using the same dataset and sorting out artifacts, the more efficient algorithm, Lymph Node Assistant (LYNA), obtained a better AUC and sensitivity with values of 0.996 and 91%, respectively. It also revised and fixed two slides the producers had incorrectly diagnosed as “natural” [35]. Finally, the detection of micro-metastases in lymph nodes was significantly improved using LYNA, with the average accuracy increased by 8% (p = 0.02) to obtain 91% instead of 83% for all samples with a slightly faster assessment period [36].
In the last decade, several studies revealed that circulating tumor cells (CTCs) could be potential determinants in estimating cancer cells’ growth and development in metastatic [37][38], even with cancer patients at the early stages [39]. CTC counts above a certain threshold are linked to serious illness, heightened metastasis, and a shorter time to relapse [40]. CTCs are intended for use as a tool to measure tumor growth and facilitate clinical treatment, along with signaling treatment success, due to the ease and limited intrusion of blood collection [41]. Nevertheless, hindrances in technical matters, including limited supply and shortage of standard assays for detection and validated markers, hinder its therapeutic use [42]. According to Zeune et al. [43], DL-based CTC detection was comparatively stable with better precision than usual human opinions. In contrast, human reviewers and counting programs differed in their manual counting of CTCs from NSCLC and prostate cancer using images with fluorescence. Considering AI’s current role in recognizing tumor areas, identifying lymph node metastasis, and detecting CTCs, as well as its ability to process vast quantities of data, AI models could assist pathologists and oncologists in the process of tumor staging.
5. Assessment of Pathological Attributes
A tumor cell’s tendency to multiply is represented by mitosis. Though, counting mitosis takes time. Therefore, an effective algorithm was generated in the Assessment of Mitosis Detection Algorithms 2013 (AMIDA13) challenge to identify the mitoses of breast cancers at high-power fields (HPFs) with an 0.611 F1 score using 1000 images that could be compared to the inter-observer agreement [44] known protein structure. The Tumor Proliferation Assessment Challenge 2016 (TUPAC 16) [45] published breast cancer proliferation scores based on WSI-level AI recognition. Tumor budding is considered an offensive behavior of tumors; therefore, its analysis is crucial. Weis et al. [46] used CNN-based models to calculate the actual figure of tumor budding in cases of colorectal carcinoma. Moreover, they could determine the association between the hotspot and lymph node conditions. The type and quantity of tumor-penetrating immune cells have been linked to immunotherapy susceptibility and diagnostic stratification in cancer patients [47][48]. In breast cancer, a DL approach with a cluster of differentiation (CD)45 marked digital images could measure immunity cells and differentiate between areas rich in immune cells and regions poor in immune cells [49]. Therefore, one of the DL-based AI advantages is the ability to identify and recognize domain-agnostic and hand-crafted attributes that could be used in different diseases and types of tissues [50].
6. Assessment of Biomarkers
The DL-based model was designed by Saha et al. [51] to identify high proliferation areas and measure the severity of cancer metastasis in breast cells using the Ki-67 scale. In contrast, an AI-based model was designed by Vandenberghe et al. [52] to segment both interstitium and normal pancreatic tissues from tumor regions on uneven Ki-67 immunoreactive WSIs to calculate the severity of pancreatic tumors accurately, especially in neuroendocrine cells using the Ki-67 index. Moreover, several biomarkers match the patient profile with the adequate therapeutic regimen. Trastuzumab is a monoclonal antibody (Herceptin) used in treating gastric and breast cancer according to the human epidermal growth factor receptor 2 (HER2) condition. A CNN-based model with pathologist assistance achieved an average accuracy of 83% in determining the status of HER2 [53]. However, the results improved after dividing the cell membranes as the natural expression position of HER2. Likewise, in gastric cancer, an AI-based model was designed to evaluate HER2-negative regions (0 and 1+), HER2-positive regions (2+ and 3+), and regions with no tumor at all with 69.9% precision [54]. An AI-based model could detect the presence of programmed death-ligand 1 (PD-L1; positive or negative) by using hematoxylin and eosin (H&E)-stained images of adenocarcinoma or squamous carcinoma lung cancers with an AUC of 0.80. The result was reasonable compared to pathologist assessments depending on PD-L1 immunohistochemistry images to identify possible patients who may have sensitivity to pembrolizumab medication [20]. A DL-based AI model evaluated biomarkers engaged in the prognosis, diagnosis, and prediction of drug interactions depending on immunohistochemical dye or fluorescent dye WSIs and HE dye WSIs.
7. Assessment of Genetic Modifications
During WSI analysis, morphological variations are examples of fundamental genetic changes. Schaumberg et al. [55] used a group of 177 patients diagnosed with prostate cancer from the The Cancer Genome Atlas (TCGA), 20 of them had mutant speckle-type POZ protein (SPOP), to train several groups of the CNN model to determine whether a mutation occurred in the SPOP gene of prostate cancer or not. Then the obtained results could be validated and confirmed based on an independent cohort from MSK-IMPACT. Furthermore, since the SPOP gene mutation and TMPRSS2-ERG gene fusion are strictly incompatible [56], the estimation of SPOP mutation status offered indirect knowledge about TMPRSS2-ERG. Thus elucidating the importance of determining the SPOP gene mutation condition and its potential contribution to targeted therapy accuracy. Using the lung adenocarcinoma pathological images from TCGA, Coudray et al. [17] developed a DL-based model to anticipate the most common ten genes that had mutated. They pointed six of these genes (AUCs = 0.733–0.856), including epidermal growth factor receptor [EGFR], serine/threonine kinase 11 [STK11], SET binding protein 1 [SETBP1], FAT atypical cadherin 1 [FAT1], Kirsten rat sarcoma two viral oncogene homolog [KRAS], and TP53. Moreover, an AI-based algorithm was designed using the images of gastrointestinal cancer stained with H&E stains to determine microsatellite instability (MSI) or microsatellite stability (MSS) without conducting assays on microsatellite instability. The model tested 185 slides from Asian patients and showed robust snap-frozen samples and endometrial cancer with elevated AUC (0.77–0.84) [57]. They found that models tested and used on FFPE performed better than those tested on frozen and FFPE samples. A similar result appeared with colorectal cancer samples. Despite the designers mentioning that Asian patients have different histological gastric cancer than non-Asian patients, this model potentially provides beneficial immunotherapy solutions to a wide range of gastrointestinal cancer patients. It could be implemented lowly and not require testing for the tissues in laboratories to efficiently determine MSI tumors [57]. Therefore, patients with particular genetic alterations were classified using these AI-based models depending on inherent genetic-histologic associations, which assisted the medical team in providing the precise therapy regime.
8. Prognosis Prediction
Bychkov et al. [58] developed a DL-dependent approach for grouping patients into high- and low-risk classes based on images of colorectal cancer tissues stained with H&E stains. The technique achieved better results when using small tissue areas as input (hazard ratio [HR] 2.3; 95% CI: 1.79–3.03; AUC 0.69) compared with human experts (HR 1.67; 95% CI: 1.28–2.19; AUC 0.58) and WSIs (HR 1.65; 95% CI: 1.30–2.15; AUC 0.57), and it was proven to be an individual prognosis element using the multivariate Cox comparative analysis to examine hazard. In multicenter samples, Kather et al. [59] found that combined interstitium features (with lymphocytes, debris, adipose, desmoplastic stroma, and muscles) that were extracted using CNN might independently predict the survival rate and survival without relapse of colorectal cancer patients (HR = 2.29 vs. HR = 1.92, respectively), despite the stage of the clinical level. In lung adenocarcinoma [60] and glioma [61], it has been shown that DL-based models could estimate the risk of prognosis by learning and understanding histological characteristics. Kather et al. [57] designed an MSI-based model to predict overall survival in gastrointestinal cancer, the model was tried, and the results were impressive. According to the mentioned findings, AI-based models are suitable to be used as a predictor of health outcomes of cancer patients in addition to pathological diagnosis.
9. Different Algorithm Models for Tumors Detection
Many ML and DL algorithms in tumor detection are based on different ML methods such as Decision Trees (DTs), Artificial Neural Networks (ANNs), K-nearest neighbor (KNN), and Support Vector Machines (SVMs) [62]. One of these models is known as Deep Transfer Learning (TL), and a study used a bunch of grained classification approaches to detect the different types of brain tumors, including glioma and meningioma, with a model accuracy of 98.9% [63]. Another designed a CNN-based model called the Bayesian-YOLOv4 and was created to detect breast tumors with a scoring accuracy exceeding 92% in many training data [64]. Furthermore, a DL model was designed to detect liver tumors using an enhanced DL method called U-Net. This model combines DL algorithms and CT images resulting in a new algorithm known as Grey Wolf-Class Topper Optimization GW-CTO with a learning ability of 85% and an accuracy exceeding 90% [65]. Designing a multi-tasking AI algorithm that functions on multiple tumors is challenging. Therefore, to obtain satisfactory results, pathologists have to use a variety of AI-based algorithms for the entire pathological study, in which the neoplasm should be diagnosed, classified, and staged by various models of the algorithm, and a separate algorithm should evaluate the characteristic high-risk tumors. A DL-based model was designed by Couture et al. [66] to conduct several studies on images of breast cancer tissues stained with H&E. The performed tasks include identifying the histological subtype (lobular or ductal) with a precision of 94%, grading based on histological characters (low-, moderate-, and high-grade), which obtained an 82% precision, and evaluating the receptor’s condition of estrogen hormone (negative or positive) with an accuracy of 77%, in addition to classifying the relapse risk (low, moderate, and high risk) with an accuracy of 76%.
References
- Bardou, D.; Zhang, K.; Ahmad, S.M. Classification of Breast Cancer Based on Histology Images Using Convolutional Neural Networks. IEEE Access 2018, 6, 24680–24693.
- LeNail, A. NN-SVG: Publication-Ready Neural Network Architecture Schematics. J. Open Source Softw. 2019, 4, 747.
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Commun. ACM 2017, 60, 84–90.
- Lecun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-Based Learning Applied to Document Recognition. Proc. IEEE 1998, 86, 2278–2324.
- Bejnordi, B.E.; Zuidhof, G.; Balkenhol, M.; Hermsen, M.; Bult, P.; van Ginneken, B.; Karssemeijer, N.; Litjens, G.; van der Laak, J. Context-Aware Stacked Convolutional Neural Networks for Classification of Breast Carcinomas in Whole-Slide Histopathology Images. J. Med. Imaging 2017, 4, 044504.
- Bejnordi, B.E.; Mullooly, M.; Pfeiffer, R.M.; Fan, S.; Vacek, P.M.; Weaver, D.L.; Herschorn, S.; Brinton, L.A.; van Ginneken, B.; Karssemeijer, N.; et al. Using Deep Convolutional Neural Networks to Identify and Classify Tumor-Associated Stroma in Diagnostic Breast Biopsies. Mod. Pathol. 2018, 31, 1502–1512.
- Kainz, P.; Pfeiffer, M.; Urschler, M. Segmentation and Classification of Colon Glands with Deep Convolutional Neural Networks and Total Variation Regularization. PeerJ 2017, 2017, 1–28.
- Awan, R.; Sirinukunwattana, K.; Epstein, D.; Jefferyes, S.; Qidwai, U.; Aftab, Z.; Mujeeb, I.; Snead, D.; Rajpoot, N. Glandular Morphometrics for Objective Grading of Colorectal Adenocarcinoma Histology Images. Sci. Rep. 2017, 7, 16852.
- Wang, L.; Ding, L.; Liu, Z.; Sun, L.; Chen, L.; Jia, R.; Dai, X.; Cao, J.; Ye, J. Automated Identification of Malignancy in Whole-Slide Pathological Images: Identification of Eyelid Malignant Melanoma in Gigapixel Pathological Slides Using Deep Learning. Br. J. Ophthalmol. 2020, 104, 318–323.
- Mercan, C.; Aksoy, S.; Mercan, E.; Shapiro, L.G.; Weaver, D.L.; Elmore, J.G. Multi-Instance Multi-Label Learning for Multi-Class Classification of Whole Slide Breast Histopathology Images. IEEE Trans. Med. Imaging 2018, 37, 316–325.
- Wang, S.; Zhu, Y.; Yu, L.; Chen, H.; Lin, H.; Wan, X.; Fan, X.; Heng, P.A. RMDL: Recalibrated Multi-Instance Deep Learning for Whole Slide Gastric Image Classification. Med. Image Anal. 2019, 58, 101549.
- Tomita, N.; Abdollahi, B.; Wei, J.; Ren, B.; Suriawinata, A.; Hassanpour, S. Attention-Based Deep Neural Networks for Detection of Cancerous and Precancerous Esophagus Tissue on Histopathological Slides. JAMA Netw. Open 2019, 2, e1914645.
- Zhang, L.; Lu, L.; Nogues, I.; Summers, R.M.; Liu, S.; Yao, J. DeepPap: Deep Convolutional Networks for Cervical Cell Classification. IEEE J. Biomed. Health Inform. 2017, 21, 1633–1643.
- Vaickus, L.J.; Suriawinata, A.A.; Wei, J.W.; Liu, X. Automating the Paris System for Urine Cytopathology—A Hybrid Deep-Learning and Morphometric Approach. Cancer Cytopathol. 2019, 127, 98–115.
- Sanghvi, A.B.; Allen, E.Z.; Callenberg, K.M.; Pantanowitz, L. Performance of an Artificial Intelligence Algorithm for Reporting Urine Cytopathology. Cancer Cytopathol. 2019, 127, 658–666.
- Guan, Q.; Wang, Y.; Ping, B.; Li, D.; Du, J.; Qin, Y.; Lu, H.; Wan, X.; Xiang, J. Deep Convolutional Neural Network VGG-16 Model for Differential Diagnosing of Papillary Thyroid Carcinomas in Cytological Images: A Pilot Study. J. Cancer 2019, 10, 4876–4882.
- Coudray, N.; Ocampo, P.S.; Sakellaropoulos, T.; Narula, N.; Snuderl, M.; Fenyö, D.; Moreira, A.L.; Razavian, N.; Tsirigos, A. Classification and Mutation Prediction from Non–Small Cell Lung Cancer Histopathology Images Using Deep Learning. Nat. Med. 2018, 24, 1559–1567.
- Gertych, A.; Swiderska-Chadaj, Z.; Ma, Z.; Ing, N.; Markiewicz, T.; Cierniak, S.; Salemi, H.; Guzman, S.; Walts, A.E.; Knudsen, B.S. Convolutional Neural Networks Can Accurately Distinguish Four Histologic Growth Patterns of Lung Adenocarcinoma in Digital Slides. Sci. Rep. 2019, 9, 1483.
- Wei, J.W.; Tafe, L.J.; Linnik, Y.A.; Vaickus, L.J.; Tomita, N.; Hassanpour, S. Pathologist-Level Classification of Histologic Patterns on Resected Lung Adenocarcinoma Slides with Deep Neural Networks. Sci. Rep. 2019, 9, 3358.
- Barbieri, A.L.; Fadare, O.; Fan, L.; Singh, H.; Parkash, V. Challenges in Communication from Referring Clinicians to Pathologists in the Electronic Health Record Era. J. Pathol. Inform. 2018, 9, 8.
- Wang, Y.; Guan, Q.; Lao, I.; Wang, L.; Wu, Y.; Li, D.; Ji, Q.; Wang, Y.; Zhu, Y.; Lu, H.; et al. Using Deep Convolutional Neural Networks for Multi-Classification of Thyroid Tumor by Histopathology: A Large-Scale Pilot Study. Ann. Transl. Med. 2019, 7, 468.
- Wu, M.; Yan, C.; Liu, H.; Liu, Q. Automatic Classification of Ovarian Cancer Types from Cytological Images Using Deep Convolutional Neural Networks. Biosci. Rep. 2018, 38, BSR20180289.
- Jiang, Y.; Chen, L.; Zhang, H.; Xiao, X. Breast Cancer Histopathological Image Classification Using Convolutional Neural Networks with Small SE-ResNet Module. PLoS ONE 2019, 14, e0214587.
- Teramoto, A.; Tsukamoto, T.; Kiriyama, Y.; Fujita, H. Automated Classification of Lung Cancer Types from Cytological Images Using Deep Convolutional Neural Networks. BioMed Res. Int. 2017, 2017, 4067832.
- Ertosun, M.G.; Rubin, D.L. Automated Grading of Gliomas Using Deep Learning in Digital Pathology Images: A Modular Approach with Ensemble of Convolutional Neural Networks. AMIA Annu. Symp. Proc. MIA Symp. 2015, 2015, 1899–1908.
- Wan, T.; Cao, J.; Chen, J.; Qin, Z. Automated Grading of Breast Cancer Histopathology Using Cascaded Ensemble with Combination of Multi-Level Image Features. Neurocomputing 2017, 229, 34–44.
- Bashashati, A.; Goldenberg, S.L. AI for Prostate Cancer Diagnosis—Hype or Today’s Reality? Nat. Rev. Urol. 2022, 19, 261–262.
- Perincheri, S.; Levi, A.W.; Celli, R.; Gershkovich, P.; Rimm, D.; Morrow, J.S.; Rothrock, B.; Raciti, P.; Klimstra, D.; Sinard, J. An Independent Assessment of an Artificial Intelligence System for Prostate Cancer Detection Shows Strong Diagnostic Accuracy. Mod. Pathol. 2021, 34, 1588–1595.
- Pantanowitz, L.; Quiroga-Garza, G.M.; Bien, L.; Heled, R.; Laifenfeld, D.; Linhart, C.; Sandbank, J.; Albrecht Shach, A.; Shalev, V.; Vecsler, M.; et al. An Artificial Intelligence Algorithm for Prostate Cancer Diagnosis in Whole Slide Images of Core Needle Biopsies: A Blinded Clinical Validation and Deployment Study. Lancet Digit. Health 2020, 2, e407–e416.
- Ström, P.; Kartasalo, K.; Olsson, H.; Solorzano, L.; Delahunt, B.; Berney, D.M.; Bostwick, D.G.; Evans, A.J.; Grignon, D.J.; Humphrey, P.A.; et al. Artificial Intelligence for Diagnosis and Grading of Prostate Cancer in Biopsies: A Population-Based, Diagnostic Study. Lancet Oncol. 2020, 21, 222–232.
- Mishra, R.; Daescu, O.; Leavey, P.; Rakheja, D.; Sengupta, A. Convolutional Neural Network for Histopathological Analysis of Osteosarcoma. J. Comput. Biol. 2018, 25, 313–325.
- Cruz-Roa, A.; Gilmore, H.; Basavanhally, A.; Feldman, M.; Ganesan, S.; Shih, N.N.C.; Tomaszewski, J.; González, F.A.; Madabhushi, A. Accurate and Reproducible Invasive Breast Cancer Detection in Whole-Slide Images: A Deep Learning Approach for Quantifying Tumor Extent. Sci. Rep. 2017, 7, 46450.
- Cruz-Roa, A.; Gilmore, H.; Basavanhally, A.; Feldman, M.; Ganesan, S.; Shih, N.; Tomaszewski, J.; Madabhushi, A.; González, F. High-Throughput Adaptive Sampling for Whole-Slide Histopathology Image Analysis (HASHI) via Convolutional Neural Networks: Application to Invasive Breast Cancer Detection. PLoS ONE 2018, 13, e0196828.
- Bejnordi, B.E.; Veta, M.; Van Diest, P.J.; Van Ginneken, B.; Karssemeijer, N.; Litjens, G.; Van Der Laak, J.A.W.M.; Hermsen, M.; Manson, Q.F.; Balkenhol, M.; et al. Diagnostic Assessment of Deep Learning Algorithms for Detection of Lymph Node Metastases in Women with Breast Cancer. JAMA J. Am. Med. Assoc. 2017, 318, 2199–2210.
- Liu, Y.; Kohlberger, T.; Norouzi, M.; Dahl, G.E.; Smith, J.L.; Mohtashamian, A.; Olson, N.; Peng, L.H.; Hipp, J.D.; Stumpe, M.C. Artificial Intelligence–Based Breast Cancer Nodal Metastasis Detection Insights into the Black Box for Pathologists. Arch. Pathol. Lab. Med. 2019, 143, 859–868.
- Steiner, D.F.; Macdonald, R.; Liu, Y.; Truszkowski, P.; Hipp, J.D.; Gammage, C.; Thng, F.; Peng, L.; Stumpe, M.C. Impact of Deep Learning Assistance on the Histopathologic Review of Lymph Nodes for Metastatic Breast Cancer. Am. J. Surg. Pathol. 2018, 42, 1636–1646.
- Cristofanilli, M. Circulating Tumor Cells, Disease Progression, and Survival in Metastatic Breast Cancer. Semin. Oncol. 2006, 33, 9–14.
- De Bono, J.S.; Scher, H.I.; Montgomery, R.B.; Parker, C.; Miller, M.C.; Tissing, H.; Doyle, G.V.; Terstappen, L.W.W.M.; Pienta, K.J.; Raghavan, D. Circulating Tumor Cells Predict Survival Benefit from Treatment in Metastatic Castration-Resistant Prostate Cancer. Clin. Cancer Res. 2008, 14, 6302–6309.
- Rhim, A.D.; Mirek, E.T.; Aiello, N.M.; Maitra, A.; Bailey, J.M.; McAllister, F.; Reichert, M.; Beatty, G.L.; Rustgi, A.K.; Vonderheide, R.H.; et al. EMT and Dissemination Precede Pancreatic Tumor Formation. Cell 2012, 148, 349–361.
- Chaffer, C.L.; Weinberg, R.A. A Perspective on Cancer Cell Metastasis. Science 2011, 331, 1559–1564.
- Pantel, K.; Alix-Panabières, C. Real-Time Liquid Biopsy in Cancer Patients: Fact or Fiction? Cancer Res. 2013, 73, 6384–6388.
- Strati, A.; Kasimir-Bauer, S.; Markou, A.; Parisi, C.; Lianidou, E.S. Comparison of Three Molecular Assays for the Detection and Molecular Characterization of Circulating Tumor Cells in Breast Cancer. Breast Cancer Res. 2013, 15, R20.
- Zeune, L.L.; de Wit, S.; Berghuis, A.M.S.; IJzerman, M.J.; Terstappen, L.W.M.M.; Brune, C. How to Agree on a CTC: Evaluating the Consensus in Circulating Tumor Cell Scoring. Cytom. Part A 2018, 93, 1202–1206.
- Veta, M.; van Diest, P.J.; Willems, S.M.; Wang, H.; Madabhushi, A.; Cruz-Roa, A.; Gonzalez, F.; Larsen, A.B.L.; Vestergaard, J.S.; Dahl, A.B.; et al. Assessment of Algorithms for Mitosis Detection in Breast Cancer Histopathology Images. Med. Image Anal. 2015, 20, 237–248.
- Veta, M.; Heng, Y.J.; Stathonikos, N.; Bejnordi, B.E.; Beca, F.; Wollmann, T.; Rohr, K.; Shah, M.A.; Wang, D.; Rousson, M.; et al. Predicting Breast Tumor Proliferation from Whole-Slide Images: The TUPAC16 Challenge. Med. Image Anal. 2019, 54, 111–121.
- Weis, C.A.; Kather, J.N.; Melchers, S.; Al-ahmdi, H.; Pollheimer, M.J.; Langner, C.; Gaiser, T. Automatic Evaluation of Tumor Budding in Immunohistochemically Stained Colorectal Carcinomas and Correlation to Clinical Outcome. Diagn. Pathol. 2018, 13, 64.
- Halama, N.; Michel, S.; Kloor, M.; Zoernig, I.; Benner, A.; Spille, A.; Pommerencke, T.; Von Knebel Doeberitz, M.; Folprecht, G.; Luber, B.; et al. Localization and Density of Immune Cells in the Invasive Margin of Human Colorectal Cancer Liver Metastases Are Prognostic for Response to Chemotherapy. Cancer Res. 2011, 71, 5670–5677.
- Savas, P.; Salgado, R.; Denkert, C.; Sotiriou, C.; Darcy, P.K.; Smyth, M.J.; Loi, S. Clinical Relevance of Host Immunity in Breast Cancer: From TILs to the Clinic. Nat. Rev. Clin. Oncol. 2016, 13, 228–241.
- Turkki, R.; Linder, N.; Kovanen, P.E.; Pellinen, T.; Lundin, J. Antibody-Supervised Deep Learning for Quantification of Tumor-Infiltrating Immune Cells in Hematoxylin and Eosin Stained Breast Cancer Samples. J. Pathol. Inform. 2016, 7, 38.
- Aprupe, L.; Litjens, G.; Brinker, T.J.; Van Der Laak, J.; Grabe, N. Robust and Accurate Quantification of Biomarkers of Immune Cells in Lung Cancer Micro-Environment Using Deep Convolutional Neural Networks. PeerJ 2019, 2019, 1–16.
- Saha, M.; Chakraborty, C.; Arun, I.; Ahmed, R.; Chatterjee, S. An Advanced Deep Learning Approach for Ki-67 Stained Hotspot Detection and Proliferation Rate Scoring for Prognostic Evaluation of Breast Cancer. Sci. Rep. 2017, 7, 3213.
- Vandenberghe, M.E.; Scott, M.L.J.; Scorer, P.W.; Söderberg, M.; Balcerzak, D.; Barker, C. Relevance of Deep Learning to Facilitate the Diagnosis of HER2 Status in Breast Cancer. Sci. Rep. 2017, 7, 45938.
- Khameneh, F.D.; Razavi, S.; Kamasak, M. Automated Segmentation of Cell Membranes to Evaluate HER2 Status in Whole Slide Images Using a Modified Deep Learning Network. Comput. Biol. Med. 2019, 110, 164–174.
- Sharma, H.; Zerbe, N.; Klempert, I.; Hellwich, O.; Hufnagl, P. Deep Convolutional Neural Networks for Automatic Classification of Gastric Carcinoma Using Whole Slide Images in Digital Histopathology. Comput. Med. Imaging Graph. 2017, 61, 2–13.
- Schaumberg, A.; Rubin, M.; Fuchs, T. H&E-Stained Whole Slide Image Deep Learning Predicts SPOP Mutation State in Prostate Cancer. bioRxiv 2016, 064279.
- Barbieri, C.E.; Baca, S.C.; Lawrence, M.S.; Demichelis, F.; Blattner, M.; Theurillat, J.P.; White, T.A.; Stojanov, P.; Van Allen, E.; Stransky, N.; et al. Exome Sequencing Identifies Recurrent SPOP, FOXA1 and MED12 Mutations in Prostate Cancer. Nat. Genet. 2012, 44, 685–689.
- Kather, J.N.; Pearson, A.T.; Halama, N.; Jäger, D.; Krause, J.; Loosen, S.H.; Marx, A.; Boor, P.; Tacke, F.; Neumann, U.P.; et al. Deep Learning Can Predict Microsatellite Instability Directly from Histology in Gastrointestinal Cancer. Nat. Med. 2019, 25, 1054–1056.
- Bychkov, D.; Linder, N.; Turkki, R.; Nordling, S.; Kovanen, P.E.; Verrill, C.; Walliander, M.; Lundin, M.; Haglund, C.; Lundin, J. Deep Learning Based Tissue Analysis Predicts Outcome in Colorectal Cancer. Sci. Rep. 2018, 8, 3395.
- Kather, J.N.; Krisam, J.; Charoentong, P.; Luedde, T.; Herpel, E.; Weis, C.A.; Gaiser, T.; Marx, A.; Valous, N.A.; Ferber, D.; et al. Predicting Survival from Colorectal Cancer Histology Slides Using Deep Learning: A Retrospective Multicenter Study. PLoS Med. 2019, 16, e1002730.
- Wang, S.; Chen, A.; Yang, L.; Cai, L.; Xie, Y.; Fujimoto, J.; Gazdar, A.; Xiao, G. Comprehensive Analysis of Lung Cancer Pathology Images to Discover Tumor Shape and Boundary Features That Predict Survival Outcome. Sci. Rep. 2018, 8, 10393.
- Mobadersany, P.; Yousefi, S.; Amgad, M.; Gutman, D.A.; Barnholtz-Sloan, J.S.; Velázquez Vega, J.E.; Brat, D.J.; Cooper, L.A.D. Predicting Cancer Outcomes from Histology and Genomics Using Convolutional Networks. Proc. Natl. Acad. Sci. USA 2018, 115, E2970–E2979.
- Shaikh, F.J.; Rao, D.S. Prediction of Cancer Disease Using Machine Learning Approach. Mater. Today Proc. 2021, 50, 40–47.
- Ullah, N.; Khan, J.A.; Khan, M.S.; Khan, W.; Hassan, I.; Obayya, M.; Negm, N.; Salama, A.S. An Effective Approach to Detect and Identify Brain Tumors Using Transfer Learning. Appl. Sci. 2022, 12, 5645.
- Zhang, Z.; Li, Y.; Wu, W.; Chen, H.; Cheng, L.; Wang, S. Tumor Detection Using Deep Learning Method in Automated Breast Ultrasound. Biomed. Signal Process. Control 2021, 68, 102677.
- Rela, M.; Suryakari, N.R.; Patil, R.R. A Diagnosis System by U-Net and Deep Neural Network Enabled with Optimal Feature Selection for Liver Tumor Detection Using CT Images. Multimed. Tools Appl. 2022.
- Couture, H.D.; Williams, L.A.; Geradts, J.; Nyante, S.J.; Butler, E.N.; Marron, J.S.; Perou, C.M.; Troester, M.A.; Niethammer, M. Image Analysis with Deep Learning to Predict Breast Cancer Grade, ER Status, Histologic Subtype, and Intrinsic Subtype. npj Breast Cancer 2018, 4, 30.
More
Information
Subjects:
Medical Informatics
Contributors
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.3K
Revisions:
2 times
(View History)
Update Date:
15 Nov 2022
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No





